Production of β-methylamino-L-alanine (BMAA) and Its Isomers by Freshwater Diatoms
Abstract
1. Introduction
1.1. Algal and Diatom Toxins
1.2. The Toxicity of BMAA, 2,4-DAB and AEG
1.3. BMAA and Its Worldwide Distribution
1.4. Producers of BMAA, 2,4-DAB & AEG
1.5. Aims of the Study
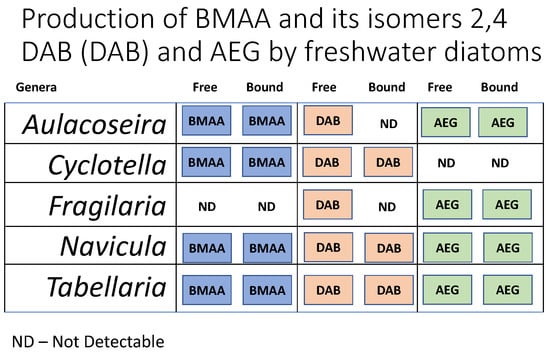
2. Results
3. Discussion
3.1. Comparison of Diatom Studies
3.2. BMAA in Freshwater Diatoms
3.3. AEG and 2,4-DAB in Freshwater Diatoms
3.4. Toxic Diatoms in Freshwater Systems
4. Conclusions
5. Materials and Methods
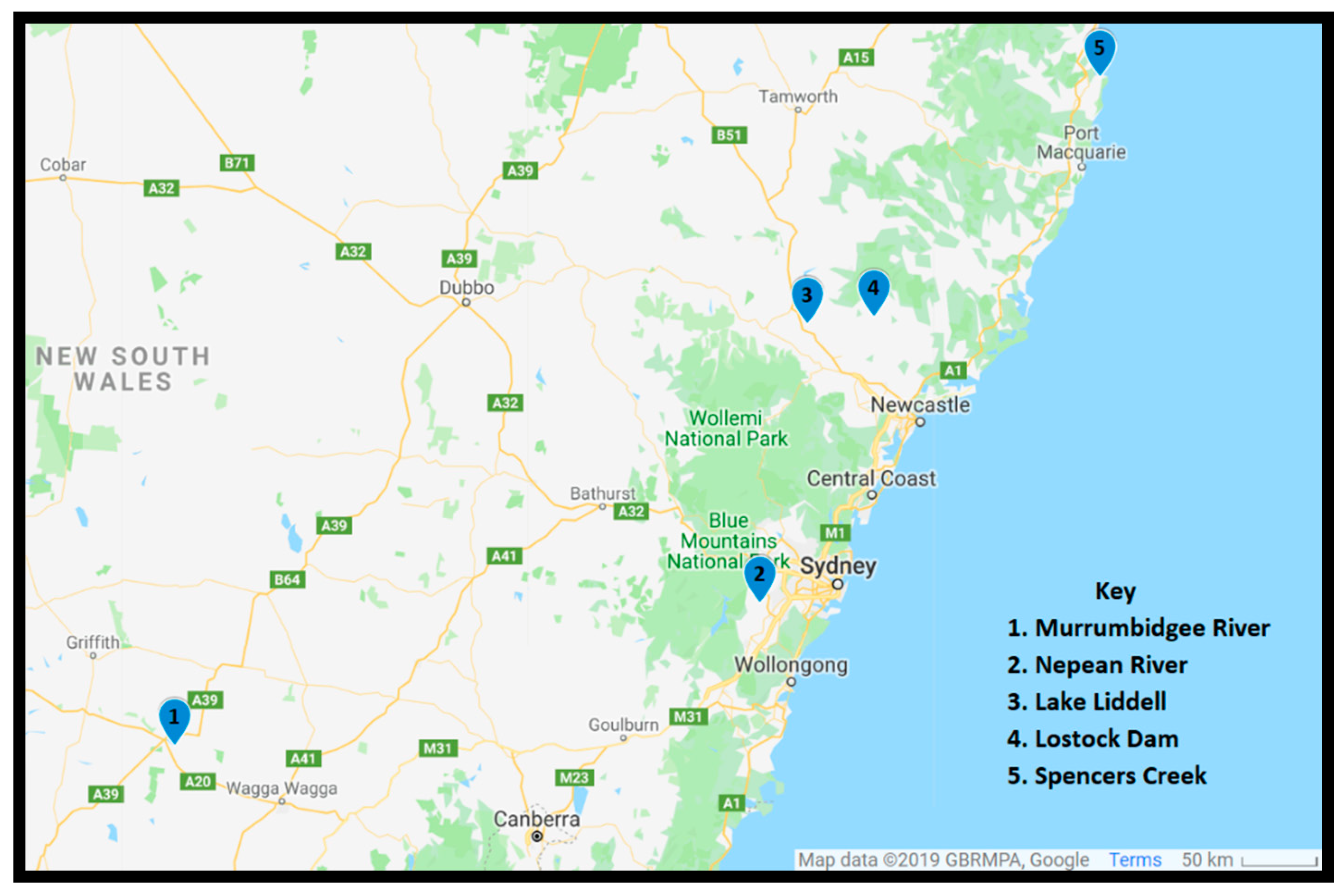
5.1. Sample Collection
5.2. Diatom Isolation, Purification & Culturing
5.2.1. Algal Media
5.2.2. Diatom Isolation
5.2.3. Diatom Purification and Culturing
5.3. Amino Acid Extraction
5.3.1. Diatom Harvesting, Lysing and Fractionation
5.3.2. Protein Hydrolysis
5.4. Amino Acid Derivatisation and LC-MS/MS
5.4.1. Propyl Chloroformate Derivatisation
5.4.2. LC-MS/MS Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sitprija, V.; Sitprija, S. Marine toxins and nephrotoxicity:Mechanism of injury. Toxicon 2019, 161, 44–49. [Google Scholar] [CrossRef]
- Merel, S.; Walker, D.; Chicana, R.; Snyder, S.; Baurès, E.; Thomas, O. State of knowledge and concerns on cyanobacterial blooms and cyanotoxins. Environ. Int. 2013, 59, 303–327. [Google Scholar] [CrossRef]
- Rao, D.V.S.; Quilliam, M.A.; Pocklington, R. Domoic Acid—A Neurotoxic Amino Acid Produced by the Marine Diatom Nitzschia pungens in Culture. Can. J. Fish. Aquat.Sci. 1988, 45, 2076–2079. [Google Scholar] [CrossRef]
- Réveillon, D.; Abadie, E.; Séchet, V.; Masseret, E.; Hess, P.; Amzil, Z. β-N-methylamino-l-alanine (BMAA) and isomers: Distribution in different food web compartments of Thau lagoon, French Mediterranean Sea. Mar. Environ. Res. 2015, 110, 8–18. [Google Scholar] [CrossRef]
- Lage, S.; Burian, A.; Rasmussen, U.; Costa, P.R.; Annadotter, H.; Godhe, A.; Rydberg, S. BMAA extraction of cyanobacteria samples: Which method to choose? Environ. Sci. Pollut. Res. Int. 2016, 23, 338–350. [Google Scholar] [CrossRef]
- Jiang, L.; Eriksson, J.; Lage, S.; Jonasson, S.; Shams, S.; Mehine, M.; Ilag, L.L.; Rasmussen, U. Diatoms: A Novel Source for the Neurotoxin BMAA in Aquatic Environments. PLoS ONE 2014, 9, e84578. [Google Scholar] [CrossRef]
- Rodgers, K.J. Non-protein amino acids and neurodegeneration: The enemy within. Exp. Neurol. 2014, 253, 192–196. [Google Scholar] [CrossRef]
- Main, B.J.; Rodgers, K.J. Assessing the Combined Toxicity of BMAA and Its Isomers 2,4-DAB and AEG In Vitro Using Human Neuroblastoma Cells. Neurotox. Res. 2017, 33, 33–42. [Google Scholar] [CrossRef]
- Metcalf, J.; Banack, S.; Richer, R.; Cox, P.; Metcalf, J. Neurotoxic amino acids and their isomers in desert environments. J. Arid. Environ. 2015, 112, 140–144. [Google Scholar] [CrossRef]
- Rao, S.D.; Banack, S.A.; Cox, P.A.; Weiss, J.H. BMAA selectively injures motor neurons via AMPA/kainate receptor activation. Exp. Neurol. 2006, 201, 244–252. [Google Scholar] [CrossRef]
- Weiss, J.H.; Christine, C.W.; Choi, D.W. Bicarbonate dependence of glutamate receptor activation by beta-N-methylamino-L-alanine: Channel recording and study with related compounds. Neuron 1989, 3, 321–326. [Google Scholar] [CrossRef]
- Murch, S.J.; Cox, P.A.; Banack, S.A.; Steele, J.C.; Sacks, O.W. Occurrence of beta-methylamino-l-alanine (BMAA) in ALS/PDC patients from Guam. Acta Neurol. Scand. 2004, 110, 267–269. [Google Scholar] [CrossRef]
- Cox, P.A.; Davis, D.A.; Mash, D.C.; Metcalf, J.S.; Banack, S.A. Do vervets and macaques respond differently to BMAA? NeuroToxicology 2016, 57, 310–311. [Google Scholar] [CrossRef]
- Dunlop, R.A.; Cox, P.A.; Banack, S.A.; Rodgers, K.J. The Non-Protein Amino Acid BMAA Is Misincorporated into Human Proteins in Place of l-Serine Causing Protein Misfolding and Aggregation. PLoS ONE 2013, 8, e75376. [Google Scholar] [CrossRef]
- Cox, P.A.; Davis, D.A.; Mash, D.C.; Metcalf, J.S.; Banack, S.A. Dietary exposure to an environmental toxin triggers neurofibrillary tangles and amyloid deposits in the brain. Proc. R. Soc. B Biol. Sci. 2016, 283, 20152397. [Google Scholar] [CrossRef]
- Murch, S.J.; Cox, P.A.; Banack, S.A. A mechanism for slow release of biomagnified cyanobacterial neurotoxins and neurodegenerative disease in Guam. Proc. Natl. Acad. Sci. USA 2004, 101, 12228–12231. [Google Scholar] [CrossRef]
- Van Onselen, R.; Downing, T. BMAA-protein interactions: A possible new mechanism of toxicity. Toxicon 2018, 143, 74–80. [Google Scholar] [CrossRef]
- Main, B.J.; Italiano, C.J.; Rodgers, K.J. Investigation of the interaction of beta-methylamino-L-alanine with eukaryotic and prokaryotic proteins. Amino Acids 2018, 50, 397–407. [Google Scholar] [CrossRef]
- Glover, W.B.; Mash, D.C.; Murch, S.J. The natural non-protein amino acid N-β-methylamino-l-alanine (BMAA) is incorporated into protein during synthesis. Amino Acids 2014, 46, 2553–2559. [Google Scholar] [CrossRef]
- Song, Y.; Zhou, H.; Vo, M.-N.; Shi, Y.; Nawaz, M.H.; Vargas-Rodriguez, O.; Diedrich, J.K.; Yates, J.R.; Kishi, S.; Musier-Forsyth, K.; et al. Double mimicry evades tRNA synthetase editing by toxic vegetable-sourced non-proteinogenic amino acid. Nat. Commun. 2017, 8, 2281. [Google Scholar] [CrossRef]
- Chernoff, N.; Hill, D.J.; Diggs, D.L.; Faison, B.D.; Francis, B.M.; Lang, J.R.; LaRue, M.M.; Le, T.-T.; Loftin, K.A.; Lugo, J.N.; et al. A critical review of the postulated role of the non-essential amino acid, β-N-methylamino-L-alanine, in neurodegenerative disease in humans. J. Toxicol. Environ. Health Part B 2017, 20, 1–47. [Google Scholar] [CrossRef]
- Okle, O.; Rath, L.; Galizia, C.G.; Dietrich, D.R. The cyanobacterial neurotoxin beta-N-methylamino-l-alanine (BMAA) induces neuronal and behavioral changes in honeybees. Toxicol. Appl. Pharmacol. 2013, 270, 9–15. [Google Scholar] [CrossRef]
- Nunn, P.B. 50 years of research on alpha-amino-beta-methylaminopropionic acid (beta-methylaminoalanine). Phytochemistry 2017, 144, 271–281. [Google Scholar] [CrossRef]
- Brody, J.A.; Stanhope, J.M.; Kurland, L.T. Patterns of Amyotrophic Lateral Sclerosis and Parkinsonism-Dementia on Guam. Contemp. Neurol. Ser. 1975, 12, 45–70. [Google Scholar]
- Mulder, D.W.; Kurland, L.T.; Iriarte, L.L. Neurologic diseases on the island of Guam. USA Armed Forces Med. J. 1954, 5, 1724–1739. [Google Scholar]
- Kurland, L.K.; Mulder, D.W. Epidemiologic Investigations of Amyotrophic Lateral Sclerosis: 1. Preliminary Report on Geographic Distribution, with Special Reference to the Mariana Islands, Including Clinical and Pathologic Observations. Neurology 1954, 4, 355. [Google Scholar] [CrossRef]
- Cox, P.A.; Sacks, O.W. Cycad neurotoxins, consumption of flying foxes, and ALS-PDC disease in Guam. Neurology 2002, 58, 956–959. [Google Scholar] [CrossRef]
- Banack, S.A.; Murch, S.J. Multiple neurotoxic items in the Chamorro diet link BMAA with ALS/PDC. Amyotroph. Lateral Scler. 2009, 10, 34–40. [Google Scholar] [CrossRef]
- Cox, P.A.; Banack, S.A.; Murch, S.J. Biomagnification of cyanobacterial neurotoxins and neurodegenerative disease among the Chamorro people of Guam. Proc. Natl. Acad. Sci. USA 2003, 100, 13380–13383. [Google Scholar] [CrossRef]
- Garruto, R.M.; Gajdusek, D.C.; Chen, K.-M.; Chen, K. Amyotrophic lateral sclerosis and parkinsonism-dementia among Filipino migrants to Guam. Ann. Neurol. 1981, 10, 341–350. [Google Scholar] [CrossRef]
- Banack, S.A.; Cox, P.A. Biomagnification of cycad neurotoxins in flying foxes: Implications for ALS-PDC in Guam. Neurology 2003, 61, 387–389. [Google Scholar] [CrossRef]
- Lage, S.; Annadotter, H.; Rasmussen, U.; Rydberg, S. Biotransfer of β-N-Methylamino-l-alanine (BMAA) in a Eutrophicated Freshwater Lake. Mar. Drugs 2015, 13, 1185–1201. [Google Scholar] [CrossRef]
- Brand, L.E.; Pablo, J.; Compton, A.; Hammerschlag, N.; Mash, D.C. Cyanobacterial Blooms and the Occurrence of the neurotoxin beta-N-methylamino-L-alanine (BMAA) in South Florida Aquatic Food Webs. Harmful Algae 2010, 9, 620–635. [Google Scholar] [CrossRef]
- Masseret, E.; Banack, S.; Boumédiene, F.; Abadie, E.; Brient, L.; Pernet, F.; Juntas-Morales, R.; Pageot, N.; Metcalf, J.; Cox, P.; et al. Dietary BMAA Exposure in an Amyotrophic Lateral Sclerosis Cluster from Southern France. PLoS ONE 2013, 8, e83406. [Google Scholar] [CrossRef]
- Jiang, L.; Kiselova, N.; Rosén, J.; Ilag, L.L. Quantification of neurotoxin BMAA (β-N-methylamino-L-alanine) in seafood from Swedish markets. Sci. Rep. 2014, 4, 6931. [Google Scholar] [CrossRef]
- Caller, T.A.; Doolin, J.W.; Haney, J.F.; Murby, A.J.; West, K.G.; Farrar, H.E.; Ball, A.; Harris, B.T.; Stommel, E.W. A cluster of amyotrophic lateral sclerosis in New Hampshire: A possible role for toxic cyanobacteria blooms. Amyotroph. Lateral Scler. 2009, 10, 101–108. [Google Scholar] [CrossRef]
- Banack, S.A.; Caller, T.; Henegan, P.; Haney, J.; Murby, A.; Metcalf, J.S.; Powell, J.; Cox, P.A.; Stommel, E. Detection of Cyanotoxins, β-N-methylamino-l-alanine and Microcystins, from a Lake Surrounded by Cases of Amyotrophic Lateral Sclerosis. Toxins 2015, 7, 322–336. [Google Scholar] [CrossRef]
- Torbick, N.; Ziniti, B.; Stommel, E.; Linder, E.; Andrew, A.; Caller, T.; Haney, J.; Bradley, W.; Henegan, P.L.; Shi, X. Assessing Cyanobacterial Harmful Algal Blooms as Risk Factors for Amyotrophic Lateral Sclerosis. Neurotox. Res. 2018, 33, 199–212. [Google Scholar] [CrossRef]
- Violi, J.P.; Mitrovic, S.M.; Colville, A.; Main, B.J.; Rodgers, K.J. Prevalence of β-methylamino-L-alanine (BMAA) and its isomers in freshwater cyanobacteria isolated from eastern Australia. Ecotoxicol. Environ. Saf. 2019, 172, 72–81. [Google Scholar] [CrossRef]
- Facciponte, D.N.; Bough, M.W.; Seidler, D.; Carroll, J.L.; Ashare, A.; Andrew, A.S.; Tsongalis, G.J.; Vaickus, L.J.; Henegan, P.L.; Butt, T.H.; et al. Identifying aerosolized cyanobacteria in the human respiratory tract: A proposed mechanism for cyanotoxin-associated diseases. Sci. Total. Environ. 2018, 645, 1003–1013. [Google Scholar] [CrossRef]
- Faassen, E.J. Presence of the Neurotoxin BMAA in Aquatic Ecosystems: What Do We Really Know? Toxins 2014, 6, 1109–1138. [Google Scholar] [CrossRef]
- Cohen, S.A. Analytical techniques for the detection of α-amino-β-methylaminopropionic acid. Analyst 2012, 137, 1991. [Google Scholar] [CrossRef] [PubMed]
- Main, B.J.; Bowling, L.C.; Padula, M.P.; Bishop, D.P.; Mitrovic, S.M.; Guillemin, G.J.; Rodgers, K.J. Detection of the suspected neurotoxin β-methylamino-l-alanine (BMAA) in cyanobacterial blooms from multiple water bodies in Eastern Australia. Harmful Algae 2018, 74, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Cox, P.A.; Banack, S.A.; Murch, S.J.; Rasmussen, U.; Tien, G.; Bidigare, R.R.; Metcalf, J.S.; Morrison, L.F.; Codd, G.A.; Bergman, B. Diverse taxa of cyanobacteria produce β-N-methylamino-l-alanine, a neurotoxic amino acid. Proc. Natl. Acad. Sci. USA 2005, 102, 5074–5078. [Google Scholar] [CrossRef] [PubMed]
- Faassen, E.J.; Gillissen, F.; Lürling, M. A Comparative Study on Three Analytical Methods for the Determination of the Neurotoxin BMAA in Cyanobacteria. PLoS ONE 2012, 7, e36667. [Google Scholar] [CrossRef] [PubMed]
- Réveillon, D.; Abadie, E.; Séchet, V.; Brient, L.; Savar, V.; Bardouil, M.; Hess, P.; Amzil, Z. Beta-N-methylamino-l-alanine: LC-MS/MS Optimization, Screening of Cyanobacterial Strains and Occurrence in Shellfish from Thau, a French Mediterranean Lagoon. Mar. Drugs 2014, 12, 5441–5467. [Google Scholar] [CrossRef] [PubMed]
- Rzymski, P.; Poniedziałek, B.; Mankiewicz-Boczek, J.; Faassen, E.J.; Jurczak, T.; Gągała-Borowska, I.; Ballot, A.; Lürling, M.; Kokociński, M. Polyphasic toxicological screening of Cylindrospermopsis raciborskii and Aphanizomenon gracile isolated in Poland. Algal Res. 2017, 24, 72–80. [Google Scholar] [CrossRef]
- Banack, S.A.; Metcalf, J.S.; Jiang, L.; Craighead, D.; Ilag, L.L.; Cox, P.A. Cyanobacteria Produce N-(2-Aminoethyl) Glycine, a Backbone for Peptide Nucleic Acids Which May Have Been the First Genetic Molecules for Life on Earth. PLoS ONE 2012, 7, e49043. [Google Scholar] [CrossRef]
- Lage, S.; Ström, L.; Godhe, A.; Rydberg, S. The effect of exogenous β-N-methylamino- l -alanine (BMAA) on the diatoms Phaeodactylum tricornutum and Thalassiosira weissflogii. Harmful Algae 2016, 58, 85–92. [Google Scholar] [CrossRef]
- Lage, S.; Ström, L.; Godhe, A.; Rydberg, S. Kinetics of β-N-methylamino-L-alanine (BMAA) and 2, 4-diaminobutyric acid (DAB) production by diatoms: The effect of nitrogen. Eur. J. Phycol. 2018, 54, 115–125. [Google Scholar] [CrossRef]
- Réveillon, D.; Séchet, V.; Hess, P.; Amzil, Z. Production of BMAA and DAB by diatoms (Phaeodactylum tricornutum, Chaetoceros sp., Chaetoceros calcitrans and, Thalassiosira pseudonana ) and bacteria isolated from a diatom culture. Harmful Algae 2016, 58, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Orsini, L.; Sarno, D.; Procaccini, G.; Poletti, R.; Dahlmann, J.; Montresor, M. Toxic Pseudo-nitzschia multistriata (Bacillariophyceae) from the Gulf of Naples: Morphology, toxin analysis and phylogenetic relationships with other Pseudo-nitzschia species. Eur. J. Phycol. 2002, 37, 247–257. [Google Scholar] [CrossRef]
- Esterhuizen, M.; Downing, T. β-N-methylamino-l-alanine (BMAA) in novel South African cyanobacterial isolates. Ecotoxicol. Environ. Saf. 2008, 71, 309–313. [Google Scholar] [CrossRef]
- Metcalf, J.S.; Banack, S.A.; Lindsay, J.; Morrison, L.F.; Cox, P.A.; Codd, G.A. Co-occurrence of β-N-methylamino-l-alanine, a neurotoxic amino acid with other cyanobacterial toxins in British waterbodies, 1990–2004. Environ. Microbiol. 2008, 10, 702–708. [Google Scholar] [CrossRef] [PubMed]
- Cervantes Cianca, R.C.; Baptista, M.S.; Lopes, V.R.; Vasconcelos, V.M. The non-protein amino acid beta-N-methylamino-L-alanine in Portuguese cyanobacterial isolates. Amino Acids 2012, 42, 2473–2479. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Tian, Z.; Li, J.; Yu, R.; Banack, S.A.; Wang, Z. Detection of the neurotoxin BMAA within cyanobacteria isolated from freshwater in China. Toxicon 2010, 55, 947–953. [Google Scholar] [CrossRef] [PubMed]
- Downing, S.; Banack, S.; Metcalf, J.; Cox, P.; Downing, T.; Metcalf, J. Nitrogen starvation of cyanobacteria results in the production of β-N-methylamino-L-alanine. Toxicon 2011, 58, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-H.; Flory, W.; Koeppe, R.E. Variation of neurotoxicity of l- and d-2,4-diaminobutyric acid with route of administration. Toxicol. Appl. Pharmacol. 1972, 23, 334–338. [Google Scholar] [CrossRef]
- Perkins, H.R.; Cummins, C.S. Chemical structure of bacterial cell walls. ornithine and 2,4-diaminobutyric acid as components of the cell walls of plant pathogenic corynebacteria. Nature 1964, 201, 1105–1107. [Google Scholar] [CrossRef]
- Suzuki, K.; Sasaki, J.; Uramoto, M.; Nakase, T.; Komagata, K. Agromyces mediolanus sp. nov., nom. rev., comb. nov., a species for “Corynebacterium mediolanum” Mamoli 1939 and for some aniline-assimilating bacteria which contain 2,4-diaminobutyric acid in the cell wall peptidoglycan. Int. J. Syst. Bacteriol. 1996, 46, 88–93. [Google Scholar] [CrossRef]
- Mitrovic, S.M.; Chessman, B.C.; Davie, A.; Avery, E.L.; Ryan, N. Development of blooms of Cyclotella meneghiniana and Nitzschia spp. (Bacillariophyceae) in a shallow river and estimation of effective suppression flows. Hydrobiologia 2008, 596, 173–185. [Google Scholar] [CrossRef]
- De Sève, M.A. Diatom bloom in the tidal freshwater zone of a turbid and shallow estuary, Rupert Bay (James Bay, Canada). Hydrobiologia 1993, 269, 225–233. [Google Scholar] [CrossRef]
- Henriques Vieira, A.A.; Coelho Ortolano, P.I.; Giroldo, D.; Dellamano Oliveira, M.J.; Bittar, T.B.; Lombardi, A.T.; Sartori, A.L.; Paulsen, B.S. Role of hydrophobic extracellular polysaccharide of Aulacoseira granulata (Bacillariophyceae) on aggregate formation in a turbulent and hypereutrophic reservoir. Limnol. Oceanogr. 2008, 53, 1887–1899. [Google Scholar] [CrossRef]
- Bolch, C.J.S.; Blackburn, S.I. Isolation and purification of Australian isolates of the toxic cyanobacteriumMicrocystis aeruginosa Kütz. Environ. Biol. Fishes 1996, 8, 5–13. [Google Scholar] [CrossRef]
- UTEX Culture Collection of Algae. BG-11 Medium Recipe. Available online: https://utex.org/products/bg-11-medium (accessed on 4 June 2019).
- UTEX Culture Collection of Algae. BG-11 Trace Metals Solution Recipe. Available online: https://utex.org/products/bg-11-trace-metals-solution-recipe (accessed on 4 June 2019).
| Genera | Location | Free BMAA (ng/g DW) | Bound BMAA (ng/g DW) | Free BMAA Per Cell (fg/Cell) | Bound BMAA Per Cell (fg/Cell) |
|---|---|---|---|---|---|
| Aulacoseira | Nepean River | ND | 40.28 ± 0.83 | ND | 0.89 ± 0.02 |
| Cyclotella | Lake Liddell | 103.58 ± 1.94 | 154.06 ± 4.94 | 2.69 ± 0.05 | 4.01 ± 0.12 |
| Fragilaria | Murrumbidgee River | ND | ND | ND | ND |
| Navicula | Lostock Dam | 151.2 ± 10.02 | 369.64 ± 11.96 | 1.82 ± 0.12 | 4.45 ± 0.14 |
| Tabellaria | Spencers Creek | ND | 19.99 ± 1.39 | ND | 0.34 ± 0.02 |
| Genera | Location | Free AEG (ng/g DW) | Bound AEG (ng/g DW) | Free AEG Per Cell (fg/Cell) | Bound AE Per Cell (fg/Cell) |
|---|---|---|---|---|---|
| Aulacoseira | Nepean River | 49.55 ± 3.29 | 196.17 ± 4.93 | 1.1 ± 0.07 | 4.38 ± 0.11 |
| Cyclotella | Lake Liddell | ND | ND | ND | ND |
| Fragilaria | Murrumbidgee River | 323.05 ± 28.75 | 1162.12 ± 43.97 | 3.64 ± 0.32 | 13.09 ± 0.49 |
| Navicula | Lostock Dam | 563.5 ± 1.57 | 1328.11 ± 25.31 | 6.78 ± 0.02 | 16 ± 0.30 |
| Tabellaria | Spencers Creek | 124.83 ± 0.7 | 150.19 ± 1.12 | 2.15 ± 0.01 | 2.59 ± 0.02 |
| Genera | Location | Free 2,4-DAB (ng/g DW) | Bound 2,4-DAB (ng/g DW) | Free 2,4-DAB Per Cell (fg/Cell) | Bound 2,4-DAB Per Cell (fg/Cell) |
|---|---|---|---|---|---|
| Aulacoseira | Nepean River | 103.27 ± 9.96 | ND | 2.3 ± 0.22 | ND |
| Cyclotella | Lake Liddell | 113.7 ± 2.13 | 169.12 ± 5.43 | 2.96 ± 0.05 | 4.4 ± 0.14 |
| Fragilaria | Murrumbidgee River | 258.57 ± 24.06 | ND | 2.91 ± 0.27 | ND |
| Navicula | Lostock Dam | 594.81 ± 53.22 | 4678.75 ± 83.2 | 7.16 ± 0.64 | 56.37 ± 1.0 |
| Tabellaria | Spencers Creek | 146.16 ± 8.8 | 248.48 ± 17.43 | 2.52 ± 0.15 | 4.28 ± 0.30 |
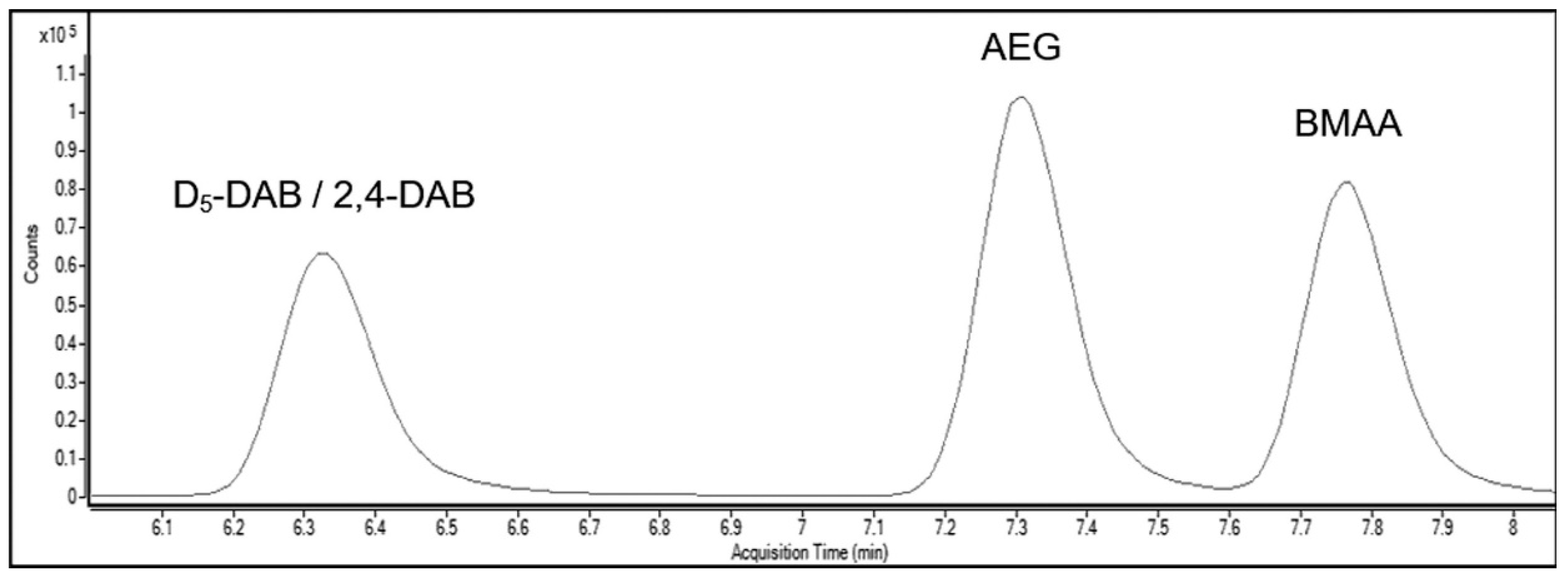
| Amino Acid | Retention Time (min) | Precursor Ion (m/z) | Product Ion (m/z) |
|---|---|---|---|
| BMAA | 7.80 | 333 | 73 99.1 159.1 187.1 * |
| 2,4-DAB | 6.30 | 333 | 56 99 187.1 273.1 * |
| AEG | 7.30 | 333 | 56.1 88* 99 187.1 |
| D5-DAB | 6.30 | 339 | 102.9 103.9 192 278.1 * |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Violi, J.P.; Facey, J.A.; Mitrovic, S.M.; Colville, A.; Rodgers, K.J. Production of β-methylamino-L-alanine (BMAA) and Its Isomers by Freshwater Diatoms. Toxins 2019, 11, 512. https://doi.org/10.3390/toxins11090512
Violi JP, Facey JA, Mitrovic SM, Colville A, Rodgers KJ. Production of β-methylamino-L-alanine (BMAA) and Its Isomers by Freshwater Diatoms. Toxins. 2019; 11(9):512. https://doi.org/10.3390/toxins11090512
Chicago/Turabian StyleVioli, Jake P., Jordan A. Facey, Simon M. Mitrovic, Anne Colville, and Kenneth J. Rodgers. 2019. "Production of β-methylamino-L-alanine (BMAA) and Its Isomers by Freshwater Diatoms" Toxins 11, no. 9: 512. https://doi.org/10.3390/toxins11090512
APA StyleVioli, J. P., Facey, J. A., Mitrovic, S. M., Colville, A., & Rodgers, K. J. (2019). Production of β-methylamino-L-alanine (BMAA) and Its Isomers by Freshwater Diatoms. Toxins, 11(9), 512. https://doi.org/10.3390/toxins11090512






