Impact of Stagnation on the Diversity of Cyanobacteria in Drinking Water Treatment Plant Sludge
Abstract
1. Introduction
2. Results and Discussions
2.1. Overview of Microbial/Cyanobacterial Diversity, Sludge Characteristics, and Microcystin Concentrations
2.2. Fate of Cyanobacterial Cells in the Sludge during Stagnation
2.3. Cyanobacteria-Laden Sludge Management
3. Conclusions
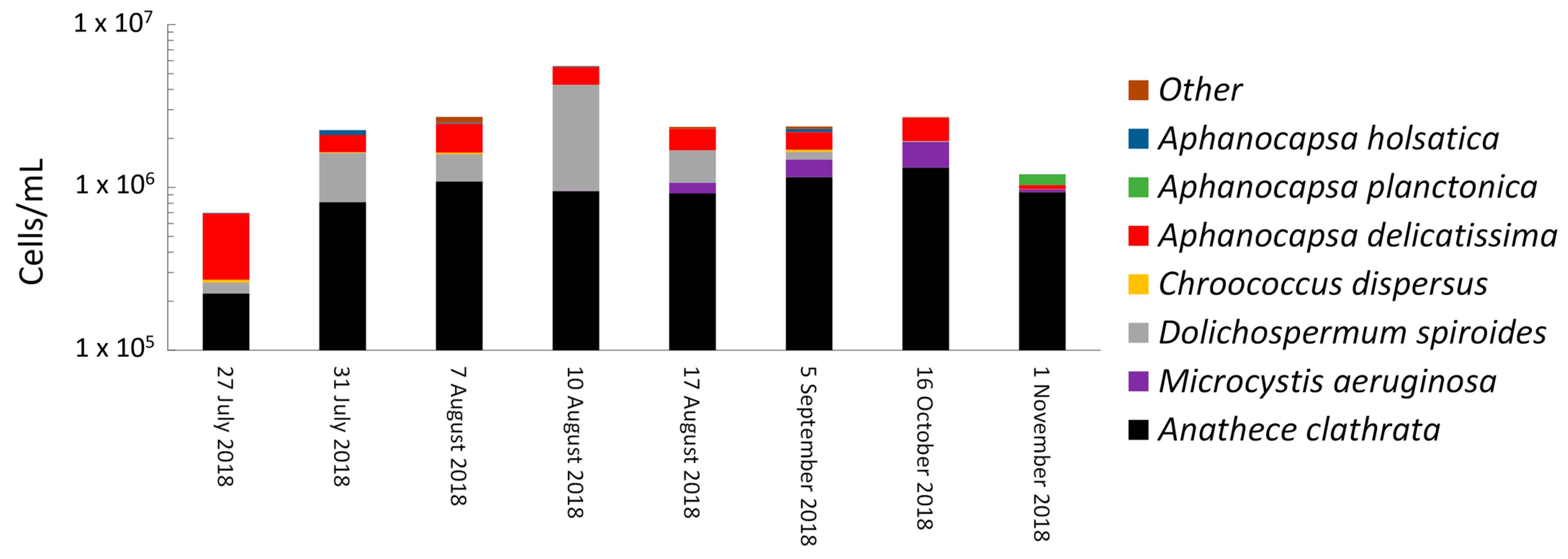
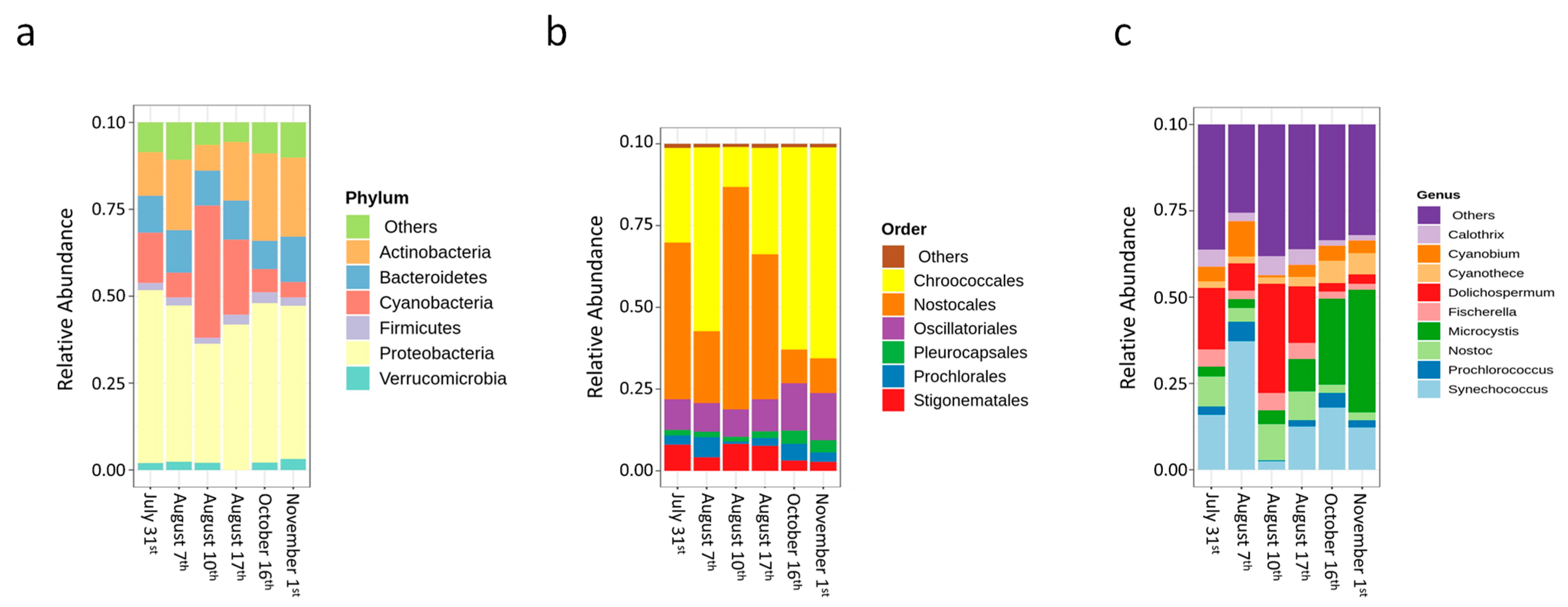
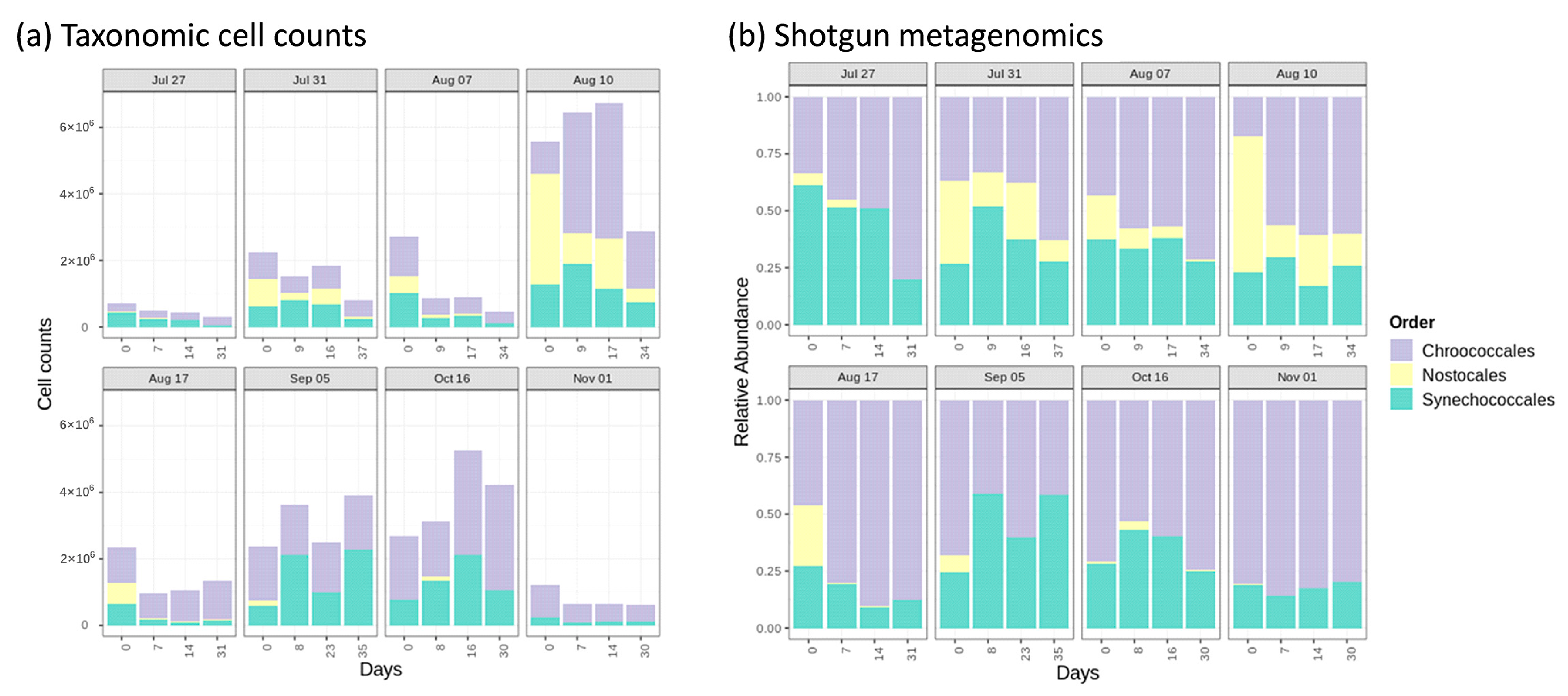
- Cyanobacterial growth, survival, and decay were quantified during sludge stagnation in the dark under controlled conditions. Longitudinal monitoring in summer and fall 2018 was conducted on sludge from a DWTP sludge holding tank. For four out of eight sampling dates, cyanobacterial cell growth was observed by total taxonomic cell counts during extended stagnation in the dark ranging from 7 to 38 days. The highest observed cell growth was 96% after 16 days of stagnation. The growth of cells was dominated by potential MC producers such as Microcystis, Aphanocapsa, Chroococcus, and Dolichospermum. Overall, up to 4200%, 1500%, 582%, and 134% cell growth was observed in potential MC-producer genera such as Aphanocapsa, Chroococcus, Dolichospermum, and Microcystis, respectively, during stagnation. Additionally, up to 35% cell growth was observed for non-toxic Anathece. Shotgun metagenomic sequencing revealed that sludge stagnation affected cyanobacterial diversity. Chroococcales (e.g., Microcystis) and Synechococcales (e.g., Synechococcus) were the most persistent orders, whereas Nostocales (e.g., Dolichospermum) was less resistant.
- Sludge characteristics including cyanobacterial cell counts and MCs dynamically changed in the sludge. Amongst studied physico-chemical parameters, only pH showed a significant correlation (p < 0.05) with the cyanobacterial community in sludge.
- Cyanobacterial biomarkers (level 4 subsystems) related to the “Circadian clock”, “rpoD-like sigma”, and “Pentose phosphate pathway” increased during stagnation, confirming cyanobacterial growth even in the dark. In contrast, the relative abundance of the “Heterocyst formation” biomarker related to filamentous genera declined in most of the stagnated samples.
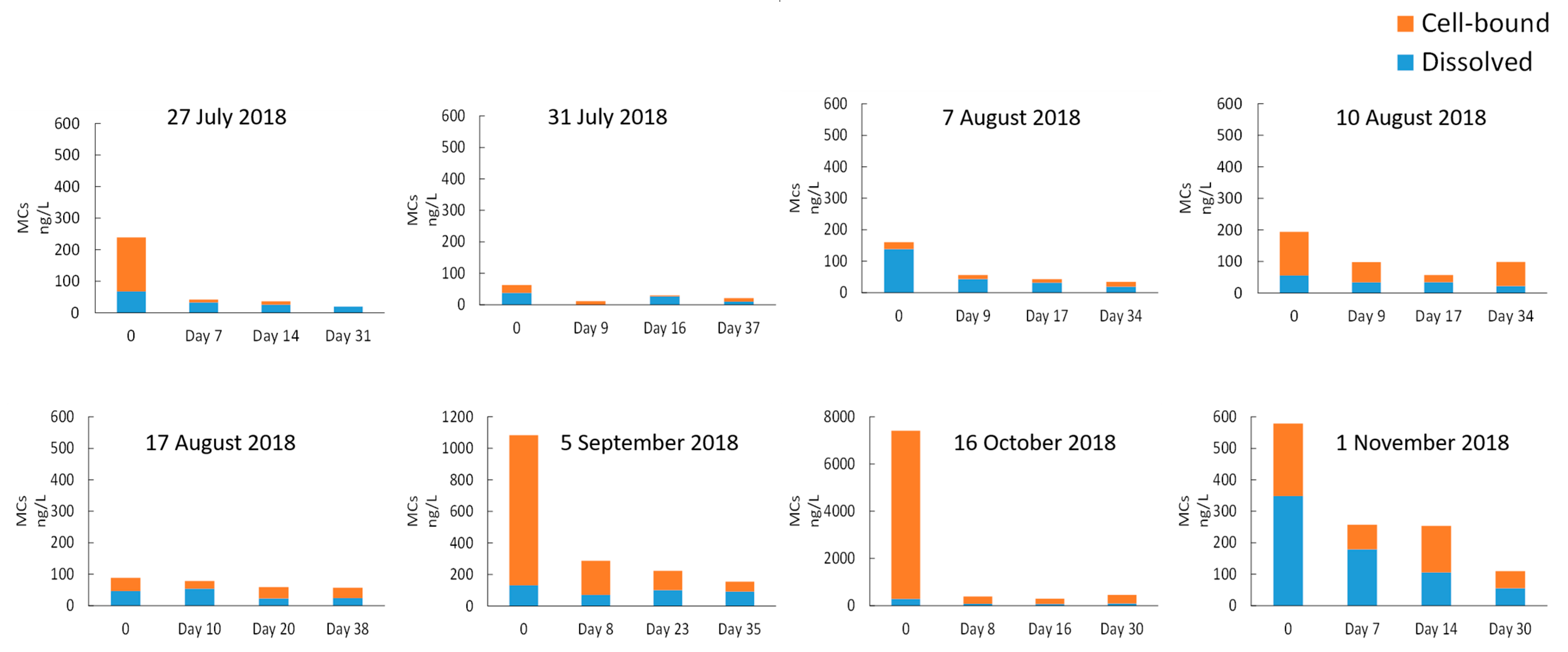
- Taxonomic cell counts, shotgun metagenomic sequencing, and cyanotoxin quantification provided consistent and complementary evidence regarding the quantification and dynamics of cyanobacteria in the stored sludge. This comprehensive investigation provides a sound basis to draft the best cyanobacteria-laden sludge management practices. Under the conditions tested, the persistence and/or growth of potential MC producers during storage raise the need to monitor cell counts and cyanotoxins in the sludge and its supernatant.
- Handling cyanobacteria-laden sludge is a challenge as the presence of cyanobacterial toxins raises limitations for their safe disposal. Furthermore, cyanotoxin-laden sludge represents a risk to the intake and potentially treated water quality if the sludge supernatant is recycled to the head of the DWTP or is discharged into the source.
4. Materials and Methods
4.1. Description of the Studied Plant, Treatment Processes, and Sampling
4.2. Sludge Stagnation
4.3. Sample Preparation
4.4. Sample Analysis
4.4.1. Taxonomic Cell Counts
4.4.2. Dissolved Organic Carbon (DOC)
4.4.3. Microcystins (MCs)
4.4.4. DNA Extraction, Shotgun Metagenomic Sequencing, Bioinformatics, and Statistical Analysis
4.4.5. Solids Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ho, J.C.; Michalak, A.M. Exploring temperature and precipitation impacts on harmful algal blooms across continental U.S. lakes. Limnol. Oceanogr. 2019, 65, 992–1009. [Google Scholar] [CrossRef]
- Ho, J.C.; Michalak, A.M.; Pahlevan, N. Widespread global increase in intense lake phytoplankton blooms since the 1980s. Nature 2019, 574, 667–670. [Google Scholar] [CrossRef] [PubMed]
- Pick, F.R. Blooming algae: A Canadian perspective on the rise of toxic cyanobacteria. Can. J. Fish. Aquat. Sci. 2016, 73, 1149–1158. [Google Scholar] [CrossRef]
- Kimambo, O.N.; Gumbo, J.R.; Chikoore, H. The occurrence of cyanobacteria blooms in freshwater ecosystems and their link with hydro-meteorological and environmental variations in Tanzania. Heliyon 2019, 5, e01312. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.G.; DeSellas, A.M.; Fletcher, R.; Heintsch, L.; Morley, A.; Nakamoto, L.; Utsumi, K. Algal blooms in Ontario, Canada: Increases in reports since 1994. Lake Reserv. Manag. 2011, 27, 107–114. [Google Scholar] [CrossRef]
- Drikas, M.; Chow, C.W.K.; House, J.; Burch, M.D. Using coagulation, flocculation, and settling to remove toxic cyanobacteria. J. Am. Water Work. Assoc. 2001, 93, 100–111. [Google Scholar] [CrossRef]
- Zamyadi, A.; Dorner, S.; Ellis, D.; Bolduc, A.; Bastien, C.; Prévost, M. Species-dependence of cyanobacteria removal efficiency by different drinking water treatment processes. Water Res. 2013, 47, 2689–2700. [Google Scholar] [CrossRef]
- Newcombe, G.; House, J.; Ho, L.; Baker, P.; Burch, M. Management Strategies for Cyanobacteria (Blue-Green Algae): A Guide for Water Utilities; The Cooperative Research Centre for Water Quality and Treatment: Adelaïde, Australia, 2010; p. 112. [Google Scholar]
- Chorus, I.; Welker, M. Toxic Cyanobacteria in Water: A Guide to Their Public Health Consequences, Monitoring and Management; CRC Press: London, UK, 2021; p. 859. [Google Scholar]
- Shang, L.; Feng, M.; Xu, X.; Liu, F.; Ke, F.; Li, W. Co-occurrence of microcystins and taste-and-odor compounds in drinking water source and their removal in a full-scale drinking water treatment plant. Toxins 2018, 10, 26. [Google Scholar] [CrossRef]
- Zamyadi, A.; MacLeod, S.; Fan, Y.; McQuaid, N.; Dorner, S.; Sauvé, S.; Prévost, M. Toxic cyanobacterial breakthrough and accumulation in a drinking water plant: A monitoring and treatment challenge. Water Res. 2012, 46, 1511–1523. [Google Scholar] [CrossRef]
- Sun, F.; Pei, H.-Y.; Hu, W.-R.; Ma, C.-X. The lysis of Microcystis aeruginosa in AlCl3 coagulation and sedimentation processes. Chem. Eng. J. 2012, 193–194, 196–202. [Google Scholar] [CrossRef]
- Teixeira, M.R.; Rosa, M.J. Comparing dissolved air flotation and conventional sedimentation to remove cyanobacterial cells of Microcystis aeruginosa. Part II. The effect of water background organics. Sep. Purif. Technol. 2007, 53, 126–134. [Google Scholar] [CrossRef]
- Zamyadi, A.; Dorner, S.; Ndong, M.; Ellis, D.; Bolduc, A.; Bastien, C.; Prévost, M. Low-risk cyanobacterial bloom sources: Cell accumulation within full-scale treatment plants. J. Am. Water Work. Assoc. 2013, 102, E651–E663. [Google Scholar] [CrossRef]
- Almuhtaram, H.; Cui, Y.; Zamyadi, A.; Hofmann, R. Cyanotoxins and cyanobacteria cell accumulations in drinking water treatment plants with a low risk of bloom formation at the source. Toxins 2018, 10, 430. [Google Scholar] [CrossRef] [PubMed]
- Ho, L.; Dreyfus, J.; Boyer, J.E.; Lowe, T.; Bustamante, H.; Duker, P.; Meli, T.; Newcombe, G. Fate of cyanobacteria and their metabolites during water treatment sludge management processes. Sci. Total Environ. 2012, 424, 232–238. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Pei, H.-Y.; Hu, W.-R.; Li, X.-Q.; Ma, C.-X.; Pei, R.-T. The cell damage of Microcystis aeruginosa in PACl coagulation and floc storage processes. Sep. Purif. Technol. 2013, 115, 123–128. [Google Scholar] [CrossRef]
- Xu, H.; Pei, H.; Xiao, H.; Jin, Y.; Li, X.; Hu, W.; Ma, C.; Sun, J.; Li, H. Behaviors of Microcystis aeruginosa cells during floc storage in drinking water treatment process. Sci. Rep. 2016, 6, 34943. [Google Scholar] [CrossRef]
- Sun, J.; Xu, H.; Pei, H.; Jin, Y.; Li, H.; Ma, C. Worse than cell lysis: The resilience of Oscillatoria sp. during sludge storage in drinking water treatment. Water Res. 2018, 142, 405–414. [Google Scholar] [CrossRef]
- Li, H.; Pei, H.; Xu, H.; Jin, Y.; Sun, J. Behavior of Cylindrospermopsis raciborskii during coagulation and sludge storage—Higher potential risk of toxin release than Microcystis aeruginosa? J. Hazard. Mater. 2018, 347, 307–316. [Google Scholar] [CrossRef]
- Water Research Foundation (WRF); Water Research Australia. Optimizing Conventional Treatment for the Removal of Cyanobacteria and Toxins; Water Research Foundation: Denver, CO, USA, 2015; p. 185. ISBN 978-1-60573-216-9. [Google Scholar]
- Pestana, C.J.; Reeve, P.J.; Sawade, E.; Voldoire, C.F.; Newton, K.; Praptiwi, R.; Collingnon, L.; Dreyfus, J.; Hobson, P.; Gaget, V.; et al. Fate of cyanobacteria in drinking water treatment plant lagoon supernatant and sludge. Sci. Total Environ. 2016, 565, 1192–1200. [Google Scholar] [CrossRef]
- Dreyfus, J.; Monrolin, Y.; Pestana, C.J.; Reeve, P.J.; Sawade, E.; Newton, K.; Ho, L.; Chow, C.W.K.; Newcombe, G. Identification and assessment of water quality risks associated with sludge supernatant recycling in the presence of cyanobacteria. J. Water Supply Res. Technol.—Aqua 2016, 65, 441–452. [Google Scholar] [CrossRef]
- Sun, F.; Hu, W.; Pei, H.; Li, X.; Xu, X.; Ma, C. Evaluation on the dewatering process of cyanobacteria-containing AlCl3 and PACl drinking water sludge. Sep. Purif. Technol. 2015, 150, 52–62. [Google Scholar] [CrossRef]
- Pei, H.; Xu, H.; Wang, J.; Jin, Y.; Xiao, H.; Ma, C.; Sun, J.; Li, H. 16S rRNA Gene amplicon sequencing reveals significant changes in microbial compositions during cyanobacteria-laden drinking water sludge storage. Environ. Sci. Technol. 2017, 51, 12774–12783. [Google Scholar] [CrossRef] [PubMed]
- Zamyadi, A.; Romanis, C.; Mills, T.; Neilan, B.; Choo, F.; Coral, L.A.; Gale, D.; Newcombe, G.; Crosbie, N.; Stuetz, R.; et al. Diagnosing water treatment critical control points for cyanobacterial removal: Exploring benefits of combined microscopy, next-generation sequencing, and cell integrity methods. Water Res. 2019, 152, 96–105. [Google Scholar] [CrossRef]
- Jalili, F.; Trigui, H.; Guerra Maldonado, J.F.; Dorner, S.; Zamyadi, A.; Shapiro, B.J.; Terrat, Y.; Fortin, N.; Sauvé, S.; Prévost, M. Can cyanobacterial diversity in the source predict the diversity in sludge and the risk of toxin release in a drinking water treatment plant? Toxins 2021, 13, 25. [Google Scholar] [CrossRef] [PubMed]
- Tromas, N.; Fortin, N.; Bedrani, L.; Terrat, Y.; Cardoso, P.; Bird, D.; Greer, C.W.; Shapiro, B.J. Characterising and predicting cyanobacterial blooms in an 8-year amplicon sequencing time course. ISME J. 2017, 11, 1746–1763. [Google Scholar] [CrossRef] [PubMed]
- Traving, S.J.; Clokie, M.R.; Middelboe, M. Increased acidification has a profound effect on the interactions between the cyanobacterium Synechococcus sp. WH7803 and its viruses. FEMS Microbiol. Ecol. 2014, 87, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Jakubowska, N.; Szeląg-Wasielewska, E. Toxic picoplanktonic cyanobacteria—A Review. Mar. Drugs 2015, 13, 1497–1518. [Google Scholar] [CrossRef]
- Mariani, M.A.; Padedda, B.M.; Kastovsky, J.; Buscarinu, P.; Sechi, N.; Virdis, T.; Luglie, A. Effects of trophic status on microcystin production and the dominance of cyanobacteria in the phytoplankton assemblage of Mediterranean reservoirs. Sci. Rep. 2015, 5, 17964. [Google Scholar] [CrossRef] [PubMed]
- Moradinejad, S.; Trigui, H.; Guerra Maldonado, J.F.; Shapiro, J.; Terrat, Y.; Zamyadi, A.; Dorner, S.; Prévost, M. Diversity assessment of toxic cyanobacterial blooms during oxidation. Toxins 2020, 12, 728. [Google Scholar] [CrossRef]
- American Water Works Association (AWWA). M37-Manual of Water Supply Practices. Operational Control of Coagulation and Filtration Processes, 3rd ed.; American Water Works Association: Denver, CO, USA, 2010. [Google Scholar]
- Park, J.; Kim, Y.; Kim, M.; Lee, W.H.; Park, J.; Kim, Y.; Kim, M.; Lee, W.H. A novel method for cell counting of Microcystis colonies in water resources using a digital imaging flow cytometer and microscope. Environ. Eng. Res. 2018, 24, 397–403. [Google Scholar] [CrossRef]
- Hawkins, P.R.; Holliday, J.; Kathuria, A.; Bowling, L. Change in cyanobacterial biovolume due to preservation by Lugol’s Iodine. Harmful Algae 2005, 4, 1033–1043. [Google Scholar] [CrossRef]
- Bag, S.; Saha, B.; Mehta, O.; Anbumani, D.; Kumar, N.; Dayal, M.; Pant, A.; Kumar, P.; Saxena, S.; Allin, K.H.; et al. An improved method for high quality metagenomics DNA extraction from human and environmental samples. Sci. Rep. 2016, 6, 26775. [Google Scholar] [CrossRef] [PubMed]
- Kuczynski, J.; Lauber, C.L.; Walters, W.A.; Parfrey, L.W.; Clemente, J.C.; Gevers, D.; Knight, R. Experimental and analytical tools for studying the human microbiome. Nat. Rev. Genet. 2011, 13, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Gevers, D.; Pop, M.; Schloss, P.D.; Huttenhower, C. Bioinformatics for the human microbiome project. PLoS Comput. Biol. 2012, 8, e1002779. [Google Scholar] [CrossRef] [PubMed]
- Teeling, H.; Glockner, F.O. Current opportunities and challenges in microbial metagenome analysis—A bioinformatic perspective. Brief. Bioinform. 2012, 13, 728–742. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.E.; Golden, S.S. Circadian Rhythms in Cyanobacteria. Microbiol. Mol. Biol. Rev. 2015, 79, 373–385. [Google Scholar] [CrossRef]
- Imamura, S.; Asayama, M. Sigma factors for cyanobacterial transcription. Gene Regul. Syst. Biol. 2009, 3, 65–87. [Google Scholar] [CrossRef]
- Kumar, K.; Mella-Herrera, R.A.; Golden, J.W. Cyanobacterial heterocysts. Cold Spring Harb. Perspect. Biol. 2010, 2, a000315. [Google Scholar] [CrossRef]
- Herrero, A.; Stavans, J.; Flores, E. The multicellular nature of filamentous heterocyst-forming cyanobacteria. FEMS Microbiol. Rev. 2016, 40, 831–854. [Google Scholar] [CrossRef]
- Rachel, R.; Pum, D.; Šmarda, J.; Šmajs, D.; Komrska, J.; Krzyzánek, V.; Rieger, G.; Stetter, K.O., II. Fine structure of S-layers. FEMS Microbiol. Rev. 1997, 20, 13–23. [Google Scholar] [CrossRef][Green Version]
- Šmarda, J.; Šmajs, D.; Komrska, J.; Krzyžánek, V. S-layers on cell walls of cyanobacteria. Micron 2002, 33, 257–277. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Messner, P.; Pum, D.; Sára, M. Crystalline Bacterial Cell Surface Layers (S Layers): From Supramolecular Cell Structure to Biomimetics and Nanotechnology. Angew. Chem. Int. Ed. 1999, 38, 1034–1054. [Google Scholar] [CrossRef]
- Callieri, C. Synechococcus plasticity under environmental changes. FEMS Microbiol. Lett. 2017, 364, fnx229. [Google Scholar] [CrossRef]
- Min, H.; Golden, S.S. A new circadian class 2 gene, opcA, whose product is important for reductant production at night in Synechococcus elongatus PCC 7942. J. Bacteriol. 2000, 182, 6214–6221. [Google Scholar] [CrossRef] [PubMed]
- Jalili, F.; Trigui, H.; Maldonado, J.F.G.; Dorner, S.; Zamyadi, A.; Shapiro, B.J.; Terrat, Y.; Fortin, N.; Sauvé, S.; Prévost, M. Oxidation to Control Cyanobacteria and Cyanotoxins in Drinking Water Treatment Plants: Challenges at the Laboratory and Full-Scale Plants. Water 2022, 14, 537. [Google Scholar] [CrossRef]
- Kormas, K.A.; Lymperopoulou, D.S. Cyanobacterial toxin degrading bacteria: Who are they? Biomed Res. Int. 2013, 2013, 463894. [Google Scholar] [CrossRef]
- Maghsoudi, E.; Fortin, N.; Greer, C.; Maynard, C.; Page, A.; Duy, S.V.; Sauve, S.; Prevost, M.; Dorner, S. Cyanotoxin degradation activity and mlr gene expression profiles of a Sphingopyxis sp. isolated from Lake Champlain, Canada. Environ. Sci. Process. Impacts 2016, 18, 1417–1426. [Google Scholar] [CrossRef]
- Zamyadi, A.; Henderson, R.K.; Stuetz, R.; Newcombe, G.; Newtown, K.; Gladman, B. Cyanobacterial management in full-scale water treatment and recycling processes: Reactive dosing following intensive monitoring. Environ. Sci. Water Res. Technol. 2016, 2, 362–375. [Google Scholar] [CrossRef]
- Jalili, F. Optimal Treatment Strategies to Prevent and Manage Cyanobacteria and Cyanotoxins in Drinking Water Sludge. Ph.D. Thesis, Polytechnique Montréal, Montréal, QC, Canada, 2022. [Google Scholar]
- Lund, J.W.G.; Kipling, C.; Le Cren, E.D. The inverted microscope method of estimating algal number and the statistical basis of estimations by counting. Hydrobiologia 1958, 11, 143–170. [Google Scholar] [CrossRef]
- Lund, J.W.G. A simple counting chamber for Nannoplankton. Limnol. Oceanogr. 1959, 4, 57–65. [Google Scholar] [CrossRef]
- Planas, D.; Desrosiers, M.; Groulx, S.R.; Paquet, S.; Carignan, R. Pelagic and benthic algal responses in eastern Canadian Boreal Shield lakes following harvesting and wildfires. Can. J. Fish. Aquat. Sci. 2000, 57, 136–145. [Google Scholar] [CrossRef]
- Zamyadi, A.; Choo, F.; Newcombe, G.; Stuetz, R.; Henderson, R.K. A review of monitoring technologies for real-time management of cyanobacteria: Recent advances and future direction. TrAC Trends Anal. Chem. 2016, 85, 83–96. [Google Scholar] [CrossRef]
- United States Environmental Protection Agency (USEPA). Method 415.1. Organic Carbon, Total (Combustion or Oxidation); United States Environmental Protection Agency: Washington, DC, USA, 1974; p. 3. [Google Scholar]
- Munoz, G.; Vo Duy, S.; Roy-Lachapelle, A.; Husk, B.; Sauve, S. Analysis of individual and total microcystins in surface water by on-line preconcentration and desalting coupled to liquid chromatography tandem mass spectrometry. J. Chromatogr. A 2017, 1516, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Roy-Lachapelle, A.; Vo Duy, S.; Munoz, G.; Dinh, Q.T.; Bahl, E.; Simon, D.F.; Sauvé, S. Analysis of multiclass cyanotoxins (microcystins, anabaenopeptins, cylindrospermopsin and anatoxins) in lake waters using on-line SPE liquid chromatography high-resolution Orbitrap mass spectrometry. Anal. Methods 2019, 11, 5289–5300. [Google Scholar] [CrossRef]
- Cox, M.P.; Peterson, D.A.; Biggs, P.J. SolexaQA: At-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinform. 2010, 11, 485. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef]
- Tatusov, R.L.; Galperin, M.Y.; Natale, D.A.; Koonin, E.V. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28, 33–36. [Google Scholar] [CrossRef]
- Moradinejad, S.; Trigui, H.; Maldonado, J.F.G.; Shapiro, B.J.; Terrat, Y.; Sauvé, S.; Fortin, N.; Zamyadi, A.; Dorner, S.; Prévost, M. Metagenomic study to evaluate functional capacity of a cyanobacterial bloom during oxidation. Chem. Eng. J. Adv. 2021, 8, 100151. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Graffelman, J. Compositional Data Analysis in Practice; Greenacre, M.J., Ed.; CRC Press: London, UK, 2018; p. 136. ISBN 978-1-138-31661-4. [Google Scholar] [CrossRef]
- Blanchet, F.G.; Legendre, P.; Borcard, D. Forward selection of explanatory variables. Ecology 2008, 89, 2623–2632. [Google Scholar] [CrossRef] [PubMed]
- American Public Health Association (APHA); American Water Works Association (AWWA); Water Environment Federation (WEF). Standard Methods for the Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 2012; Volume 5, p. 1360. [Google Scholar]


| Sampling Date | Shotgun Metagenomic Sequencing | Taxonomic Cell Counts | MCs (ng/L) | DOC (mg/L) | pH | Turbidity (NTU) | TSS (mg/L) | TVS (mg/L) | Sludge Storage Time (d) | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Cells/mL ×106 | mm3/L | Cell-Bound | Dissolved | ||||||||
| 27 July 2018 | - | 0.70 | 7.11 | 170.8 | 67.9 | 4.10 | - | - | - | - | 3 |
| 31 July 2018 | * | 2.25 | 147.4 | 24.9 | 37.9 | 3.60 | 7.05 | 201 | 716 | 367 | 7 |
| 7 August 2018 | * | 2.71 | 96.20 | 22.0 | 138.5 | 3.19 | 7.54 | 171 | 728 | 456 | 5 |
| 10 August 2018 | * | 5.57 | 608.9 | 138.2 | 55.8 | 5.20 | 6.8 | 258 | 1022 | 409 | 8 |
| 17 August 2018 | * | 2.35 | 138.3 | 41.6 | 46.6 | 3.35 | 7.12 | 327 | 1092 | 434 | 3 |
| 5 September 2018 | - | 2.37 | 52.76 | 951.8 | 131.2 | 9.80 | 6.81 | 701 | 1957 | 1230 | 6 |
| 16 October 2018 | * | 2.70 | 25.95 | 7129.0 | 284.2 | 3.46 | 6.87 | 2300 | 3394 | 1084 | 8 |
| 1 November 2018 | * | 1.21 | 4.42 | 230.8 | 348.2 | 2.76 | 6.74 | 225 | 740 | 546 | 6 |
| Treatment Step | Parameters | July | August | September | October | Specifications |
|---|---|---|---|---|---|---|
| Raw water (RW) | Turbidity (NTU) | 2.1–153.7 | 9.4–153.1 | 11.6–152.7 | 17.5–152.9 | - |
| pH | 6.1–8.1 | 5.8–8.3 | 6.2–9.0 | 5.9–8.8 | - | |
| Clarifier (CW) | Turbidity (NTU) | 0.01–20.1 | 0.30–20.0 | 0.33–10.2 | 0.01–10.3 | PAC (wood-based): 1.5–27.0 mg/L, Coagulant: PAXL, 49–410 mg/L, Polymer: Hydrex (silicate), 0.05–0.1 mg/L Effective clarifier depth: 4.90 m, Max. sludge bed: 2.95 m, Hydraulic retention time: 1 h, Solid retention time: 48 h |
| pH | 6.1–7.2 | 6.7–7.2 | 6.2–7.1 | 6.6–7.2 | ||
| Dual sand-antrachite filter (FW) | Turbidity (NTU) | 0.14–0.4 | 0.16–0.4 | 0.11–0.6 | 0.17–0.6 | Retention time: 2 h |
| Treated water (TW) | Turbidity (NTU) | 0.21–0.60 | 0.25–0.49 | 0.21–0.43 | 0.23–0.48 | Injected chlorine: NaOCl, 1.3–6.0 mg/L |
| pH | 6.6–8.1 | 6.9–8.0 | 7.1–8.6 | 7.1–8.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jalili, F.; Trigui, H.; Maldonado, J.F.G.; Dorner, S.; Zamyadi, A.; Shapiro, B.J.; Terrat, Y.; Fortin, N.; Sauvé, S.; Prévost, M. Impact of Stagnation on the Diversity of Cyanobacteria in Drinking Water Treatment Plant Sludge. Toxins 2022, 14, 749. https://doi.org/10.3390/toxins14110749
Jalili F, Trigui H, Maldonado JFG, Dorner S, Zamyadi A, Shapiro BJ, Terrat Y, Fortin N, Sauvé S, Prévost M. Impact of Stagnation on the Diversity of Cyanobacteria in Drinking Water Treatment Plant Sludge. Toxins. 2022; 14(11):749. https://doi.org/10.3390/toxins14110749
Chicago/Turabian StyleJalili, Farhad, Hana Trigui, Juan Francisco Guerra Maldonado, Sarah Dorner, Arash Zamyadi, B. Jesse Shapiro, Yves Terrat, Nathalie Fortin, Sébastien Sauvé, and Michèle Prévost. 2022. "Impact of Stagnation on the Diversity of Cyanobacteria in Drinking Water Treatment Plant Sludge" Toxins 14, no. 11: 749. https://doi.org/10.3390/toxins14110749
APA StyleJalili, F., Trigui, H., Maldonado, J. F. G., Dorner, S., Zamyadi, A., Shapiro, B. J., Terrat, Y., Fortin, N., Sauvé, S., & Prévost, M. (2022). Impact of Stagnation on the Diversity of Cyanobacteria in Drinking Water Treatment Plant Sludge. Toxins, 14(11), 749. https://doi.org/10.3390/toxins14110749







