Removal of Aflatoxin B1 by Edible Mushroom-Forming Fungi and Its Mechanism
Abstract
:1. Introduction
2. Results
2.1. Time Course of Fungal Growth, pH, and Removal of AFB1 Using Five Edible Mushroom-Forming Fungi
2.2. Test for Degradation of Remazol Brilliant Blue R (RBBR) and Coumarin by Mushroom-Forming Fungi
2.3. AFB1 Removal by Cell-Free Supernatants and Cell Lysates
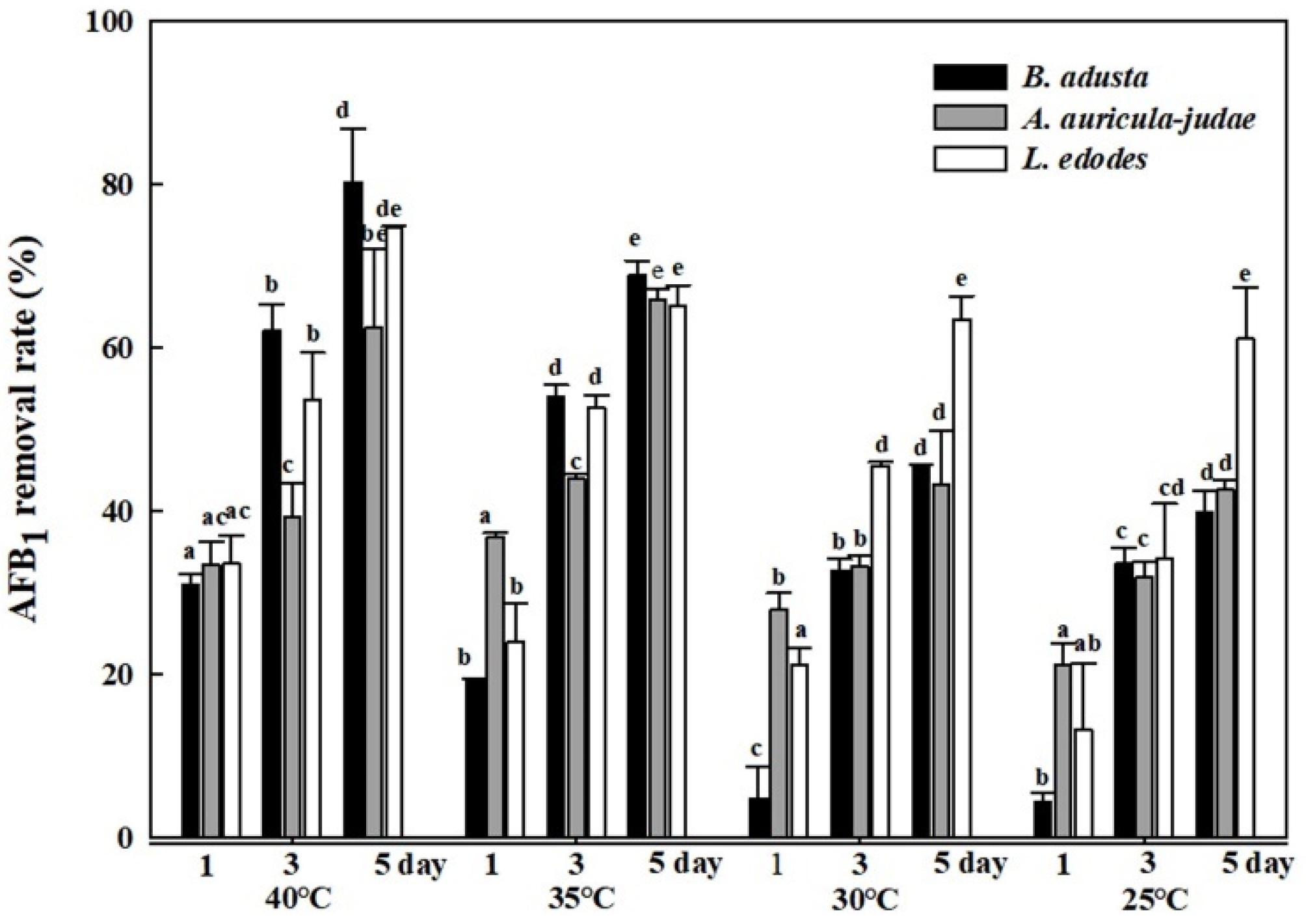
2.4. Effects of Different Reaction Temperatures on AFB1 Removal by Cell Lysates
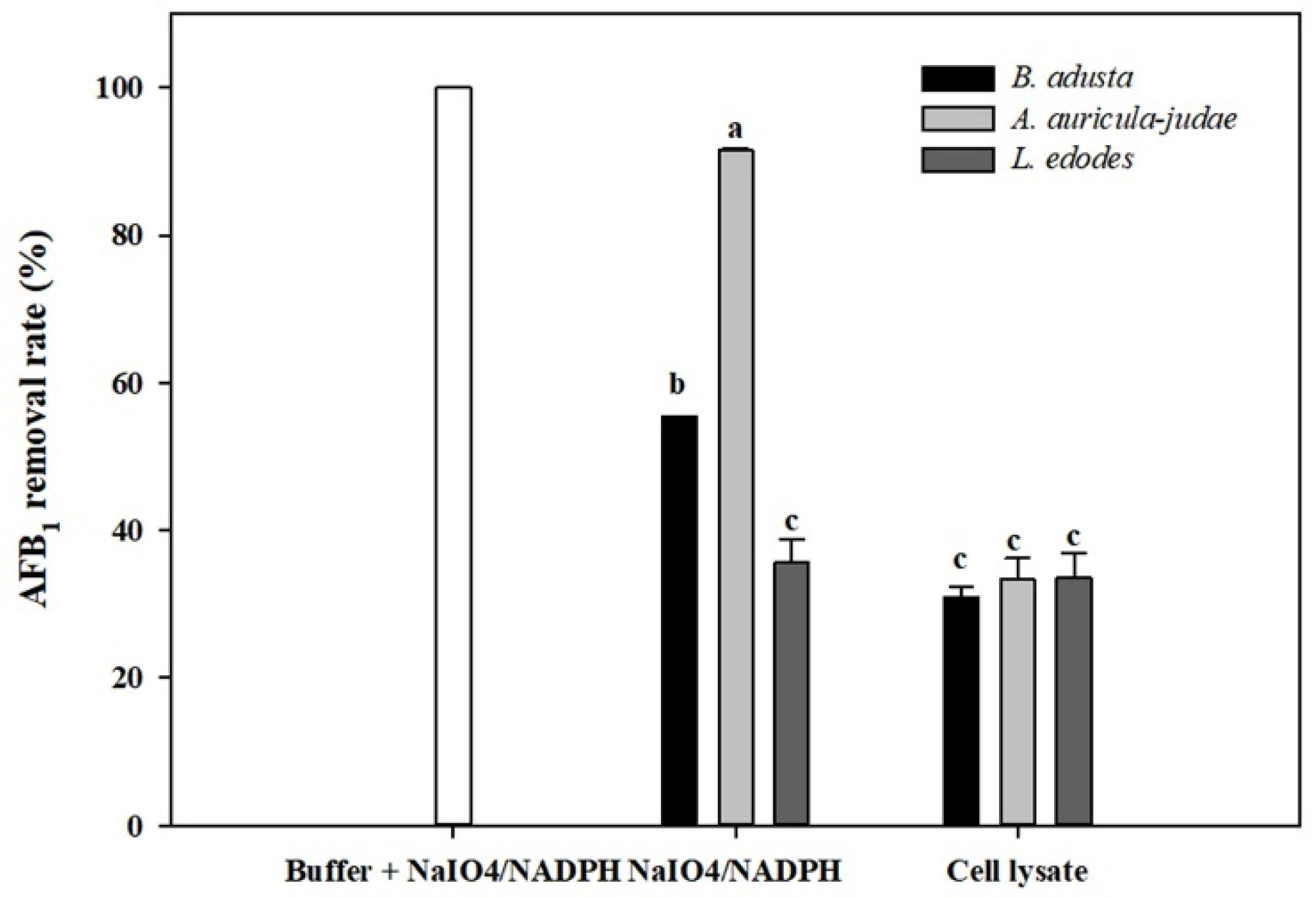
2.5. Effects of NADPH and NaIO4 on AFB1 Removal by Cell Lysates
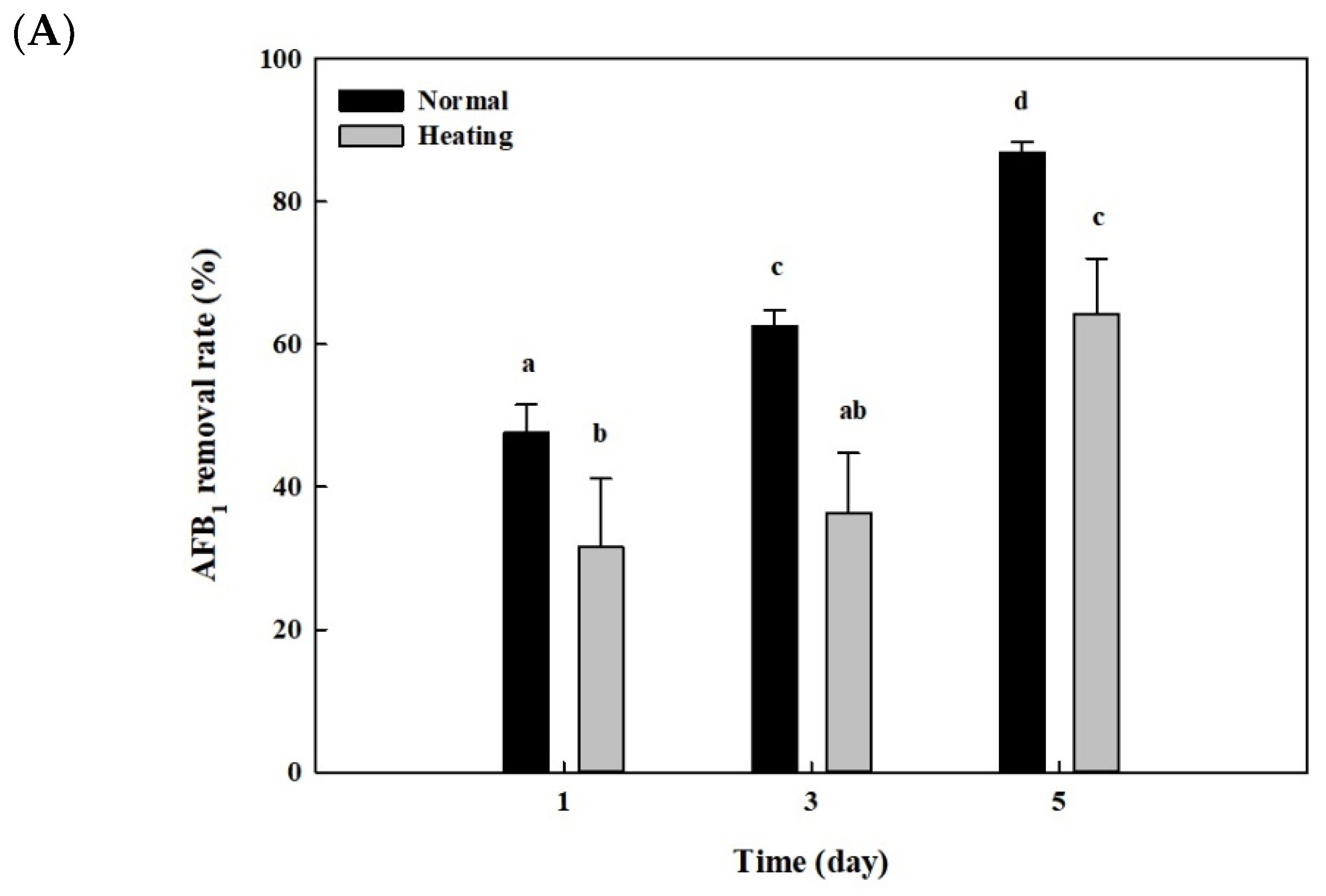
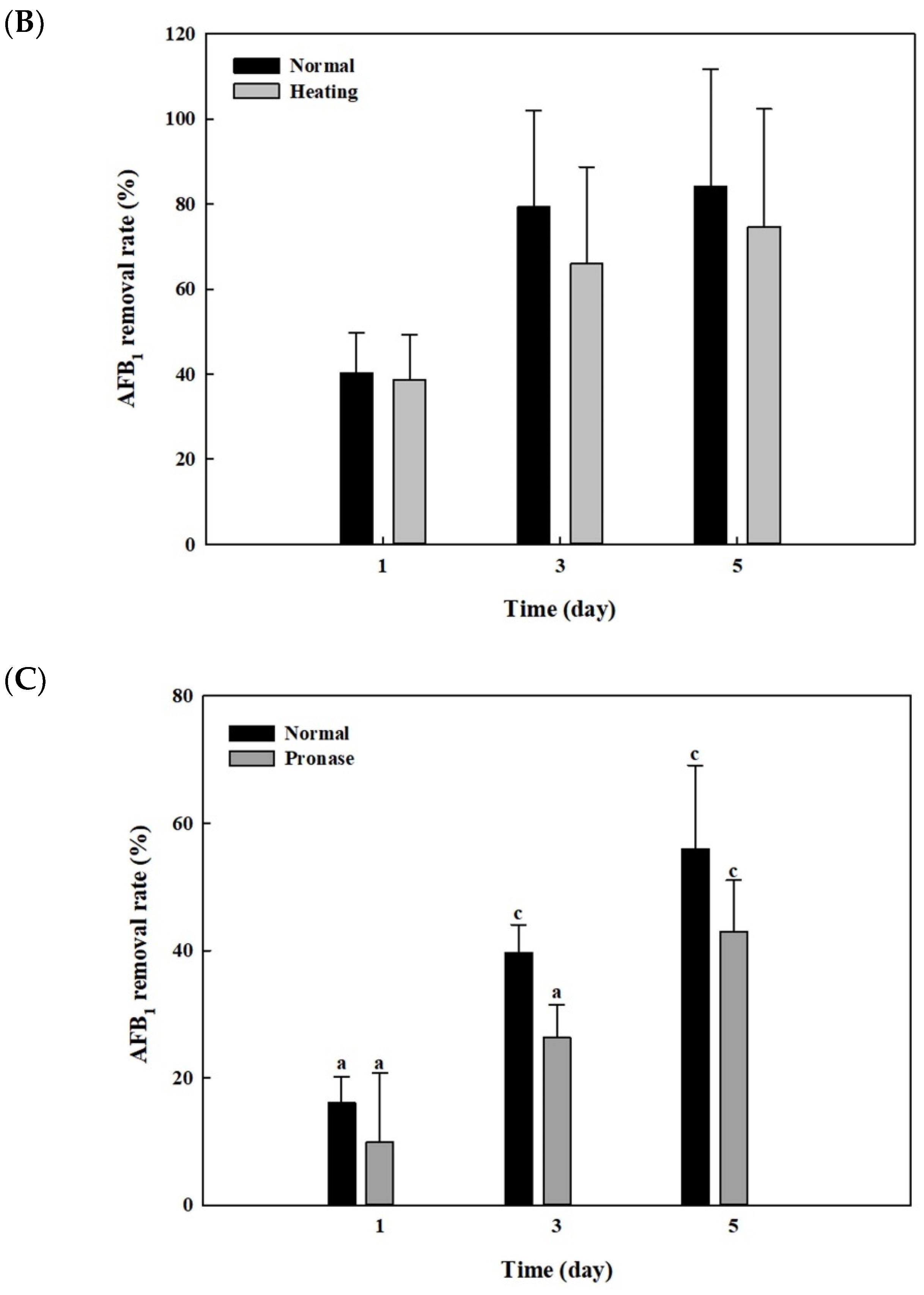
2.6. Effects of Heat or Proteinase Treatment on AFB1 Removal by Whole Cells and Cell Lysates from B. adusta Cultures
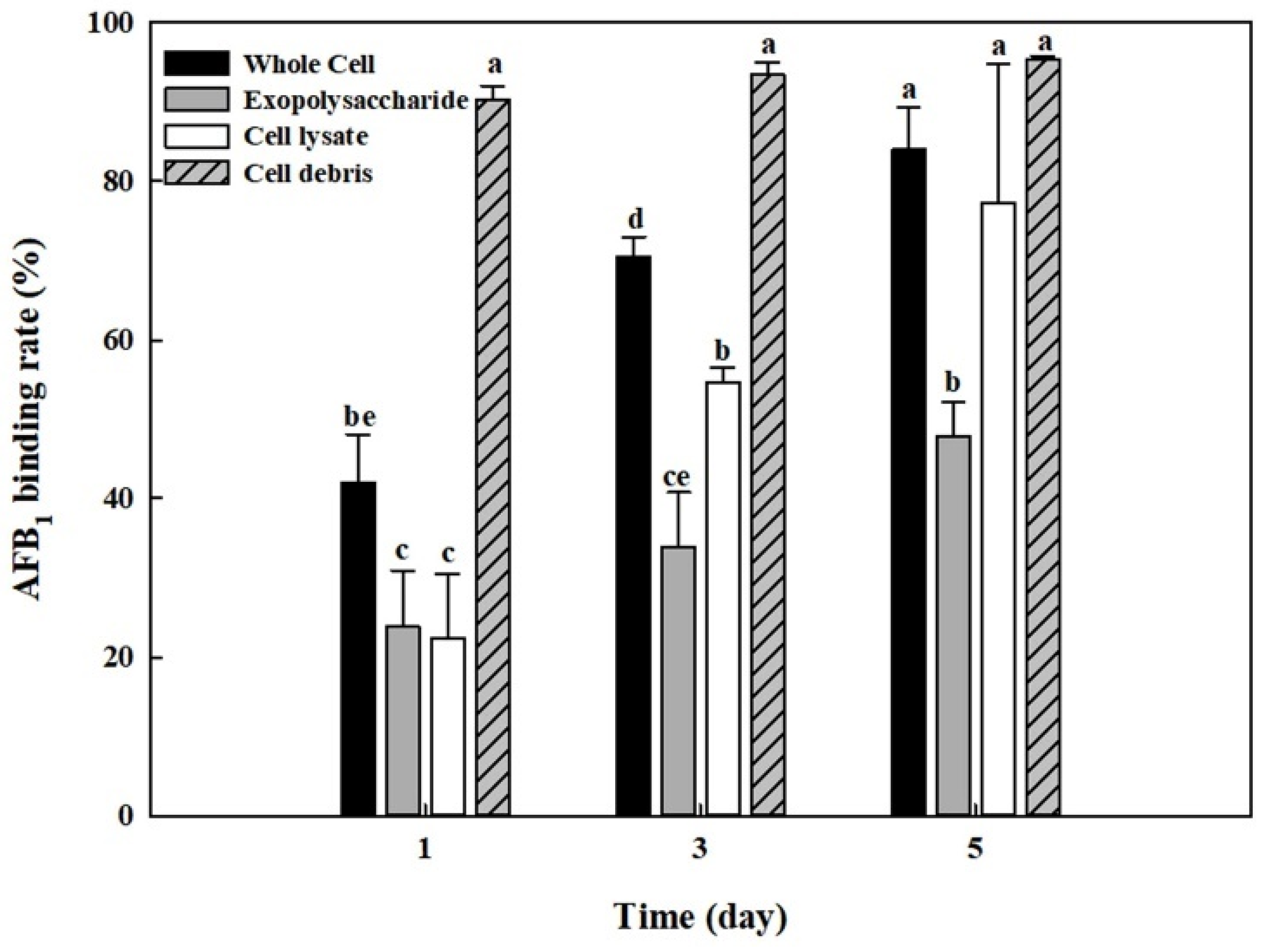
2.7. AFB1 Removal by Whole Cells, Cell Lysates, Cell Debris, and Exopolysaccharides from B. adusta Cultures
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Chemicals and Reagents
5.2. Fungal Strains and Culture Conditions
5.3. Test for Degradation of RBBR and Coumarin by Mushrooms
5.4. Time Course of Fungal Growth, Change in pH, and Removal of AFB1
5.5. AFB1 Extraction from Fungal Culture Media
5.6. AFB1 Assays Using Whole Cells, Cell Lysates after Cell Disruption, Cell-Free Supernatants, and Cell Debris
5.7. Effects of 3 mM NaIO4 and 0.2 mM NADHP on AFB1 Removal by Cell Lysates
5.8. AFB1 Assays Using Heat- or Pronase-Treated Whole Cells and Cell Lysates
5.9. AFB1 Analysis by HPLC-FLD
5.10. Determination of Total Carbohydrates, Protein, and Glucosamine in Cell-Free Supernatants, Cell Lysates, and Cell Debris
5.11. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Cotty, P.J.; Bayman, P.; Egel, D.S.; Elias, D.S. Agriculture, aflatoxins, and Asperigiilus. In The Genus Aspergillus; Powell, K.A., Fenwick, A., Peberdy, J.F., Eds.; Plenum Press: New York, NY, USA, 1994; pp. 1–27. [Google Scholar]
- Gourama, H.; Bullerman, L.B. Aspergillus flavus and Aspergillus parasiticus: Aflatoxigenic fungi of concern in foods and feeds: A review. J. Food Prot. 1995, 58, 1395–1404. [Google Scholar] [CrossRef] [PubMed]
- Eaton, D.L.; Gallagher, E.P. Mechanism of aflatoxin carcinogenesis. Annu. Rev. Pharm. 1994, 34, 135–172. [Google Scholar] [CrossRef]
- IARC. Summaries and Evaluations: Aflatoxins; IARC: Lyon, France, 2003; p. 171. [Google Scholar]
- Herzallah, S.; Alshawabkeh, K.; Al Fataftah, A. Aflatoxin decontamination of artificially contaminated feeds by sunlight, gamma-radiation, and microwave heating. J. Appl. Poult. Res. 2008, 17, 515–521. [Google Scholar] [CrossRef]
- Diaz, D.E.; Hagler, W.M.; Blackwelder, J.T.; Eve, J.A.; Hopkins, B.A.; Anderson, K.L.; Jones, F.T.; Whitlow, L.W. Aflatoxin Binders II: Reduction of aflatoxin M1 in milk by sequestering agents of cows consuming aflatoxin in feed. Mycopathologia 2004, 157, 233–241. [Google Scholar] [CrossRef]
- Grant, P.G.; Phillips, T.D. Isothermal adsorption of aflatoxin B1 on HSCAS clay. J. Agr. Food Chem. 1998, 46, 599–605. [Google Scholar] [CrossRef]
- Mishra, H.N.; Das, C. A review on biological control and metabolism of aflatoxin. Crit. Rev. Food Sci. 2003, 43, 245–264. [Google Scholar] [CrossRef]
- Dwarakanath, C.T.; Rayner, E.T.; Mann, G.E.; Dollear, F.G. Reduction of aflatoxin levels in cottonseed and peanut meals by ozonization. J. Am. Oil Chem. Soc. 1968, 45, 93–95. [Google Scholar] [CrossRef] [PubMed]
- Doyle, M.P.; Marth, E.H. Bisulfite degrades aflatoxin—Effect of temperature and concentration of bisulfite. J. Food Prot. 1978, 41, 774–780. [Google Scholar] [CrossRef] [PubMed]
- Park, D.L.; Lee, L.; Koltun, S.A. Distribution of ammonia-related aflatoxin reaction-products in cottonseed meal. J. Am. Oil Chem. Soc. 1984, 61, 1071–1074. [Google Scholar] [CrossRef]
- Kabak, B.; Dobson, A.D.W.; Var, I. Strategies to prevent mycotoxin contamination of food and animal feed: A review. Crit. Rev. Food Sci. 2006, 46, 593–619. [Google Scholar] [CrossRef]
- Huwig, A.; Freimund, S.; Kappeli, O.; Dutler, H. Mycotoxin detoxication of animal feed by different adsorbents. Toxicol. Lett. 2001, 122, 179–188. [Google Scholar] [CrossRef]
- Samuel, S.M.; Aiko, V.; Panda, P.; Mehta, A. Aflatoxin B1 occurrence, biosynthesis and its degradation. J. Pure Appl. Microbio. 2013, 7, 965–971. [Google Scholar]
- Kolosova, A.; Stroka, J. Substances for reduction of the contamination of feed by mycotoxins: A review. World Mycotoxin J. 2011, 4, 225–256. [Google Scholar] [CrossRef]
- Hormisch, D.; Brost, I.; Kohring, G.W.; Giffhorn, E.; Kroppenstedt, R.M.; Stackebrandt, E.; Farber, P.; Holzapfel, W.H. Mycobacterium fluoranthenivorans sp nov., a fluoranthene and aflatoxin B1 degrading bacterium from contaminated soil of a former coal gas plant. Syst. Appl. Microbiol. 2004, 27, 653–660. [Google Scholar] [CrossRef]
- Teniola, O.D.; Addo, P.A.; Brost, I.M.; Farber, P.; Jany, K.D.; Alberts, J.F.; van Zyl, W.H.; Steyn, P.S.; Holzapfel, W.H. Degradation of aflatoxin B1 by cell-free extracts of Rhodococcus erythropolis and Mycobacterium fluoranthenivorans sp nov DSM44556(T). Int. J. Food Microbiol. 2005, 105, 111–117. [Google Scholar] [CrossRef]
- Shantha, T. Fungal degradation of aflatoxin B1. Nat. Toxins 1999, 7, 175–178. [Google Scholar] [CrossRef]
- Doyle, M.P.; Applebaum, R.S.; Brackett, R.E.; Marth, E.H. Physical, chemical and biological degradation of mycotoxins in foods and agricultural commodities. J. Food Prot. 1982, 45, 964–971. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Q.; Ogata, M.; Hirai, H.; Kawagishi, H. Detoxification of aflatoxin B1 by manganese peroxidase from the white-rot fungus Phanerochaete sordida YK-624. FEMS Microbiol. Lett. 2011, 314, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Arora, D.S.; Sharma, R.K. Ligninolytic fungal laccases and their biotechnological applications. Appl. Biochem. Biotech. 2010, 160, 1760–1788. [Google Scholar] [CrossRef]
- Vyas, B.R.M.; Molitoris, H.P. Involvement of an extracellular H2O2-dependent ligninolytic activity of the white-rot fungus Pleurotus ostreatus in the decolorization of remazol-brilliant-blue-R. Appl. Environ. Microbiol. 1995, 61, 3919–3927. [Google Scholar] [CrossRef] [Green Version]
- Pickard, M.A.; Roman, R.; Tinoco, R.; Vazquez-Duhalt, R. Polycyclic aromatic hydrocarbon metabolism by white rot fungi and oxidation by Coriolopsis gallica UAMH 8260 laccase. Appl. Environ. Microbiol. 1999, 65, 3805–3809. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alberts, J.F.; Gelderblom, W.C.A.; Botha, A.; van Zyl, W.H. Degradation of aflatoxin B1 by fungal laccase enzymes. Int. J. Food Microbiol. 2009, 135, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Motomura, M.; Toyomasu, T.; Mizuno, K.; Shinozawa, T. Purification and characterization of an aflatoxin degradation enzyme from Pleurotus ostreatus. Microbiol. Res. 2003, 158, 237–242. [Google Scholar] [CrossRef] [PubMed]
- Yehia, R.S. Aflatoxin detoxification by manganese peroxidase purified from Pleurotus ostreatus. Braz. J. Microbiol. 2014, 45, 127–133. [Google Scholar] [CrossRef] [Green Version]
- Wu, Q.; Jezkova, A.; Yuan, Z.; Pavlikova, L.; Dohnal, V.; Kuca, K. Biological degradation of aflatoxins. Drug Metab. Rev. 2009, 41, 1–7. [Google Scholar] [CrossRef]
- Adebo, O.A.; Njobeh, P.B.; Gbashi, S.; Nwinyi, O.C.; Mavumengwana, V. Review on microbial degradation of aflatoxins. Crit. Rev. Food Sci. 2017, 57, 3208–3217. [Google Scholar] [CrossRef]
- Gratz, S.; Wu, Q.K.; El-Nezami, H.; Juvonen, R.O.; Mykkanen, H.; Turner, P.C. Lactobacillus rhamnosus strain GG reduces aflatoxin B1 transport, metabolism, and toxicity in caco-2 cells. Appl. Environ. Microbiol. 2007, 73, 3958–3964. [Google Scholar] [CrossRef] [Green Version]
- El Khoury, A.; Atoui, A.; Yaghi, J. Analysis of aflatoxin M1 in milk and yogurt and AFM1 reduction by lactic acid bacteria used in Lebanese industry. Food Control 2011, 22, 1695–1699. [Google Scholar] [CrossRef]
- El-Nezami, H.; Mykkanen, H.; Kankaanpaa, P.; Salminen, S.; Ahokas, J. Ability of Lactobacillus and Propionibacterium strains to remove aflatoxin B1 from the chicken duodenum. J. Food Prot. 2000, 63, 549–552. [Google Scholar] [CrossRef]
- Oluwafemi, F.; Kumar, M.; Bandyopadhyay, R.; Ogunbanwo, T.; Ayanwande, K.B. Bio-detoxification of aflatoxin B1 in artificially contaminated maize grains using lactic acid bacteria. Toxin Rev. 2010, 29, 115–122. [Google Scholar] [CrossRef]
- Peltonen, K.; El-Nezami, H.; Haskard, C.; Ahokas, J.; Salminen, S. Aflatoxin B1 binding by dairy strains of lactic acid bacteria and bifidobacteria. J. Dairy Sci. 2001, 84, 2152–2156. [Google Scholar] [CrossRef]
- El-Shiekh, H.H.; Mahdy, H.M.; El-Aaser, M.M. Bioremediation of aflatoxins by some reference fungal strains. Pol. J. Microbiol. 2007, 56, 215–223. [Google Scholar] [PubMed]
- Goncalves, B.L.; Rosim, R.E.; de Oliveira, C.A.F.; Corassin, C.H. The in vitro ability of different Saccharomyces cerevisiae—Based products to bind aflatoxin B1. Food Control 2015, 47, 298–300. [Google Scholar] [CrossRef]
- Kusumaningtyas, E.; Widiastuti, R.; Maryam, R. Reduction of aflatoxin B1 in chicken feed by using Saccharomyces cerevisiae, Rhizopus oligosporus and their combination. Mycopathologia 2006, 162, 307–311. [Google Scholar] [CrossRef] [PubMed]
- Shetty, P.H.; Hald, B.; Jespersen, L. Surface binding of aflatoxin B1 by Saccharomyces cerevisiae strains with potential decontaminating abilities in indigenous fermented foods. Int. J. Food Microbiol. 2007, 113, 41–46. [Google Scholar] [CrossRef]
- Shetty, P.H.; Jespersen, L. Saccharomyces cerevisiae and lactic acid bacteria as potential mycotoxin decontaminating agents. Trends Food Sci. Technol. 2006, 17, 48–55. [Google Scholar] [CrossRef]
- Asgher, M.; Bhatti, H.N.; Ashraf, M.; Legge, R.L. Recent developments in biodegradation of industrial pollutants by white rot fungi and their enzyme system. Biodegradation 2008, 19, 771–783. [Google Scholar] [CrossRef]
- Mester, T.; Field, J.A. Characterization of a novel manganese peroxidase-lignin peroxidase hybrid isozyme produced by Bjerkandera species strain BOS55 in the absence of manganese. J. Biol. Chem. 1998, 273, 15412–15417. [Google Scholar] [CrossRef] [Green Version]
- Pogni, R.; Baratto, M.C.; Giansanti, S.; Teutloff, C.; Verdin, J.; Valderrama, B.; Lendzian, F.; Lubitz, W.; Vazquez-Duhalt, R.; Basosi, R. Tryptophan-based radical in the catalytic mechanism of versatile peroxidase from Bjerkandera adusta. Biochemistry 2005, 44, 4267–4274. [Google Scholar] [CrossRef]
- Tinoco, R.; Verdin, J.; Vazquez-Duhalt, R. Role of oxidizing mediators and tryptophan 172 in the decoloration of industrial dyes by the versatile peroxidase from Bjerkandera adusta. J. Mol. Catal. B-Enzym. 2007, 46, 1–7. [Google Scholar] [CrossRef]
- Boer, C.G.; Obici, L.; de Souza, C.G.M.; Peralta, R.M. Decolorization of synthetic dyes by solid state cultures of Lentinula (Lentinus) edodes producing manganese peroxidase as the main ligninolytic enzyme. Bioresour. Technol. 2004, 94, 107–112. [Google Scholar] [CrossRef]
- Bergot, B.J.; Stanley, W.L.; Masri, M.S. Reaction of coumarin with aqua ammonia—implications in detoxification of aflatoxin. J. Agr. Food Chem. 1977, 25, 965–966. [Google Scholar] [CrossRef]
- Grove, M.D.; Plattner, R.D.; Weisleder, D. Ammoniation products of an aflatoxin model coumarin. J. Agr. Food Chem. 1981, 29, 1161–1164. [Google Scholar] [CrossRef]
- Hamid, A.B.; Smith, J.E. Degradation of aflatoxin by Aspergillus flavus. J. Gen. Microbiol. 1987, 133, 2023–2029. [Google Scholar] [CrossRef] [Green Version]
- Taheur, F.B.; Fedhila, K.; Chaieb, K.; Kouidhi, B.; Bakhrouf, A.; Abrunhosa, L. Adsorption of aflatoxin B1, zearalenone and ochratoxin A by microorganisms isolated from Kefir grains. Int. J. Food Microbiol. 2017, 251, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Serrano-Nino, J.C.; Cavazos-Garduno, A.; Hernandez-Mendoza, A.; Applegate, B.; Ferruzzi, M.G.; San Martin-Gonzalez, M.F.; Garcia, H.S. Assessment of probiotic strains ability to reduce the bioaccessibility of aflatoxin M1 in artificially contaminated milk using an in vitro digestive model. Food Control 2013, 31, 202–207. [Google Scholar] [CrossRef]
- Bueno, D.J.; Casale, C.H.; Pizzolitto, R.P.; Salvano, M.A.; Oliver, G. Physical adsorption of aflatoxin B1 by lactic acid bacteria and Saccharomyces cerevisiae: A theoretical model. J. Food Prot. 2007, 70, 2148–2154. [Google Scholar] [CrossRef]
- Pizzolitto, R.P.; Bueno, D.J.; Armando, M.R.; Cavaglieri, L.; Dalcero, A.M.; Salvano, M.A. Binding of aflatoxin B1 to lactic acid bacteria and Saccharomyces cerevisiae in vitro: A useful model to determine the most efficient microorganism. In Aflatoxins-Biochemistry and Molecular Biology, Guevara-Gonzalez; Guevara-Gonzalez, R.G., Ed.; InTech: Rijeka, Croatia, 2011; pp. 323–346. [Google Scholar] [CrossRef] [Green Version]
- Haskard, C.A.; El-Nezami, H.S.; Kankaanpaa, P.E.; Salminen, S.; Ahokas, J.T. Surface binding of aflatoxin B1 by lactic acid bacteria. Appl. Environ. Microbiol. 2001, 67, 3086–3091. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rahaie, S.; Emam-Djomeh, Z.; Razavi, S.H.; Mazaheri, M. Evaluation of aflatoxin decontaminating by two strains of Saccharomyces cerevisiae and Lactobacillus rhamnosus strain GG in pistachio nuts. Int. J. Food Sci. Technol. 2012, 47, 1647–1653. [Google Scholar] [CrossRef]
- Yiannikouris, A.; Francois, J.; Poughon, L.; Dussap, C.G.; Bertin, G.; Jeminet, G.; Jouany, J.P. Adsorption of zearalenone by beta-D-glucans in the Saccharomyces cerevisiae cell wall. J. Food Prot. 2004, 67, 1195–1200. [Google Scholar] [CrossRef] [PubMed]
- Yiannikouris, A.; Andre, G.; Poughon, L.; Francois, J.; Dussap, C.G.; Jeminet, G.; Bertin, G.; Jouany, J.P. Chemical and conformational study of the interactions involved in mycotoxin complexation with beta-D-glucans. Biomacromolecules 2006, 7, 1147–1155. [Google Scholar] [CrossRef] [PubMed]
- Karaman, M.; Basmacioglu, H.; Ortatatli, M.; Oguz, H. Evaluation of the detoxifying effect of yeast glucomannan on aflatoxicosis in broilers as assessed by gross examination and histopathology. Brit. Poult. Sci. 2005, 46, 394–400. [Google Scholar] [CrossRef]
- Lahtinen, S.J.; Haskard, C.A.; Ouwehand, A.C.; Salminen, S.J.; Ahokas, J.T. Binding of aflatoxin B1 to cell wall components of Lactobacillus rhamnosus strain GG. Food Addit. Contam. A 2004, 21, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Herrera, J. Fungal Cell Wall: Structure, Synthesis and Assembly; CRC Press: Boca Raton, FL, USA, 1992. [Google Scholar]
- Ha, H.-C. Screening and production of lignocellulolytic enzymes secreted by the edible basidiomycete Pleurotus ostreatus. J. Mushrooms 2012, 10, 74–82. [Google Scholar] [CrossRef]
- Guan, S.; Ji, C.; Zhou, T.; Li, J.X.; Ma, Q.G.; Niu, T.G. Aflatoxin B1 degradation by Stenotrophomonas maltophilia and other microbes selected using coumarin medium. Int. J. Mol. Sci. 2008, 9, 1489–1503. [Google Scholar] [CrossRef] [Green Version]
- Abbas, H.K.; Zablotowicz, R.M.; Bruns, H.A.; Abel, C.A. Biocontrol of aflatoxin in corn by inoculation with non-aflatoxigenic Aspergillus flavus isolates. Biocontrol Sci. Technol. 2006, 16, 437–449. [Google Scholar] [CrossRef]
- Dubois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.A.; Smith, F. Colorimetric Method for Determination of Sugars and Related Substances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Rondle, C.J.; Morgan, W.T. The determination of glucosamine and galactosamine. Biochem. J. 1955, 61, 586–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Cell Fraction | Total Carbohydrate (mg/mL) | Protein (mg/mL) | Glucosamine (mg/mL) |
|---|---|---|---|
| Cell-free supernatant | 42.75 ± 2.64 a | 10.52 ± 1.16 a | N.D. 1 |
| Cell lysate | 11.15 ± 3.63 b | 11.78 ± 0.03 a | N.D. |
| Cell debris | 2.41 ± 0.03 c | 5.12 ± 0.20 b | 0.30 ± 0.01 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choo, M.-J.; Hong, S.-Y.; Chung, S.-H.; Om, A.-S. Removal of Aflatoxin B1 by Edible Mushroom-Forming Fungi and Its Mechanism. Toxins 2021, 13, 668. https://doi.org/10.3390/toxins13090668
Choo M-J, Hong S-Y, Chung S-H, Om A-S. Removal of Aflatoxin B1 by Edible Mushroom-Forming Fungi and Its Mechanism. Toxins. 2021; 13(9):668. https://doi.org/10.3390/toxins13090668
Chicago/Turabian StyleChoo, Min-Jung, Sung-Yong Hong, Soo-Hyun Chung, and Ae-Son Om. 2021. "Removal of Aflatoxin B1 by Edible Mushroom-Forming Fungi and Its Mechanism" Toxins 13, no. 9: 668. https://doi.org/10.3390/toxins13090668
APA StyleChoo, M.-J., Hong, S.-Y., Chung, S.-H., & Om, A.-S. (2021). Removal of Aflatoxin B1 by Edible Mushroom-Forming Fungi and Its Mechanism. Toxins, 13(9), 668. https://doi.org/10.3390/toxins13090668




