Detoxification of Mycotoxins through Biotransformation
Abstract
1. Introduction
2. Reaction Types of Mycotoxin Biotransformation
2.1. Hydroxylation
2.2. Oxido-Reduction between Alcohols and Ketones
2.3. Hydrogenation of the Carbon-Carbon Double Bond
2.4. De-Epoxidation
2.5. Other Oxido-Reductions
2.6. Methylation
2.7. Glycosylation and Glucuronidation
2.8. Esterification
2.9. Hydrolysis
2.10. Sulfation
2.11. Demethylation
2.12. Deamination
2.13. Miscellaneous Reactions
3. Detoxification of Important Mycotoxins by Biotransformation
3.1. Detoxification of Aflatoxins
3.2. Detoxification of Alternaria Toxins
3.3. Detoxification of Citrinin
3.4. Detoxification of Fomannoxin
3.5. Detoxification of Fumonisins
3.6. Detoxification of Ochratoxins
3.7. Detoxification of Patulin
3.8. Detoxification of Trichothecenes
3.9. Detoxification of Zearalenone Analogues
4. Conclusions and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Var, I.; Kabak, B.; Erginkaya, Z. Reduction in ochratoxin A levels in white wine, following treatment with activated carbon and sodium bentonite. Food Control 2008, 19, 592–598. [Google Scholar] [CrossRef]
- Diao, E.; Li, X.; Zhang, Z.; Ma, W.; Ji, N.; Dong, H. Ultraviolet irradiation detoxification of aflatoxins: A review. Trends Food Sci. Technol. 2015, 42, 64–69. [Google Scholar] [CrossRef]
- Freitas-Silva, O.; Venancio, A. Ozone applications to prevent and degrade mycotoxin: A review. Drug Metab. Rev. 2010, 42, 612–620. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S.P. Microbial detoxification of mycotoxins. J. Chem. Ecol. 2013, 39, 907–918. [Google Scholar] [CrossRef] [PubMed]
- Hathout, A.S.; Aly, S.E. Biological detoxification of mycotoxins: A review. Ann. Microbiol. 2014, 64, 905–919. [Google Scholar] [CrossRef]
- Ji, C.; Fan, Y.; Zhao, L. Review on biological degradation of mycotoxins. Anim. Nutr. 2016, 2, 127–133. [Google Scholar] [CrossRef]
- Loi, M.; Fanelli, F.; Liuzzi, V.C.; Logrieco, A.F.; Mule, G. Mycotoxin biotransformation by native and commercial enzymes: Present and future perspectives. Toxins 2017, 9, 111. [Google Scholar] [CrossRef]
- Taheur, F.B.; Kouidhi, B.; Al Qurashi, Y.M.A.; Salah-Abbes, J.B.; Chaieb, K. Review: Biotechnology of mycotoxins detoxification using microorganisms and enzymes. Toxicon 2019, 160, 12–22. [Google Scholar] [CrossRef]
- He, J.; Zhou, T.; Young, J.C.; Boland, G.J.; Scott, P.M. Chemical and biological transformations for detoxification of trichothecene mycotoxins in human and animal food chains: A review. Trends Food Sci. Technol. 2010, 21, 67–76. [Google Scholar] [CrossRef]
- Zhu, Y.; Hassan, Y.I.; Watts, C.; Zhou, T. Innovative technologies for the mitigation of mycotoxins in animal feed and ingredients—A review of recent patents. Anim. Feed Sci. Technol. 2016, 216, 19–29. [Google Scholar] [CrossRef]
- Zhu, Y.; Hassan, Y.I.; Lepp, D.; Shao, S.; Zhou, T. Strategies and methodologies for developing microbial detoxification systems to mitigate mycotoxins. Toxins 2017, 9, 130. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Wu, W.; Pan, J.; Long, M. Detoxification strategies for zearalenone using microorganism: A review. Microorgansims 2019, 7, 208. [Google Scholar] [CrossRef] [PubMed]
- Karlovsky, P. Biological detoxification of fungal toxins and its use in plant breeding, feed and food production. Nat. Toxins 1999, 7, 1–23. [Google Scholar] [CrossRef]
- Wu, Q.; Jezkova, A.; Yuan, Z.; Pavlikova, L.; Dohnal, V.; Kuca, K. Biological degradation of aflatoxins. Drug Metab. Dev. 2009, 41, 1–7. [Google Scholar] [CrossRef]
- Wu, Q.; Dohnal, V.; Huang, L.; Kuca, K.; Yuan, Z. Metabolic pathways of trichothecenes. Drug Metab. Rev. 2010, 42, 250–267. [Google Scholar] [CrossRef]
- Wu, Q.; Dohnal, V.; Huang, L.; Kuca, K.; Wang, X.; Chen, G.; Yuan, Z. Metabolic pathways of ochratoxin A. Curr. Drug Metab. 2011, 12, 1–10. [Google Scholar] [CrossRef]
- Vanhoutte, I.; Audenaert, K.; De Gelder, L. Biodegradation of mycotoxins: Tales from known and unexplored worlds. Front. Microbiol. 2016, 7, 561. [Google Scholar] [CrossRef]
- Peng, Z.; Chen, L.; Zhu, Y.; Huang, Y.; Hu, X.; Wu, Q.; Nussler, A.K.; Liu, L.; Yang, W. Current major degradation methods for aflatoxins: A review. Trends Food Sci. Technol. 2018, 80, 155–166. [Google Scholar] [CrossRef]
- Bryla, M.; Waskiewicz, A.; Ksieniewicz-Wozniak, E.; Szymczyk, K.; Jedrzejczak, R. Modified Fusarium mycotoxins and their products-metabolism, occurrence, and toxicity: An updated review. Molecules 2018, 23, 963. [Google Scholar] [CrossRef]
- Gallagher, E.P.; Eaton, D.L. In vitro biotransformation of aflatoxin B1 (AFB1) in channel catfish liver. Toxicol. Appl. Pharmacol. 1995, 132, 82–90. [Google Scholar] [CrossRef]
- Halvorson, M.R.; Safe, S.H.; Parkinson, A.; Phillips, T.D. Aflatoxin B1 hydroxylation by the pregnenolone-16α-carbonitrile-inducible form of rat liver microsomal cytochrome P-450. Carcinogenesis 1988, 9, 2103–2108. [Google Scholar] [CrossRef] [PubMed]
- Krieger, R.I.; Salhab, A.S.; Dalezios, J.I.; Hseh, D.P.H. Aflatoxin B1 hydroxylation by hepatic microsomal preparations from the rhesus money. Food Cosmet. Toxicol. 1975, 13, 211–219. [Google Scholar] [CrossRef]
- Pfeiffer, E.; Schebb, N.H.; Podlech, J.; Metzler, M. Novel oxidative in vitro metabolites of the mycotoxins alternariaol and alternariol methyl ether. Mol. Nutr. Food Res. 2007, 51, 307–316. [Google Scholar] [CrossRef] [PubMed]
- Pedras, M.S.C.; Zaharia, I.L.; Gai, Y.; Zhou, Y.; Ward, D.E. In planta sequential hydroxylation and glycosylation of a fungal phytotoxin: Avoiding cell death and overcoming the fungal invader. Proc. Nat. Acad. Sci. USA 2001, 98, 747–752. [Google Scholar] [CrossRef]
- Pedras, M.S.C.; Khallaf, I. Molecular interactions of the phytotoxins destruxin B and sirodesmin PL with crucifers and cereals: Metabolism and elicitation of plant defenses. Phytochemistry 2012, 77, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Ng, T.B. Pharmacological activities of fusaric acid (5-butylpicolinic acid). Life Sci. 1999, 65, 849–856. [Google Scholar] [CrossRef]
- Crutcher, F.K.; Puckhaber, L.S.; Bell, A.A.; Liu, J.; Duke, S.E.; Stipanovic, R.D.; Nichols, R.L. Detoxification of fusaric acid by the soil microbe Mucor rouxii. J. Agric. Food Chem. 2017, 65, 4989–4992. [Google Scholar] [CrossRef]
- Wegst, W.; Lingens, F. Bacterial degradation of ochratoxin A. FEMS Microbiol. Lett. 1983, 17, 341–344. [Google Scholar] [CrossRef]
- Stormer, F.C.; Pedersen, J.I. Formation of 4-hydroxyochratoxin A from ochratoxin A by rat liver microsomes. Appl. Environ. Microbiol. 1980, 39, 971–975. [Google Scholar] [CrossRef]
- Stormer, F.C.; Storen, O.; Hansen, C.E.; Pedersen, J.I.; Aasen, A.J. Formation of (4R)- and (4S)-4-hydroxyochratoxin A and 10-hydroxyochratoxin A from ochratoxin A by rabbit liver microsomes. Appl. Environ. Microbiol. 1983, 45, 1183–1187. [Google Scholar] [CrossRef]
- Ruhland, M.; Engelhardt, G.; Wallnofer, P.R.; Schafer, W. Transformation of the mycotoxin ochratoxin A in wheat and maize cell suspension cultures. Naturwissenschaften 1994, 81, 453–454. [Google Scholar] [CrossRef] [PubMed]
- Mally, A.; Zepnik, H.; Wanek, P.; Eder, E.; Dingley, K.; Ihmels, H.; Volkel, W.; Dekant, W. Ochratoxin A: Lack of formation of covalent DNA adducts. Chem. Res. Toxicol. 2004, 17, 234–242. [Google Scholar] [CrossRef]
- Nieto, C.H.D.; Granero, A.M.; Zon, M.A.; Fernandez, H. Sterigmatocystin: A mycotoxin to be seriously considered. Food Chem. Toxicol. 2018, 118, 460–470. [Google Scholar] [CrossRef] [PubMed]
- Pfeiffer, E.; Fleck, S.C.; Metzler, M. Catechol formation: A novel pathway in the metabolism of sterigmatocystin and 11-methoxysterigmatocystin. Chem. Res. Toxicol. 2014, 27, 2093–2099. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Zhou, X.; Yang, J.; Li, M.; Qiu, X. T-2 toxin is hydroxylated by chicken CYP3A37. Food Chem. Toxicol. 2013, 62, 622–627. [Google Scholar] [CrossRef]
- Shang, S.; Jiang, J.; Deng, Y. Chicken cytochrome P4501A5 is the key enzyme for metabolizing T-2 toxin to 3′OH-T-2. Int. J. Mol. Sci. 2013, 14, 10809–10818. [Google Scholar] [CrossRef]
- Yoshizawa, T.; Sakamoto, T.; Okamoto, K. In vitro formation of 3′-hydroxy T-2 and 3′-hydroxy HT-2 toxins from T-2 toxin by liver homogenates from mice and monkeys. Appl. Environ. Microbiol. 1984, 47, 130–134. [Google Scholar] [CrossRef]
- El-Sharkawy, S.H.; Abul-Hajj, Y.J. Microbial transformation of zearalenone. 2. Reduction, hydroxylation, and methylation products. J. Org. Chem. 1988, 53, 515–519. [Google Scholar] [CrossRef]
- Hildebrand, A.A.; Pfeiffer, E.; Rapp, A.; Metzler, M. Hydroxylation of the mycotoxin zearalenone at aliphatic positions: Novel mammalian metabolites. Mycotoxin Res. 2012, 28, 1–8. [Google Scholar] [CrossRef]
- Pfeiffer, E.; Hildebrand, A.; Damm, G.; Rapp, A.; Cramer, B.; Uumpf, H.-U.; Metzler, M. Aromatic hydroxylation is a major metabolic pathway of the mycotoxin zearalenone in vitro. Mol. Nutr. Food Res. 2009, 53, 1123–1133. [Google Scholar] [CrossRef]
- Nakazato, M.; Morozumi, S.; Saito, K.; Fujinuma, K.; Nishima, T.; Kasai, N. Interconversion of aflatoxin B1 and aflatoxicol by several fungi. Appl. Environ. Microbiol. 1990, 56, 1465–1470. [Google Scholar] [CrossRef] [PubMed]
- Carere, J.; Hassan, Y.I.; Lepp, D.; Zhou, T. The enzymatic detoxification of the mycotoxin deoxynivalenol: Identification of DepA from the DON Epimerization pathway. Microb. Biotechnol. 2018, 11, 1106–1111. [Google Scholar] [CrossRef] [PubMed]
- Heslin, M.C.; Stuart, M.R.; Murchu, P.O.; Donnelly, D.M.X. Fomannoxin, a phytotoxic metabolite of Fomes annosus: In vitro production, host toxicity and isolation from naturally infected Sitka spruce heartwood. Eur. J. For. Pathol. 1983, 13, 11–23. [Google Scholar] [CrossRef]
- Zweimuller, M.; Antus, S.; Kovacs, T.; Sonnenbichler, J. Biotransformation of the fungal toxin formannoxin by conifer cell cultures. Biol. Chem. 1997, 378, 915–921. [Google Scholar] [CrossRef] [PubMed]
- Palyusik, M.; Hagler, W.M.; Horvath, L.; Microcha, C.J. Biotransformation of zearalenone to zearalenol by Candida tropicalis. Acta Vet. Acad. Sci. Hugaricae 1980, 28, 159–166. [Google Scholar]
- Boswald, C.; Engelhardt, G.; Vogel, H.; Wallnofer, P.R. Metabolism of the Fusarium mycotoxins zearalenone and deoxynivalenol by yeast strains of technological relevance. Nat. Toxins 1995, 3, 138–144. [Google Scholar] [CrossRef]
- Brodehl, A.; Moller, A.; Kunte, H.-J.; Koch, M.; Maul, R. Biotransformation of the mycotoxin zearalenone by fungi of the genera Rhizopus and Aspergillus. FEMS Microbiol. Lett. 2014, 359, 124–130. [Google Scholar] [CrossRef]
- Horlacher, N.; Nachtigall, J.; Schulz, D.; Sussmuth, R.D.; Hampp, R.; Fiedler, H.-P.; Schrey, S.D. Biotransformation of the fungal phytotoxin fomannoxin by soil streptomycetes. J. Chem. Ecol. 2013, 39, 931–941. [Google Scholar] [CrossRef]
- Miles, C.O.; Erasmuson, A.F.; Wilkins, A.L.; Towers, N.R.; Smith, B.L.; Garthwaite, I.; Scahill, B.G.; Hansen, R.P. Ovine metabolism of zearalenone to α-zearalanol (zeranol). J. Agric. Food Chem. 1996, 44, 3244–3250. [Google Scholar] [CrossRef]
- Binder, J.; Horvath, E.M.; Schatzmayr, G.; Ellend, N.; Danner, H.; Krska, R.; Braun, R. Screening for deoxynivalenol-detoxifying anaerobic rumen microorganisms. Cereal Res. Commun. 1997, 25, 343–346. [Google Scholar] [CrossRef]
- Fuchs, E.; Binder, E.M.; Heidler, D.; Krska, R. Characterisation of metabolites after the microbial degradation of A- and B-trichothecenes by BBSH797. Mycotoxin Res. 2000, 16, 66–69. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; He, J.; Gong, J. Microbial transformation of trichothecene mycotoxins. World Mycotoxin J. 2008, 1, 23–30. [Google Scholar] [CrossRef]
- Onji, Y.; Dohi, Y.; Aoki, Y.; Moriyama, T.; Nagami, H.; Uno, M.; Tanaka, T.; Yamazoe, Y. Deepoxynivalenol: A new metabolite of nivalenol found in the excreta of orally administered rats. J. Agric. Food Chem. 1989, 37, 478–481. [Google Scholar] [CrossRef]
- Wang, J.; Ogata, M.; Hirai, H.; Kawagishi, H. Detoxification of aflatoxin B1 by manganese peroxidase from the white-rot fungus Phanerochaete sordida YK-624. FEMS Microbiol. Lett. 2011, 314, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Fleck, S.C.; Pfeiffer, E.; Podlech, J.; Metzler, M. Epoxide reduction to an alcohol: A novel metabolic pathway for perylene quinone-type Alternaria mycotoxins in mammalian cells. Chem. Res. Toxicol. 2014, 27, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Daoubi, M.; Duran-Patron, R.; Hernandez-Galan, R.; Benharref, A.; Hanson, J.R.; Collado, I.G. The role of botrydienediol in the biodegradation of the sesquiterpenoids phytotoxin botrydial by Botrytis cinerea. Tetrahedron 2006, 62, 8256–8261. [Google Scholar] [CrossRef]
- Dunn, B.B.; Stack, M.E.; Park, D.L.; Joshi, A.; Friedman, L.; King, R.L. Isolation and identification of dihydrocitrinone, a urinary metabolite of citrinin in rats. J. Toxicol. Environ. Health 1983, 12, 283–289. [Google Scholar] [CrossRef]
- Follmann, W.; Behm, C.; Degen, G.H. Toxicity of the mycotoxin citrinin and its metabolite dihydrocitrinoine and of mixtures of citrinin and ochratoxin A in vitro. Arch. Toxicol. 2014, 88, 1097–1107. [Google Scholar] [CrossRef]
- Crutcher, F.K.; Liu, J.; Puckhaber, L.S.; Stipanovic, R.D.; Duke, S.E.; Bell, A.A.; Williams, H.J.; Nichols, R.L. Conversion of fusaric acid to fusarinol by Aspergillus tubingensis: A detoxification reaction. J. Chem. Ecol. 2014, 40, 84–89. [Google Scholar] [CrossRef]
- Dong, X.; Jiang, W.; Li, C.; Ma, N.; Xu, Y.; Meng, X. Patulin biodegradation by marine yeast Kodameae ohmeri. Food Addit. Contam. A 2015, 32, 352–360. [Google Scholar]
- Tannous, J.; Snini, S.P.; El Khoury, R.; Canlet, C.; Pinton, P.; Lippi, Y.; Alassane-Kpembi, I.; Gauthier, T.; El Khoury, A.; Atoui, A.; et al. Patulin transformation products and last intermediates in its biosynthetic pathway, E- and Z-ascladiol, are not toxic to human cells. Arch. Toxicol. 2016, 91, 2455–2467. [Google Scholar] [CrossRef] [PubMed]
- Hawar, S.; Vevers, W.; Karieb, S.; Ali, B.K.; Billington, R.; Beal, J. Biotransformation of patulin to hydroascladiol by Lactobacillus plantarum. Food Control 2013, 34, 502–508. [Google Scholar] [CrossRef]
- Stinson, E.E.; Moreau, R.A. Partial purification and some properties of an alternariol-O-methyltransferase from Alternaria tenuis. Phytochemistry 1986, 25, 2721–2724. [Google Scholar]
- Hiltunen, M.; Soderhall, K. Inhibition of polyketide synthesis in Alternaria alternata by the fatty acid synthesis inhibitor cerulenin. Appl. Environ. Microbiol. 1992, 58, 1043–1045. [Google Scholar] [CrossRef] [PubMed]
- Wetterhorn, K.M.; Newmister, S.A.; Caniza, R.K.; Busman, M.; McCormick, S.P.; Berthiller, F.; Adam, G.; Rayment, I. Crystal structure of Os79 (Os04g0206600) from Oryza sativa: A UDP-glucosyltransferase involved in the detoxification of deoxynivalenol. Biochemistry 2016, 55, 6175–6186. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, A.A.; Kohn, B.N.; Pfeiffer, E.; Wefers, D.; Metzler, M.; Bunzel, M. Conjugation of the mycotoxins alternariol and alternariol monomethyl ether in tobacco suspension cells. J. Agric. Food Chem. 2015, 63, 4728–4736. [Google Scholar] [CrossRef] [PubMed]
- Soukup, S.T.; Kohn, B.N.; Pfeiffer, E.; Geisen, R.; Metzler, M.; Bunzel, M.; Kulling, S.E. Sulfoglucosides as novel modified forms of the mycotoxins alternariol and alternariol monomethyl ether. J. Agric. Food Chem. 2016, 64, 8892–8901. [Google Scholar] [CrossRef]
- Meepagala, K.M.; Johnson, R.D.; Duke, S.O. Curvularin and dehydrocurvularin as phytotoxic constituents from Curvularia intermedia infecting Pandaus amryllifolius. J. Agric. Chem. Environ. 2016, 5, 12–22. [Google Scholar]
- Zhan, J.; Gunatilak, A.A.L. Microbial transformation of curvularin. J. Nat. Prod. 2005, 68, 1271–1273. [Google Scholar] [CrossRef]
- Michlmayr, H.; Malachova, A.; Varga, E.; Kleinova, J.; Lemmens, M.; Newmister, S.; Rayment, I.; Berthiller, F.; Adam, G. Biochemical characterization of a recombinant UDP-glucosltransferase from rice and enzymatic production of deoxynivalenol-3-O-β-d-glucoside. Toxins 2015, 7, 2685–2700. [Google Scholar] [CrossRef]
- Schwartz-Zimmermann, H.E.; Hametner, C.; Nagl, V.; Fiby, I.; Macheiner, L.; Winkler, J.; Danicke, S.; Clark, E.; Pestka, J.J.; Berthiller, F. Glucuronidation of deoxynivalenol (DON) by different animal species: Identification of iso-DON glucuronides and iso-deepoxy-DON glucuronides as novel DON metabolites in pigs, rats, mice, and cows. Arch. Toxcol. 2017, 91, 3857–3872. [Google Scholar] [CrossRef] [PubMed]
- Schwartz-Zimmermann, H.E.; Hametner, C.; Nagl, V.; Fiby, I.; Macheiner, L.; Winkler, J.; Danicke, S.; Clark, E.; Pestka, J.J.; Berthiller, F. Correction to: Glucuronidation of deoxynivalenol (DON) by different animal species: Identification of iso-DON glucuronides and iso-deepoxy-DON glucuronides as novel DON metabolites in pigs, rats, mice, and cows. Arch. Toxcol. 2018, 92, 3245–3246. [Google Scholar] [CrossRef] [PubMed]
- Schollenberger, M.; Muller, H.-M.; Liebscher, M.; Schlecker, C.; Berger, M.; Hermann, W. Accumulation kinetics of three scirpentriol-based toxins in oats inoculated in vitro with isolates of Fusairum sporotrichioides and Fusarium poae. Toxins 2011, 3, 442–452. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, H.; Sakamoto, S.; Sago, Y.; Kushiro, M.; Nagashima, H. Detection of masked mycotoxins derived from type A trichothecenes in corn by high-resolution LC-Orbitrap mass spectrometer. Food Addit. Contam. A 2013, 30, 1407–1414. [Google Scholar] [CrossRef]
- Nakagawa, H. Research on mycotoxin glucosides (masked mycotoxins). JSM Mycotoxins 2016, 66, 21–25. [Google Scholar] [CrossRef]
- Gorst-Allman, C.P.; Steyn, P.S.; Vleggaar, R.; Rabie, C.J. Structure elucidation of a novel trichothecene glycoside using 1H and 13C nuclear magnetic resonance spectroscopy. J. Chem. Soc. Perkin Trans. 1 1985, 1985, 1553–1555. [Google Scholar] [CrossRef]
- Busan, M.; Poling, S.M.; Maragos, C.M. Observation of T-2 toxin and HT-2 toxin glucosides from Fusarium sporotrichioides by liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS). Toxins 2011, 3, 1554–1568. [Google Scholar] [CrossRef]
- Roush, W.R.; Marletta, M.A.; Russo-Rodriguez, S.; Recchia, J. Trichotecene metabolism studies: Isolation and structure determination of 15-acetyl-3α-(1′β-d-glucopyranosiduronyl)-scirpen-3,4β,15-triol. J. Am. Chem. Soc. 1985, 107, 3354–3355. [Google Scholar] [CrossRef]
- McCormick, S.P.; Price, N.P.J.; Kurtzman, C.P. Glucosylation and other biotransformations of T-2 toxin by yeasts of the Trichomonascus Clade. Appl. Environ. Microbiol. 2012, 78, 8694–8702. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, Y.; Wang, Y.; Sun, L.; Gooneratne, R. Preparation of T-2-glucoronide with rat hepatic microsomes and its use along with T-2 for activation of the JAK/STAT signaling pathway in RAW264.7 cells. J. Agric. Food Chem. 2017, 65, 4811–4818. [Google Scholar] [CrossRef] [PubMed]
- Poppenberger, B.; Berthiller, F.; Bachmann, H.; Lucyshyn, D.; Peterbauer, C.; Mitterbauer, R.; Schuhmacher, R.; Krska, R.; Glossl, J.; Adam, G. Heterologous expression of Arabidopsis UDP-glucosyltransferases in Saccharomyces cerevisiae for production of zearalenone-4-O-glucoside. Appl. Environ. Microbiol. 2006, 72, 4404–4410. [Google Scholar] [CrossRef] [PubMed]
- El-Sharkawy, S.H.; Abul-Hajj, Y.J. Microbial transformation of zearalenone. 1. Formation of zearalenone-4-O-β-glucoside. J. Nat. Prod. 1987, 50, 520–521. [Google Scholar] [CrossRef]
- Paris, M.P.K.; Schweiger, W.; Hametner, C.; Stuckler, R.; Muehlbauer, G.J.; Varga, E.; Krska, R.; Berthiller, F.; Adam, G. Zearalenone-16-O-glucoside: A new masked mycotoxin. J. Agric. Food Chem. 2014, 62, 1181–1189. [Google Scholar] [CrossRef] [PubMed]
- Michlayr, H.; Varga, E.; Lupi, F.; Malachova, A.; Hametner, C.; Berthiller, F.; Adam, G. Synthesis of mono- and di-glucosides of zearalenone and α/β-zearalenol by recombinant barley glucosyltransferase HvUGT14077. Toxins 2017, 9, 58. [Google Scholar] [CrossRef] [PubMed]
- Munger, C.E.; Ivie, G.W.; Christopher, R.J.; Hammock, B.D.; Phillips, T.D. Acetylation/deacetylation reactions of T-2, acetyl T-2, HT-2, and acetyl HT-2 toxins in bovine rumen fluid in vitro. J. Agric. Food Chem. 1987, 35, 354–358. [Google Scholar] [CrossRef]
- Fuchs, E.; Binder, E.M.; Heiler, D.; Krska, R. Structural characterization of metabolites after the microbial degradation of type A trichothecenes by the bacterial strain BBSH 797. Food Addi. Contam. 2002, 19, 379–386. [Google Scholar] [CrossRef]
- Kim, S.H.; Vujanovic, V. Biodegradation and biodetoxification of Fusarium mycotoxins by Sphaerodes mycoparasitica. AMB Expr. 2017, 7, 145. [Google Scholar] [CrossRef]
- Udell, M.N.; Dewick, P.M. Metabolic conversions of trichothecene mycotoxins: De-esterification reactions using cell-free extracts of Fusarium. Z. Naturforsch. C 1989, 44, 660–668. [Google Scholar] [CrossRef]
- Duvick, J.; Rood, T.; Maddox, J.; Gilliam, J. Detoxification of mycotoxins in planta as a strategy for improving grain quality and disease resistance: Identification of fumonisin-degrading microbes from maize. Dev. Plant Pathol. 1998, 13, 369–381. [Google Scholar]
- Heinl, S.; Hartinger, D.; Thamhesl, M.; Vekiru, E.; Krska, R.; Schatzmayr, G.; Moll, W.-D.; Grabherr, R. Degradation of fumonisin B1 by the consecutive action of two bacterial enzymes. J. Biotechnol. 2010, 145, 120–129. [Google Scholar] [CrossRef]
- Hartinger, D.; Schwartz, H.; Hametner, C.; Schatzmayr, G.; Haltrich, D.; Moll, W.-D. Enzyme characteristics of aminotransferase FumI of Sphingopyxis sp. MTA144 for deamination of hydrolyzed fumonisin B1. Appl. Microbiol. Biotechnol. 2011, 91, 757–768. [Google Scholar] [CrossRef] [PubMed]
- Poapolathep, A.; Singhasem, S.; Noonpugdee, C.; Sugita-Konishi, Y.; Doi, K.; Kumagai, S. The fate and transmission of fusarenon-X (FX), a trichothecene mycotoxin in mice. Toxicol. Appl. Pharmacol. 2004, 197, 367. [Google Scholar]
- Phruksawan, W.; Poapolathep, S.; Giorgi, M.; Imsilp, K.; Sakulthaew, C.; Owen, H.; Poapolathep, A. Toxicokinetic profile of fusarenon-X and its metabolite nivalenol in the goat (Capra hircus). Toxicon 2018, 153, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Stander, M.A.; Bornscheuer, U.T.; Henke, E.; Steyn, P.S. Screening of commercial hydrolases for the degradation of ochratoxin A. J. Agric. Food Chem. 2000, 48, 5736–5739. [Google Scholar] [CrossRef]
- Abrunhosa, L.; Santos, L.; Verancio, A. Degradation of ochratoxin by proteases and by a crude enzyme of Aspergillus niger. Food Biotechnol. 2006, 20, 231–240. [Google Scholar] [CrossRef]
- Pitout, M.J. The hydrolysis of ochratoxin A by some proteolytic enzymes. Biochem. Pharmacol. 1969, 18, 485–491. [Google Scholar] [CrossRef]
- Varga, J.; Rigo, K.; Teren, J. Degradation of ochratoxin A by Aspergillus species. Int. J. Food Microbiol. 2000, 59, 1–7. [Google Scholar] [CrossRef]
- Chang, X.; Wu, Z.; Wu, S.; Dai, Y.; Sun, C. Degradation of ochratoxin A by Bacillus amyloliquefaciens ASAG1. Food Addit. Contam. A 2015, 32, 564–571. [Google Scholar] [CrossRef]
- Li, S.; Marquardt, R.R.; Frohlich, A.A.; Vitti, T.G.; Crow, G. Pharmacokinetics of ochratoxin A and its metabolites in rats. Toxicol. Appl. Pharmacol. 1997, 145, 82–90. [Google Scholar] [CrossRef]
- Fuchs, R.; Hult, K.; Peraica, M.; Razica, R.; Plestina, R. Conversion of ochratoxin C into ochratoxin A in vivo. Appl. Environ. Microbiol. 1984, 48, 41–42. [Google Scholar] [CrossRef]
- Ohta, M.; Ishii, K.; Ueno, Y. Metabolism of trichothecene mycotoxins I. J. Biochem. 1977, 82, 1591–1598. [Google Scholar] [CrossRef] [PubMed]
- Yoshizawa, T.; Swanson, S.P.; Mirocha, C.J. In vitro metabolism of T-2 toxin in rats. Appl. Environ. Microbiol. 1980, 40, 901–906. [Google Scholar] [CrossRef] [PubMed]
- Catteuw, A.; Broekaert, N.; De Baere, S.; Lauwers, M.; Gasthuys, E.; Huybrechts, B.; Callebaut, A.; Ivanova, L.; Uhlig, S.; De Boevre, M.; et al. Insights into in vivo absolute oral bioavailability, biotransformation, and toxicokinetics of zearalenone, α-zearalenol, β-zearalenol, zearalenone-14-glucosdie, and zearalenone-14-sulfate in pigs. J. Agric. Food Chem. 2019, 67, 3448–3458. [Google Scholar] [CrossRef] [PubMed]
- Olsen, M.; Visconti, A. Metabolism of alternariol monomethylether by porcine liver and intestinal mucosa in vitro. Toxicol. In Vitro 1988, 2, 27–29. [Google Scholar] [CrossRef]
- Niu, G.; Wen, Z.; Rupasinghe, S.G.; Zeng, R.S.; Berenbaum, M.R.; Schuler, M.A. Aflatoxin B1 detoxification by CYP321A1 in Helicomverpa zea. Arch. Insect biochem. Physiol. 2008, 69, 32–45. [Google Scholar] [CrossRef]
- Burgess, K.M.N.; Renaud, J.B.; McDowell, T.; Sumarah, M.W. Mechanistic insight into the biosynthesis and detoxification of fumonisin mycotoxins. ACS Chem. Biol. 2016, 11, 2618–2625. [Google Scholar] [CrossRef]
- Blackwell, B.A.; Gilliam, J.T.; Savard, M.E.; Miller, D.; Duvick, J.P. Oxidative deamination of hydrolyzed fumonisin B1 (AP1) by cultures of Exophiala spinifera. Nat. Toxins 1999, 7, 31–38. [Google Scholar] [CrossRef]
- Das, A.; Bhattacharya, S.; Palaniswamy, M.; Angayarkanni, J. Biodegradation of aflatoxin B1 incontaminated rice straw by Pleurotus ostreatus MTCC142 and Pleurotus ostreatus GHBBF10 in the presence of metal salts and surfactants. World J. Microbiol. Biotechnol. 2014, 30, 2315–2324. [Google Scholar] [CrossRef]
- Eshelli, M.; Harvey, L.; Edrada-Ebel, R.; McNeil, B. Metabolomics of the bio-degradation process of aflatoxin B1 by actinomycetes at an initial pH of 6.0. Toxins 2015, 7, 439–456. [Google Scholar] [CrossRef]
- Samuel, M.S.; Sivaramakrishna, A.; Mehta, A. Degradation and detoxification of aflatoxin B1 by Pseudomonas putida. Int. Biodeterior. Biodegr. 2014, 86, 202–209. [Google Scholar] [CrossRef]
- Devi, P.; Naik, C.G.; Rodrigues, C. Biotransformation of citrinin to decarboxycitrinin using an organic solvent-tolerant marine bacterium, Moraxella sp. MB1. Mar. Biotechnol. 2006, 8, 129–138. [Google Scholar] [CrossRef]
- Jackson, L.K.; Ciegler, A. Production and analysis of citrinin in corn. Appl. Environ. Microbiol. 1978, 36, 408–411. [Google Scholar] [CrossRef]
- Ikunaga, Y.; Sato, I.; Grond, S.; Numaziri, N.; Yoshida, S.; Yamaya, H.; Hiradate, S.; Hasegawa, M.; Toshima, H.; Koitabashi, M. Nocardioides sp. strain WSN05-2, isolated from a wheat field, degrades deoxynivalenol, producing the novel intermediate 3-epi-deoxynivalenol. Appl. Microbiol. Biotechnol. 2011, 89, 419–427. [Google Scholar] [CrossRef]
- Hassan, Y.I.; Zhou, T. Addressing the mycotoxin deoxynivalenol contamination with soil-derive bacterial and enzymatic transformations targeting the C3 carbon. World Mycotoxin J. 2018, 11, 101–111. [Google Scholar] [CrossRef]
- Hassan, Y.; He, J.W.; Perilla, N.; Tang, K.J.; Karlovsky, P.; Zhou, T. The enzymatic epimerization of deoxynivalenol by Devosia mutans proceeds through the formation of 3-keto-DON intermediate. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef]
- Stadler, D.; Lambertini, F.; Bueschl, C.; Wiesenberger, G.; Hametner, C.; Shwartz-Zimmermann, H.; Hellinger, R.; Sulyok, M.; Lemmens, M.; Schuhmacher, R.; et al. Untargeted LC-MS based 13C labelling provides a full mass balance of deoxynivalenol and its degradation products formed during baking of crackers, biscuits and bread. Food Chem. 2019, 279, 303–311. [Google Scholar] [CrossRef]
- Adlouni, C.E.; Pinelli, E.; Azemar, B.; Zaoui, D.; Beaune, P.; Pfohl-Leszkowicz, A. Phenobarbital increases DNA adduct and metabolites formed by ochratoxin A: Role of CYP 2C9 and microsomal glutathione-S-transferase. Environ. Mol. Mutagen. 2000, 35, 123–131. [Google Scholar] [CrossRef]
- Pinedo, C.; Wright, S.A.I.; Collado, I.G.; Goss, R.J.M.; Castoria, R.; Hrelia, P.; Maffei, F.; Duran-Patron, R. Isotopic labeling studies reveal the patulin detoxification pathway by the biocontrol yeast Rhodotorula kratochvilovae LS11. J. Nat. Prod. 2018, 81, 2692–2699. [Google Scholar] [CrossRef]
- Zhu, R.; Feusner, K.; Wu, T.; Yan, F.; Karlovsky, P.; Zheng, X. Detoxification of mycotoxin patulin by the yeast Rhodosporidium paludigenum. Food Chem. 2015, 179, 1–5. [Google Scholar] [CrossRef]
- Castoria, R.; Mannina, L.; Duran-Patron, R.; Maffei, F.; Sobolev, A.P.; De Felice, D.V.; Pinedo-Rivilla, C.; Ritieni, A.; Ferracane, R.; Wright, S.A.I. Conversion of the mycotoxin patulin to the less toxic desoxypatulinic acid by the biocontrol yeast Rhodosporidium kratochvilovae strain LS11. J. Agric. Food Chem. 2011, 59, 11571–11578. [Google Scholar] [CrossRef]
- Belhassen, H.; Jimenez-Diaz, I.; Ghali, R.; Ghorbel, H.; Molina-Molina, J.M. Validation of a UHPLC-MS/MS method for quantification of zearaenone, α-zearalenol, β-zearalenol, α-zearalanol, β-zearalanol and zearalanone in human urine. J. Chromatogr. B 2014, 962, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Yu, M.; Dong, F.; Shi, J.; Xu, J. Esterase activity inspired selection and characterization of zearalenone degrading bacteria Bacillus pumilus ES-21. Food Control 2017, 77, 57–64. [Google Scholar] [CrossRef]
- Xu, Z.; Liu, W.; Chen, C.-C.; Li, Q.; Huang, J.-W.; Ko, T.-P.; Liu, G.; Liu, W.; Peng, W.; Cheng, Y.-S.; et al. Enhanced α-zearlenol hydrolyzing activity of a mycoestrogen-detoxifying lactonase by structure-based engineering. ACS Catal. 2016, 6, 7657–7663. [Google Scholar] [CrossRef]
- Fruhauf, S.; Novak, B.; Nagl, V.; Hackl, M.; Hartinger, D.; Rainer, V.; Labudova, S.; Adam, G.; Aleschko, M.; Moll, W.-D.; et al. Biotransformation of the mycotoxin zearalenone to its metabolites hydrolyzed zearalenone (HZEN) and decarboxylated hydrolyzed zearaleneone (DHEN) diminishes its estrogenicity in vitro and in vivo. Toxins 2019, 11, 481. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [PubMed]
- Samuel, M.S.; Aiko, V.; Panda, P.; Metha, A. Aflatoxin B1 occurrence, biosynthesis and its degradation. J. Pure Appl. Microbiol. 2013, 7, 965–971. [Google Scholar]
- Essigmann, J.M.; Croy, R.G.; Nadzan, A.M.; Busby, W.F.; Reinhold, V.N.; Buchi, G.; Wogan, G.N. Structural identification of the major DNA adduct formed by aflatoxin B1 in vitro. Proc. Nat. Acad. Sci. USA 1977, 74, 1870–1874. [Google Scholar] [CrossRef]
- Eaton, D.L.; Gallagher, E.P. Mechanisms of aflatoxin carcinogenesis. Annu. Rev. Pharmacol. Toxicol. 1994, 34, 135–172. [Google Scholar] [CrossRef]
- Schroeder, T.U.; Zweifel, P.; Sagelsdorff, U.; Friederich, J.; Luthy, C.; Schlatter, J. Ammoniation of aflatoxin-containing corn: Distribution, in vivo covalent deoxyribonucleic acid binding, and mutagenicity of reaction products. Agric. Food Chem. 1985, 33, 311–316. [Google Scholar] [CrossRef]
- Verheecke, C.; Liboz, T.; Mathieu, F. Microbial degradation of aflatoxin B1: Current status and future advances. Int. J. Food Microbiol. 2016, 237, 1–9. [Google Scholar] [CrossRef]
- Singh, V.P. Aflatoxin biotransformations: Biodetoxification aspects. Prog. Ind. Microbiol. 1995, 32, 51–63. [Google Scholar]
- Mishra, H.N.; Das, C. A review on biological control and metabolism of aflatoxin. Crit. Rev. Food Sci. Nutr. 2003, 43, 245–264. [Google Scholar] [CrossRef] [PubMed]
- Adebo, O.A.; Njobeh, P.B.; Gbashi, S.; Nwinyi, O.C.; Mavumengwana, V. Review on microbial degradation of aflatoxins. Crit. Rev. Food Sci. Nutr. 2017, 57, 3208–3217. [Google Scholar] [CrossRef] [PubMed]
- Rushing, B.R.; Selim, M.I. Aflatoxin B1: A review on metabolism, toxicity, occurrence in food, occupational exposure, and detoxification methods. Food Chem. Toxicol. 2019, 124, 81–100. [Google Scholar] [CrossRef] [PubMed]
- Lou, J.; Fu, L.; Peng, Y.; Zhou, L. Metabolites from Alternaria fungi and their bioactivities. Molecules 2013, 18, 5891–5935. [Google Scholar] [CrossRef]
- Liu, G.T.; Qian, Y.Z.; Zhang, P.; Dong, Z.M.; Shi, Z.Y.; Zhen, Y.Z.; Miao, J.; Xu, Y.M. Relationships between Alternaria alternata and oeophageal cancer. IARC Sci. Publ. 1991, 105, 258–262. [Google Scholar]
- Brugger, E.-M.; Wagner, J.; Schumacher, D.M.; Koch, K.; Podlech, J.; Metzler, M.; Lehmann, L. Mutagenicity of the mycotoxin alternariol in cultured mammalian cells. Toxicol. Lett. 2006, 164, 221–230. [Google Scholar] [CrossRef]
- Lehmann, L.; Wagner, J.; Metzler, M. Estrogenic and clastogenic potential of the mycotoxin alternariol in cultured mammalian cells. Food Chem. Toxicol. 2006, 44, 398–408. [Google Scholar] [CrossRef]
- Ponce-Garcia, N.; Serna-Saldivar, S.O.; Garcia-Lara, S. Fumonisins and their analogues in contaminated corn and its processed foods—A review. Food Addit. Contam. A 2018, 35, 2183–2203. [Google Scholar] [CrossRef]
- Soriano, J.M.; Gonzalez, L.; Catala, A.I. Mechanism of action of sphingolipids and their metabolites in the toxicity of fumonisin B1. Progr. Lipid Res. 2005, 44, 345–356. [Google Scholar] [CrossRef]
- Lumsangkul, C.; Chiang, H.; Lo, N.-W.; Fan, Y.-K.; Ju, J.-C. Developmental toxicity of mycototoxin fumonisin B1 in animal embryogenesis: An overview. Toxins 2019, 11, 114. [Google Scholar] [CrossRef]
- Voss, K.A.; Smith, G.W.; Haschek, W.M. Fumonisins: Toxicokinetics, mechanism of action and toxicity. Anim. Feed Sci. Technol. 2007, 137, 299–325. [Google Scholar] [CrossRef]
- Jestoi, M. Emerging Fusarium-mycotoxins fusaproliferin, beauvericin, enniatins, and moniliformin-a review. Crit. Rev. Food Sci. Nutr. 2008, 48, 21–49. [Google Scholar] [CrossRef]
- Krogh, P. Role of chratoxin in disease causation. Food Chem. Toxicol. 1992, 30, 213–224. [Google Scholar] [CrossRef]
- Zhang, H.H.; Wang, Y.; Zhao, C.; Wang, J.; Zhang, X.L. Biodegradation of ochratoxin A by Alcaligenes faecalis isolated from soil. J. Appl. Microbiol. 2017, 123, 661–668. [Google Scholar] [CrossRef]
- Rodriguez, H.; Reveron, I.; Doria, F.; Costantini, A.; De Las Rivas, B.; Munoz, R.; Garcia-Moruno, E. Degradation of ochratoxin A by Brevibacterium species. J. Agric. Food Chem. 2011, 59, 10755–10760. [Google Scholar] [CrossRef]
- Cho, S.M.; Jeong, S.E.; Lee, K.R.; Sudhani, H.P.K.; Kim, M.; Hong, S.-Y.; Chung, S.H. Biodegradation of ochratoxin A by Aspergillus tubingensis isolated form meju. J. Microbiol. Biotechnol. 2016, 26, 1687–1695. [Google Scholar] [CrossRef]
- Glaser, N.; Stopper, H. Paulin: Mechanism of genotoxicity. Food Chem. Toxicol. 2012, 50, 1796–1801. [Google Scholar] [CrossRef]
- Sajid, M.; Mehmood, S.; Yuan, Y.; Yue, T. Mycotoxin patulin in food matrices: Occurrence and its biological degradation strategies. Drug Metab. Rev. 2019, 51, 105–120. [Google Scholar] [CrossRef]
- Saleh, I.; Goktepe, I. The characteristics, occurrence, and toxicological effects of patulin. Food Chem. Toxicol. 2019, 129, 301–311. [Google Scholar] [CrossRef]
- Diao, E.; Hou, H.; Hu, W.; Dong, H.; Li, X. Removing and detoxifying methods of patulin: A review. Trends Food Sci. Technol. 2018, 81, 139–145. [Google Scholar] [CrossRef]
- Zheng, X.; Li, Y.; Zhang, H.; Apaliya, M.T.; Zhang, X.; Zhao, L.; Jiang, Z.; Yang, Q.; Gu, X. Identification and toxicological analysis of products of patulin degradation by Pichia caribbica. Biol. Control 2018, 123, 127–136. [Google Scholar] [CrossRef]
- Li, X.; Tang, H.; Yang, C.; Meng, X.; Liu, B. Detoxification of mycotoxin patulin by the yeast Rhodotorula mucilaginosa. Food Control 2019, 96, 47–52. [Google Scholar] [CrossRef]
- Moss, M.; Long, M. Fate of patulin in the presence of the yeast Saccharomyces cerevisiae. Food Addit. Contam. 2002, 19, 387–399. [Google Scholar] [CrossRef]
- Ianiri, G.; Idnurm, A.; Wright, S.A.; Duran-Patron, R.; Mannina, L.; Ferracane, R.; Ritieni, A.; Castoria, R. Searching for genes responsible for patulin degradation in abiocontrol yeast provides insight into the basis for resistance to this mycotoxin. Appl. Environ. Microbiol. 2013, 79, 3101–3115. [Google Scholar] [CrossRef]
- Sudakin, D.L. Trichoghecenes in the environment: Relevance to human health. Toxicol. Lett. 2003, 143, 97–107. [Google Scholar] [CrossRef]
- Kimura, M.; Tokai, T.; Takahashi-Ando, N.; Ohsato, S.; Fujimura, M. Molecular and genetic studies of Fusarium trichothecene biosynthesis: Pathways, genes, and evolution. Biosci. Biotechnol. Biochem. 2007, 71, 2105–2123. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Z.; Beier, R.C.; Shen, J.Z.; DeSmet, D.; De Saeger, S.; Zhang, S. T-2 toxin, a trichothecene mycotoxin: Review of toxicity, metabolism, and analytical methods. J. Agric. Food Chem. 2011, 59, 3441–3453. [Google Scholar] [CrossRef]
- Wu, Q.; Dohnal, V.; Kuca, K.; Yuan, Z. Trichothecenes: Structure-toxic activity relationships. Curr. Drug Metab. 2013, 14, 641–660. [Google Scholar] [CrossRef]
- Payros, D.; Alassane-Kpembi, I.; Pierron, A.; Loiseau, N.; Pinton, P.; Oswald, I.P. Toxicology of deoxynivalenol and its acetylated and modified forms. Arch. Toxicol. 2016, 90, 2931–2957. [Google Scholar] [CrossRef]
- Karlovsky, P. Biological detoxification of the mycotoxin deoxynivalenol and its use in genetically engineered crops and feed additives. Appl. Microbiol. Biotechnol. 2011, 91, 491–504. [Google Scholar] [CrossRef] [PubMed]
- Pestka, J.J. Deoxynivalenol: Mechanisms of action, human exposure, and toxicological relevance. Arch. Toxicol. 2010, 84, 663–679. [Google Scholar] [CrossRef]
- Baldwin, N.C.P.; Bycroft, B.W.; Dewick, P.M. Metabolic conversions of trichothecene mycotoxins: Biotransformation of 3-acetyldeoynivalenol into fusarenon-X. Z. Naturforsch. C 1986, 41, 845–850. [Google Scholar] [CrossRef] [PubMed]
- Islam, R.; Zhou, T.; Young, J.C.; Goodwin, P.H.; Pauls, K.P. Aerobic and anaerobic de-epoxidation of mycotoxin deoxynivalenol by bacteria originating from agricultural soil. World J. Microbiol. Biotechnol. 2012, 28, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Turner, P.C.; Hopton, R.P.; White, K.L.M.; Fisher, J.; Cade, J.E.; Wild, C.P. Assessment of deoxynivalenol metabolite profiles in UK adults. Food Chem. Toxicol. 2011, 49, 132–135. [Google Scholar] [CrossRef] [PubMed]
- Zinedine, A.; Soriano, J.M.; Molto, J.C.; Mañes, J. Review on the toxicity, occurrence, metabolism, detoxification, regulations and intake of zearalenone: An oestrogenic mycotoxin. Food Chem. Toxicol. 2007, 45, 1–18. [Google Scholar] [CrossRef]
- Dai, M.; Jiang, S.; Yuan, X.; Yang, W.; Yang, Z.; Huang, L. Effects of zearalenone-diet on expression of ghrelin and PCNA genes in ovaries of post-weaning piglets. Anim. Reprod. Sci. 2016, 168, 126–137. [Google Scholar] [CrossRef]
- Lee, J.T.; Jessen, K.A.; Beltran, R.; Starkl, V.; Schatzmayr, G.; Borutova, R.; Caldwell, D.J. Effects of mycotoxin-contaminated diets and deactivating compound in laying hens: 2. Effects on white shell egg quality and characteristics. Poult. Sci. 2012, 91, 2096–2104. [Google Scholar] [CrossRef]
- Masching, S.; Naehrer, K.; Schwartz-Zimmermann, H.E.; Sarandan, M.; Schaumberger, S.; Dohnal, I.; Nagl, V.; Schatzmayr, D. Gastrointestinal degradation of fumonisin B1 by carboxylesterase fumD prevents fumonisin induced alteration of sphingolipid metabolism in turkey and swine. Toxins 2016, 8, 84. [Google Scholar] [CrossRef]
- Karabulut, S.; Paytakov, G.; Leszczynski, J. Reduction of alftoxin B1 to afatoxicol: A comprehensive DFT study provides clues to its toxicity. J. Sci. Food Agric. 2014, 94, 3134–3140. [Google Scholar] [CrossRef]
- Hahn, I.; Kunz-Vekiru, E.; Twarzek, M.; Grajewski, J.; Krska, R.; Berthiller, F. Aerobic and anaerobic in vitro testing of feed additives claiming to detoxify deoxynivalenol and zearalenone. Food Addit. Contam. Part A 2015, 32, 922–933. [Google Scholar] [CrossRef] [PubMed]
- Berthiller, F.; Crews, C.; Dall’Asta, C.; Saeger, S.D.; Haesaert, G.; Karlovsky, P.; Oswald, I.P.; Seefelder, W.; Speijiers, G.; Stroka, J. Masked mycotoxins: A review. Mol. Nutr. Food Res. 2013, 57, 165–186. [Google Scholar] [CrossRef] [PubMed]
- Rolli, E.; Righetti, L.; Galaverna, G.; Suman, M.; Dall’Asta, C.; Bruni, R. Zearalenone uptake and biotransformation in micropropgated Triticum durum Desf. plants: A xenobolomic approach. J. Agric. Food Chem. 2018, 66, 1523–1532. [Google Scholar] [CrossRef]
- Meena, M.; Samal, S. Alternaria host-specific (HSTs) toxins: An overview of chemical characterization, target sites, regulation and their toxic effects. Toxicol. Rep. 2019, 6, 745–758. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Wang, W.; Zhang, S. Purification and identification of an aflatoxin B1 degradation enzyme form Pantoea sp. T6. Toxicon 2019, 157, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Alexander, N.J. The TRI101 story: Engineering wheat and barley to resist Fusarium head blight. World Mycotoxin J. 2008, 1, 31–37. [Google Scholar] [CrossRef]
- Igawa, T.; Takahashi-Ando, N.; Ochiai, N.; Ohsato, S.; Shimizu, T.; Kudo, T.; Yamaguchi, I.; Kimura, M. Reduced contamination by the Fusarium mycotoxin zearalenone in maize kernels through genetic modification with a detoxification gene. Appl. Environ. Microbiol. 2007, 73, 1622–1629. [Google Scholar] [CrossRef]
- Xu, L.; Liu, Y.; Zhou, L.; Wu, J. Enhanced beauvericin production with in situ adsorption in mycelial liquid culture of Fusarium redolens Dzf2. Process Biochem. 2009, 44, 1063–1067. [Google Scholar] [CrossRef]
- Xu, L.; Wang, J.; Zhao, J.; Li, P.; Shan, T.; Wang, J.; Li, X.; Zhou, L. Beauvericin from the endophytic fungus, Fusarium redolens, isolated from Dioscorea zingiberensis and its antibacterial activity. Nat. Prod. Commun. 2010, 5, 811–814. [Google Scholar] [CrossRef]
- Firakova, S.; Proksa, B.; Sturdikova, M. Biosynthesis and biological activity of enniatins. Pharmazie 2007, 62, 563–568. [Google Scholar]
- Wang, X.; Wang, J.; Lai, D.; Wang, W.; Dai, J.; Zhou, L.; Liu, Y. Ustiloxin G, a new cyclopeptide mycotoxin from rice false smut balls. Toxins 2017, 9, 54. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Sun, W.; Meng, J.; Wang, A.; Wang, X.; Tian, J.; Fu, X.; Dai, J.; Liu, Y.; Lai, D.; et al. Bioactive bis-naphtho-γ-pyrones from rice false smut pathogen Ustilaginoidea virens. J. Agric. Food Chem. 2015, 63, 3501–3508. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Wang, A.; Xu, D.; Wang, W.; Meng, J.; Dai, J.; Liu, Y.; Lai, D.; Zhou, L. New ustilaginoidins from rice false smut balls caused by Villosiclava virens and their phytotoxic and cytotoxic activities. J. Agric. Food Chem. 2017, 65, 5151–5160. [Google Scholar] [CrossRef] [PubMed]
- Lai, D.; Meng, J.; Xu, D.; Zhang, X.; Liang, Y.; Han, Y.; Jiang, C.; Liu, H.; Wang, C.; Zhou, L.; et al. Determination of the absolute configurations of the stereogenic centers of ustilaginoidins by studying the biosynthetic monomers from a gene knockout mutant of Villosiclava virens. Sci. Rep. 2019, 9, 1855. [Google Scholar] [CrossRef]
- Meng, J.; Zhao, S.; Dang, P.; Zhou, Z.; Lai, D.; Zhou, L. Ustilaginoidin M1, a new bis-naphtho-γ-pyrone from the fungus Villosiclava virens. Nat. Prod. Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Lai, D.; Meng, J.; Zhang, X.; Xu, D.; Dai, J.; Zhou, L. Ustilobisorbicillinol A, a cytotoxic sorbyl-containing aromatic polyketide from Ustilaginoidea virens. Org. Lett. 2019, 21, 1311–1314. [Google Scholar] [CrossRef]
- Meng, J.; Gu, G.; Dang, P.; Zhang, X.; Wang, W.; Dai, J.; Liu, Y.; Lai, D.; Zhou, L. Sorbicillinoids from the fungus Ustilaginoidea virens and their phytotoxic, cytotoxic, and antimicrobial activities. Front. Chem. 2019, 7, 435. [Google Scholar] [CrossRef]
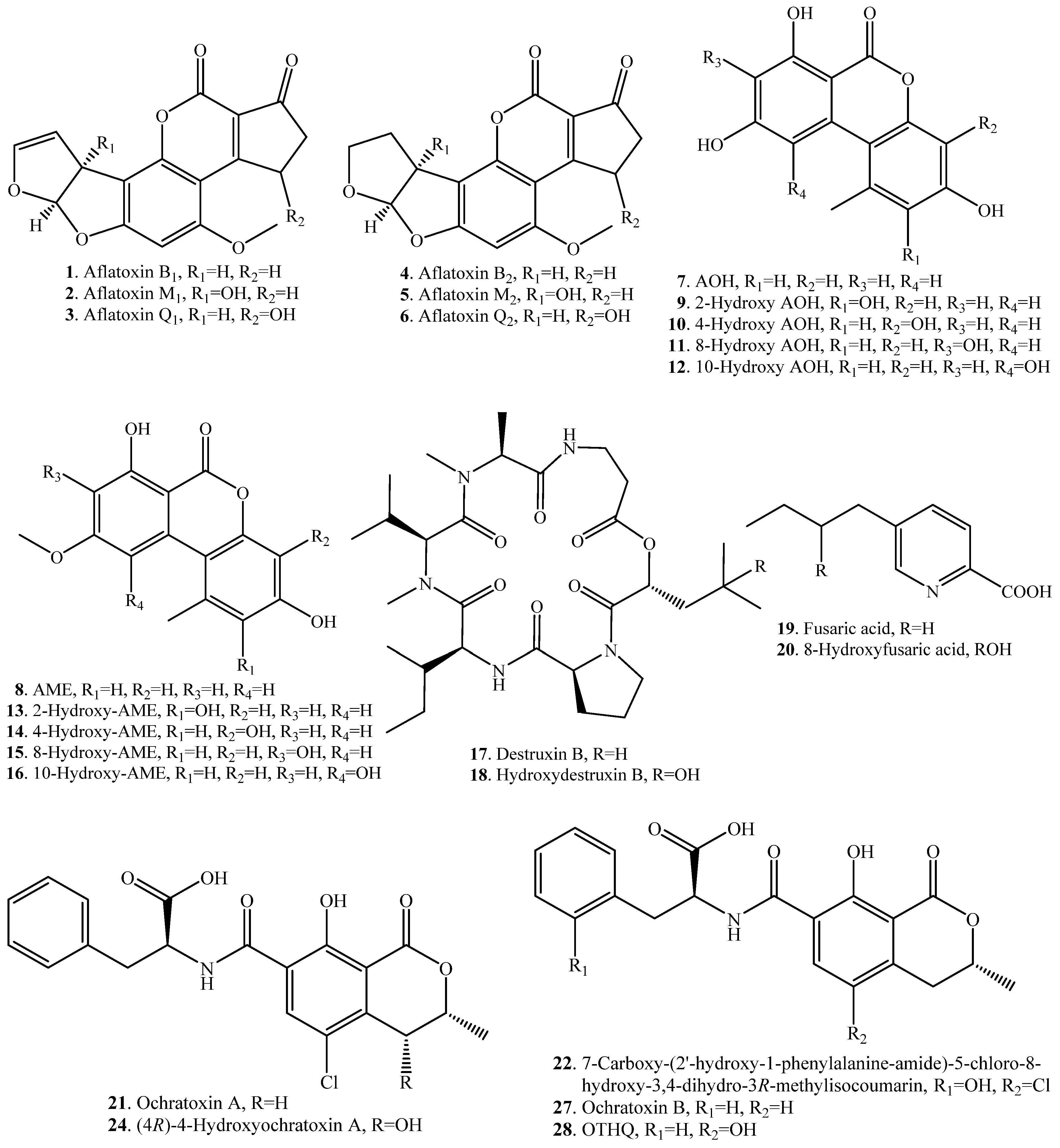
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
| Aflatoxin B1 (AFB1, 1) | Aflatoxins M1 (2) and Q1 (3) | Hepatic microsomal mixed-function oxidase of rhesus monkey | [22] |
| Aflatoxin M1 (2) | Channel catfish liver | [20] | |
| Aflatoxin Q1 (3) | Rat liver microsomal cytochrome P450p | [21] | |
| Aflatoxin B2 (AFB2, 4) | Aflatoxins M2 (5) and Q2 (6) | Animal liver microsomes | [14] |
| Alternariol (AOH, 7) | 2-Hydroxy AOH (9) | Microsomes from rat, human and porcine liver | [23] |
| 4-Hydroxy AOH (10) | Microsomes from rat, human and porcine liver | [23] | |
| 8-Hydroxy AOH (11) | Microsomes from rat, human and porcine liver | [23] | |
| 10-Hydroxy AOH (12) | Microsomes from rat, human and porcine liver | [23] | |
| Alternariol 9-O-methyl ether (AME, 8) | 2-Hydroxy AME (13) | Microsomes from rat, human and porcine liver | [23] |
| 4-Hydroxy AME (14) | Microsomes from rat, human and porcine liver | [23] | |
| 8-Hydroxy AME (15) | Microsomes from rat, human and porcine liver | [23] | |
| 10-Hydroxy AME (16) | Microsomes from rat, human and porcine liver | [23] | |
| Destruxin B (17) | Hydroxydestruxin B (18) | Crucifers such as Brassica napus | [24] |
| Fusaric acid (19) | 8-Hydroxyfusaric acid (20) | Mucor rouxii (fungus) | [27] |
| Ochratoxin A(OTA, 21) | 7-Carboxy-(2′-hydroxy-1- phenylalanine-amide)- 5-chloro-8-hydroxy-3,4-dihydro-3R- methylisocoumarin (22) | Phenylobacterium immobile (bacterium) | [28] |
| Dihydrodiol derivative of ochratoxin A (23) | Phenylobacterium immobile (bacterium) | [28] | |
| (4R)-4-Hydroxyochratoxin A (24) | Rat liver microsomes | [29] | |
| Cell cultures of wheat and maize | [29] | ||
| Rabbit liver microsomes | [30] | ||
| (4S)-4-Hydroxyochratoxin A (25) | Cell cultures of wheat and maize | [31] | |
| Rabbit liver microsomes | [30] | ||
| 10-Hydroxyochratoxin A (26) | Rabbit liver microsomes | [30] | |
| Ochratoxin B (OTB, 27) | Hydroquinone metabolite of ochratoxin (OTHQ, 28) | Horse radish peroxidase (HPR) | [32] |
| Sterigmatocystin (29) | 9-Hydroxy sterigmatocystin (30) | Human and rat hepatic microsomes | [34] |
| T-2 toxin (31) | 19-OH T-2 toxin = 3‘-OH T-2 toxin (32) | Chicken CYP3A37 (enzyme) | [35] |
| Zearalenone (ZEN, 33) | (5S)-5-Hydroxy ZEN (34) | Cunninghamella bainieri (fungus) | [38] |
| 13-Hydroxy ZEN (35) | Human liver microsomes | [40] | |
| 15-Hydroxy ZEN (36) | Human liver microsomes | [40] |
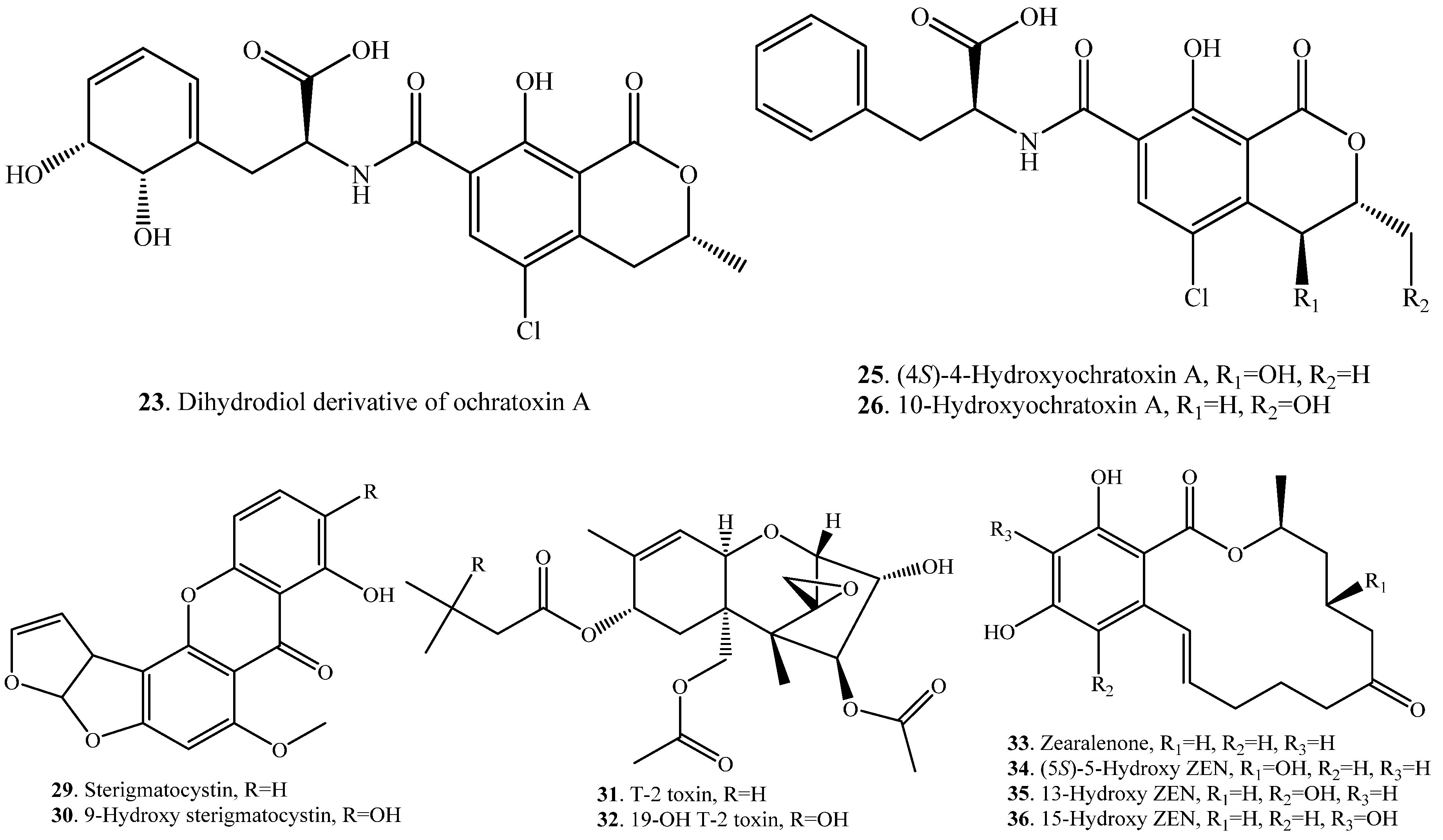
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
| Aflatoxin B1 (AFB1, 1) | Aflatoxicol (37) | Fungi Aspergillus niger, Eurotium herbariorum, Rhizopus sp. | [41] |
| Deoxynivalenol (DON, 38) | 3-Keto-DON (39) | Devosia mutans (bacterium) | [42] |
| Fomannoxin (40) | Fomannoxin alcohol (41) | Pinus sylvestris cell cultures | [44] |
| Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] | ||
| Zearalenone(ZEN, 33) | α-Zearalenol (42) | Candida tropicalis (fungus) | [45] |
| Fungi: Saccharomyces cerevisae, Torulaspora delbruckii, Zygosaccharomyces rouxii, Pichia fermentans, and several yeast strains of the genera Candida, Hansenula, Brettanomyces, Schizosaccharornyces and Saccharomycopsis | [46] | ||
| Fungi Rhizopus sp. and Aspergillus sp. | [47] | ||
| β-Zearalenol (43) | Candida tropicalis (fungus) | [45] | |
| Fungi: Saccharomyces cerevisae, Torulaspora delbruckii and Zygosaccharomyces rouxii | [46] |
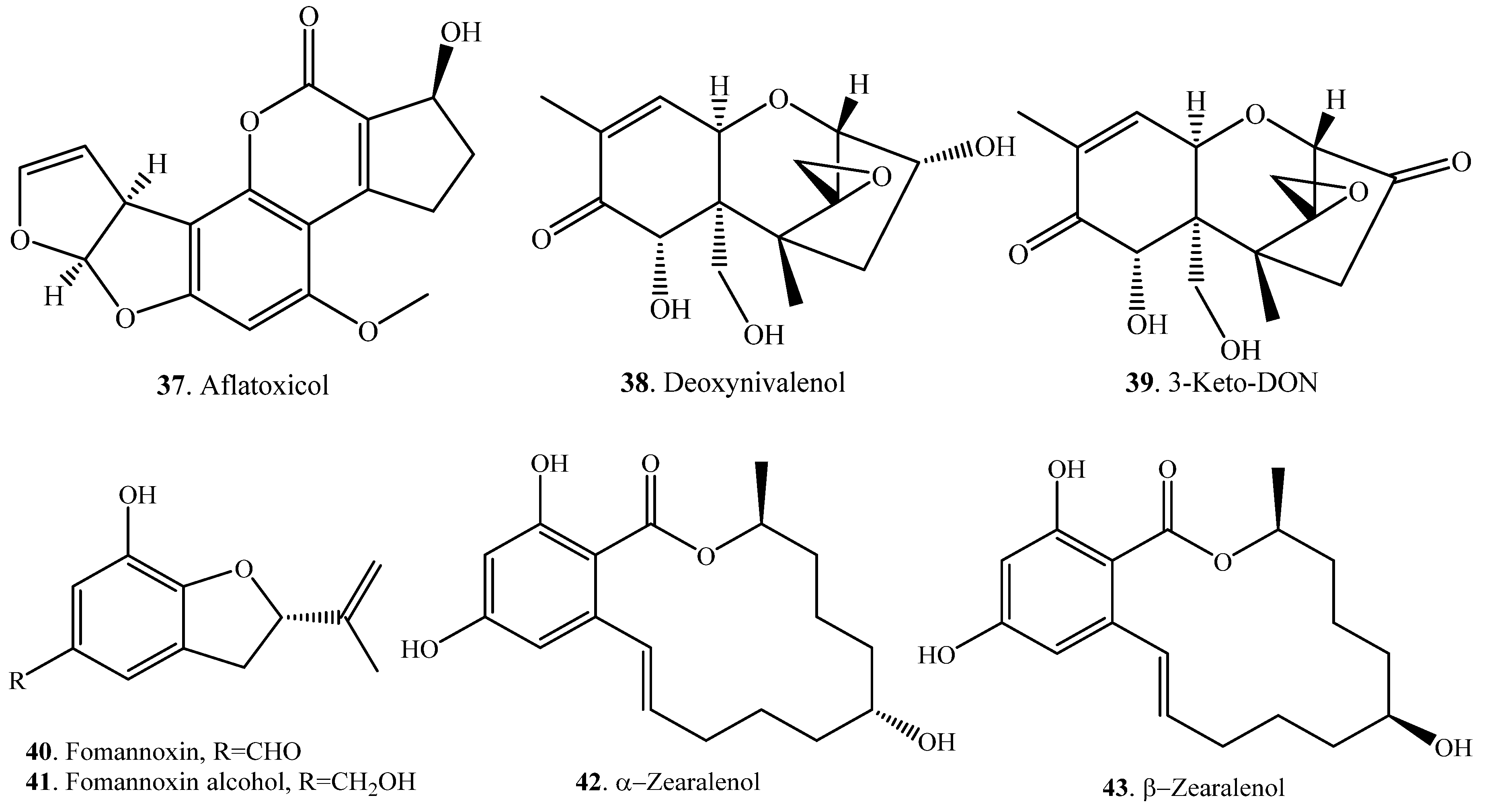
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
| Aflatxin B1 (AFB1, 1) | Aflatoxin B2 (AFB2, 4) | Penicillium raistrickii (fungus) | [14] |
| Zearalenone (ZEN, 33) | Zearalanone (ZAN, 44) | Ovine | [49] |
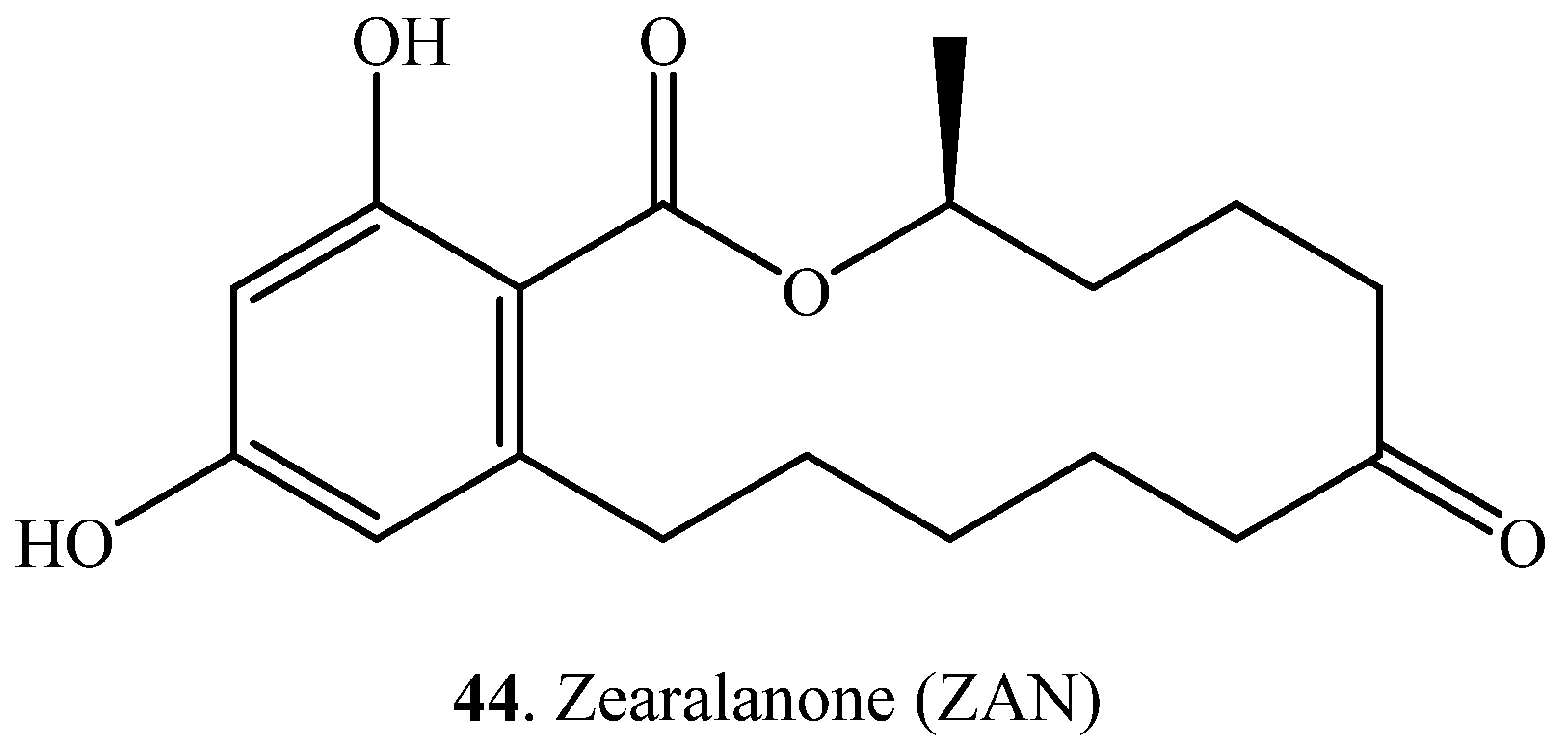
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
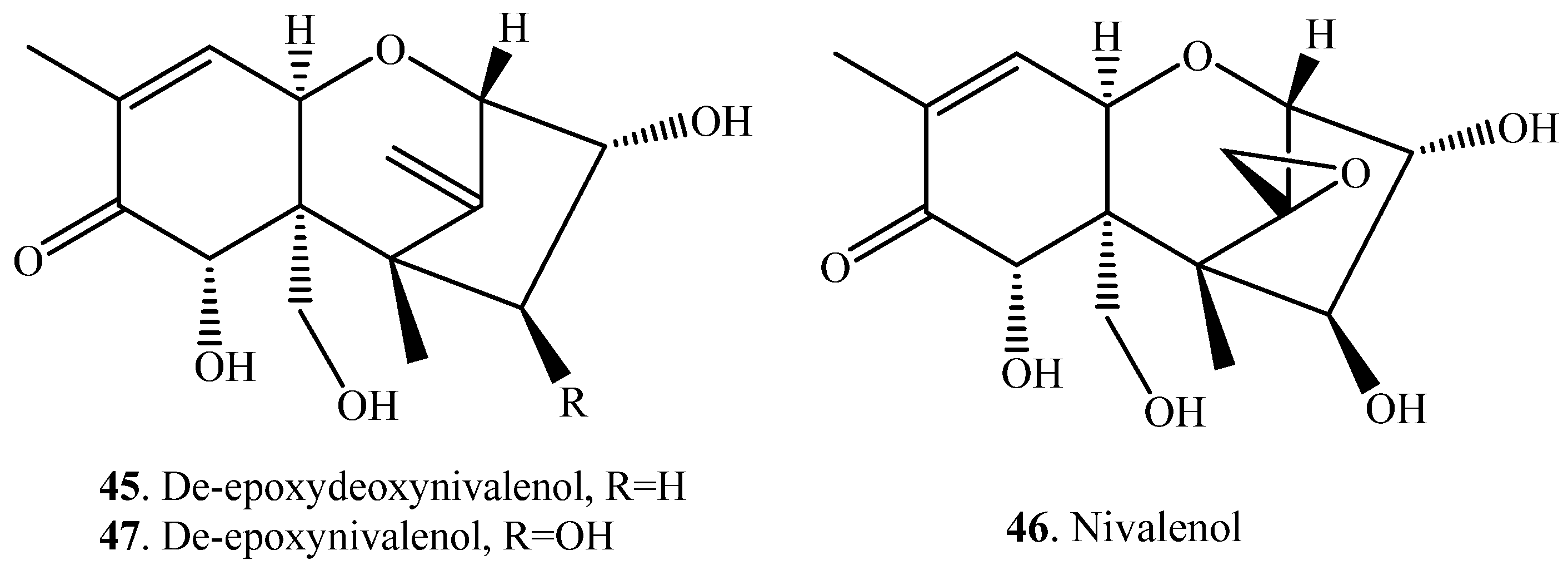
| Deoxynivalenol (DON, 38) | Deepoxydeoxynivalenol (DOM, 45) | Eubacterium sp. DSM 11,798 (bacterium) | [50] |
| Nivalenol (NIV, 46) | De-epoxy NIV (47) | Euacterium sp. BBSH 797 (bacterium) | [51] |
| Wistar rats | [53] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
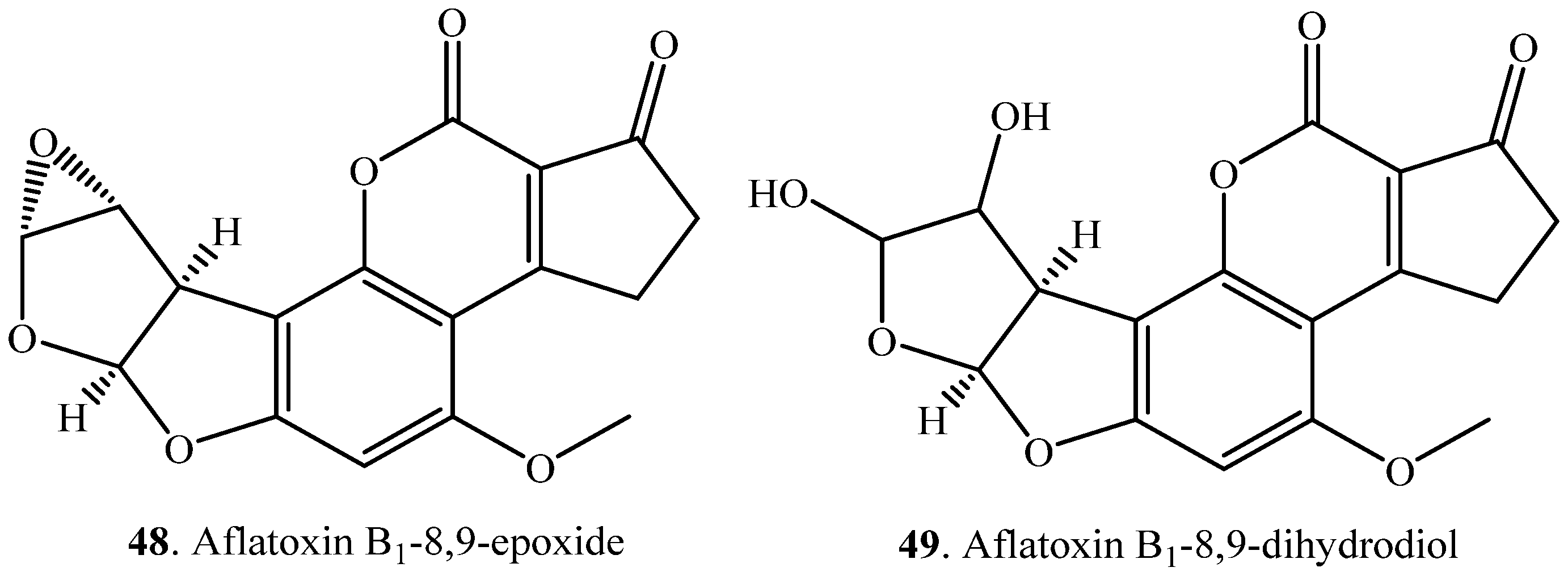
| Aflatoxin B1 (AFB1, 1) | AFB1-8,9-epoxide (48) | Channel catfish liver | [20] |
| AFB1-8,9-dihydrodiol (49) | Phanerochaete sordida YK-624 (fungus) | [54] | |
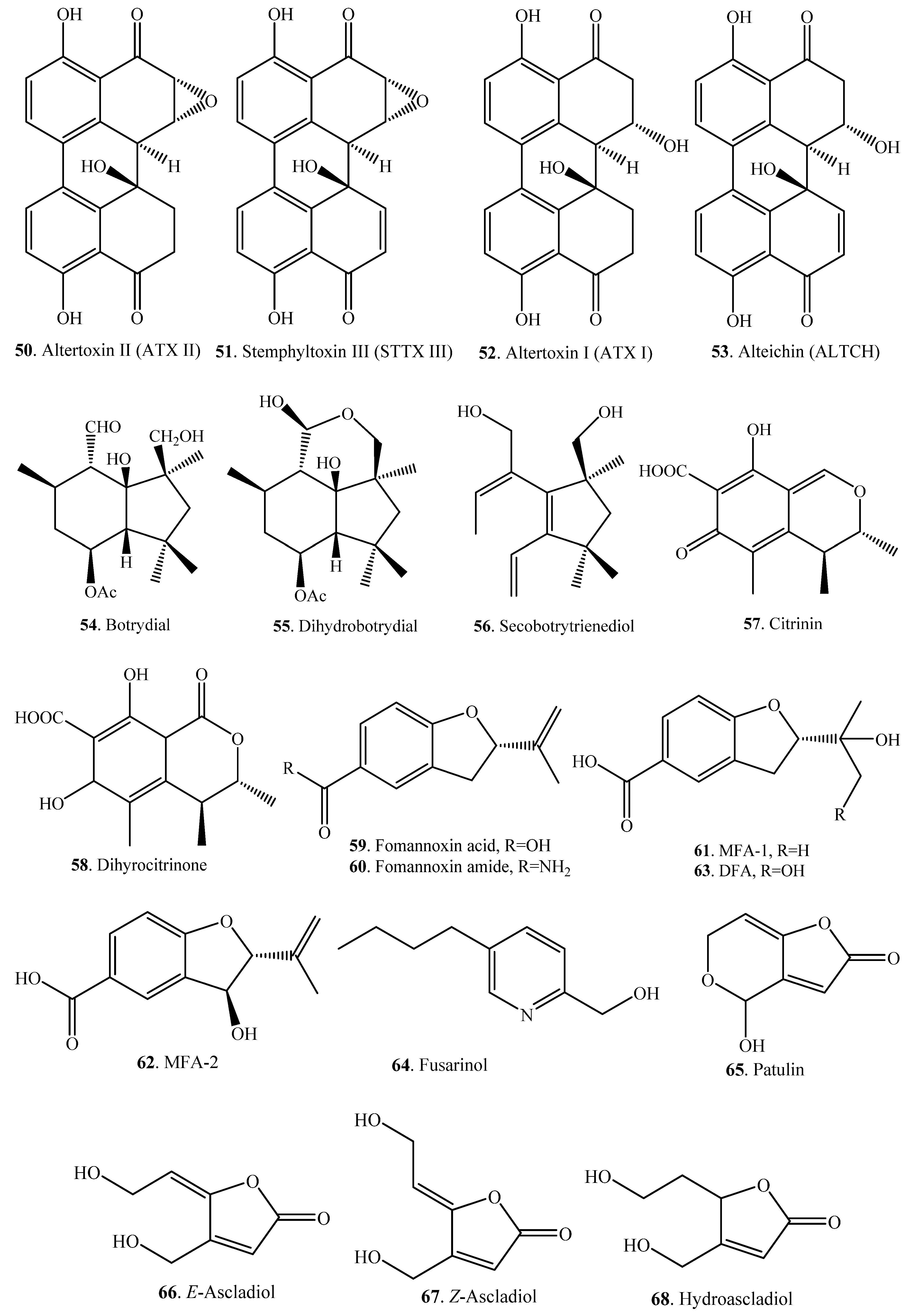
| Altertoxin II (50) | Altertoxin I (52) | Mammalian cell lines Caco-2, HCT 116, HepG2, V79 | [55] |
| Stemphyltoxin III (51) | Alteichin (53) | Mammalian cell line Caco-2 | [55] |
| Botrydial (54) | Dihydrobotrydial (55) | Botrytis cinerea (fungus) | [56] |
| Secobotrytrienediol (56) | Botrytis cinerea (fungus) | [56] | |
| Citrinin (57) | Dihydrocitrinone (58) | Rats and humans | [57] |
| Fomannoxin (40) | Fomannoxin acid (59) | Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] |
| Fomannoxin amide (60) | Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] | |
| MFA-1 (61) | Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] | |
| MFA-2 (62) | Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] | |
| DFA (63) | Rhizosphere-associated bacterium Streptomyces sp. AcH 505 | [48] | |
| Fusaric acid (19) | Fusarinol (64) | Aspergillus tubingensis (fungus) | [59] |
| Patulin (65) | E-Ascladiol (66), Z-ascladiol (67) | Kodameae ohmeri (fungus) | [60] |
| E-Ascladiol (66), Z-ascladiol (67), hydroascladiol (68) | Lactobacillus plantarum (bacterium) | [62] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
| Alternariol (AOH, 7) | Alternariol 9-O-methyl ether (AME, 8) | Methyltransferase | [64] |
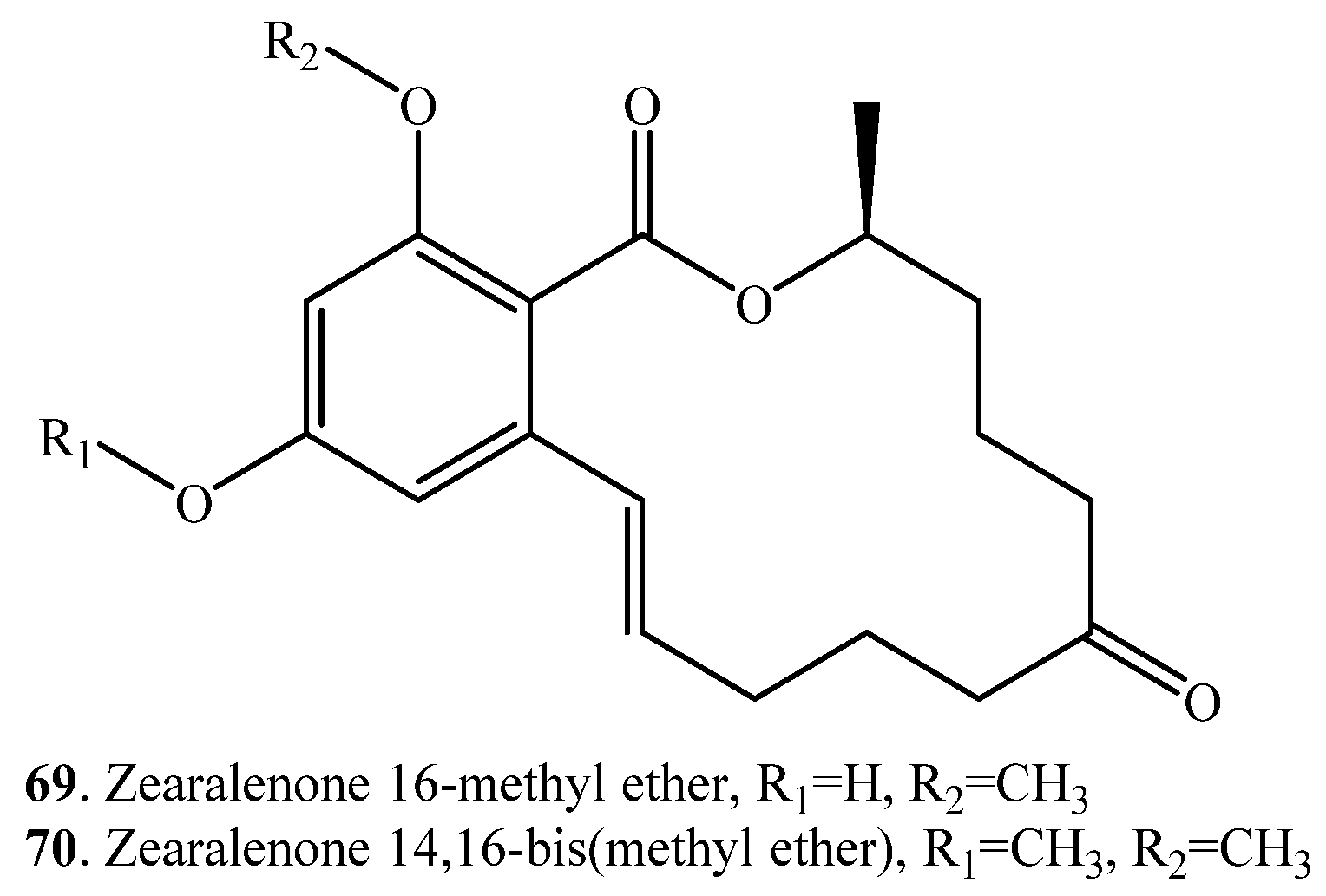
| Zearalenone (ZEN, 33) | Zearalenone 16-methyl ether (69) | Cunninghamella bainieri (fungus) | [38] |
| Zearalenone 14,16-bis (methyl ether) (70) | Cunninghamella bainieri (fungus) | [38] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
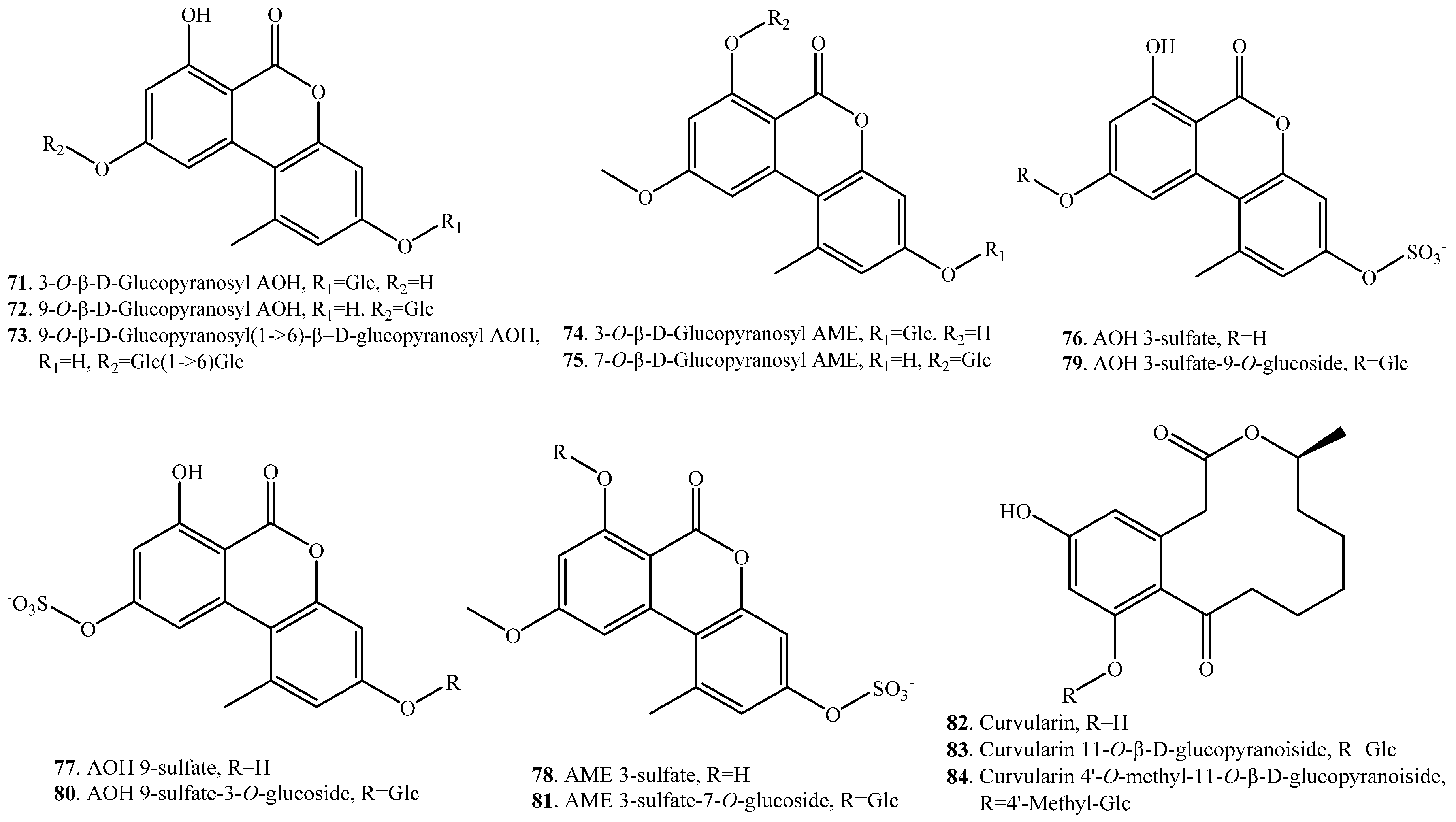
| Alternariol (AOH, 7) | 3-O-β-d-Glucopyranosyl alternariol (71) | Suspension cell cultures of Nicotiana tabacum | [66] |
| 9-O-β-d-Glucopyranosyl alternariol (72) | Suspension cell cultures of Nicotiana tabacum | [66] | |
| 9-O-β-d-Glucopyranosyl (1→6)-β-d-glucopyranosyl alternariol (73) | Suspension cell cultures of Nicotiana tabacum | [66] | |
| Alternariol 9-O-methyl ether (AME, 8) | 3-O-β-d-Glucopyranosyl AME (74) | Suspension cell cultures of Nicotiana tabacum | [66] |
| 7-O-β-d-Glucopyranosyl AME (75) | Suspension cell cultures of Nicotiana tabacum | [66] | |
| AOH 3-sulfate (76) | AOH 3-sulfate 9-O-glucoside (79) | Tomato tissues and cultured tobacco cells | [67] |
| AOH 9-sulfate (77) | AOH 9-sulfate 3-O-glucoside (80) | Tomato tissues and cultured tobacco cells | [67] |
| AME 3-sulfate (78) | AME 3-sulfate 7-O-glucoside (81) | Tomato tissues and cultured tobacco cells | [67] |
| Curvularin (82) | Curvularin 11-O-β-d-glucopyranoside (83) | Beauveria bassiana (fungus) | [69] |
| Curvularin 4‘-O-methyl-11-O-β-d-glucopyranoside (84) | Beauveria bassiana (fungus) | [69] | |
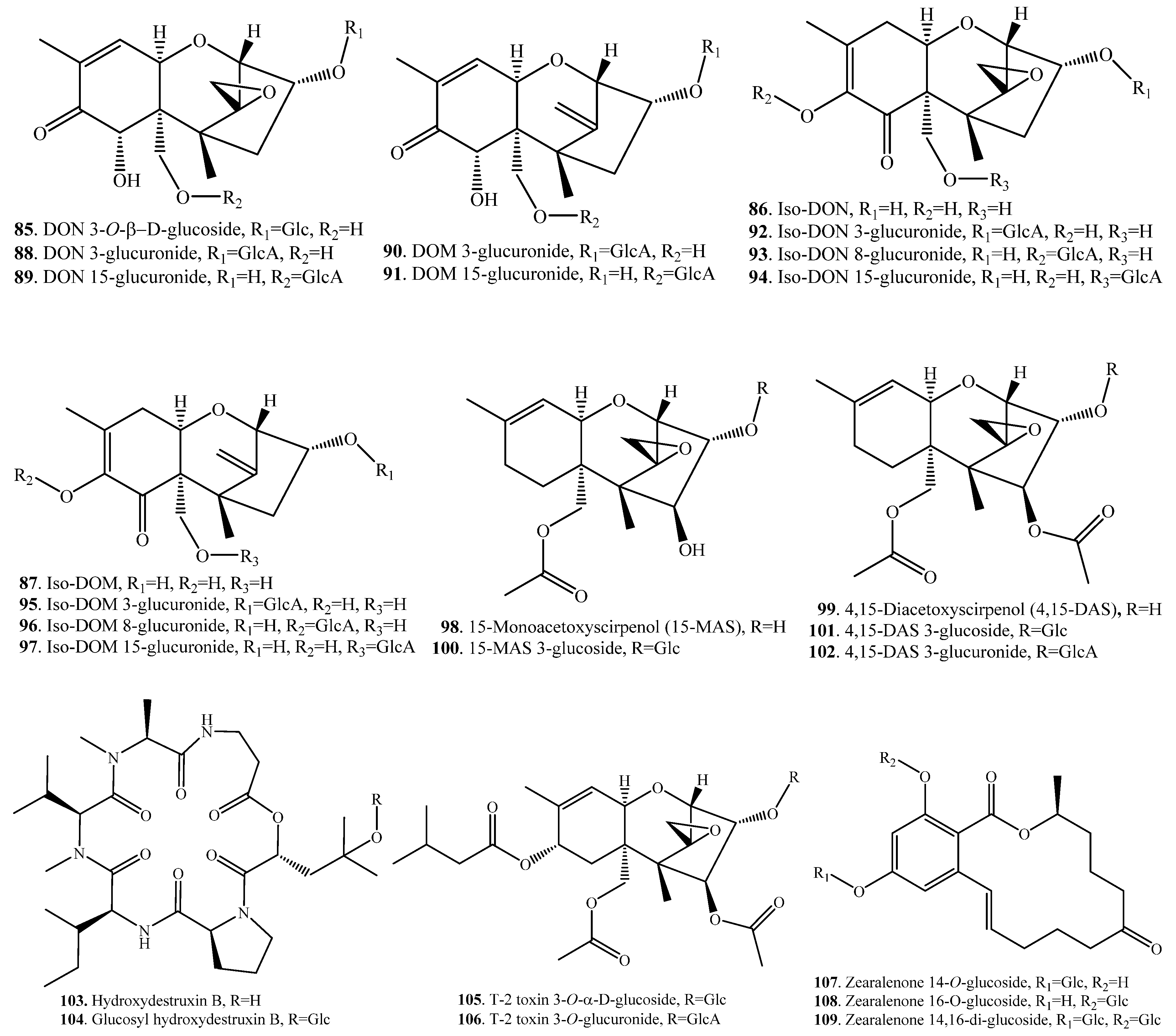
| Deoxynivalenol (DON, 38) | DON 3-O-β-d-glucoside (85) | A combinant UDP-gluosyltransferase from rice | [70] |
| DON-3-GlcA (88) | Rat liver microsomes (RLM), human liver microsomes (HLM) | [71] | |
| DON-15-GlcA (89) | RLM, HLM | [71] | |
| Deepoxy-deoxynivalenol (DOM, 45) | DOM-3-GlcA (90) | RLM, HLM | [71] |
| DOM-15-GlcA (91) | RLM, HLM | [71] | |
| Iso-DON (86) | Iso-DON-3-GlcA (92) | RLM, HLM | [71] |
| Iso-DON-8-GlcA (93) | RLM | [71] | |
| Iso-DON-15-GlcA (94) | RLM, HLM | [71] | |
| Iso-DOM (87) | Iso-DOM-3-GlcA (95) | RLM, HLM | [71] |
| Iso-DOM-8-GlcA (96) | RLM | [71] | |
| Iso-DOM-15-GlcA (97) | RLM, HLM | [71] | |
| 15-Monoacetoxyscirpenol (15-MAS, 98) | 15-MAS 3-glucoside (100) | Corn (Zea Mays) plants | [75] |
| 4,15-Diacetoxyscirpenol (4,15-DAS, 99) | 4,15-DAS 3-glucoside (101) | Corn (Zea Mays) plants | [75] |
| 4,15-DAS 3-Glucuronide (102) | Rats | [78] | |
| Hydroxydestruxin B (103) | Glucosyl hydroxydestruxin B (104) | Crucifers such as Brassica napus | [24] |
| T-2 toxin (31) | T-2 toxin 3-O-α-d-glucoside (105) | Fungi Blastobotrys muscicola, B. robertii | [79] |
| T-2 toxin 3-O-glucuronide (T-2 GlcA, 106) | Rat hepatic microsomes | [80] | |
| Zearalenone (ZEN, 33) | ZEN 14-O-glucoside (107) | Arabidopis UDP-glucosyltransferase | [81] |
| Mucor bainieri (fungus) | [82] | ||
| Thamnidium elegans (fungus) | [82] | ||
| Barley UDP-glucosyltransferase | [83] | ||
| ZEN 16-O-glucoside (108) | Barley UDP-glucosyltransferase | [83] | |
| ZEN 14,16-di-glucoside (109) | Recombinant barley glucosyltransferase | [84] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
| Ochratoxin A (21) | Ochratoxin A methyl ester (110) | Cell cultures of wheat and maize | [31] |
| T-2 toxin (31) | 3-Acetyl T-2 toxin (111) | Bovine rumen fluid in vitro | [85] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
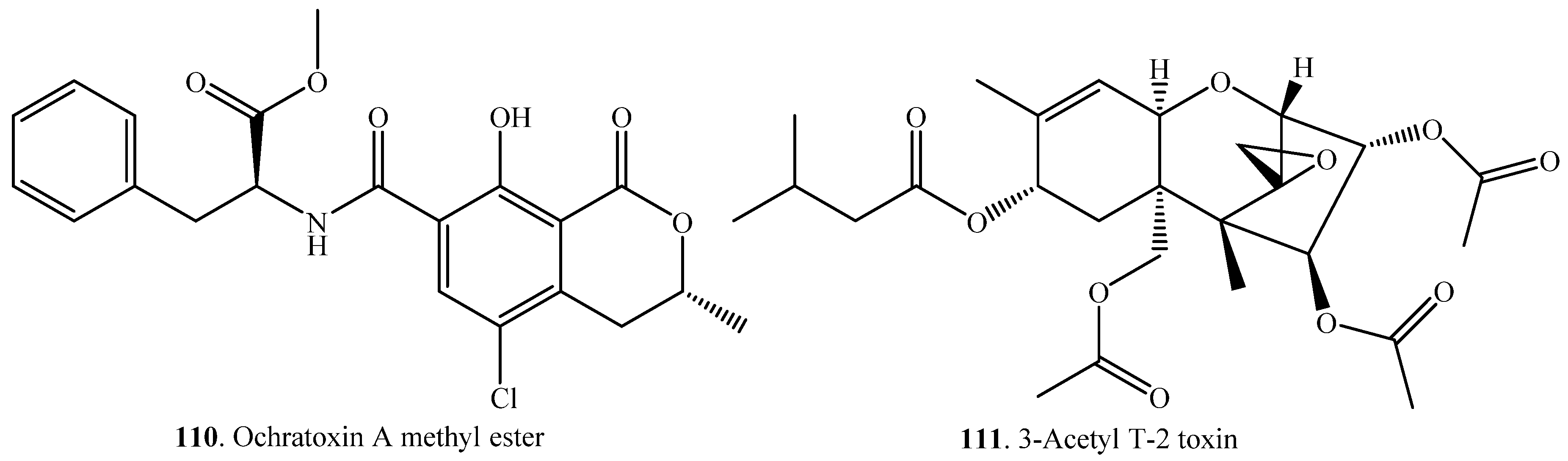
| 3-Acetyl DON (112) | DON (38) | Cell-free extracts of the fungus Fusarium sp. | [88] |
| Sphaerodes mycoparasitica (fungus) | [87] | ||
| 4,15-Diacetoxyscirpenol (4,15-DAS, 99) | 4-Monoacetoxyscirpenol (4-MAS, 113) | Rats | [78] |
| 15-Monoacetoxyscirpenol (15-MAS, 98) | Rats | [78] | |
| Scirpentriol (SCP, 114) | Rats | [78] | |
| Fumonisin B1 (115) | Hydrolyzed fumonisin B1 = Aminopentol 1 (AP1, 116) | Exophiala spinifera 2141.10 (fungus) | [89] |
| Hydroxylase from the bacterium Sphingopyxis sp. MTA144 | [90] | ||
| Carboxylesterase FumD | [91] | ||
| Fusarenon-X (FX, (117) | Nivalenol (NIV, 46) | Mice | [92] |
| Goat (Capra hircus) | [93] | ||
| Ochratoxin A (OTA, 21) | Ochratoxin α (118) and L-β-phenylalanine (119) | Crude lipase from Aspergillus niger | [94] |
| Protease A prolyve PAC and pancreatin | [95] | ||
| Carboxypeptidase A | [96] | ||
| Fungus: Aspergillus niger (fungus) | [97] | ||
| Bacillus amyloliquefaciens (bacterium) | [98] | ||
| Lactone-opened ochratoxin A (OP-OTA, 120) | Rats | [99] | |
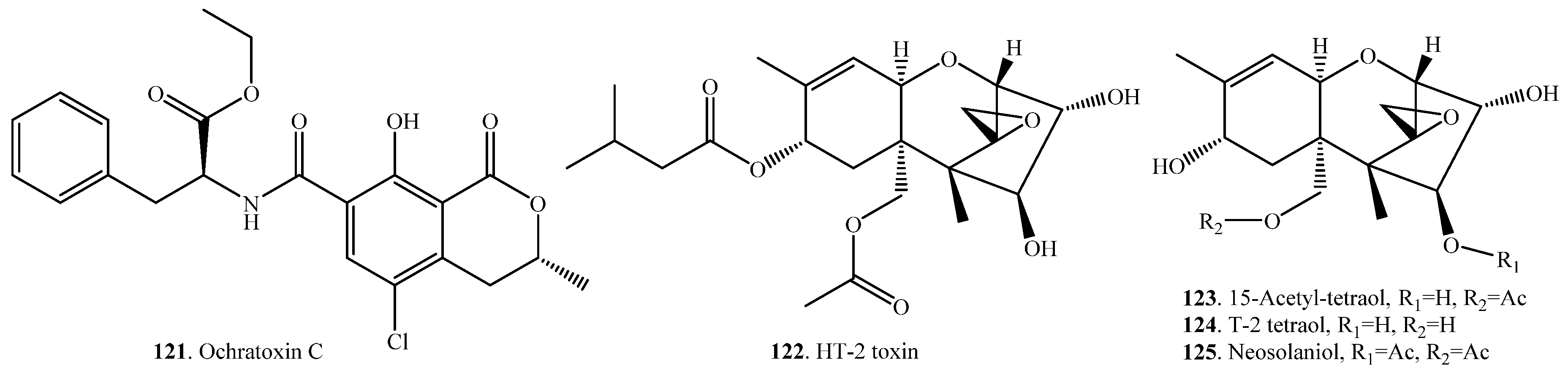
| Ochratoxin C (OTC) = Ochratoxin A ethyl ester (121) | Ochratoxin A (OTA, 21) | Rats | [100] |
| T-2 toxin (31) | HT-2 toxin (122) | Eubacterium BBSH 797 (bacterium) | [86] |
| HT-2 toxin (122),15-acetyl-tetraol (123),T-2 tetraol (124) | Liver and intestines of rats | [102] | |
| Neosolaniol (125) | Blastobotrys capitulate (fungus) | [79] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
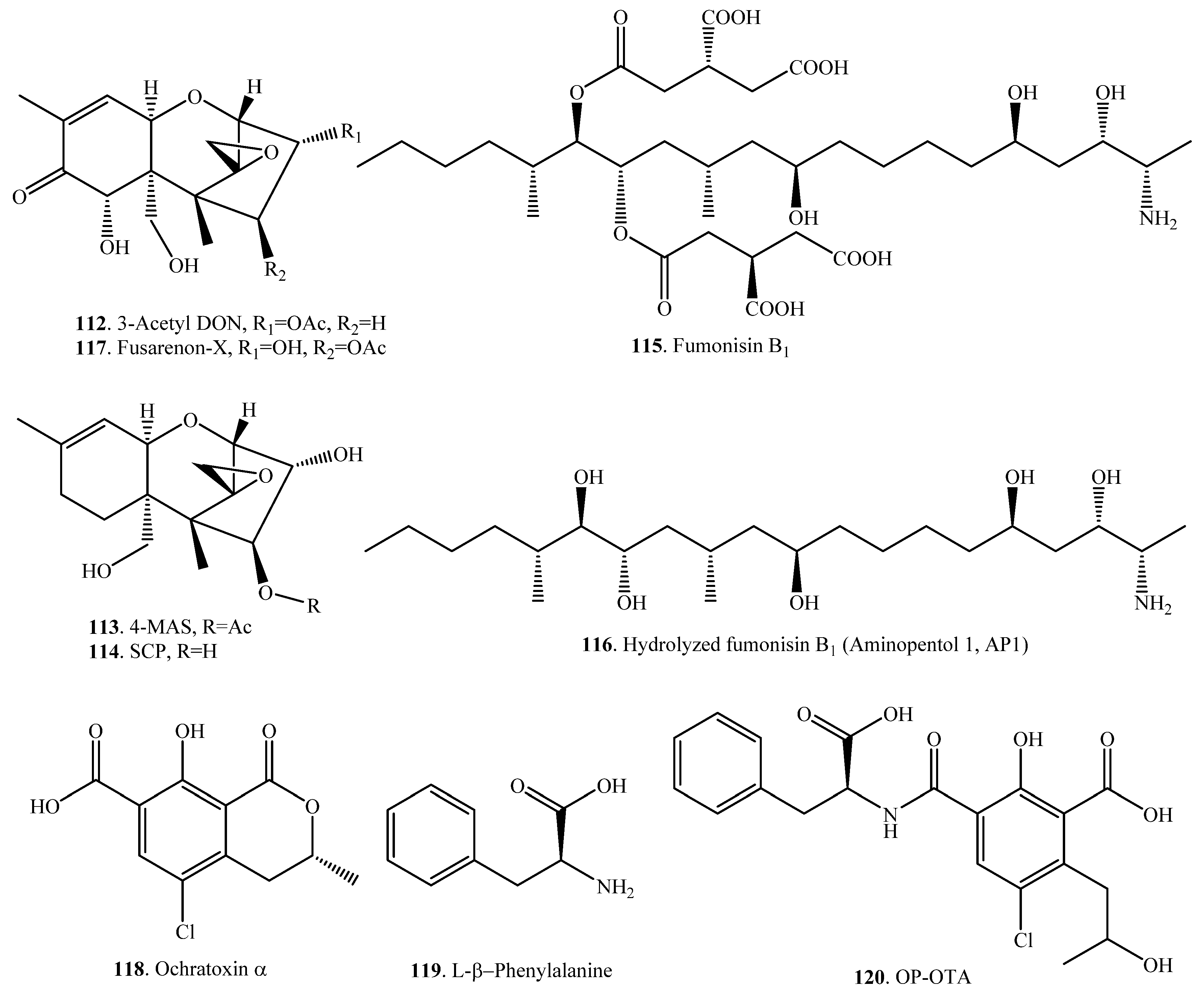
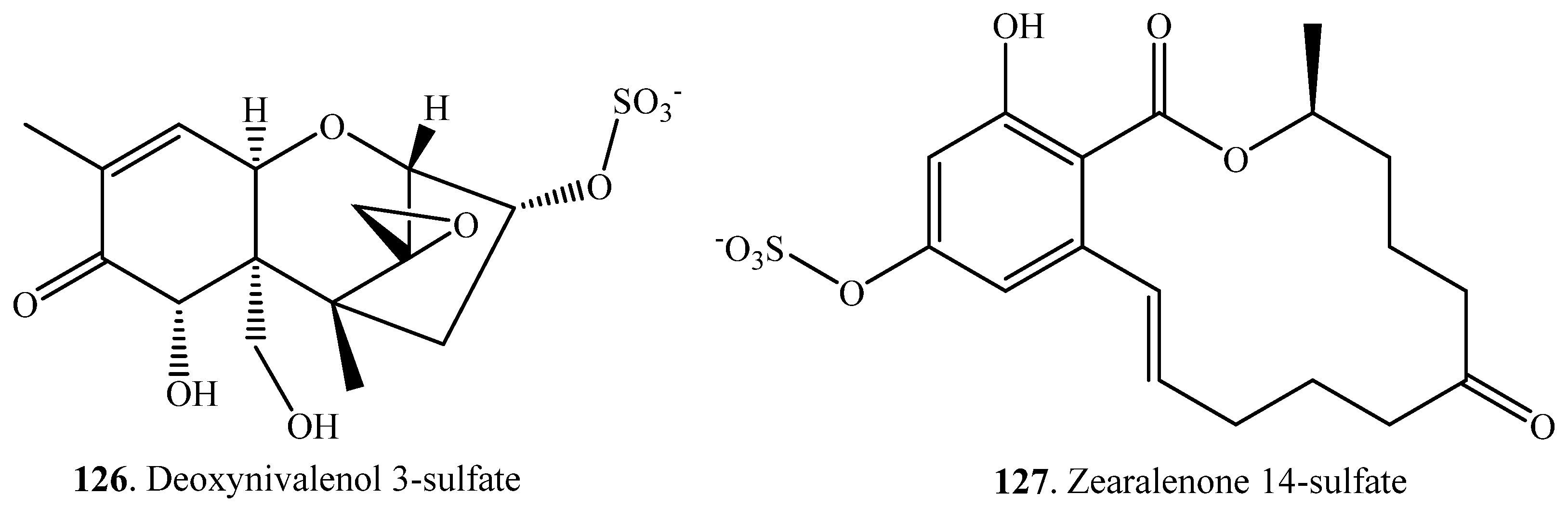
| Deoxynivalenol (38) | Deoxynivlenol 3-sulfate (126) | Sphaerodes mycoparasitica (fungus) | [87] |
| Zearalenone (33) | Zearalenone 14-sulfate (127) | Sphaerodes mycoparasitica (fungus) | [87] |
| Pigs | [103] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
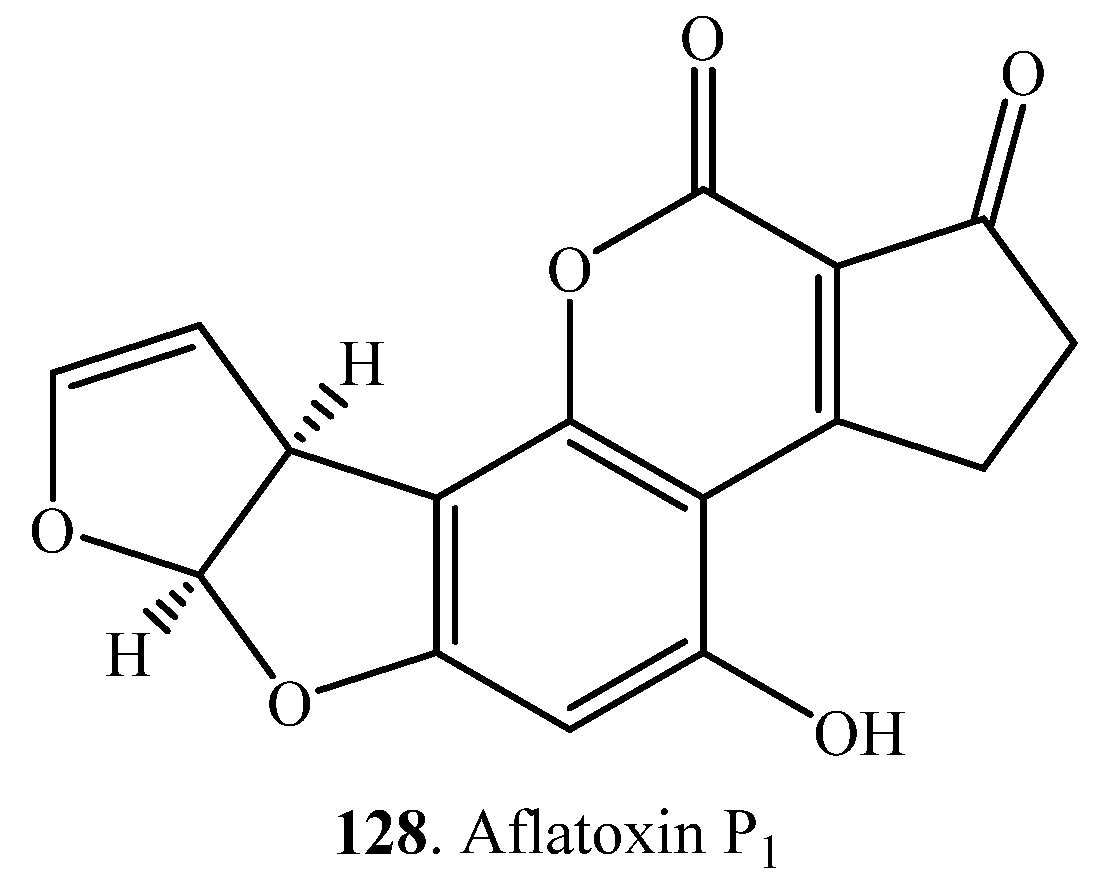
| AME (8) | AOH (7) | Homogenate of porcine liver in the presence of NADPH | [104] |
| AFB1 (1) | Aflatoxin P1 (AFP1, 128) | Enzyme CYP321A1 from Helicoverpa zea | [105] |
| Substrate | Product | Biotransformation System | Ref. |
|---|---|---|---|
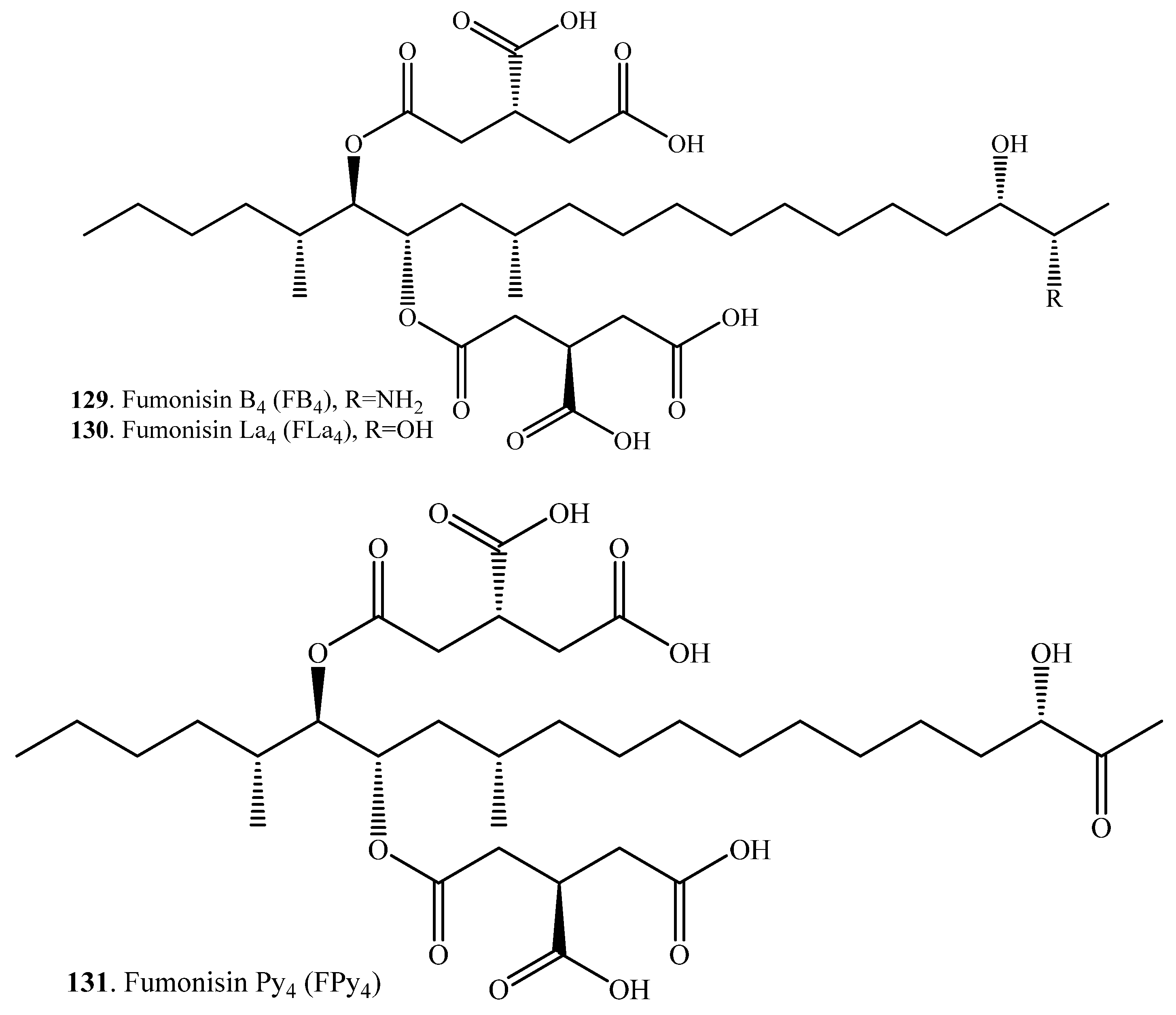
| Fumonisin B4 (129) | Fumonisin La4 (FLa4, 130) | Aspergillus sp. (fungus) | [106] |
| Fumonisin Py4 (FPy4, 131) | Aspergillus sp. (fungus) | [106] | |
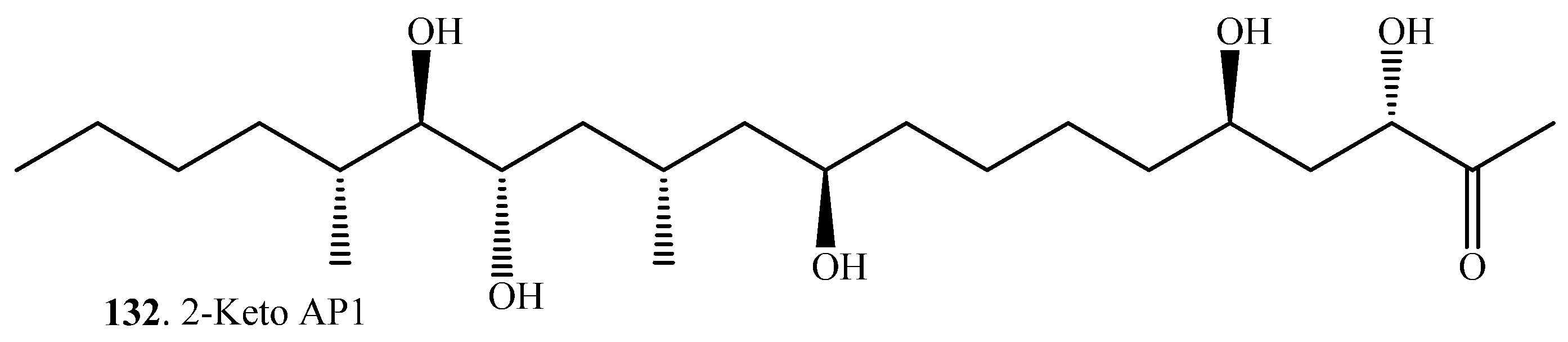
| Hydrolyzed fumonisin B1 = Aminopentol 1 (AP1, 116) | 2-Keto HFB1 = 2-keto AP1 (132) | Exophiala spinifera (fungus) | [107] |
| Substrate | Product | Type | Biotransformation System | Ref. |
|---|---|---|---|---|
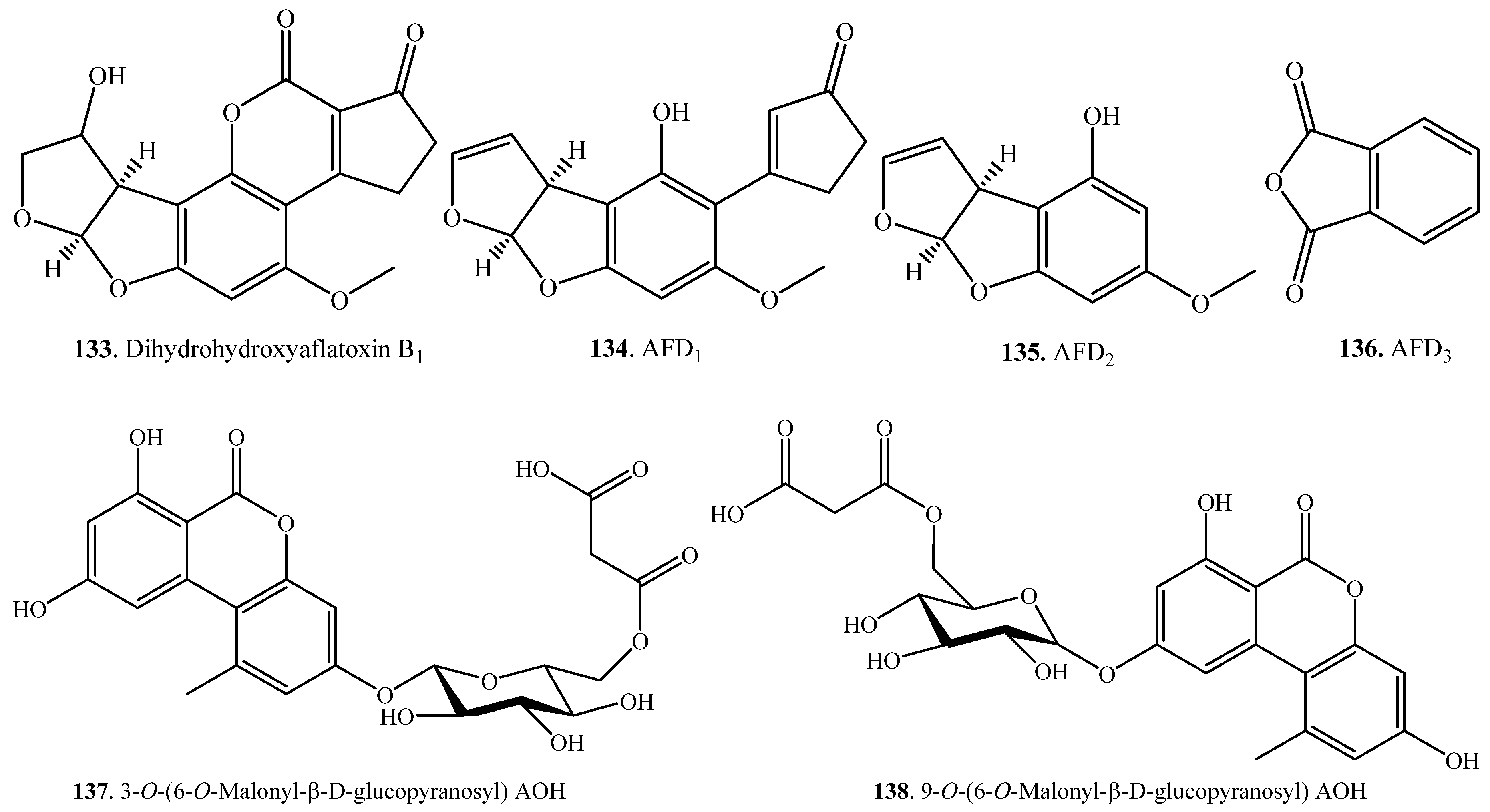
| Aflatoxin B1 (AFB1, 1) | Dihydrohydroxyalfatoxin B1 (AFB2a, 133) | Reduction and oxidation | Pleurotus ostreatus (fungus) | [86] |
| AFD1 (134), AFD2 (135), and AFD3 (136) | Hydolysis, decarboxylation, oxidation-reduction | Pseudomonas putida (bacterium) | [110] | |
| Alternariol (AOH, 7) | 3-O-(6-O-Malonyl-β-d-glucopyranosyl) AOH (137) | Glycosylation and Esterification | Suspension cell cultures of Nicotiana tabacum | [66] |
| 9-O-(6-O-Malonyl-β-d-glucopyranosyl) AOH (138) | Glycosylation and Esterification | Suspension cell cultures of Nicotiana tabacum | [66] | |
| Alternariol 9-O-methyl ether = AME (8) | 3-O-(4-O-Malonyl-β-d-glucopyranosyl) AME (139) | Glycosylation and Esterification | Suspension cell cultures of Nicotiana tabacum | [66] |
| 3-O-(6-O-Malonyl-β-d-glucopyranosyl) AME (140) | Glycosylation and Esterification | Suspension cell cultures of Nicotiana tabacum | [66] | |
| Botrydial (54) | Botryenedial (141) | Dehydration | Botrytis cinerea (fungus) | [56] |
| Citrinin (57) | Decarboxycitrinin (142) | Decarboxylation | Moraxella sp. MB1 (bacterium) | [111] |
| Deoxynivalenol (DON, 38) | 3-epi-Deoxynivalenol (143) | Epimerization | Nocardioides sp. WSN05-2 (bacterium) | [113] |
| Devosia sp. (bacterium) | [114] | |||
| DON-8,15-hemiketal-8-GlcA (144) | Oxidation and glucuronidation | Rat liver microsomes (RLM) | [71] | |
| Iso-DON (86) | Isomerization | RLM | [71] | |
| Iso-DON-3-GlcA (92) and iso-DON-8-GlcA (93) | Isomerization and glucuronization | RLM | [71] | |
| Deepoxy-deoxynivalenol (DOM, 45) | Iso-DOM (87) | Isomerization | RLM | [71] |
| Iso-DOM-3-GlcA (95) and iso-DOM-8-GlcA (96) | Isomerization and glucuronization | RLM | [71] | |
| Fomannoxin (40) | Fomannoxin acid (59) and fomannoxin acid β-glucoside (145) | Oxidation and glycosylation | Cell cultures of Pinus sylvestris | [44] |
| Fumonisin B1 (115) | Hydrolyzed fumonisin B1 = Aminopentol 1 (AP1, 116) and 2-keto AP1 (132) | Hydolysis and deamination | Recombinant enzymes from the bacterium Sphingopyxis sp. | [87] |
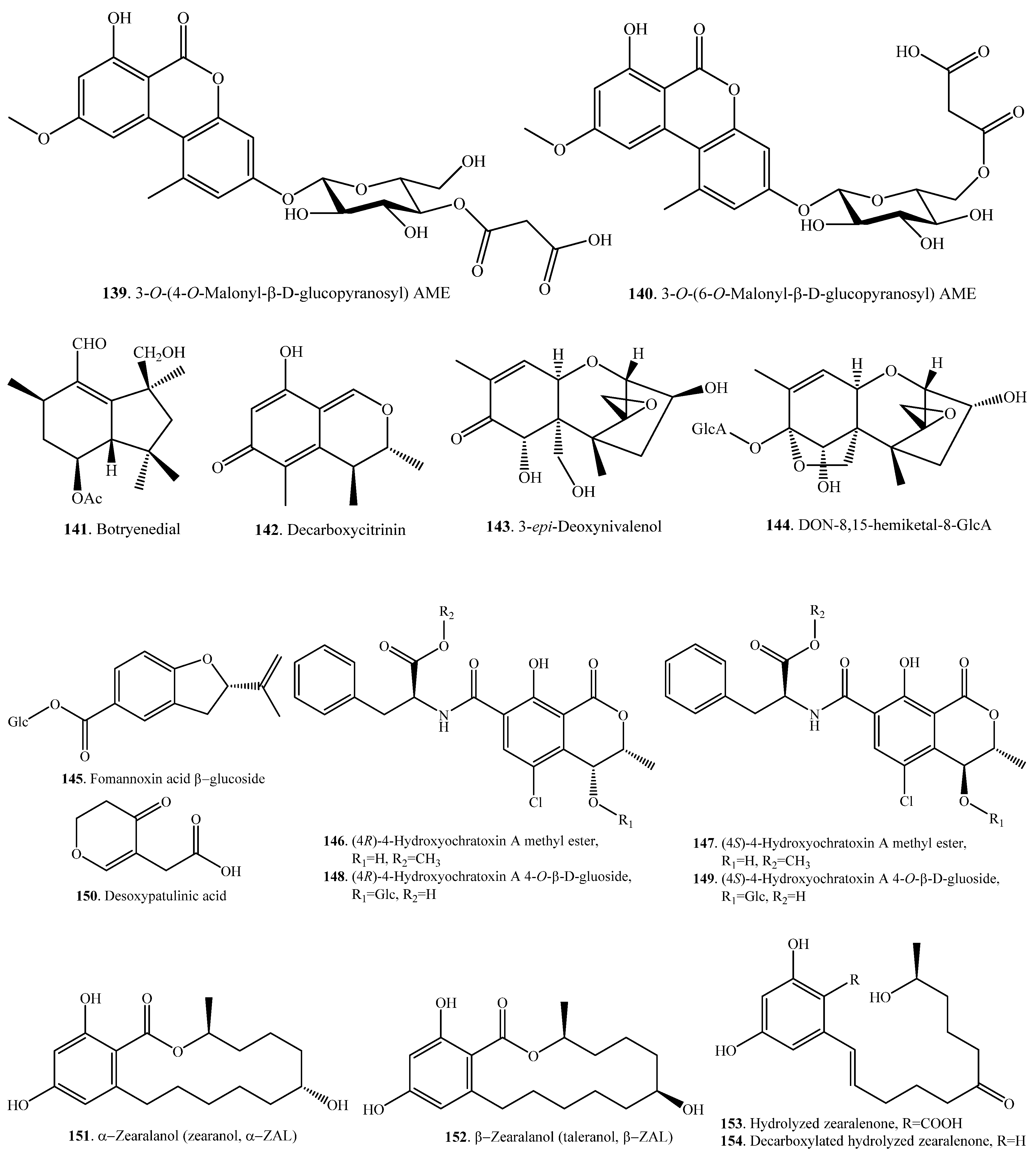
| Ochratoxin A (OTA, 21) | (4R)-4-Hydroxyochratoxin A methyl ester (146) | Hydroxylation and esterification | Cell cultures of wheat and maize | [31] |
| (4S)-4-Hydroxyochratoxin A methyl ester (147) | Hydroxylation and esterification | Cell cultures of wheat and maize | [31] | |
| (4R)-4-Hydroxyochratoxin A 4-O–β-d-glucoside (148) | Hydroxylation and glycosylation | Cell cultures of wheat and maize | [31] | |
| (4S)-4-Hydroxyochratoxin A 4-O-β-d-glucoside (149) | Hydroxylation and glycosylation | Cell cultures of wheat and maize | [31] | |
| Ochratoxin B (OTB, 27) | Dechlorination | Renal microsomes | [117] | |
| Patulin (65) | Desoxypatulinic acid (150) | Hydolysis, reduction and dehydration | Rhodotorula kratochvilovae (fungus) | [118] |
| Hydolysis, reduction and dehydration | Rhodosporidium paludigenum (fungus) | [119] | ||
| Zearalenone (ZEN, 33) | α-Zearalenol (42), β-zearalenol (43), zearalanone (44), α-zearalanol (151), and β-zearalanol (152) | Reduction and oxidation | Human | [121] |
| Hydrolyzed ZEN (153) and decarboxylated hydrolyzed ZEN (154) | Hydrolysis, spontaneous decarboxylation | Bacillus pumilus (bacterium) | [122] | |
| Hydrolysis, spontaneous decarboxylation | Lactonase | [123] | ||
| Hydrolysis, spontaneous decarboxylation | Lactonase | [124] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, P.; Su, R.; Yin, R.; Lai, D.; Wang, M.; Liu, Y.; Zhou, L. Detoxification of Mycotoxins through Biotransformation. Toxins 2020, 12, 121. https://doi.org/10.3390/toxins12020121
Li P, Su R, Yin R, Lai D, Wang M, Liu Y, Zhou L. Detoxification of Mycotoxins through Biotransformation. Toxins. 2020; 12(2):121. https://doi.org/10.3390/toxins12020121
Chicago/Turabian StyleLi, Peng, Ruixue Su, Ruya Yin, Daowan Lai, Mingan Wang, Yang Liu, and Ligang Zhou. 2020. "Detoxification of Mycotoxins through Biotransformation" Toxins 12, no. 2: 121. https://doi.org/10.3390/toxins12020121
APA StyleLi, P., Su, R., Yin, R., Lai, D., Wang, M., Liu, Y., & Zhou, L. (2020). Detoxification of Mycotoxins through Biotransformation. Toxins, 12(2), 121. https://doi.org/10.3390/toxins12020121








