Effects of Heme Modulation on Ovophis and Trimeresurus Venom Activity in Human Plasma
Abstract
1. Introduction
2. Results
2.1. Ovophis monticola
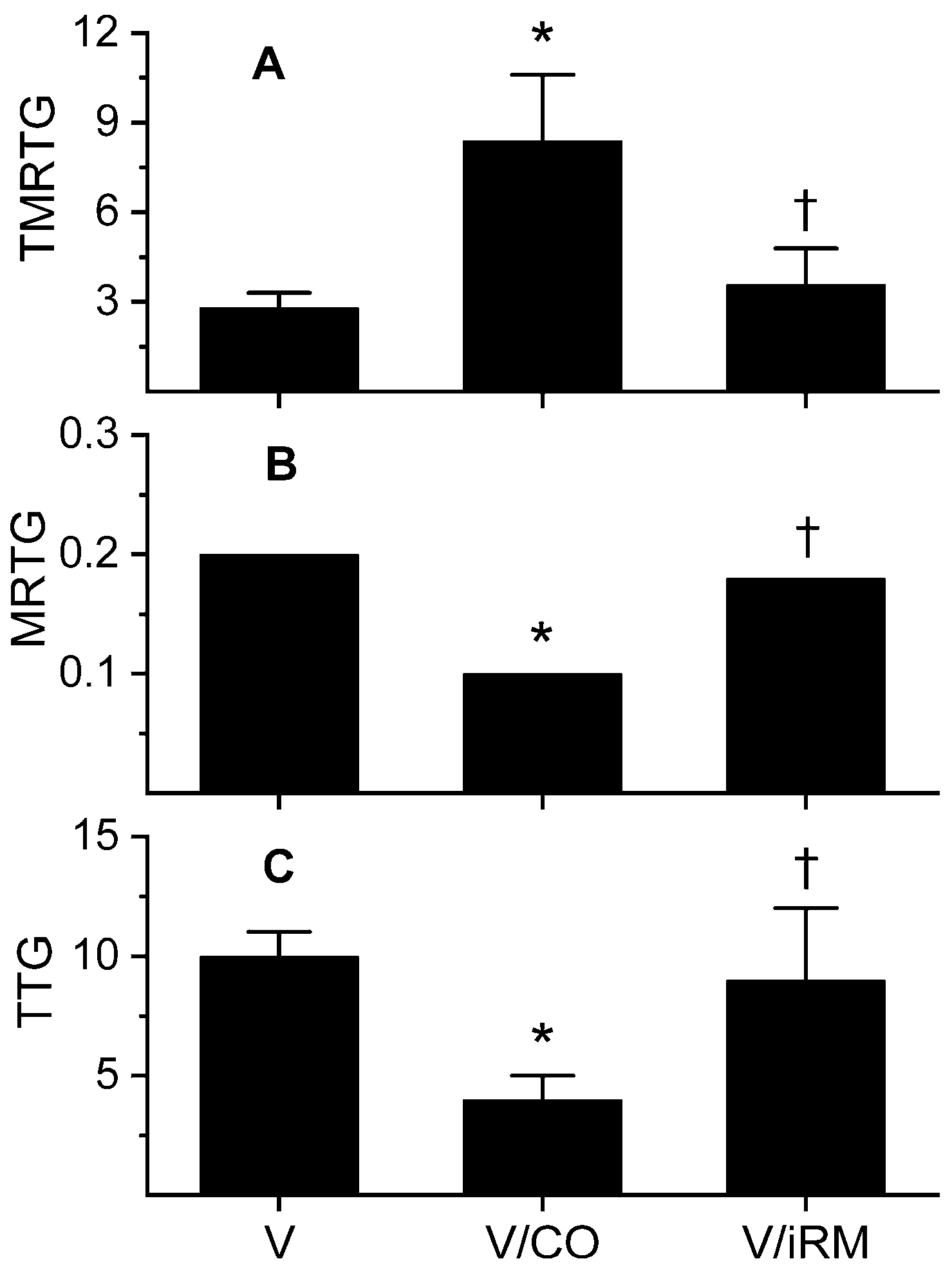
2.2. Ovophis okinavensis
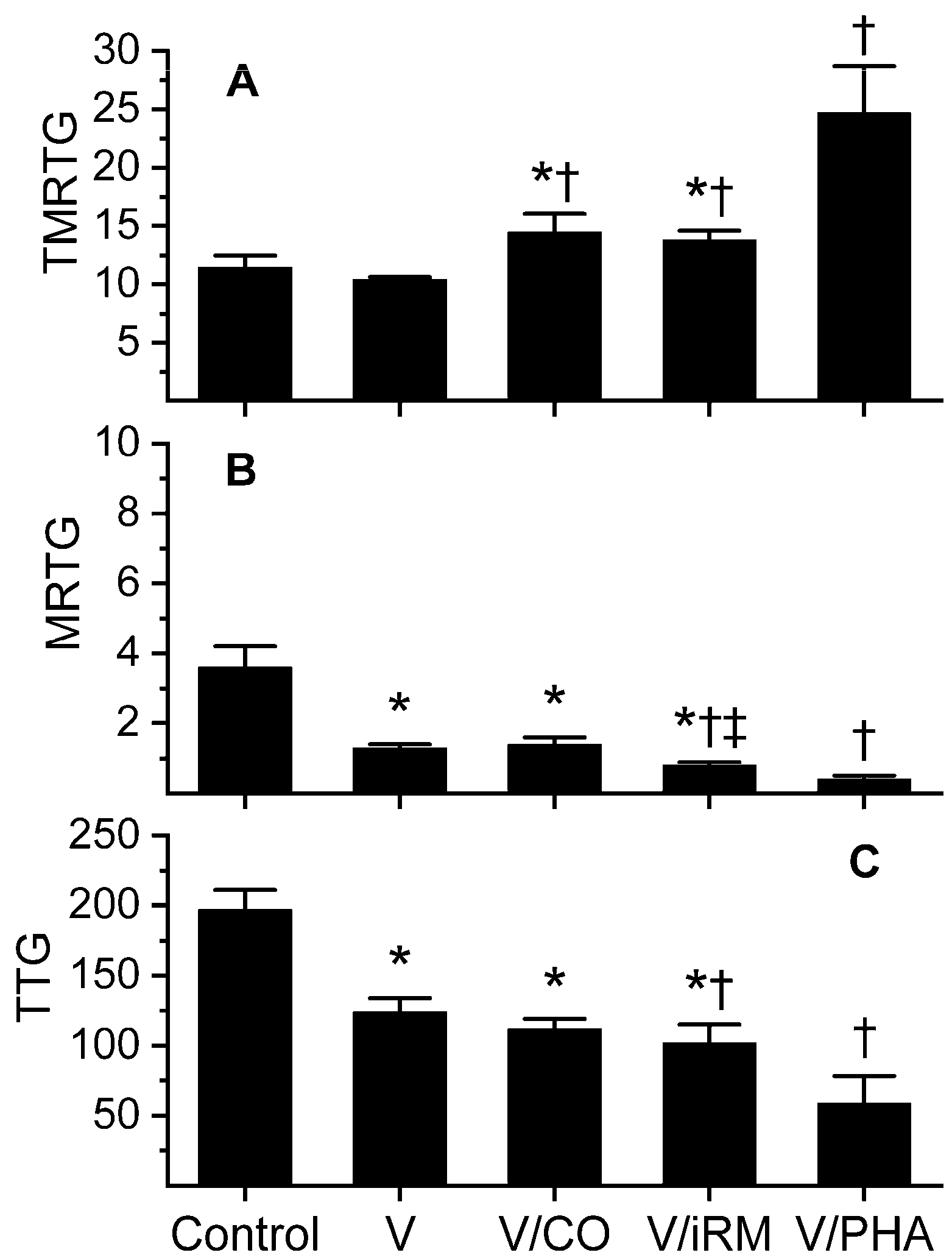
2.3. Trimeresurus borneensis
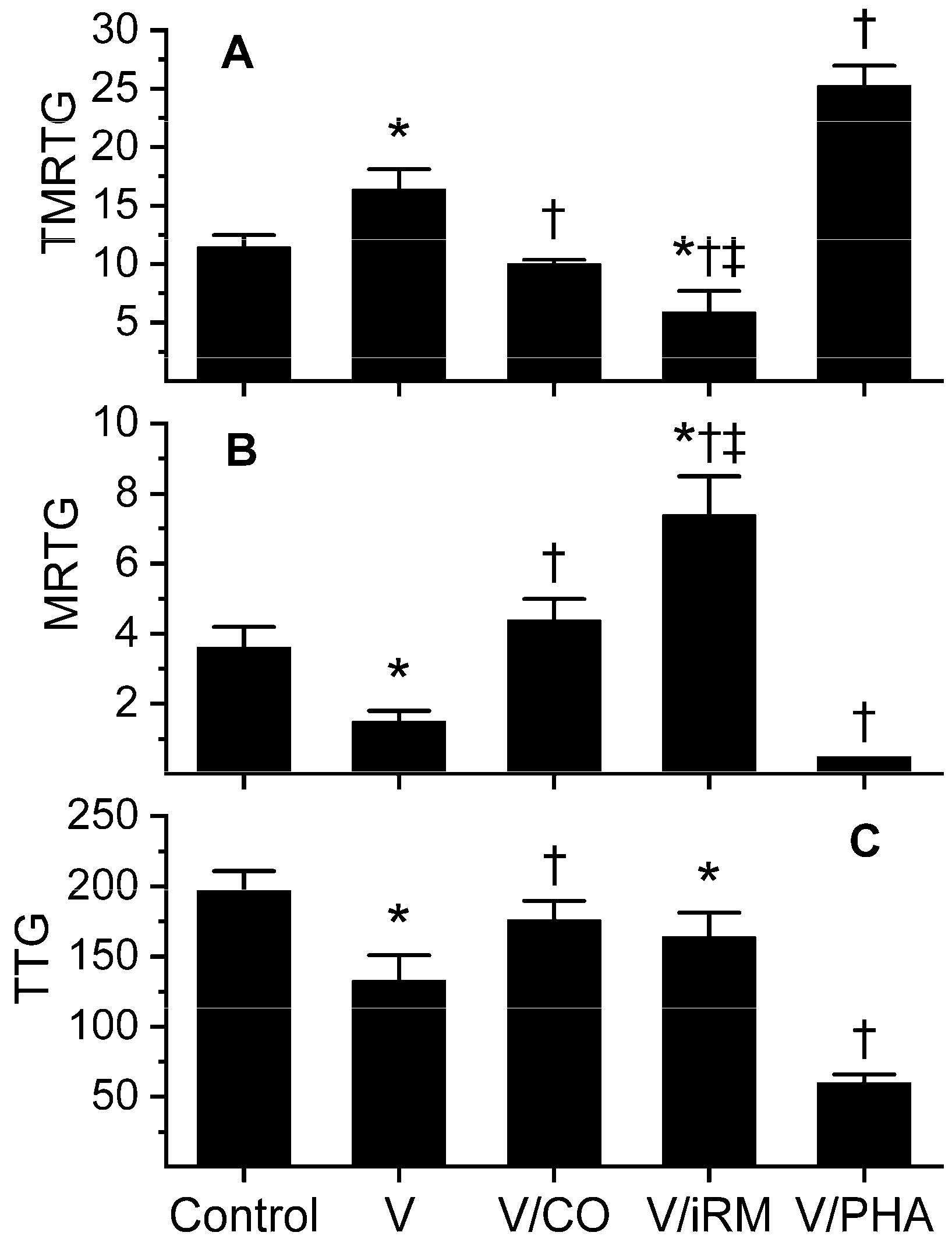
2.4. Trimeresurus fasciatus
2.5. Trimeresurus purpureomaculatus
3. Discussion
4. Materials and Methods
4.1. Human Plasma, Venoms, and Other Chemicals
4.2. Thrombelastographic Analyses
4.3. Generation of Venom Concentration Experimentation and Effects of Isolated CORM-2 or PHA Venom Exposures
4.4. Separation of Venom-Mediated Thrombin Generation from Thrombinlike Activity
4.5. Statistical Analyses
Author Contributions
Funding
Conflicts of Interest
References
- Núñez, V.; Cid, P.; Sanz, L.; De La Torre, P.; Angulo, Y.; Lomonte, B.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics and antivenomics of Bothrops atrox venoms from Colombia and the Amazon regions of Brazil, Perú and Ecuador suggest the occurrence of geographic variation of venom phenotype by a trend towards paedomorphism. J. Proteom. 2009, 73, 57–78. [Google Scholar] [CrossRef] [PubMed]
- Barlow, A.; Pook, C.E.; Harrison, R.A.; Wüster, W. Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc. Biol. Sci. 2009, 276, 2443–2449. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves-Machado, L.; Pla, D.; Sanz, L.; Jorge, R.J.B.; Leitão-De-Araújo, M.; Alves, M.L.M.; Alvares, D.J.; De Miranda, J.; Nowatzki, J.; de Morais-Zani, K.; et al. Combined venomics, venom gland transcriptomics, bioactivities, and antivenomics of two Bothrops jararaca populations from geographic isolated regions within the Brazilian Atlantic rainforest. J. Proteom. 2016, 135, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, A.; Thorpe, R.S. A phylogeny of four mitochondrial gene regions suggests a revised taxonomy for Asian pitvipers (Trimeresurus and Ovophis). Mol. Phylogenet. Evol. 2004, 32, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Berling, I.; Isbister, G.K. Hematologic effects and complications of snake envenoming. Transfus. Med. Rev. 2015, 29, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Cerruti, M.A.; Valencia, O.M.; Amos, Q. Decreased snake venom metalloproteinase effects via inhibition of enzyme and modification of fibrinogen. Biometals 2016, 29, 913–919. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Bazzell, C.M. Carbon monoxide attenuates the effects of snake venoms containing metalloproteinases with fibrinogenase or thrombin-like activity on plasmatic coagulation. MedChemComm 2016, 7, 1973–1979. [Google Scholar] [CrossRef]
- Nielsen, V.G.; Bazzell, C.M. Carbon monoxide releasing molecule-2 inhibition of snake venom thrombin-like activity: Novel biochemical “brake”? J. Thromb. Thrombolysis 2017, 43, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Losada, P.A. Direct inhibitory effects of carbon monoxide on six venoms containing fibrinogenolytic metalloproteinases. Basic Clin. Pharmacol. Toxicol. 2017, 120, 207–212. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Frank, N.; Matika, R.W. Carbon monoxide inhibits hemotoxic activity of Elapidae venoms: Potential role of heme. Biometals 2018, 31, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Matika, R.W. Effects of iron and carbon monoxide on Lachesis muta muta venom-mediated degradation of plasmatic coagulation. Hum. Exp. Toxicol. 2017, 36, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Kühl, T.; Imhof, D. Regulatory Fe(II/III) heme: The reconstruction of a molecule’s biography. Chembiochem 2014, 15, 2024–2035. [Google Scholar] [CrossRef] [PubMed]
- Suntravat, M.; Langlais, P.R.; Sánchez, E.E.; Nielsen, V.G. CatroxMP-II: A heme-modulated fibrinogenolytic metalloproteinase isolated from Crotalus atrox venom. Biometals 2018, 31, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Aird, S.D.; Watanabe, Y.; Villar-Briones, A.; Roy, M.C.; Terada, K.; Mikheyev, A.S. Quantitative high-throughput profiling of snake venom gland transcriptomes and proteomes (Ovophis okinavensis and Protobothrops flavoviridis). BMC Genomics 2013, 14, 790. [Google Scholar] [CrossRef] [PubMed]
- Komori, Y.; Murakami, E.; Uchiya, K.; Nonogaki, T.; Nikai, T. Okinalysin, a novel P-I metalloproteinase from Ovophis okinavensis: Biological properties and effect on vascular endothelial cells. Toxins 2014, 6, 2594–2604. [Google Scholar] [CrossRef] [PubMed]
- Nget, H.T.; Chon, S.T. Biological properties of Trimeresurus purpureomaculatus (shore pit viper) venom and its fractions. Toxicon 1988, 26, 989–996. [Google Scholar]
- Patterson, E.K.; Fraser, D.D.; Capretta, A.; Potter, R.F.; Cepinskas, G. Carbon monoxide-releasing molecule 3 inhibits myeloperoxidase (MPO) and protects against MPO-induced vascular endothelial cell activation/dysfunction. Free Radic. Biol. Med. 2014, 70, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Wilson, J.L.; Wareham, L.K.; McLean, S.; Begg, R.; Greaves, S.; Mann, B.E.; Sanguinetti, G.; Poole, R.K. CO-Releasing Molecules Have Nonheme Targets in Bacteria: Transcriptomic, Mathematical Modeling and Biochemical Analyses of CORM-3 [Ru(CO)3Cl(glycinate)] Actions on a Heme-Deficient Mutant of Escherichia coli. Antioxid. Redox Signal. 2015, 23, 148–162. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Garza, J.I. Comparison of the effects of CORM-2, CORM-3 and CORM-A1 on coagulation in human plasma. Blood Coagul. Fibrinol. 2014, 25, 801–805. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G. Crotalus atrox venom exposed to carbon monoxide has decreased fibrinogenolytic activity in vivo in rabbits. Basic Clin. Pharmacol. Toxicol. 2018, 122, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Kirklin, J.K.; George, J.F. Carbon monoxide-releasing molecule-2 decreases fibrinolysis in human plasma. Blood Coagul. Fibrinolysis 2009, 20, 448–455. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Arkebauer, M.R.; Vosseller, K. Redox-based thrombelastographic method to detect carboxyhemefibrinogen-mediated hypercoagulability. Blood Coagul. Fibrinolysis 2011, 22, 657–661. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Hafner, D.T.; Steinbrenner, E.B. Tobacco smoke-induced hypercoagulation in human plasma: Role of carbon monoxide. Blood Coagul Fibrinolysis 2013, 24, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, V.G.; Boyer, L.V.; Redford, D.T.; Ford, P. Thrombelastographic characterization of the thrombin-like activity of Crotalus simus and Bothrops asper venom. Blood Coagul. Fibrinolysis 2017, 28, 211–217. [Google Scholar] [CrossRef] [PubMed]
| Species | Common Name | Location |
|---|---|---|
| Ovophis monticola | Mountain Pit Viper | Indian Subcontinent, Southeast Asia |
| Ovophis okinavensis | Okinawa Pit Viper | Ryukyu Islands of Japan |
| Trimeresurus borneensis | Bornean Pit Viper | Brunei, Indonesia, Malaysia, Thailand |
| Trimeresurus fasciatus | Banded Pit Viper | Djampea Island (Indonesia) |
| Trimeresurus purpureomaculatus | Mangrove Pit Viper | Indian Subcontinent, Southeast Asia |
| Control | V | V/CO | V/iRM | V/PHA | |
|---|---|---|---|---|---|
| Trimeresurus borneensis (1 µg/mL) | |||||
| TMRTG | 11.5 ± 1.0 | 17.8 ± 2.1 * | 13.6 ± 3.6 | 25.2 ± 4.9 * † ‡ | 23.7 ± 2 † |
| MRTG | 3.6 ± 0.6 | 1.1 ± 0.3 * | 3.2 ± 1.4 † | 0.2 ± 0.2 * ‡ | 0.7 ± 0.1 † |
| TTG | 197 ± 14 | 144 ± 22 * | 186 ± 38 † | 28 ± 29 * † ‡ | 93 ± 19 † |
| Trimeresurus fasciatus (10 µg/mL) | |||||
| TMRTG | --- | 23.2 ± 3.6 * | 14.7 ± 3.4 † | 24.2 ± 6.5 * ‡ | 20.0 ± 1.9 |
| MRTG | --- | 1.0 ± 0.4 * | 3.0 ± 0.9 † | 0.5 ± 0.6 * ‡ | 1.8 ± 0.3 † |
| TTG | --- | 100 ± 26 * | 205 ± 16 † | 42 ± 52 * † ‡ | 166 ± 11 † |
| Trimeresurus purpureomaculatus (5 µg/mL) | |||||
| TMRTG | --- | 22.6 ± 2.2 * | 13.9 ± 3.2 † | 20.8 ± 1.8 * ‡ | 21.6 ± 4.1 |
| MRTG | --- | 0.4 ± 0.2 * | 2.7 ± 1.4 † | 0.5 ± 0.2 * ‡ | 0.6 ± 0.2 |
| TTG | --- | 57 ± 12 * | 190 ± 15 † | 70 ± 19 * ‡ | 87 ± 25 † |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nielsen, V.G.; Frank, N.; Matika, R.W. Effects of Heme Modulation on Ovophis and Trimeresurus Venom Activity in Human Plasma. Toxins 2018, 10, 322. https://doi.org/10.3390/toxins10080322
Nielsen VG, Frank N, Matika RW. Effects of Heme Modulation on Ovophis and Trimeresurus Venom Activity in Human Plasma. Toxins. 2018; 10(8):322. https://doi.org/10.3390/toxins10080322
Chicago/Turabian StyleNielsen, Vance G., Nathaniel Frank, and Ryan W. Matika. 2018. "Effects of Heme Modulation on Ovophis and Trimeresurus Venom Activity in Human Plasma" Toxins 10, no. 8: 322. https://doi.org/10.3390/toxins10080322
APA StyleNielsen, V. G., Frank, N., & Matika, R. W. (2018). Effects of Heme Modulation on Ovophis and Trimeresurus Venom Activity in Human Plasma. Toxins, 10(8), 322. https://doi.org/10.3390/toxins10080322





