Dianthin and Its Potential in Targeted Tumor Therapies
Abstract
1. Introduction
2. Structure and Function
2.1. Purification and Expression of Dianthins
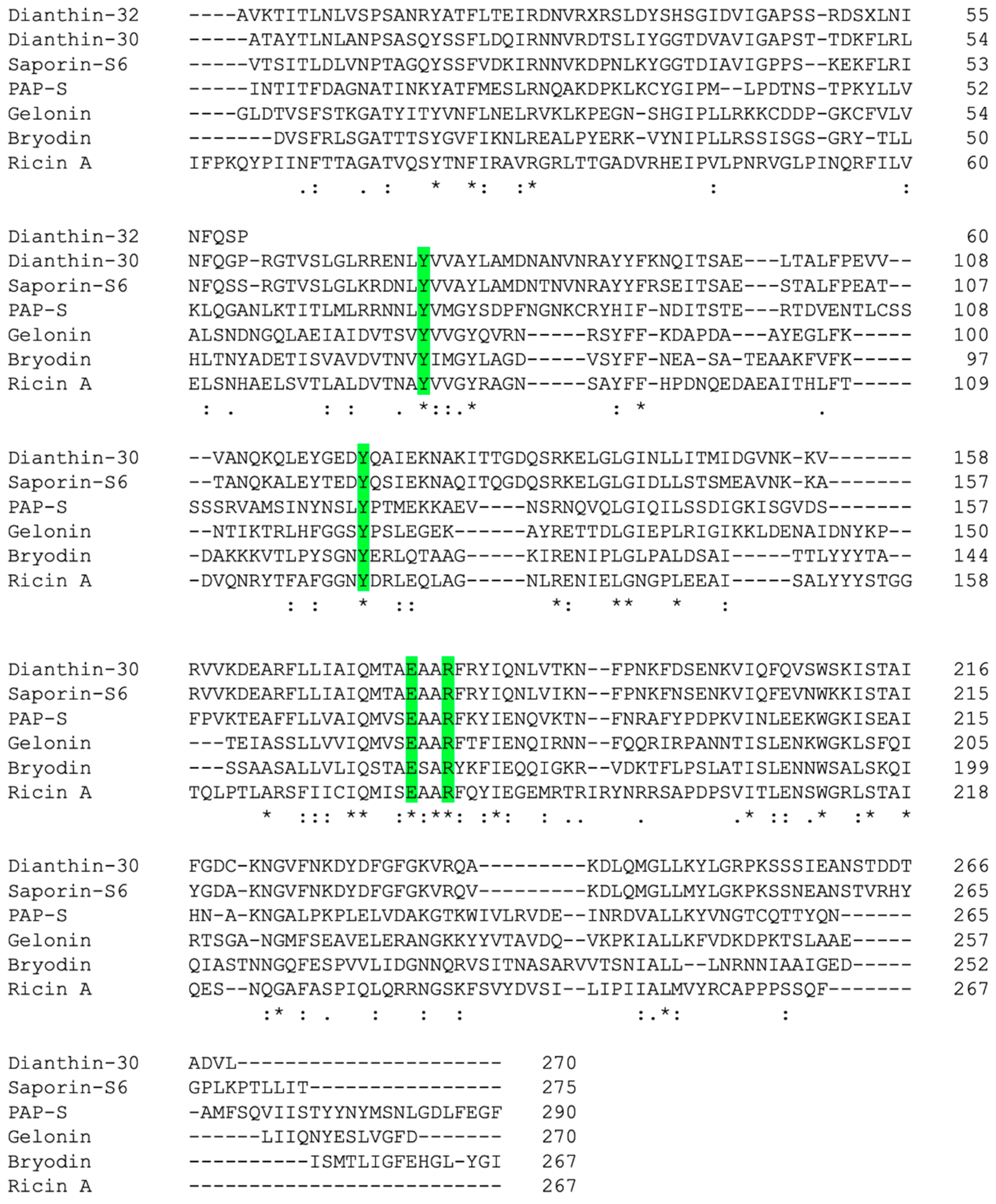
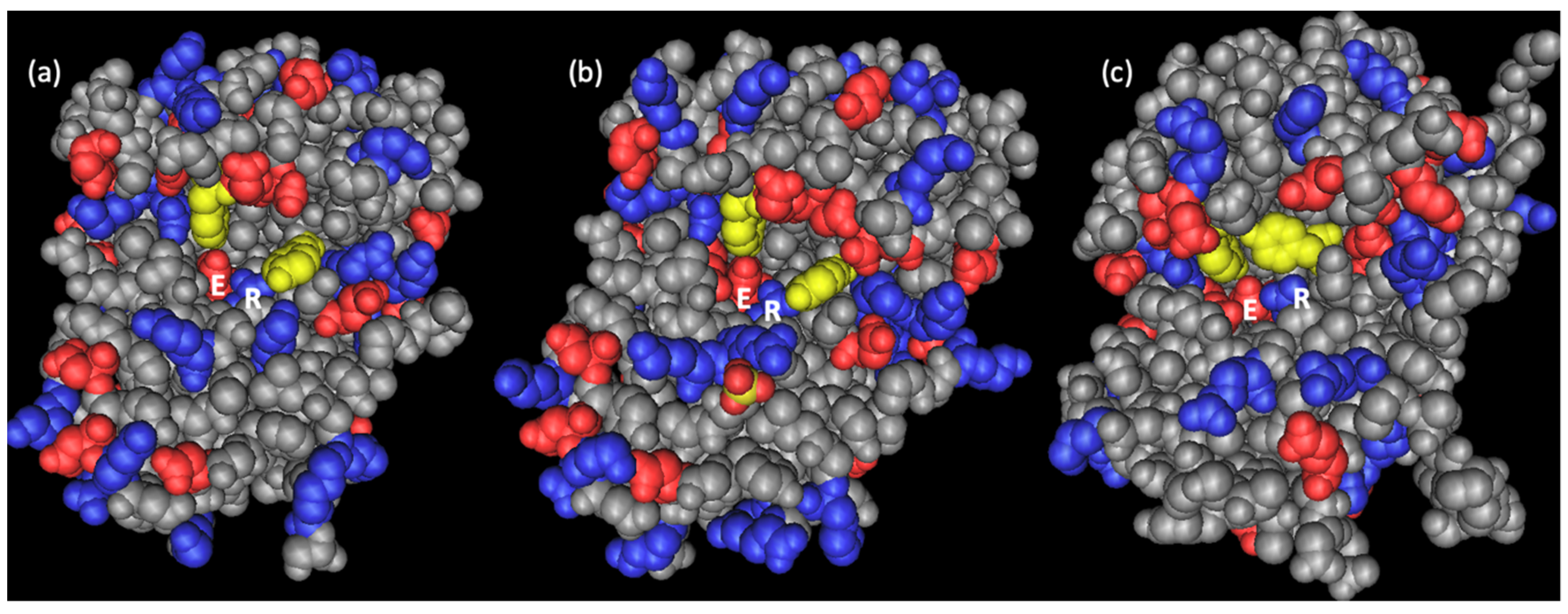
2.2. Primary and Spatial Structure of Dianthin-30 and Its Catalytic Center
2.3. Enzymatic Activity and Biological Function
3. Dianthin Conjugates and Fusion Proteins
4. Endosomal Escape
5. Mouse Tumor Models
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Stirpe, F.; Williams, D.G.; Onyon, L.J.; Legg, R.F.; A Stevens, W. Dianthins, ribosome-damaging proteins with anti-viral properties from Dianthus caryophyllus L. (carnation). Biochem. J. 1981, 195, 399–405. [Google Scholar] [CrossRef]
- Prestle, J.; Hornung, E.; Schönfelder, M.; Mundry, K.-W. Mechanism and site of action of a ribosome-inactivating protein type 1 from Dianthus barbatus which inactivates Escherichia coli ribosomes. FEBS Lett. 1992, 297, 250–252. [Google Scholar] [CrossRef]
- Wang, Y.-C.; Tan, N.-H.; Zhou, J.; Wu, H.-M. Cyclopeptides from dianthus superbus. Phytochemistry 1998, 49, 1453–1456. [Google Scholar] [CrossRef]
- Hsieh, P.-W.; Chang, F.-R.; Wu, C.-C.; Wu, K.-Y.; Li, C.-M.; Chen, S.-L.; Wu, Y.-C. New Cytotoxic Cyclic Peptides and Dianthramide from Dianthus superbus. J. Nat. Prod. 2004, 67, 1522–1527. [Google Scholar] [CrossRef]
- Amso, Z.; Kowalczyk, R.; Park, Y.-E.; Watson, M.; Lin, J.-M.; Musson, D.S.; Cornish, J.; Brimble, M.A. Synthesis and in vitro bone cell activity of analogues of the cyclohexapeptide dianthin G. Org. Biomol. Chem. 2016, 14, 6231–6243. [Google Scholar] [CrossRef] [PubMed]
- Tong, Y.; Luo, J.-G.; Wang, R.; Wang, X.-B.; Kong, L.-Y. New cyclic peptides with osteoblastic proliferative activity from Dianthus superbus. Bioorg. Med. Chem. Lett. 2012, 22, 1908–1911. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Huang, M.; Wang, Z.; Zheng, Y.; Zeng, G.; He, W.; Tan, N. Cyclopentapeptides from Dianthus chinensis. J. Pept. Sci. 2015, 21, 550–553. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Amso, Z.; Rodriguez, L.M.D.L.; Kaur, H.; Brimble, M.A.; Rodriguez, L.M.D.L. Synthesis of Natural Cyclopentapeptides Isolated from Dianthus chinensis. J. Nat. Prod. 2016, 79, 1769–1774. [Google Scholar] [CrossRef] [PubMed]
- Endo, Y.; Mitsui, K.; Motizuki, M.; Tsurugi, K. The mechanism of action of ricin and related toxic lectins on eukaryotic ribosomes. The site and the characteristics of the modification in 28 S ribosomal RNA caused by the toxins. J. Boil. Chem. 1987, 262, 5908–5912. [Google Scholar]
- Endo, Y.; Tsurugi, K.; Lambert, J.M. The site of action of six different ribosome-inactivating proteins from plants on eukaryotic ribosomes: The RNA N-glycosidase activity of the proteins. Biochem. Biophys. Res. Commun. 1988, 150, 1032–1036. [Google Scholar] [CrossRef]
- Cho, H.-J.; Lee, S.-J.; Kim, S.; Kim, B.-D. Isolation and Characterization of cDNAs Encoding Ribosome Inactivating Protein from Dianthus sinensis L. Mol. Cells 2000, 10, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Ferreras, J.M.; Iglesias, R.; Barbieri, L.; Alegre, C.; Bolognesi, A.; Rojo, M.A.; Carbajales, M.L.; Escarmis, C.; Girbes, T. Effects and molecular action of ribosome-inactivating proteins on ribosomes from Streptomyces lividans. Biochim. Biophys. Acta (BBA) Gen. Subj. 1995, 1243, 85–93. [Google Scholar] [CrossRef]
- Gilabert-Oriol, R.; Weng, A.; Trautner, A.; Weise, C.; Schmid, D.; Bhargava, C.; Niesler, N.; Wookey, P.J.; Fuchs, H.; Thakur, M. Combinatorial approach to increase efficacy of Cetuximab, Panitumumab and Trastuzumab by dianthin conjugation and co-application of SO1861. Biochem. Pharmacol. 2015, 97, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Legname, G.; Fossati, G.; Monzini, N.; Gromo, G.; Marcucci, F.; Mascagni, P.; Modena, D. Heterologous expression, purification, activity and conformational studies of different forms of dianthin 30. Biomed. Pept. Proteins Nucleic Acids Struct. Synth. Boil. Act. 1995, 1, 61–68. [Google Scholar]
- Girbés, T.; Barbieri, L.; Ferreras, M.; Arias, F.J.; Rojo, M.A.; Alegre, C.; Stirpe, F.; Iglesias, R.; Escarmis, C. Effects of ribosome-inactivating proteins on Escherichia coli and Agrobacterium tumefaciens translation systems. J. Bacteriol. 1993, 175, 6721–6724. [Google Scholar] [CrossRef] [PubMed]
- Schrot, J.; Weng, A.; Melzig, M.F. Ribosome-Inactivating and Related Proteins. Toxins 2015, 7, 1556–1615. [Google Scholar] [CrossRef]
- Gilabert-Oriol, R.; Weng, A.; Von Mallinckrodt, B.; Melzig, M.F.; Fuchs, H.; Thakur, M. Immunotoxins Constructed with Ribosome-Inactivating Proteins and their Enhancers: A Lethal Cocktail with Tumor Specific Efficacy. Curr. Pharm. Des. 2014, 20, 6584–6643. [Google Scholar] [CrossRef]
- Rust, A.; Partridge, L.J.; Davletov, B.; Hautbergue, G.M. The Use of Plant-Derived Ribosome Inactivating Proteins in Immunotoxin Development: Past, Present and Future Generations. Toxins 2017, 9, 344. [Google Scholar] [CrossRef]
- Bortolotti, M.; Bolognesi, A.; Polito, L. Bouganin, an Attractive Weapon for Immunotoxins. Toxins 2018, 10, 323. [Google Scholar] [CrossRef]
- Polito, L.; Bortolotti, M.; Mercatelli, D.; Battelli, M.G.; Bolognesi, A. Saporin-S6: A Useful Tool in Cancer Therapy. Toxins 2013, 5, 1698–1722. [Google Scholar] [CrossRef]
- Fabbrini, M.S.; Rappocciolo, E.; Carpani, D.; Solinas, M.; Valsasina, B.; Breme, U.; Cavallaro, U.; Nykjaer, A.; Rovida, E.; Legname, G.; et al. Characterization of a saporin isoform with lower ribosome-inhibiting activity. Biochem. J. 1997, 322, 719–727. [Google Scholar] [CrossRef] [PubMed]
- Olsnes, S.; Pihl, A. Different biological properties of the two constituent peptide chains of ricin a toxic protein inhibiting protein synthesis. Biochemistry 1973, 12, 3121–3126. [Google Scholar] [CrossRef] [PubMed]
- Falasca, A.; Gasperi-Campani, A.; Abbondanza, A.; Barbieri, L.; Stirpe, F. Properties of the ribosome-inactivating proteins gelonin, Momordica charantia inhibitor, and dianthins. Biochem. J. 1982, 207, 505–509. [Google Scholar] [CrossRef] [PubMed]
- Reisbig, R.R.; Bruland, Ø.S. Dianthin 30 and 32 from Dianthus caryophyllus: Two inhibitors of plant protein synthesis and their tissue distribution. Arch. Biochem. Biophys. 1983, 224, 700–706. [Google Scholar] [CrossRef]
- Strocchi, P.; Barbieri, L.; Stirpe, F. Immunological properties of ribosome-inactivating proteins and a saporin immunotoxin. J. Immunol. Methods 1992, 155, 57–63. [Google Scholar] [CrossRef]
- Di Maro, A.; Valbonesi, P.; Bolognesi, A.; Stirpe, F.; De Luca, P.; Gigliano, G.S.; Gaudio, L.; Bovi, P.D.; Ferranti, P.; Malorni, A.; et al. Isolation and characterization of four type-1 ribosome-inactivating proteins, with polynucleotide:adenosine glycosidase activity, from leaves of Phytolacca dioica L. Planta 1999, 208, 125–131. [Google Scholar] [CrossRef]
- Bolognesi, A.; Olivieri, F.; Battelli, M.G.; Barbieri, L.; Falasca, A.I.; Parente, A.; Blanco, F.V.; Stirpe, F. Ribosome-Inactivating Proteins (RNA N-glycosidases) from the Seeds of Saponaria ocymoides and Vaccaria pyramidata. JBIC J. Boil. Inorg. Chem. 1995, 228, 935–940. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, A.; Polito, L.; Olivieri, F.; Valbonesi, P.; Barbieri, L.; Battelli, M.G.; Carusi, M.V.; Benvenuto, E.; Del Vecchio Blanco, F.; Di Maro, A.; et al. New ribosome-inactivating proteins with polynucleotide:Adenosine glycosidase and antiviral activities from Basella rubra L. And bougainvillea spectabilis willd. Planta 1997, 203, 422–429. [Google Scholar] [CrossRef]
- Barbieri, L.; Polito, L.; Bolognesi, A.; Ciani, M.; Pelosi, E.; Farini, V.; Jha, A.; Sharma, N.; Vivanco, J.; Chambery, A. Ribosome-inactivating proteins in edible plants and purification and characterization of a new ribosome-inactivating protein from Cucurbita moschata. Biochim. Biophys. Acta (BBA) Gen. Subj. 2006, 1760, 783–792. [Google Scholar] [CrossRef]
- Lappi, D.A.; Esch, F.S.; Barbieri, L.; Stirpe, F.; Soria, M. Characterization of a Saponaria officinalis seed ribosome-inactivating protein: Immunoreactivity and sequence homologies. Biochem. Biophys. Res. Commun. 1985, 129, 934–942. [Google Scholar] [CrossRef]
- Porro, G.; Bonardi, M.; Giovanetti, E.; Lento, P.; Modena, D. Production and Characterization of Monoclonal Antibodies Against the Ribosome Inactivating Proteins Dianthin32 and Momochin. Hybridoma 1994, 13, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Szalai, K.; Schöll, I.; Polito, L.; Bolognesi, A.; Untersmayr, E.; Riemer, A.B.; Stirpe, F.; Forster-Waldl, E.; Riemer, A.; Boltz-Nitulescu, G.; et al. Occupational sensitization to ribosome-inactivating proteins in researchers. Clin. Exp. Allergy 2005, 35, 1354–1360. [Google Scholar] [CrossRef] [PubMed]
- Legname, G.; Gromo, G.; Lord, J.; Monzini, N.; Modena, D. Expression and Activity of Pre-dianthin 30 and Dianthin 30. Biochem. Biophys. Res. Commun. 1993, 192, 1230–1237. [Google Scholar] [CrossRef] [PubMed]
- Hartley, M.R.; Legname, G.; Osborn, R.; Chen, Z.; Lord, J. Single-chain ribosome inactivating proteins from plants depurinate Escherichia coli 23S ribosomal RNA. FEBS Lett. 1991, 290, 65–68. [Google Scholar] [CrossRef]
- Legname, G.; Bellosta, P.; Gromo, G.; Modena, D.; Keen, J.N.; Roberts, L.M.; Lord, J.M. Nucleotide sequence of cDNA coding for dianthin 30, a ribosome inactivating protein from Dianthus caryophyllus. Biochim. Biophys. Acta (BBA) Gene Struct. Expr. 1991, 1090, 119–122. [Google Scholar] [CrossRef]
- Kokorin, A.; Weise, C.; Sama, S.; Weng, A. A new type 1 ribosome-inactivating protein from the seeds of Gypsophila elegans M.Bieb. Phytochemistry 2019, 157, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Bednarek, S.Y.; Wilkins, T.A.; Dombrowski, J.E.; Raikhel, N.V. A Carboxyl-Terminal Propeptide Is Necessary for Proper Sorting of Barley Lectin to Vacuoles of Tobacco. Plant Cell 1990, 2, 1145. [Google Scholar] [PubMed]
- Fermani, S.; Falini, G.; Ripamonti, A.; Polito, L.; Stirpe, F.; Bolognesi, A. The 1.4 anstroms structure of dianthin 30 indicates a role of surface potential at the active site of type 1 ribosome inactivating proteins. J. Struct. Biol. 2005, 149, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef]
- Bravi, G.; Legname, G.; Chan, A. Substrate recognition by ribosome-inactivating protein studied by molecular modeling and molecular electrostatic potentials. J. Mol. Graph. 1995, 13, 83–88. [Google Scholar] [CrossRef]
- Kurinov, I.V.; Rajamohan, F.; Uckun, F.M. High resolution x-ray structure and potent anti-hiv activity of recombinant dianthin antiviral protein. Arzneimittelforschung 2004, 54, 692–702. [Google Scholar] [CrossRef] [PubMed]
- Fermani, S.; Falini, G.; Ripamonti, A.; Bolognesi, A.; Polito, L.; Stirpe, F. Crystallization and preliminary X-ray diffraction analysis of two ribosome-inactivating proteins: Lychnin and dianthin 30. Acta Crystallogr. Sect. D Boil. Crystallogr. 2003, 59, 1227–1229. [Google Scholar] [CrossRef] [PubMed]
- Fermani, S.; Tosi, G.; Farini, V.; Polito, L.; Falini, G.; Ripamonti, A.; Barbieri, L.; Chambery, A.; Bolognesi, A. Structure/function studies on two type 1 ribosome inactivating proteins: Bouganin and lychnin. J. Struct. Boil. 2009, 168, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Gawlak, S.L.; Neubauer, M.; Klei, H.E.; Chang; Einspahr, H.M.; Siegall, C.B. Molecular, Biological, and Preliminary Structural Analysis of Recombinant Bryodin 1, a Ribosome-Inactivating Protein from the PlantBryonia dioica. Biochemistry 1997, 36, 3095–3103. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.C.; Kung, H.F.; Huang, P.L.; I Huang, H.; Lee-Huang, S. Anti-HIV plant proteins catalyze topological changes of DNA into inactive forms. BioFactors 1992, 4, 37–41. [Google Scholar]
- Lee-Huang, S.; Kung, H.-F.; Huang, P.L.; Huang, P.L.; Li, B.-Q.; Huang, P.; Huang, H.I.; Chen, H.-C. A new class of anti-HIV agents: GAP31, DAPs 30 and 32. FEBS Lett. 1991, 291, 139–144. [Google Scholar] [CrossRef]
- Savino, C.; Federici, L.; Ippoliti, R.; Lendaro, E.; Tsernoglou, D. The crystal structure of saporin SO6 from Saponaria officinalis and its interaction with the ribosome. FEBS Lett. 2000, 470, 239–243. [Google Scholar] [CrossRef]
- Mlsna, D.; Monzingo, A.F.; Katzin, B.J.; Ernst, S.; Robertus, J.D. Structure of recombinant ricin a chain at 2.3 a. Protein Sci. 1993, 2, 429–435. [Google Scholar] [CrossRef]
- Xu, Y.-Z.; Liu, W.-Y. Effects of the Active Aldehyde Group Generated by RNA N-Glycosidase in the Sarcin/Ricin Domain of Rat 28S Ribosomal RNA on Peptide Elongation. Boil. Chem. 2000, 381, 113–119. [Google Scholar] [CrossRef]
- Harley, S.M.; Beevers, H. Ricin inhibition of in vitro protein synthesis by plant ribosomes. Proc. Natl. Acad. Sci. USA 1982, 79, 5935–5938. [Google Scholar] [CrossRef]
- Lappi, D.A.; Ying, W.; Barthelemy, I.; Martineau, D.; Prieto, I.; Benatti, L.; Soria, M.; Baird, A. Expression and activities of a recombinant basic fibroblast growth factor-saporin fusion protein. J. Boil. Chem. 1994, 269, 12552–12558. [Google Scholar]
- Taylor, S.; Massiah, A.; Lomonossoff, G.; Roberts, L.M.; Lord, J.M.; Hartley, M. Correlation between the activities of five ribosome-inactivating proteins in depurination of tobacco ribosomes and inhibition of tobacco mosaic virus infection. Plant J. 1994, 5, 827–835. [Google Scholar] [CrossRef] [PubMed]
- Zarovni, N.; Vago, R.; Solda, T.; Monaco, L.; Fabbrini, M.S. Saporin as a novel suicide gene in anticancer gene therapy. Cancer Gene Ther. 2007, 14, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, A.; Tazzari, P.L.; Legname, G.; Olivieri, F.; Modena, D.; Conte, R.; Stirpe, F.; Legname, G. Anti-CD30 immunotoxins with native and recombinant dianthin 30. Cancer Immunol. Immunother. 1995, 40, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Lorenzetti, I.; Meneguzzi, A.; Fracasso, G.; Potrich, C.; Costantini, L.; Chiesa, E.; Legname, G.; Menestrina, G.; Tridente, G.; Colombatti, M. Genetic grafting of membrane-acting peptides to the cytotoxin dianthin augments its ability to de-stabilize lipid bilayers and enhances its cytotoxic potential as the component of transferrin-toxin conjugates. Int. J. Cancer 2000, 86, 582–589. [Google Scholar] [CrossRef]
- Cenini, P.; Bolognesi, A.; Stirpe, F. Ribosome-inactivating proteins from plants inhibit ribosome activity of Trypanosoma and Leishmania. J. Protozool. 1988, 35, 384–387. [Google Scholar] [CrossRef]
- Bolognesi, A.; Tazzari, P.L.; Tassi, C.; Gromo, G.; Gobbi, M.; Stirpe, F. A comparison of anti-lymphocyte immunotoxins containing different ribosome-inactivating proteins and antibodies. Clin. Exp. Immunol. 1992, 89, 341–346. [Google Scholar] [CrossRef]
- Cenini, P.; Battelli, M.G.; Bolognesi, A.; Stirpe, F.; Villemez, C.L. Effect of ribosome-inactivating proteins on ribosomes from Tetrahymena pyriformis and Acanthamoeba castellanii. Biochem. Biophys. Res. Commun. 1987, 148, 521–527. [Google Scholar] [CrossRef]
- Schlegel, R.; Wade, M. A synthetic peptide corresponding to the NH2 terminus of vesicular stomatitis virus glycoprotein is a pH-dependent hemolysin. J. Boil. Chem. 1984, 259, 4691–4694. [Google Scholar]
- Wagner, E.; Plank, C.; Zatloukal, K.; Cotten, M.; Birnstiel, M.L. Influenza virus hemagglutinin HA-2 N-terminal fusogenic peptides augment gene transfer by transferrin-polylysine-DNA complexes: Toward a synthetic virus-like gene-transfer vehicle. Proc. Natl. Acad. Sci. USA 1992, 89, 7934–7938. [Google Scholar] [CrossRef]
- Pécheur, E.I.; Sainte-Marie, J.; Bienvene, A.; Hoekstra, D. Peptides and membrane fusion: Towards an understanding of the molecular mechanism of protein-induced fusion. J. Membr. Biol. 1999, 167, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, L.; Gorini, P.; Valbonesi, P.; Castiglioni, P.; Stirpe, F. Unexpected activity of saporins. Nature 1994, 372, 624. [Google Scholar] [CrossRef] [PubMed]
- Roncuzzi, L.; Gasperi-Campani, A. DNA-nuclease activity of the single-chain ribosome-inactivating proteins dianthin 30, saporin 6 and gelonin. FEBS Lett. 1996, 392, 16–20. [Google Scholar] [CrossRef]
- Panayotatos, N.; Wells, R.D. Cruciform structures in supercoiled DNA. Nature 1981, 289, 466–470. [Google Scholar] [CrossRef] [PubMed]
- De Benito, F.M.; Citores, L.; Iglesias, R.; Ferreras, J.; Camafeita, E.; Méndez, E.; Girbés, T. Isolation and partial characterization of a novel and uncommon two-chain 64-kDa ribosome-inactivating protein from the bark of elder (Sambucus nigra L.). FEBS Lett. 1997, 413, 85–91. [Google Scholar] [CrossRef]
- De Benito, F.M.; Iglesias, R.; Ferreras, J.M.; Citores, L.; Camafeita, E.; Méndez, E.; Girbés, T. Constitutive and inducible type 1 ribosome-inactivating proteins (RIPs) in elderberry (Sambucus nigra L.). FEBS Lett. 1998, 428, 75–79. [Google Scholar] [CrossRef]
- Heisler, I.; Keller, J.; Tauber, R.; Sutherland, M.; Fuchs, H. A Colorimetric Assay for the Quantitation of Free Adenine Applied to Determine the Enzymatic Activity of Ribosome-Inactivating Proteins. Anal. Biochem. 2002, 302, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Weng, A. A novel adenine-releasing assay for ribosome-inactivating proteins. J. Chromatogr. B 2018, 1072, 300–304. [Google Scholar] [CrossRef] [PubMed]
- Weng, A.; Thakur, M.; Beceren-Braun, F.; Bachran, D.; Bachran, C.; Riese, S.B.; Jenett-Siems, K.; Gilabert-Oriol, R.; Melzig, M.F.; Fuchs, H. The toxin component of targeted anti-tumor toxins determines their efficacy increase by saponins. Mol. Oncol. 2012, 6, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Lubelli, C.; Chatgilialoglu, A.; Bolognesi, A.; Strocchi, P.; Colombatti, M.; Stirpe, F. Detection of ricin and other ribosome-inactivating proteins by an immuno-polymerase chain reaction assay. Anal. Biochem. 2006, 355, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, C.; Dürkop, H.; Zhao, X.; Weng, A.; Melzig, M.F.; Fuchs, H. Targeted dianthin is a powerful toxin to treat pancreatic carcinoma when applied in combination with the glycosylated triterpene SO1861. Mol. Oncol. 2017, 11, 1527–1543. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, L.; Bolognesi, A.; Valbonesi, P.; Polito, L.; Olivieri, F.; Stirpe, F. Polynucleotide: Adenosine Glycosidase Activity of Immunotoxins Containing Ribosome-Inactivating Proteins. J. Drug Target. 2000, 8, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Von Mallinckrodt, B.; Thakur, M.; Weng, A.; Gilabert-Oriol, R.; Dürkop, H.; Brenner, W.; Lukas, M.; Beindorff, N.; Melzig, M.F.; Fuchs, H. Dianthin-EGF is an effective tumor targeted toxin in combination with saponins in a xenograft model for colon carcinoma. Futur. Oncol. 2014, 10, 2161–2175. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, L.; Brigotti, M.; Perocco, P.; Carnicelli, D.; Ciani, M.; Mercatali, L.; Stirpe, F. Ribosome-inactivating proteins depurinate poly(ADP-ribosyl)ated poly(ADP-ribose) polymerase and have transforming activity for 3T3 fibroblasts. FEBS Lett. 2003, 538, 178–182. [Google Scholar] [CrossRef]
- Duggar, B.M.; Armstrong, J.K. The Effect of Treating the Virus of Tobacco Mosaic with the Juices of Various Plants. Ann. Mol. Bot. Gard. 1925, 12, 359. [Google Scholar] [CrossRef]
- Wyatt, S.D.; Shepherd, R.J. Isolation and characterization of a virus inhibitor from Phytolacca americana. Phytopathology 1969, 59, 1787–1794. [Google Scholar] [PubMed]
- Obrig, T.G.; Irvin, J.D.; Hardesty, B. The effect of an antiviral peptide on the ribosomal reactions of the peptide elongation enzymes, EF-I and EF-II. Arch. Biochem. Biophys. 1973, 155, 278–289. [Google Scholar] [CrossRef]
- Campadelli-Fiume, G.; Barbieri, L.; Stirpe, F.; Foà-Tomasi, L. Effect of ribosome-inactivating proteins on virus-infected cells. Inhibition of virus multiplication and of protein synthesis. Arch. Virol. 1982, 71, 323–332. [Google Scholar]
- Battelli, M.G.; Montacuti, V.; Stirpe, F. High sensitivity of cultured human trophoblasts to ribosome-inactivating proteins. Exp. Cell Res. 1992, 201, 109–112. [Google Scholar] [CrossRef]
- Hong, Y.; Saunders, K.; Hartley, M.R.; Stanley, J. Resistance to Geminivirus Infection by Virus-Induced Expression of Dianthin in Transgenic Plants. Virology 1996, 220, 119–127. [Google Scholar] [CrossRef]
- Hong, Y.; Saunders, K.; Stanley, J. Transactivation of Dianthin Transgene Expression by African Cassava Mosaic Virus AC2. Virology 1997, 228, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Shah, J.M.; Veluthambi, K. DIANTHIN, a negative selection marker in tobacco, is non-toxic in transgenic rice and confers sheath blight resistance. Boil. Plant. 2010, 54, 443–450. [Google Scholar] [CrossRef]
- Polito, L.; Djemil, A.; Bortolotti, M. Plant Toxin-Based Immunotoxins for Cancer Therapy: A Short Overview. Biomedicines 2016, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- Worbs, S.; Köhler, K.; Pauly, D.; Avondet, M.-A.; Schaer, M.; Dorner, M.B.; Dorner, B.G. Ricinus communis Intoxications in Human and Veterinary Medicine-A Summary of Real Cases. Toxins 2011, 3, 1332–1372. [Google Scholar] [CrossRef] [PubMed]
- Gilabert-Oriol, R.; Furness, S.G.B.; Stringer, B.W.; Weng, A.; Fuchs, H.; Day, B.W.; Kourakis, A.; Boyd, A.W.; Hare, D.L.; Thakur, M.; et al. Dianthin-30 or gelonin versus monomethyl auristatin e, each configured with an anti-calcitonin receptor antibody, are differentially potent in vitro in high-grade glioma cell lines derived from glioblastoma. Cancer Immunol. Immunother. 2017, 66, 1217–1228. [Google Scholar] [CrossRef] [PubMed]
- Bachran, D.; Schneider, S.; Bachran, C.; Urban, R.; Weng, A.; Melzig, M.F.; Hoffmann, C.; Kaufmann, A.M.; Fuchs, H. Epidermal growth factor receptor expression affects the efficacy of the combined application of saponin and a targeted toxin on human cervical carcinoma cells. Int. J. Cancer 2010, 127, 1453–1461. [Google Scholar] [CrossRef]
- Gilabert-Oriol, R.; Thakur, M.; Weise, C.; Dernedde, J.; Von Mallinckrodt, B.; Fuchs, H.; Weng, A. Small structural differences of targeted anti-tumor toxins result in strong variation of protein expression. Protein Expr. Purif. 2013, 91, 54–60. [Google Scholar] [CrossRef]
- Vago, R.; Marsden, C.J.; Lord, J.M.; Ippoliti, R.; Flavell, D.J.; Flavell, S.-U.; Ceriotti, A.; Fabbrini, M.S. Saporin and ricin A chain follow different intracellular routes to enter the cytosol of intoxicated cells. FEBS J. 2005, 272, 4983–4995. [Google Scholar] [CrossRef]
- Kaul, P.; Silverman, J.; Shen, W.H.; Blanke, S.R.; Huynh, P.D.; Finkelstein, A.; Collier, R.J. Roles of glu 349 and asp 352 in membrane insertion and translocation by diphtheria toxin. Protein Sci. 1996, 5, 687–692. [Google Scholar] [CrossRef]
- Fuchs, H.; Bachran, C.; Li, T.; Heisler, I.; Dürkop, H.; Sutherland, M. A cleavable molecular adapter reduces side effects and concomitantly enhances efficacy in tumor treatment by targeted toxins in mice. J. Control. Release 2007, 117, 342–350. [Google Scholar] [CrossRef]
- Fuchs, H.; Niesler, N.; Trautner, A.; Sama, S.; Jerz, G.; Panjideh, H.; Weng, A. Glycosylated Triterpenoids as Endosomal Escape Enhancers in Targeted Tumor Therapies. Biomedicines 2017, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Bachran, C.; Dürkop, H.; Sutherland, M.; Bachran, D.; Müller, C.; Weng, A.; Melzig, M.F.; Fuchs, H. Inhibition of Tumor Growth by Targeted Toxins in Mice is Dramatically Improved by Saponinum Album in a Synergistic Way. J. Immunother. 2009, 32, 713–725. [Google Scholar] [CrossRef] [PubMed]
- Sama, S.; Jerz, G.; Thakur, M.; Melzig, M.F.; Weng, A. Structure-Activity Relationship of Transfection-Modulating Saponins—A Pursuit for the Optimal Gene Trafficker. Planta Med. 2019, 85, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Sama, S.; Woith, E.; Walther, W.; Jerz, G.; Chen, W.; Hart, S.; Melzig, M.F.; Weng, A. Targeted suicide gene transfections reveal promising results in nu/nu mice with aggressive neuroblastoma. J. Control. Release 2018, 275, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Bachran, C.; Weng, A.; Bachran, D.; Riese, S.B.; Schellmann, N.; Melzig, M.F.; Fuchs, H. The distribution of saponins in vivo affects their synergy with chimeric toxins against tumours expressing human epidermal growth factor receptors in mice. Br. J. Pharmacol. 2010, 159, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Thakur, M.; Mergel, K.; Weng, A.; von Mallinckrodt, B.; Gilabert-Oriol, R.; Durkop, H.; Melzig, M.F.; Fuchs, H. Targeted tumor therapy by epidermal growth factor appended toxin and purified saponin: An evaluation of toxicity and therapeutic potential in syngeneic tumor bearing mice. Mol. Oncol. 2013, 7, 475–483. [Google Scholar] [CrossRef] [PubMed]

| RIP | Type | Plant | Plant Source | Molecular Mass (kDa) |
|---|---|---|---|---|
| dianthin-29 | 1 | Dianthus barbatus L. | leaves | 29.0 |
| dianthin-30 | 1 | Dianthus caryophyllus L. | leaves | 29.5 |
| dianthin-32 | 1 | Dianthus caryophyllus L. | leaves | 31.7 |
| saporin-S3 | 1 | Saponaria officinalis L. | seeds | 28.6 |
| saporin-S6 | 1 | Saponaria officinalis L. | seeds | 28.6 |
| saporin-S9 | 1 | Saponaria officinalis L. | seeds | 28.5 |
| saporin-R1 | 1 | Saponaria officinalis L. | roots | 30.2 |
| saporin-R2 | 1 | Saponaria officinalis L. | roots | 30.9 |
| PAP 1 | 1 | Phytolacca americana L. | leaves | 29–30 |
| PAP-S 1 | 1 | Phytolacca americana L. | seeds | 30 |
| PAP-R 1 | 1 | Phytolacca americana L. | roots | 29.8 |
| PD-S2 2 | 1 | Phytolacca dioica L. | seeds | 29.6 |
| gelonin | 1 | Gelonium multiflorum A.Juss | seeds | 30–31 |
| bryodin | 1 | Bryonia dioica Jacq. | roots | 29 |
| momordin | 1 | Momordica charantia L. | seeds | 31 |
| momorcochin-S | 1 | Momordica cochinchinensis Spreng. | seeds | 30 |
| trichokirin | 1 | Trichosanthes kirilowii Maxim. | seeds | 27 |
| tritin | 1 | Triticum aestivum L. | germ | 30 |
| crotin-3 | 1 | Croton tiglium L. | seeds | n. a. 3 |
| lychnin | 1 | Lychnis chalcedonica L. | seeds | 26.1 |
| bouganin | 1 | Bougainvillea spectabilis Willd. | leaves | 26.2 |
| colocin-1 | 1 | Citrullus colocynthis (L.) Schrad. | seeds | 26.3 |
| asparin | 1 | Asparagus officinalis L. | seeds | 29.8–30.5 |
| barley RIP | 1 | Hordeum vulgare L. | seeds | 30 |
| ricin A-chain | 2 | Ricinus communis L. | seeds | 32 |
| ricin | 2 | Ricinus communis L. | seeds | 62.8 |
| abrin-c | 2 | Abrus precatorius L. | seeds | 60.1–62.5 |
| modeccin | 2 | Adenia digitata (Harv.) Engl. | roots | 57–63 |
| viscumin | 2 | Viscum album L. | leaves | 115–125 |
| volkensin | 2 | Adenia volkensii Harms | roots | 62 |
| RIP or RIP Fusion Proteins/Conjugates | Translation Inhibition IC50 (nM) | Ribosomal Source | Reference |
|---|---|---|---|
| dianthin-30 | 0.133 | rabbit reticulocyte lysate | [54] |
| 0.18 | rabbit reticulocyte lysate | [21] | |
| 0.227 2 | rabbit reticulocyte lysate | [54] | |
| 0.29 1,2 | rabbit reticulocyte lysate | [14] | |
| 0.31 | rabbit reticulocyte lysate | [1] | |
| 0.37 2 | rabbit reticulocyte lysate | [14] | |
| 0.41 2 | rabbit reticulocyte lysate | [55] | |
| 0.61 | rabbit reticulocyte lysate | [14] | |
| 0.35–0.70 | wheat germ extracts | [24] | |
| 110 | Leishmania infantum | [56] | |
| 153 | Trypanosoma brucei rhodesiense | [56] | |
| 6478 | Streptomyces lividans | [12] | |
| dianthin-30-anti-CD30 3 | 0.1 | rabbit reticulocyte lysate | [54] |
| 1.054 2 | rabbit reticulocyte lysate | [54] | |
| KFT25-dianthin-30 4 | 0.47 2 | rabbit reticulocyte lysate | [55] |
| pHA2-dianthin-30 4 | 0.53 2 | rabbit reticulocyte lysate | [55] |
| pJVE-dianthin-30 4 | 0.54 2 | rabbit reticulocyte lysate | [55] |
| dianthin-32 | 0.02 | Saccharomyces cerevisiae | [52] |
| 0.11 | rabbit reticulocyte lysate | [1] | |
| 0.12 | rabbit reticulocyte lysate | [15] | |
| 0.12 | rabbit reticulocyte lysate | [57] | |
| 0.35–0.70 | wheat germ extracts | [24] | |
| 0.63 | Nicotiana tabacum | [52] | |
| 7 | Acanthamoeba castellanii | [58] | |
| 27 | Leishmania infantum | [56] | |
| 30 | Tetrahymena pyriformis | [58] | |
| 66 | Trypanosoma brucei rhodesiense | [56] | |
| 331 | Streptomyces lividans | [12] | |
| 991 | Agrobacterium tumefaciens | [15] | |
| >1000 | Escherichia coli | [15] | |
| dianthin-32-F(ab’)2 | 0.167 | rabbit reticulocyte lysate | [57] |
| saporin-S3 | 0.01–0.02 | rabbit reticulocyte lysate | [21] |
| saporin-S6 | 0.01–0.02 | rabbit reticulocyte lysate | [21] |
| 0.033 | rabbit reticulocyte lysate | [57] | |
| 17 | Acanthamoeba castellanii | [58] | |
| 33 | Leishmania infantum | [56] | |
| 116 | Trypanosoma brucei rhodesiense | [56] | |
| 2630 | Tetrahymena pyriformis | [58] | |
| >7133 | Streptomyces lividans | [12] | |
| saporin-S6-F(ab’)2 | 0.086 | rabbit reticulocyte lysate | [57] |
| saporin-S9 | 26 | Trypanosoma brucei rhodesiense | [56] |
| 31 | Leishmania infantum | [56] | |
| saporin-R1 | 0.86 | rabbit reticulocyte lysate | [15] |
| 927 | Escherichia coli | [15] | |
| >1000 | Escherichia coli | [15] | |
| >1000 | Agrobacterium tumefaciens | [15] | |
| saporin-R2 | 0.47 | rabbit reticulocyte lysate | [15] |
| 423 | Escherichia coli | [15] | |
| >1000 | Escherichia coli | [15] | |
| >1000 | Agrobacterium tumefaciens | [15] | |
| PAP | 0.14 | Nicotiana tabacum | [52] |
| 0.29 | Saccharomyces cerevisiae | [52] | |
| PAP-S | 0.037 | rabbit reticulocyte lysate | [57] |
| 17 | Trypanosoma brucei rhodesiense | [56] | |
| 21 | Leishmania infantum | [56] | |
| 1040 | Streptomyces lividans | [12] | |
| 1230 | Acanthamoeba castellanii | [58] | |
| 1570 | Tetrahymena pyriformis | [58] | |
| PAP-S-F(ab’)2 | 0.41 | rabbit reticulocyte lysate | [57] |
| PAP-R | 40 | Trypanosoma brucei rhodesiense | [56] |
| 53 | Leishmania infantum | [56] | |
| 2005 | Streptomyces lividans | [12] | |
| gelonin | 0.4 | rabbit reticulocyte lysate | [57] |
| 2590 | Leishmania infantum | [56] | |
| 3330 | Trypanosoma brucei rhodesiense | [56] | |
| >3330 | Tetrahymena pyriformis | [58] | |
| >3300 | Acanthamoeba castellanii | [58] | |
| >11,000 | Streptomyces lividans | [12] | |
| gelonin-F(ab’)2 | 6.2 | rabbit reticulocyte lysate | [57] |
| bryodin | 0.12 | rabbit reticulocyte lysate | [57] |
| 430 | Acanthamoeba castellanii | [58] | |
| 3330 | Trypanosoma brucei rhodesiense | [56] | |
| >3330 | Tetrahymena pyriformis | [58] | |
| >3330 | Leishmania infantum | [56] | |
| >10,932 | Streptomyces lividans | [12] | |
| bryodin-F(ab’)2 | 0.247 | rabbit reticulocyte lysate | [57] |
| momordin | 0.06 | rabbit reticulocyte lysate | [57] |
| 0.06 | rabbit reticulocyte lysate | [15] | |
| 72 | Streptomyces lividans | [12] | |
| 190 | Escherichia coli | [15] | |
| 190 | Agrobacterium tumefaciens | [15] | |
| 300 | Tetrahymena pyriformis | [58] | |
| 857 | Escherichia coli | [15] | |
| 1130 | Acanthamoeba castellanii | [58] | |
| 3330 | Trypanosoma brucei rhodesiense | [56] | |
| >3330 | Leishmania infantum | [56] | |
| momordin-F(ab’)2 | 0.44 | rabbit reticulocyte lysate | [57] |
| trichokirin | 0.087 | rabbit reticulocyte lysate | [57] |
| 1265 | Trypanosoma brucei rhodesiense | [56] | |
| 1747 | Streptomyces lividans | [12] | |
| >3330 | Leishmania infantum | [56] | |
| trichokirin-F(ab’)2 | 0.29 | rabbit reticulocyte lysate | [57] |
| tritin | 22 | Saccharomyces cerevisiae | [52] |
| >2200 | Nicotiana tabacum | [52] | |
| crotin-3 | 0.2 | rabbit reticulocyte lysate | [15] |
| 13 | Escherichia coli | [15] | |
| 19 | Streptomyces lividans | [12] | |
| 40 | Escherichia coli | [15] | |
| barley RIP | 3 | Saccharomyces cerevisiae | [52] |
| 1598 | Trypanosoma brucei rhodesiense | [56] | |
| >2200 | Nicotiana tabacum | [52] | |
| 3330 | Leishmania infantum | [56] | |
| 4922 | Streptomyces lividans | [12] | |
| ricin A-chain | 0.066 | rabbit reticulocyte lysate | [57] |
| 0.17 | Saccharomyces cerevisiae | [52] | |
| 0.3 | rabbit reticulocyte lysate | [55] | |
| 21 | Nicotiana tabacum | [52] | |
| 800 | wheat germ extracts | [50] | |
| ricin A-chain-F(ab’)2 | 0.078 | rabbit reticulocyte lysate | [57] |
| ricin | 1030 | Acanthamoeba castellanii | [58] |
| >1700 | Tetrahymena pyriformis, | [58] | |
| >1700 | Trypanosoma brucei rhodesiense | [56] | |
| >1700 | Leishmania infantum | [56] | |
| >3300 | Streptomyces lividans | [12] | |
| abrin | 5 | Leishmania infantum | [56] |
| 49 | Trypanosoma brucei rhodesiense | [56] | |
| 100 | Acanthamoeba castellanii | [58] | |
| >1700 | Tetrahymena pyriformis | [58] | |
| modeccin | 1700 | Leishmania infantum | [56] |
| >1700 | Tetrahymena pyriformis, | [58] | |
| >1700 | Acanthamoeba castellanii, | [58] | |
| >1700 | Trypanosoma brucei rhodesiense | [56] | |
| viscumin | 1700 | Acanthamoeba castellanii | [58] |
| >1700 | Tetrahymena pyriformis, | [58] | |
| >1700 | Trypanosoma brucei rhodesiense | [56] | |
| >1700 | Leishmania infantum | [56] | |
| volkensin | 830 | Acanthamoeba castellanii | [58] |
| 1700 | Leishmania infantum | [56] | |
| >1700 | Tetrahymena pyriformis, | [58] | |
| >1700 | Trypanosoma brucei rhodesiense | [56] | |
| >3300 | Streptomyces lividans | [12] |
| RIP or RIP Fusion Proteins/Conjugates | Enzymatic Activity Adenine Release (pmol/pmol/h) | Substrate | Reference |
|---|---|---|---|
| dianthin-30 | 0.54 | poly(A) | [43] |
| 0.57 | rat ribosomes | [43] | |
| <10 1 | 60S ribosomal subunit | [67] | |
| <10 1 | poly(A) | [67] | |
| 23.99 | herring sperm DNA | [43] | |
| 67 1 | herring sperm DNA | [71] | |
| 110 1 | (dA)30 | [68] | |
| 140 1 | 28S-rRNA | [67] | |
| 330 1 | mitochondrial DNA | [67] | |
| 644 | herring sperm DNA | [72] | |
| 775 1 | herring sperm DNA | [67] | |
| dianthin-30-EGF 3 | 42 1 | herring sperm DNA | [71] |
| 67 1 | (dA)30 | [68] | |
| 116 1 | herring sperm DNA | [73] | |
| 218 1 | herring sperm DNA | [69] | |
| dianthin-30-anti-CD30 4 | 247 1 | herring sperm DNA | [72] |
| 346 | herring sperm DNA | [72] | |
| saporin-S3 | <10 1 | 60S ribosomal subunit | [67] |
| <10 1 | poly(A) | [67] | |
| 50 1 | mitochondrial DNA | [67] | |
| 125 1 | (dA)30 | [68] | |
| 140 1 | 28S-rRNA | [67] | |
| 312 1 | herring sperm DNA | [69] | |
| 397 | herring sperm DNA | [69] | |
| 670 1 | herring sperm DNA | [67] | |
| saporin-S3-EGF 3 | 70 1 | (dA)30 | [68] |
| 80 1 | herring sperm DNA | [73] | |
| 240 1 | herring sperm DNA | [69] | |
| saporin-S6 | 1.91 | rat ribosomes | [43] |
| 18.3 | herring sperm DNA | [74] | |
| >30 | poly(A) | [43] | |
| 37.61 | herring sperm DNA | [43] | |
| 439 1 | herring sperm DNA | [72] | |
| saporin-S6-anti-CD30 4 | 176 | herring sperm DNA | [72] |
| saporin-L1 | 861 | herring sperm DNA | [72] |
| saporin-L1-anti-CD30 4 | 429 | herring sperm DNA | [72] |
| saporin-L2 | 934 | herring sperm DNA | [74] |
| PAP-S | 519 | herring sperm DNA | [72] |
| PAP-S-anti-CD22 4 | 115 | herring sperm DNA | [72] |
| PAP-R | <d. l. 2 | poly(A) | [43] |
| 0.51 | rat ribosomes | [43] | |
| 50.32 | herring sperm DNA | [43] | |
| gelonin | 12.9 | herring sperm DNA | [74] |
| 579 | herring sperm DNA | [72] | |
| gelonin-anti-CD30 4 | 130 | herring sperm DNA | [72] |
| momordin | <d. l. 2 | poly(A) | [43] |
| 0.39 | rat ribosomes | [43] | |
| 0.75 | herring sperm DNA | [74] | |
| 2.71 | herring sperm DNA | [43] | |
| 18 | herring sperm DNA | [72] | |
| momordin-anti-CD30 4 | 1.8 | herring sperm DNA | [72] |
| lychnin | <d. l. 2 | poly(A) | [43] |
| 0.72 | rat ribosomes | [43] | |
| 2.95 | herring sperm DNA | [43] | |
| bouganin | <d. l. 2 | poly(A) | [43] |
| 0.48 | rat ribosomes | [43] | |
| 37.77 | herring sperm DNA | [43] | |
| ricin A chain | <d. l. 2 | poly(A) | [43] |
| 0.62 | rat ribosomes | [43] | |
| 4.85 | herring sperm DNA | [43] | |
| <10 | 60S ribosomal subunit | [67] | |
| <10 | poly(A) | [67] | |
| 22 | herring sperm DNA | [69] | |
| 27.6 1 | herring sperm DNA | [72] | |
| 43 | herring sperm DNA | [67] | |
| 185 | 28S-rRNA | [67] | |
| ricin A chain-EGF 3 | 17 1 | herring sperm DNA | [69] |
| ricin A chain-anti-CD30 4 | 7.2 1 | herring sperm DNA | [72] |
| ricin | 7.5 | herring sperm DNA | [74] |
| <10 | mitochondrial DNA | [67] | |
| <10 | 60S ribosomal subunit | [67] | |
| 12 | 28S-rRNA | [67] | |
| 70 | herring sperm DNA | [67] |
| RIP | Concentration (µg/mL) | Virus 1 | Target | Inhibition(%) | Reference |
|---|---|---|---|---|---|
| dianthin-30 | 0.5 | TMV | N. glutinosa L. | 63 | [1] |
| 1 | TMV | N. glutinosa L. | 91 | [1] | |
| 5 | TMV | N. glutinosa L. | 96 | [1] | |
| 10 | TMV | N. glutinosa L. | 99 | [1] | |
| dianthin-32 | 0.01 | TMV | N. tabacum L. | 0 | [52] |
| 0.05 | TMV | N. tabacum L. | 0 | [52] | |
| 0.1 | TMV | N. tabacum L. | 57 | [52] | |
| 0.5 | TMV | N. tabacum L. | 76 | [52] | |
| 0.5 | TMV | N. glutinosa L. | 33 | [1] | |
| 1 | TMV | N. tabacum L. | 91 | [52] | |
| 1 | TMV | N. glutinosa L. | 84 | [1] | |
| 2 | TMV | N. tabacum L. | 91 | [52] | |
| 5 | TMV | N. glutinosa L. | 86 | [1] | |
| 10 | TMV | N. tabacum L. | 100 | [52] | |
| 10 | TMV | N. glutinosa L. | 98 | [1] | |
| 100 | TMV | N. tabacum L. | 100 | [52] | |
| 100 | HSV-1 | HEp-2 cells | 77 | [78] | |
| 200 | HSV-1 | HEp-2 cells | 85 | [78] | |
| PAP | 0.01 | TMV | N. tabacum L. | 0 | [52] |
| 0.05 | TMV | N. tabacum L. | 0 | [52] | |
| 0.1 | TMV | N. tabacum L. | 57 | [52] | |
| 0.5 | TMV | N. tabacum L. | 76 | [52] | |
| 1 | TMV | N. tabacum L. | 91 | [52] | |
| 2 | TMV | N. tabacum L. | 91 | [52] | |
| 10 | TMV | N. tabacum L. | 100 | [52] | |
| 100 | TMV | N. tabacum L. | 100 | [52] | |
| PAP-S | 100 | HSV-1 | HEp-2 cells | 81 | [78] |
| 200 | HSV-1 | HEp-2 cells | 90 | [78] | |
| gelonin | 100 | HSV-1 | HEp-2 cells | 31 | [78] |
| 200 | HSV-1 | HEp-2 cells | 83 | [78] | |
| momordin | 100 | HSV-1 | HEp-2 cells | 70 | [78] |
| 200 | HSV-1 | HEp-2 cells | 81 | [78] | |
| tritin | 10 | TMV | N. tabacum L. | 36 | [52] |
| 100 | TMV | N. tabacum L. | 23 | [52] | |
| barley RIP | 10 | TMV | N. tabacum L. | 33 | [52] |
| 100 | TMV | N. tabacum L. | 16 | [52] | |
| ricin A-chain | 10 | TMV | N. tabacum L. | 20 | [52] |
| 100 | TMV | N. tabacum L. | 25 | [52] |
| RIP or RIP Fusion Proteins/Conjugates | Cytotoxicity IC50 (nM) | Cells/Cell Line | Reference |
|---|---|---|---|
| dianthin-30 | >100 | D430B | [54] |
| >100 1 | D430B | [54] | |
| ~30 | L540 | [54] | |
| ~100 1 | L540 | [54] | |
| >100 | L428 | [54] | |
| ~100 1 | L428 | [54] | |
| 712 1 | Jurkat | [55] | |
| 203 1,2 | Jurkat | [55] | |
| 910 | H9 | [46] | |
| dianthin-30-EGF 3 | 0.45 1 | HER14 4 | [69] |
| <0.0000001 1,2 | HER14 4 | [69] | |
| dianthin-30-anti-CD30 5 | 0.479 | D430B | [54] |
| 0.162 1 | D430B | [54] | |
| 0.324 | L540 | [54] | |
| 0.045 1 | L540 | [54] | |
| 0.355 | L428 | [54] | |
| 0.182 1 | L428 | [54] | |
| 7.6 | K562 | [54] | |
| >50 1 | K562 | [54] | |
| dianthin-30-cetuximab | >10 1 | HCT-116 | [13] |
| 0.0053 1,2 | HCT-116 | [13] | |
| dianthin-30-panitumumab | >10 1 | HCT-116 | [13] |
| 0.0015 1,2 | HCT-116 | [13] | |
| dianthin-30-trastuzumab | >10 1 | BT-474 | [13] |
| 0.023 1,2 | BT-474 | [13] | |
| dianthin-30-mAb2C4 | 10–20 | SB2b | [85] |
| 0.01 2 | SB2b | [85] | |
| 0.01 2 | JK2 | [85] | |
| 0.02 2 | U87MG | [85] | |
| transferrin-dianthin-30 | 1.5 1 | Jurkat | [55] |
| 0.018 1,2 | Jurkat | [55] | |
| KFT25-dianthin-30 6 | >1000 1 | Jurkat | [55] |
| 712 1,2 | Jurkat | [55] | |
| pHA2-dianthin-30 6 | >1000 1 | Jurkat | [55] |
| 610 1,2 | Jurkat | [55] | |
| pJVE-dianthin-30 6 | 814 1 | Jurkat | [55] |
| 183 1,2 | Jurkat | [55] | |
| dianthin-32 | >3200 | H9 | [46] |
| dianthin-32-F(ab’)2 | 0.011 | PBMC 7 | [57] |
| saporin-S3 | 611 1 | HeLa | [86] |
| 0.24 1,2 | HeLa | [86] | |
| >1000 1 | CaSki | [86] | |
| 0.26 1,2 | CaSki | [86] | |
| 53 1 | HER14 4 | [86] | |
| 0.08 1,2 | HER14 4 | [86] | |
| saporin-S3-EGF 3 | 53 1 | HeLa | [86] |
| 0.00070 1,2 | HeLa | [86] | |
| 5 1 | CaSki | [86] | |
| 0.00013 1,2 | CaSki | [86] | |
| 2.5 1 | HER14 4 | [86] | |
| 0.00090 1,2 | HER14 4 | [86] | |
| 57 1 | HER14 4 | [69] | |
| <0.0000001 1,2 | HER14 4 | [69] | |
| saporin-S6-F(ab’)2 | 0.0055 | PBMC 7 | [57] |
| PAP-S-F(ab’)2 | 0.0026 | PBMC 7 | [57] |
| gelonin-mAb2C4 | 0.02 2 | U87MG | [85] |
| gelonin-F(ab’)2 | 10 | PBMC 7 | [57] |
| bryodin-F(ab’)2 | 0.0288 | PBMC 7 | [57] |
| momordin-F(ab’)2 | 0.02 | PBMC 7 | [57] |
| trichokirin-F(ab’)2 | 0.0323 | PBMC 7 | [57] |
| ricin A-chain | 280 | Jurkat | [55] |
| 140 2 | Jurkat | [55] | |
| ricin A-chain-EGF 3 | >1000 1 | HER14 4 | [69] |
| 61 1,2 | HER14 4 | [69] | |
| ricin A-chain-F(ab’)2 | 5 | PBMC 7 | [57] |
| RIP | Antibody Titer (Arbitrary Units) |
|---|---|
| saporin-S6 | 44.0 |
| lychnin | 21.0 |
| momordin | 15.0 |
| colocin-1 | 7.0 |
| momorcochin-S | 6.50 |
| dianthin-32 | 4.18 |
| PAP-R | 2.83 |
| trichokirin | 2.16 |
| bryodin-R | 1.50 |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fuchs, H. Dianthin and Its Potential in Targeted Tumor Therapies. Toxins 2019, 11, 592. https://doi.org/10.3390/toxins11100592
Fuchs H. Dianthin and Its Potential in Targeted Tumor Therapies. Toxins. 2019; 11(10):592. https://doi.org/10.3390/toxins11100592
Chicago/Turabian StyleFuchs, Hendrik. 2019. "Dianthin and Its Potential in Targeted Tumor Therapies" Toxins 11, no. 10: 592. https://doi.org/10.3390/toxins11100592
APA StyleFuchs, H. (2019). Dianthin and Its Potential in Targeted Tumor Therapies. Toxins, 11(10), 592. https://doi.org/10.3390/toxins11100592





