Diversity Assessment of Toxic Cyanobacterial Blooms during Oxidation
Abstract
1. Introduction
2. Results and Discussion
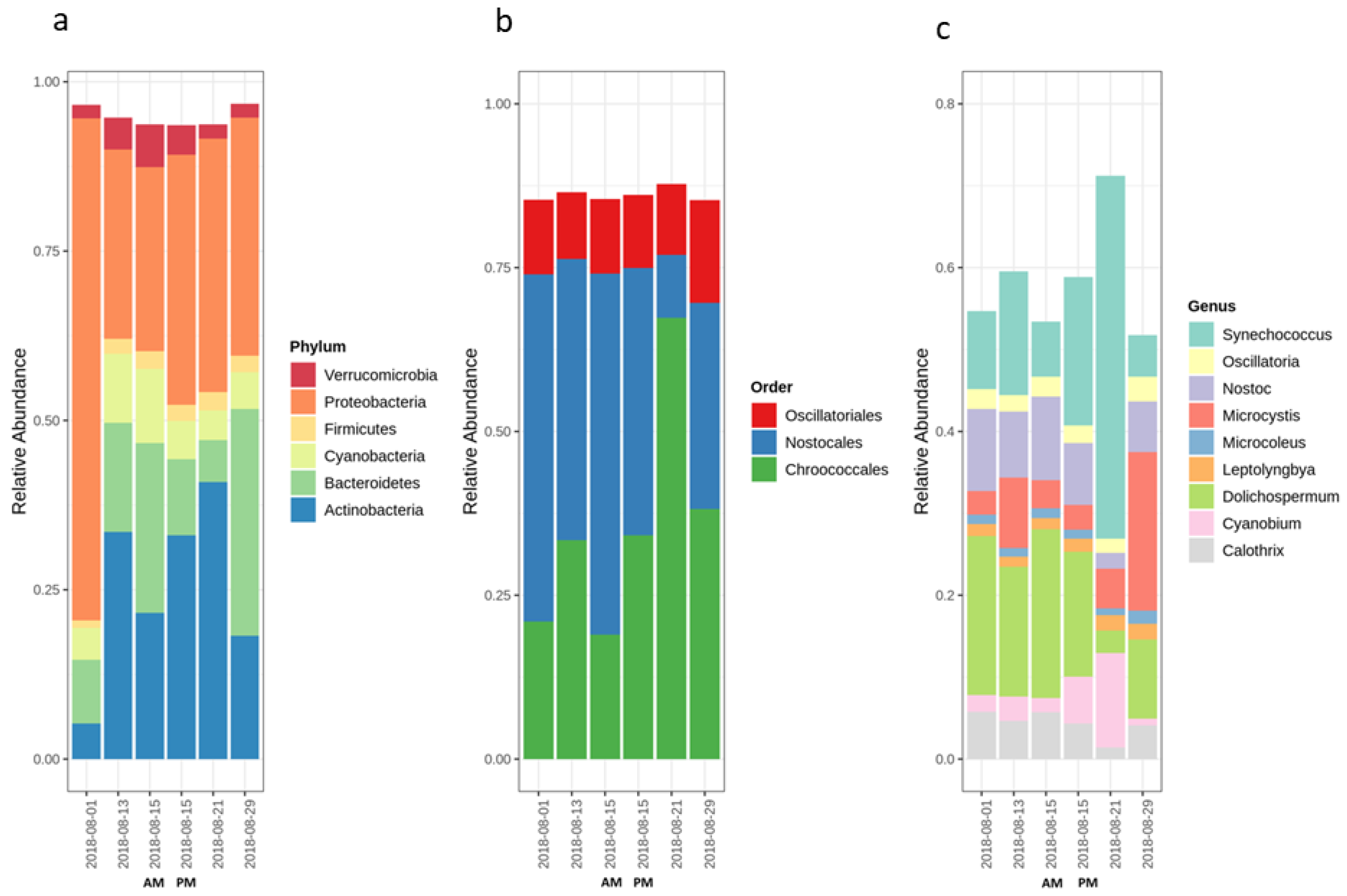
2.1. Cyanobacterial Bloom Characteristics Throughout Sampling
2.2. Variation of the Cyanobacterial Bloom Composition
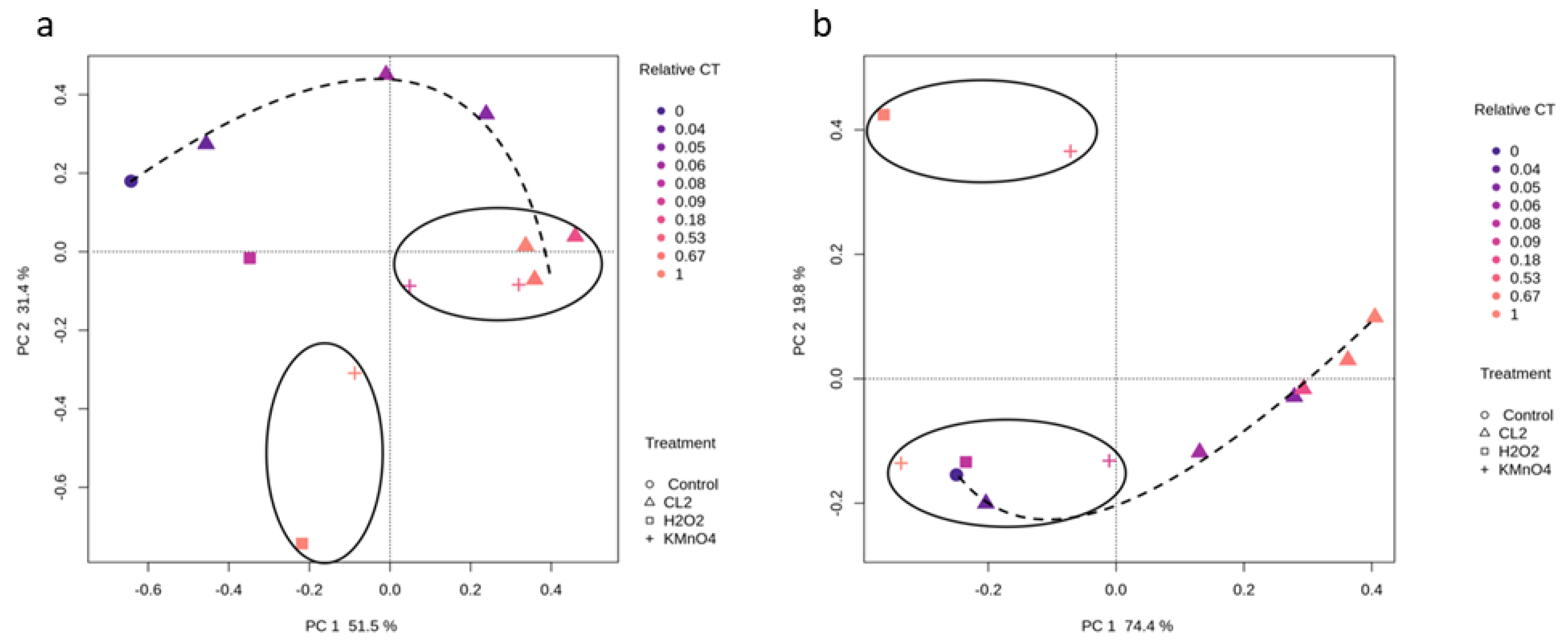
2.3. Impact of Oxidation on Cyanobacterial Diversity
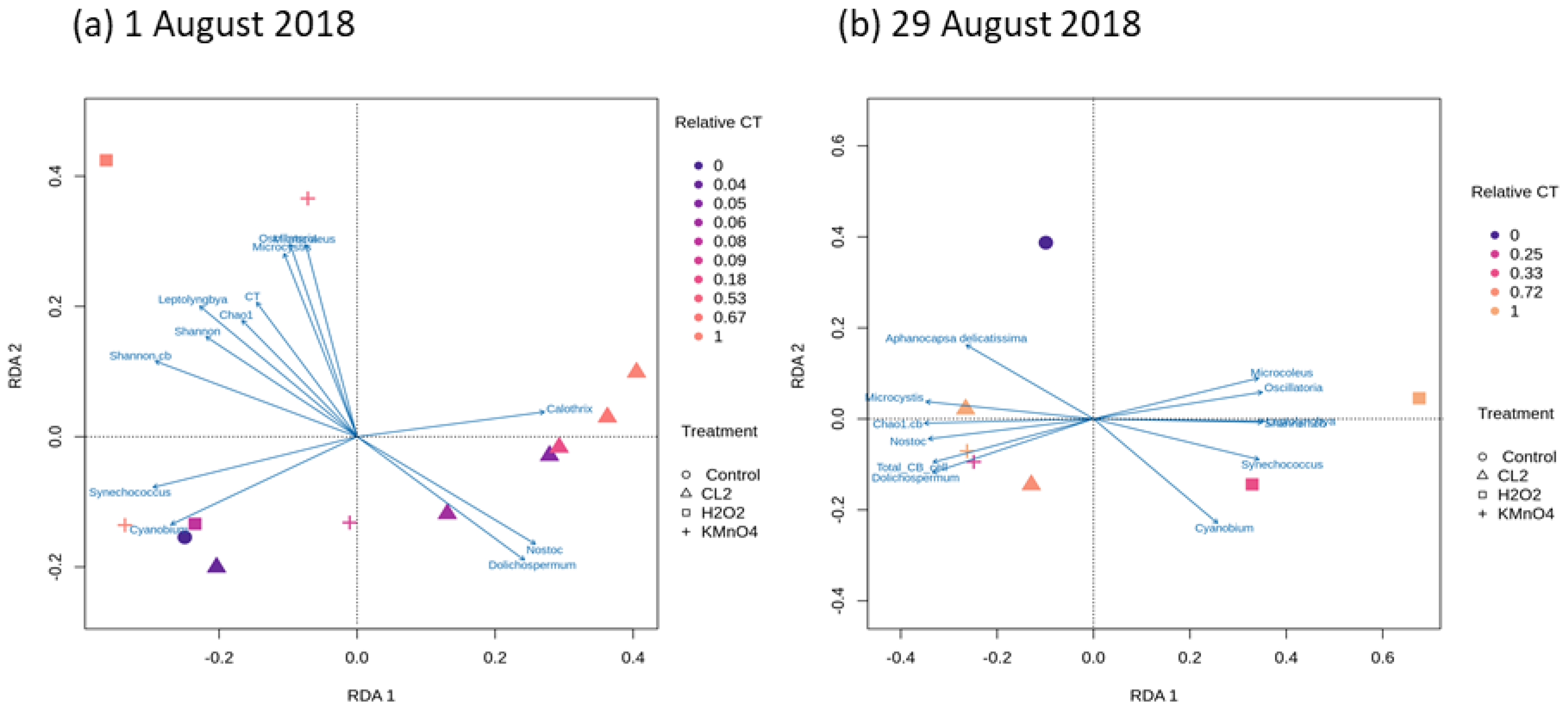
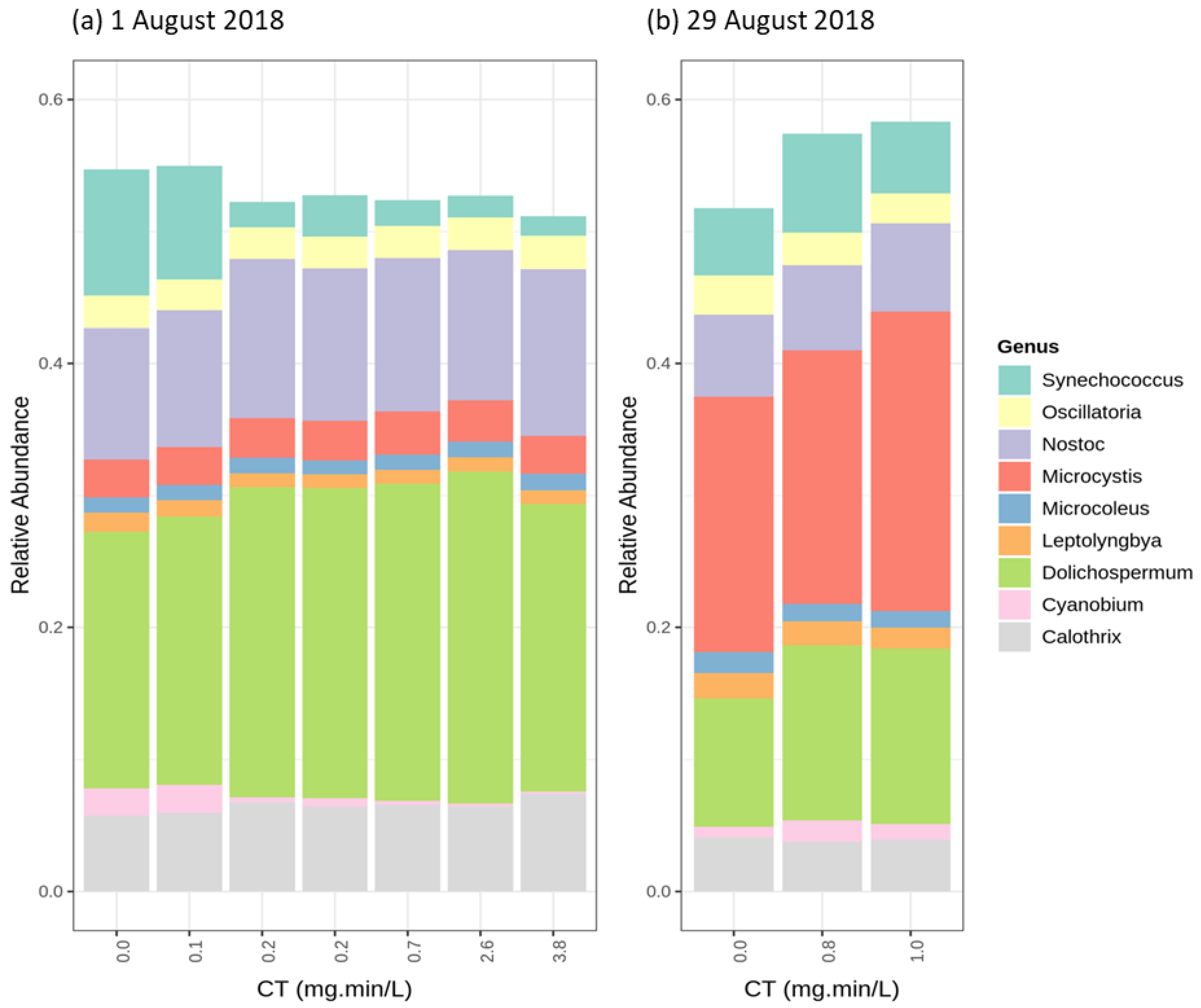
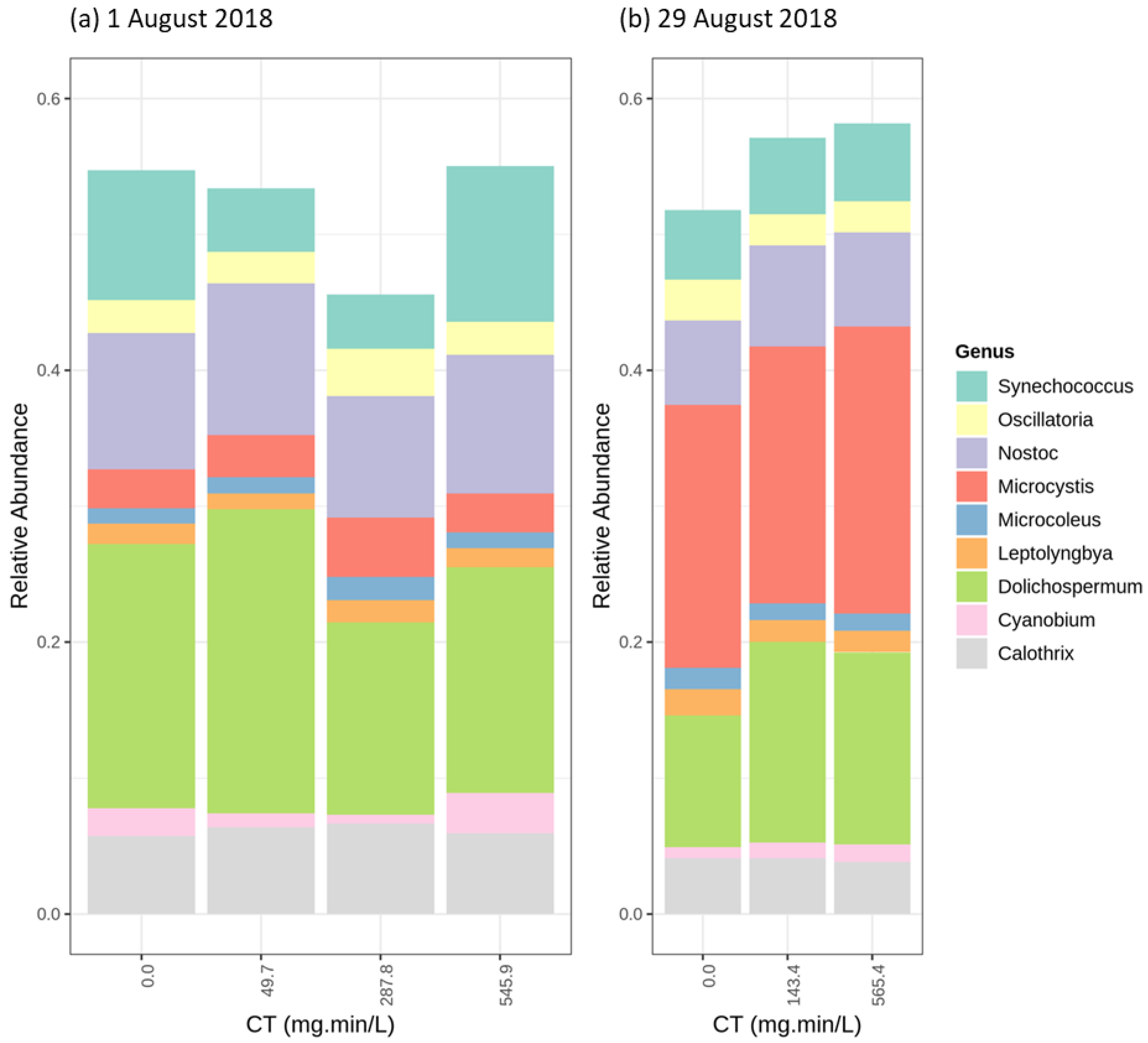
2.3.1. Cyanobacterial Composition
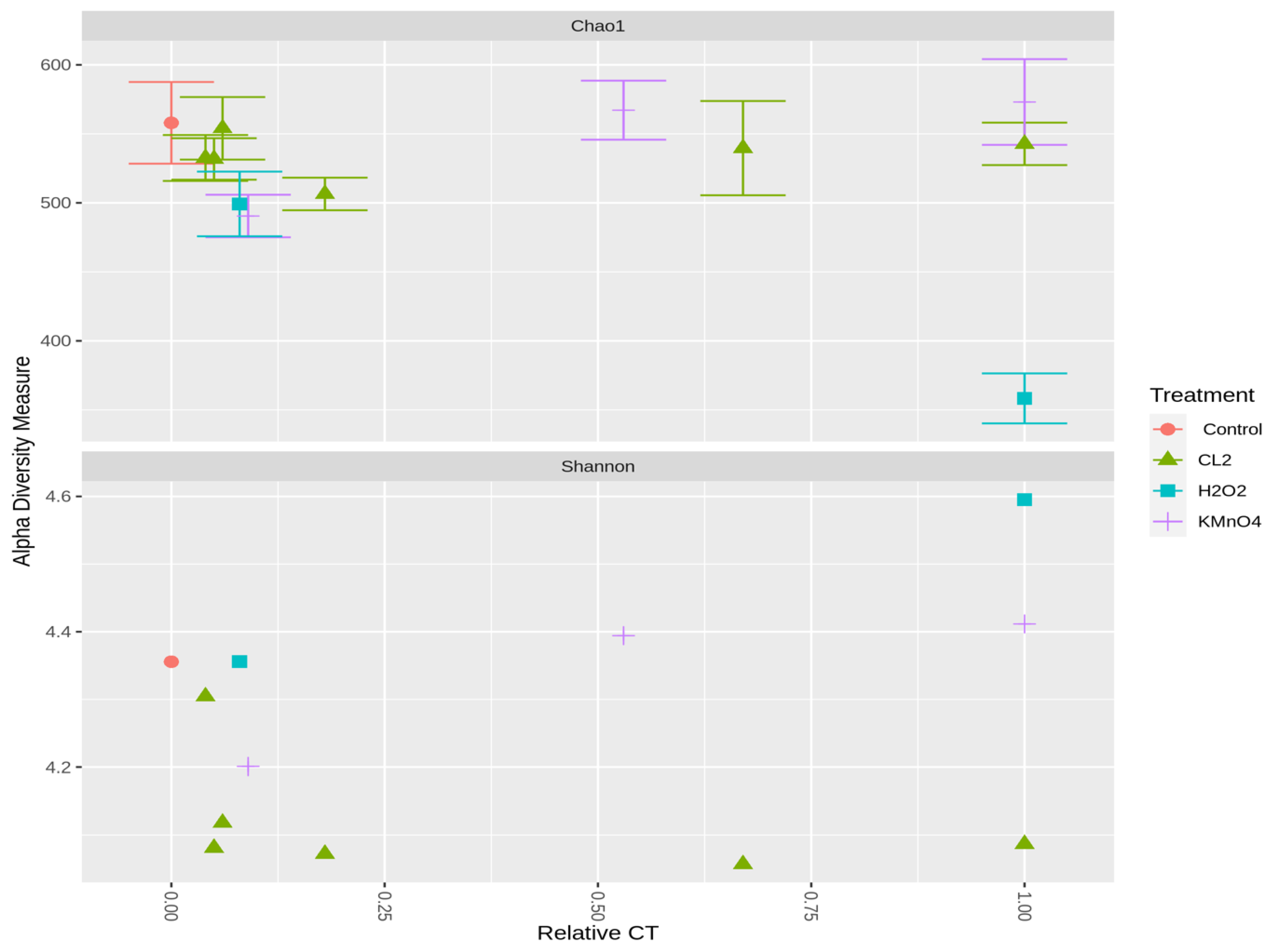
2.3.2. Effect of Oxidation on Cyanobacterial Community Richness and Diversity
2.4. Cyanobacterial Community Assessment Following Oxidation; Longitudinal Study
2.4.1. Chlorination (Cl2)
2.4.2. Potassium Permanganate (KMnO4)
2.4.3. Ozonation (O3)
2.4.4. Hydrogen Peroxide (H2O2)
2.5. Comparison of the Microscopic Cell Count vs. High Throughput Sequencing Results
3. Conclusions
4. Material and Methods
4.1. Sampling Site Description
4.2. Chemicals and Reagents
4.3. DNA Extraction, Metagenomic Preparation, Sequencing and Bioinformatic Analysis
4.4. DOC and Cyanobacteria Cell Count
4.5. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wells, M.L.; Trainer, V.L.; Smayda, T.J.; Karlson, B.S.; Trick, C.G.; Kudela, R.M.; Ishikawa, A.; Bernard, S.; Wulff, A.; Anderson, D.M.; et al. Harmful algal blooms and climate change: Learning from the past and present to forecast the future. Harmful Algae 2015, 49, 68–93. [Google Scholar] [CrossRef] [PubMed]
- Paerl, H.W.; Paul, V.J. Climate change: Links to global expansion of harmful cyanobacteria. Water Res. 2012, 46, 1349–1363. [Google Scholar] [CrossRef]
- Al-Sammak, M.A.; Hoagland, K.D.; Cassada, D.; Snow, D.D. Co-occurrence of the cyanotoxins BMAA, DABA and anatoxin-a in Nebraska reservoirs, fish, and aquatic plants. Toxins 2014, 6, 488–508. [Google Scholar] [CrossRef] [PubMed]
- Westrick, J.A.; Szlag, D.C.; Southwell, B.J.; Sinclair, J. A review of cyanobacteria and cyanotoxins removal/inactivation in drinking water treatment. Anal. Bioanal. Chem. 2010, 397, 1705–1714. [Google Scholar] [CrossRef]
- Casero, M.C.; Velázquez, D.; Medina-Cobo, M.; Quesada, A.; Cirés, S. Unmasking the identity of toxigenic cyanobacteria driving a multi-toxin bloom by high-throughput sequencing of cyanotoxins genes and 16S rRNA metabarcoding. Sci. Total Environ. 2019, 665, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Zamyadi, A.; Ho, L.; Newcombe, G.; Bustamante, H.; Prévost, M. Fate of toxic cyanobacterial cells and disinfection by-products formation after chlorination. Water Res. 2012, 46, 1524–1535. [Google Scholar] [CrossRef] [PubMed]
- Pazouki, P.; Prévost, M.; McQuaid, N.; Barbeau, B.; De Boutray, M.-L.; Zamyadi, A.; Dorner, S. Breakthrough of cyanobacteria in bank filtration. Water Res. 2016, 102, 170–179. [Google Scholar] [CrossRef] [PubMed]
- Zamyadi, A.; Dorner, S.; Ndong, M.; Ellis, D.; Bolduc, A.; Bastien, C.; Prévost, M. Low-risk cyanobacterial bloom sources: Cell accumulation within full-scale treatment plants. J. Am. Water Work. Assoc. 2013, 105, E651–E663. [Google Scholar] [CrossRef]
- Almuhtaram, H.; Cui, Y.; Zamyadi, A.; Hofmann, R. Cyanotoxins and Cyanobacteria Cell Accumulations in Drinking Water Treatment Plants with a Low Risk of Bloom Formation at the Source. Toxins 2018, 10, 430. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Daly, R.; Hobson, P.; Ho, L.; Brookes, J. Impact of potassium permanganate on cyanobacterial cell integrity and toxin release and degradation. Chemosphere 2013, 92, 529–534. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Ho, L.; Hobson, P.; Daly, R.; Brookes, J.D. Application of Various Oxidants for Cyanobacteria Control and Cyanotoxin Removal in Wastewater Treatment. J. Environ. Eng. 2014, 140, 04014022. [Google Scholar] [CrossRef]
- Coral, L.A.; Zamyadi, A.; Barbeau, B.; Bassetti, F.J.; Lapolli, F.R.; Prevost, M. Oxidation of M. aeruginosa and A. flos-aquae by ozone: Impacts on cell integrity and chlorination by-product formation. Water Res. 2013, 47, 2983–2994. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Shi, H.; Timmons, T.; Adams, C. Release and Removal of Microcystins from Microcystis during Oxidative-, Physical-, and UV-Based Disinfection. J. Environ. Eng. 2010, 136, 2–11. [Google Scholar] [CrossRef]
- Zamyadi, A.; Coral, L.A.; Barbeau, B.; Dorner, S.; Lapolli, F.R.; Prévost, M. Fate of toxic cyanobacterial genera from natural bloom events during ozonation. Water Res. 2015, 73, 204–215. [Google Scholar] [CrossRef]
- Zamyadi, A.; Greenstein, K.E.; Glover, C.M.; Adams, C.; Rosenfeldt, E.; Wert, E.C. Impact of Hydrogen Peroxide and Copper Sulfate on the Delayed Release of Microcystin. Water 2020, 12, 1105. [Google Scholar] [CrossRef]
- Zamyadi, A.; Ho, L.; Newcombe, G.; Daly, R.I.; Burch, M.; Baker, P.; Prévost, M. Release and Oxidation of Cell-Bound Saxitoxins during Chlorination of Anabaena circinalis Cells. Environ. Sci. Technol. 2010, 44, 9055–9061.e9. [Google Scholar] [CrossRef]
- Vlad, S.; Anderson, W.B.; Peldszus, S.; Huck, P.M. Removal of the cyanotoxin anatoxin-a by drinking water treatment processes: A review. J. Water Health 2014, 12, 601–617. [Google Scholar] [CrossRef]
- He, X.; Wert, E.C. Colonial cell disaggregation and intracellular microcystin release following chlorination of naturally occurring Microcystis. Water Res. 2016, 101, 10–16. [Google Scholar] [CrossRef]
- Wert, E.C.; Dong, M.M.; Rosario-Ortiz, F.L. Using digital flow cytometry to assess the degradation of three cyanobacteria species after oxidation processes. Water Res. 2013, 47, 3752–3761. [Google Scholar] [CrossRef]
- Merel, S.; Walker, D.; Chicana, R.; Snyder, S.A.; Baurès, E.; Thomas, O. State of knowledge and concerns on cyanobacterial blooms and cyanotoxins. Environ. Int. 2013, 59, 303–327. [Google Scholar] [CrossRef]
- Zamyadi, A.; Dorner, S.; Sauvé, S.; Ellis, D.; Bolduc, A.; Bastien, C.; Prévost, M. Species-dependence of cyanobacteria removal efficiency by different drinking water treatment processes. Water Res. 2013, 47, 2689–2700. [Google Scholar] [CrossRef]
- Zhu, L.; Zuo, J.; Song, L.; Gan, N. Microcystin-degrading bacteria affect mcyD expression and microcystin synthesis in Microcystis spp. J. Environ. Sci. 2016, 41, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Zamyadi, A.; Romanis, C.; Mills, T.; Neilan, B.; Choo, F.; Coral, L.A.; Gale, D.; Newcombe, G.; Crosbie, N.D.; Stuetz, R.M.; et al. Diagnosing water treatment critical control points for cyanobacterial removal: Exploring benefits of combined microscopy, next-generation sequencing, and cell integrity methods. Water Res. 2019, 152, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, P.R.; Holliday, J.; Kathuria, A.; Bowling, L. Change in cyanobacterial biovolume due to preservation by Lugol’s Iodine. Harmful Algae 2005, 4, 1033–1043. [Google Scholar] [CrossRef]
- American Water Works Association (AWWA). M57-Algae Source to Treatment; American Water Works Association: Denver, CO, USA, 2010. [Google Scholar]
- Berry, M.A.; Davis, T.W.; Cory, R.M.; Duhaime, M.B.; Johengen, T.H.; Kling, G.W.; Marino, J.A.; Uyl, P.A.D.; Gossiaux, D.; Dick, G.J.; et al. Cyanobacterial harmful algal blooms are a biological disturbance to Western Lake Erie bacterial communities. Environ. Microbiol. 2017, 19, 1149–1162. [Google Scholar] [CrossRef] [PubMed]
- Lezcano, M.Á.; Velázquez, D.; Quesada, A.; El-Shehawy, R. Diversity and temporal shifts of the bacterial community associated with a toxic cyanobacterial bloom: An interplay between microcystin producers and degraders. Water Res. 2017, 125, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Yoon, Y.; Hong, W.-Y.; Kim, J.; Cho, Y.-C.; Hwang, S.-J. Application of metagenome analysis to characterize the molecular diversity and saxitoxin-producing potentials of a cyanobacterial community: A case study in the North Han River, Korea. Appl. Biol. Chem. 2018, 61, 153–161. [Google Scholar] [CrossRef]
- Eldridge, S.L.C.; Wood, T.M. Annual variations in microcystin occurrence in Upper Klamath Lake, Oregon, based on high-throughput DNA sequencing, qPCR, and environmental parameters. Lake Reserv. Manag. 2019, 36, 31–44. [Google Scholar] [CrossRef]
- Casero, M.C.; Ballot, A.; Agha, R.; Quesada, A.; Cirés, S. Characterization of saxitoxin production and release and phylogeny of sxt genes in paralytic shellfish poisoning toxin-producing Aphanizomenon gracile. Harmful Algae 2014, 37, 28–37. [Google Scholar] [CrossRef]
- Woodhouse, J.N.; Kinsela, A.S.; Collins, R.N.; Bowling, L.C.; Honeyman, G.L.; Holliday, J.K.; Neilan, B.A. Microbial communities reflect temporal changes in cyanobacterial composition in a shallow ephemeral freshwater lake. ISME J. 2016, 10, 1337–1351. [Google Scholar] [CrossRef]
- Pessi, I.S.; Maalouf, P.D.C.; Laughinghouse, H.D.; Baurain, D.; Wilmotte, A. On the use of high-throughput sequencing for the study of cyanobacterial diversity in Antarctic aquatic mats. J. Phycol. 2016, 52, 356–368. [Google Scholar] [CrossRef] [PubMed]
- Scherer, P.I.; Millard, A.D.; Miller, A.; Schoen, R.; Raeder, U.; Geist, J.; Zwirglmaier, K. Temporal Dynamics of the Microbial Community Composition with a Focus on Toxic Cyanobacteria and Toxin Presence during Harmful Algal Blooms in Two South German Lakes. Front. Microbiol. 2017, 8, 2387. [Google Scholar] [CrossRef] [PubMed]
- Willis, A.; Woodhouse, J.N. Defining Cyanobacterial Species: Diversity and Description Through Genomics. Crit. Rev. Plant Sci. 2020, 39, 101–124. [Google Scholar] [CrossRef]
- Xu, H.; Pei, H.; Jin, Y.; Ma, C.; Wang, Y.; Sun, J.; Li, H. High-throughput sequencing reveals microbial communities in drinking water treatment sludge from six geographically distributed plants, including potentially toxic cyanobacteria and pathogens. Sci. Total Environ. 2018, 634, 769–779. [Google Scholar] [CrossRef] [PubMed]
- Pei, H.; Xu, H.; Wang, J.; Jin, Y.; Xiao, H.; Ma, C.; Sun, J.; Li, H. 16S rRNA Gene Amplicon Sequencing Reveals Significant Changes in Microbial Compositions during Cyanobacteria-Laden Drinking Water Sludge Storage. Environ. Sci. Technol. 2017, 51, 12774–12783. [Google Scholar] [CrossRef]
- Lusty, M.W.; Gobler, C.J. The Efficacy of Hydrogen Peroxide in Mitigating Cyanobacterial Blooms and Altering Microbial Communities across Four Lakes in NY, USA. Toxins 2020, 12, 428. [Google Scholar] [CrossRef]
- Newcombe, G.; House, J.; Ho, L.; Baker, P.; Burch, M. Management Strategies for Cyanobacteria (Blue-Green Algae): A Guide for Water Utilities; The Cooperative Research Centre for Water Quality and Treatment: Adelaïde, Australia, 2010; p. 112. [Google Scholar]
- Lee, C.; Marion, J.; Cheung, M.; Lee, C.S.; Lee, J. Associations among Human-Associated Fecal Contamination, Microcystis aeruginosa, and Microcystin at Lake Erie Beaches. Int. J. Environ. Res. Public Health 2015, 12, 11466–11485. [Google Scholar] [CrossRef]
- Vadde, K.K.; Feng, Q.; Wang, J.; McCarthy, A.J.; Sekar, R. Next-generation sequencing reveals fecal contamination and potentially pathogenic bacteria in a major inflow river of Taihu Lake. Environ. Pollut. 2019, 254, 113108. [Google Scholar] [CrossRef]
- Ballesté, E.; Blanch, A.R. Persistence of Bacteroides Species Populations in a River as Measured by Molecular and Culture Techniques. Appl. Environ. Microbiol. 2010, 76, 7608–7616. [Google Scholar] [CrossRef]
- Zamyadi, A.; MacLeod, S.L.; Fan, Y.; McQuaid, N.; Dorner, S.; Sauvé, S.; Prévost, M. Toxic cyanobacterial breakthrough and accumulation in a drinking water plant: A monitoring and treatment challenge. Water Res. 2012, 46, 1511–1523. [Google Scholar] [CrossRef]
- McQuaid, N.; Zamyadi, A.; Prévost, M.; Bird, D.F.; Dorner, S. Use of in vivophycocyanin fluorescence to monitor potential microcystin-producing cyanobacterial biovolume in a drinkingwater source. J. Environ. Monit. 2011, 13, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Tromas, N.; Fortin, N.; Bedrani, L.; Terrat, Y.; Cardoso, P.; Bird, D.; Greer, C.W.; Shapiro, B.J. Characterising and predicting cyanobacterial blooms in an 8-year amplicon sequencing time course. ISME J. 2017, 11, 1746–1763. [Google Scholar] [CrossRef] [PubMed]
- Andersson, A.; Höglander, H.; Karlsson, C.; Huseby, S. Key role of phosphorus and nitrogen in regulating cyanobacterial community composition in the northern Baltic Sea. Estuar. Coast. Shelf Sci. 2015, 164, 161–171. [Google Scholar] [CrossRef]
- Fortin, N.; Munoz-Ramos, V.; Bird, D.; Lévesque, B.; Whyte, L.G.; Greer, C.W. Toxic Cyanobacterial Bloom Triggers in Missisquoi Bay, Lake Champlain, as Determined by Next-Generation Sequencing and Quantitative PCR. Life 2015, 5, 1346–1380. [Google Scholar] [CrossRef]
- Harke, M.J.; Davis, T.W.; Watson, S.B.; Gobler, C.J. Nutrient-Controlled Niche Differentiation of Western Lake Erie Cyanobacterial Populations Revealed via Metatranscriptomic Surveys. Environ. Sci. Technol. 2016, 50, 604–615. [Google Scholar] [CrossRef]
- Li, Q.; Lin, F.; Yang, C.; Wang, J.; Lin, Y.; Shen, M.; Park, M.S.; Li, T.; Zhao, J. A Large-Scale Comparative Metagenomic Study Reveals the Functional Interactions in Six Bloom-Forming Microcystis-Epibiont Communities. Front. Microbiol. 2018, 9, 746. [Google Scholar] [CrossRef]
- Shi, L.; Cai, Y.; Li, P.; Yang, H.; Liu, Z.; Kong, L.; Yü, Y.; Kong, F. Molecular Identification of the Colony-Associated Cultivable Bacteria of the CyanobacteriumMicrocystis aeruginosaand Their Effects on Algal Growth. J. Freshw. Ecol. 2009, 24, 211–218. [Google Scholar] [CrossRef]
- Maruyama, T.; Kato, K.; Yokoyama, A.; Tanaka, T.; Hiraishi, A.; Park, H.-D. Dynamics of microcystin-degrading bacteria in mucilage of Microcystis. Microb. Ecol. 2003, 46, 279–288. [Google Scholar] [CrossRef]
- Wacklin, P.; Hoffmann, L.; Komarek, J. Nomenclatural validation of the genetically revised cyanobacterial genus Dolichospermum (RALFS ex BORNET et FLAHAULT) comb. nova. Fottea 2009, 9, 59–64. [Google Scholar] [CrossRef]
- Van Wichelen, J.; Vanormelingen, P.; Codd, G.A.; Vyverman, W. The common bloom-forming cyanobacterium Microcystis is prone to a wide array of microbial antagonists. Harmful Algae 2016, 55, 97–111. [Google Scholar] [CrossRef]
- Li, Y.; Li, D. Competition between toxic Microcystis aeruginosa and nontoxic Microcystis wesenbergii with Anabaena PCC. Environ. Boil. Fishes 2012, 24, 69–78. [Google Scholar]
- Kardinaal, W.; Janse, I.; Agterveld, M.K.-V.; Meima, M.; Snoek, J.; Mur, L.; Huisman, J.; Zwart, G.; Visser, P. Microcystis genotype succession in relation to microcystin concentrations in freshwater lakes. Aquat. Microb. Ecol. 2007, 48, 1–12. [Google Scholar] [CrossRef]
- Zhang, X.-W.; Fu, J.; Song, S.; Zhang, P.; Yang, X.-H.; Zhang, L.-R.; Luo, Y.; Liu, C.-H.; Zhu, H.-L. Interspecific competition between Microcystis aeruginosa and Anabaena flos-aquae from Taihu Lake, China. Z. für Nat. C 2014, 69, 53–60. [Google Scholar] [CrossRef]
- Chia, M.A.; Jankowiak, J.G.; Kramer, B.J.; Goleski, J.A.; Huang, I.-S.; Zimba, P.V.; Bittencourt-Oliveira, M.D.C.; Gobler, C.J. Succession and toxicity of Microcystis and Anabaena (Dolichospermum) blooms are controlled by nutrient-dependent allelopathic interactions. Harmful Algae 2018, 74, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Ndong, M.; Bird, D.; Nguyen-Quang, T.; De Boutray, M.-L.; Zamyadi, A.; Vinçon-Leite, B.; Lemaire, B.J.; Prévost, M.; Dorner, S. Estimating the risk of cyanobacterial occurrence using an index integrating meteorological factors: Application to drinking water production. Water Res. 2014, 56, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Hobson, P.; Ho, L.; Daly, R.; Brookes, J.D. The effects of various control and water treatment processes on the membrane integrity and toxin fate of cyanobacteria. J. Hazard. Mater. 2014, 264, 313–322. [Google Scholar] [CrossRef]
- Zamyadi, A.; Fan, Y.; Daly, R.I.; Prévost, M. Chlorination of Microcystis aeruginosa: Toxin release and oxidation, cellular chlorine demand and disinfection by-products formation. Water Res. 2013, 47, 1080–1090. [Google Scholar] [CrossRef]
- Fan, J.; Ho, L.; Hobson, P.; Brookes, J.D. Evaluating the effectiveness of copper sulphate, chlorine, potassium permanganate, hydrogen peroxide and ozone on cyanobacterial cell integrity. Water Res. 2013, 47, 5153–5164. [Google Scholar] [CrossRef]
- Matthijs, H.C.; Visser, P.M.; Reeze, B.; Meeuse, J.; Slot, P.C.; Wijn, G.; Talens, R.; Huisman, J. Selective suppression of harmful cyanobacteria in an entire lake with hydrogen peroxide. Water Res. 2012, 46, 1460–1472. [Google Scholar] [CrossRef]
- Zamyadi, A.; Henderson, R.K.; Stuetz, R.; Newcombe, G.; Newtown, K.; Gladman, B. Cyanobacterial management in full-scale water treatment and recycling processes: Reactive dosing following intensive monitoring. Environ. Sci. Water Res. Technol. 2016, 2, 362–375. [Google Scholar] [CrossRef]
- Moradinejad, S.; Glover, C.M.; Mailly, J.; Seighalani, T.Z.; Peldszus, S.; Barbeau, B.; Dorner, S.; Prévost, M.; Zamyadi, A. Using Advanced Spectroscopy and Organic Matter Characterization to Evaluate the Impact of Oxidation on Cyanobacteria. Toxins 2019, 11, 278. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Shao, Y.; Gao, N.; Deng, Y.; Qiao, J.; Ou, H.; Deng, J. Effects of different algaecides on the photosynthetic capacity, cell integrity and microcystin-LR release of Microcystis aeruginosa. Sci. Total Environ. 2013, 463, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Kim, Y.; Kim, M.; Lee, W.H. A novel method for cell counting of Microcystis colonies in water resources using a digital imaging flow cytometer and microscope. Environ. Eng. Res. 2018, 24, 397–403. [Google Scholar] [CrossRef]
- Xiao, X.; Sogge, H.; Lagesen, K.; Tooming-Klunderud, A.; Jakobsen, K.S.; Rohrlack, T. Use of High Throughput Sequencing and Light Microscopy Show Contrasting Results in a Study of Phytoplankton Occurrence in a Freshwater Environment. PLoS ONE 2014, 9, e106510. [Google Scholar]
- American Public Health Association (APHA); American Water Works Association (AWWA); Water Environment Federation (WEF). Standard Methods for the Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 2012; Volume 5, p. 1360. [Google Scholar]
- Cox, M.P.; Peterson, D.A.; Biggs, P.J. SolexaQA: At-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinform. 2010, 11, 1–6. [Google Scholar] [CrossRef]
- Kim, D.; Hahn, A.S.; Wu, S.-J.; Hanson, N.W.; Konwar, K.M.; Hallam, S.J. FragGeneScan-Plus for Scalable High-Throughput Short-Read Open Reading Frame Prediction; IEEE CIBCB Conference: Niagara Falls, ON, Canada, 2015; pp. 1–8. [Google Scholar]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Lund, J.W.G. A Simple Counting Chamber for Nannoplankton. Limnol. Oceanogr. 1959, 4, 57–65. [Google Scholar] [CrossRef]
- Planas, D.; Desrosiers, M.; Groulx, S.; Paquet, S.; Carignan, R. Pelagic and benthic algal responses in eastern Canadian Boreal Shield lakes following harvesting and wildfires. Can. J. Fish. Aquat. Sci. 2000, 57, 136–145. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S.P. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar]
- Raivo, K. Pheatmap: Pretty Heatmaps, R package version 1.0.12; GitHub, Inc.: San Francisco, CA, USA, 2019. [Google Scholar]
- Greenacre, M. Compositional Data Analysis in Practice, 1st ed.; CRC Press: Cleveland, OH, USA, 2018. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Community Ecology Package, R Package ‘Vegan’, version 2.5-0; GitHub, Inc.: San Francisco, CA, USA, 2016; p. 285. [Google Scholar]



| Sampling Date | DOC (mg/L) | pH | Cell Count (cells/mL) | Biovolume (mm3/L) |
|---|---|---|---|---|
| 1 August 2018 | 5.9 | 7.6 | 3.3 × 105 | 30.6 |
| 13 August 2018 | 5.8 | 7.3 | 7.8 × 104 | 4.6 |
| 15 August 2018 | 5.5 | 7.4 | 1.4 × 105 | 9.4 |
| 21 August 2018 | 4.9 | 7.4 | 6.8 × 104 | 0.3 |
| 29 August 2018 | 5.6 | 7.5 | 5.4 × 104 | 2.1 |
| Oxidant | Water Type | Oxidant Dose (mg/L) | Contact Time |
|---|---|---|---|
| Cl2 | Real Bloom, Missisquoi Bay, Quebec, Canada | 0.2 0.6 | 1 min 2 min 5 min 10 min 60 min 120 min |
| O3 | 0.1 0.3 | 1 min 2 min 5 min 10 min | |
| KMnO4 | 5 | 30 min 60 min 120 min | |
| H2O2 | 10 | 6 h 24 h |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moradinejad, S.; Trigui, H.; Guerra Maldonado, J.F.; Shapiro, J.; Terrat, Y.; Zamyadi, A.; Dorner, S.; Prévost, M. Diversity Assessment of Toxic Cyanobacterial Blooms during Oxidation. Toxins 2020, 12, 728. https://doi.org/10.3390/toxins12110728
Moradinejad S, Trigui H, Guerra Maldonado JF, Shapiro J, Terrat Y, Zamyadi A, Dorner S, Prévost M. Diversity Assessment of Toxic Cyanobacterial Blooms during Oxidation. Toxins. 2020; 12(11):728. https://doi.org/10.3390/toxins12110728
Chicago/Turabian StyleMoradinejad, Saber, Hana Trigui, Juan Francisco Guerra Maldonado, Jesse Shapiro, Yves Terrat, Arash Zamyadi, Sarah Dorner, and Michèle Prévost. 2020. "Diversity Assessment of Toxic Cyanobacterial Blooms during Oxidation" Toxins 12, no. 11: 728. https://doi.org/10.3390/toxins12110728
APA StyleMoradinejad, S., Trigui, H., Guerra Maldonado, J. F., Shapiro, J., Terrat, Y., Zamyadi, A., Dorner, S., & Prévost, M. (2020). Diversity Assessment of Toxic Cyanobacterial Blooms during Oxidation. Toxins, 12(11), 728. https://doi.org/10.3390/toxins12110728






