Characterization of the Venom of C. d. cumanesis of Colombia: Proteomic Analysis and Antivenomic Study
Abstract
:1. Introduction
2. Results
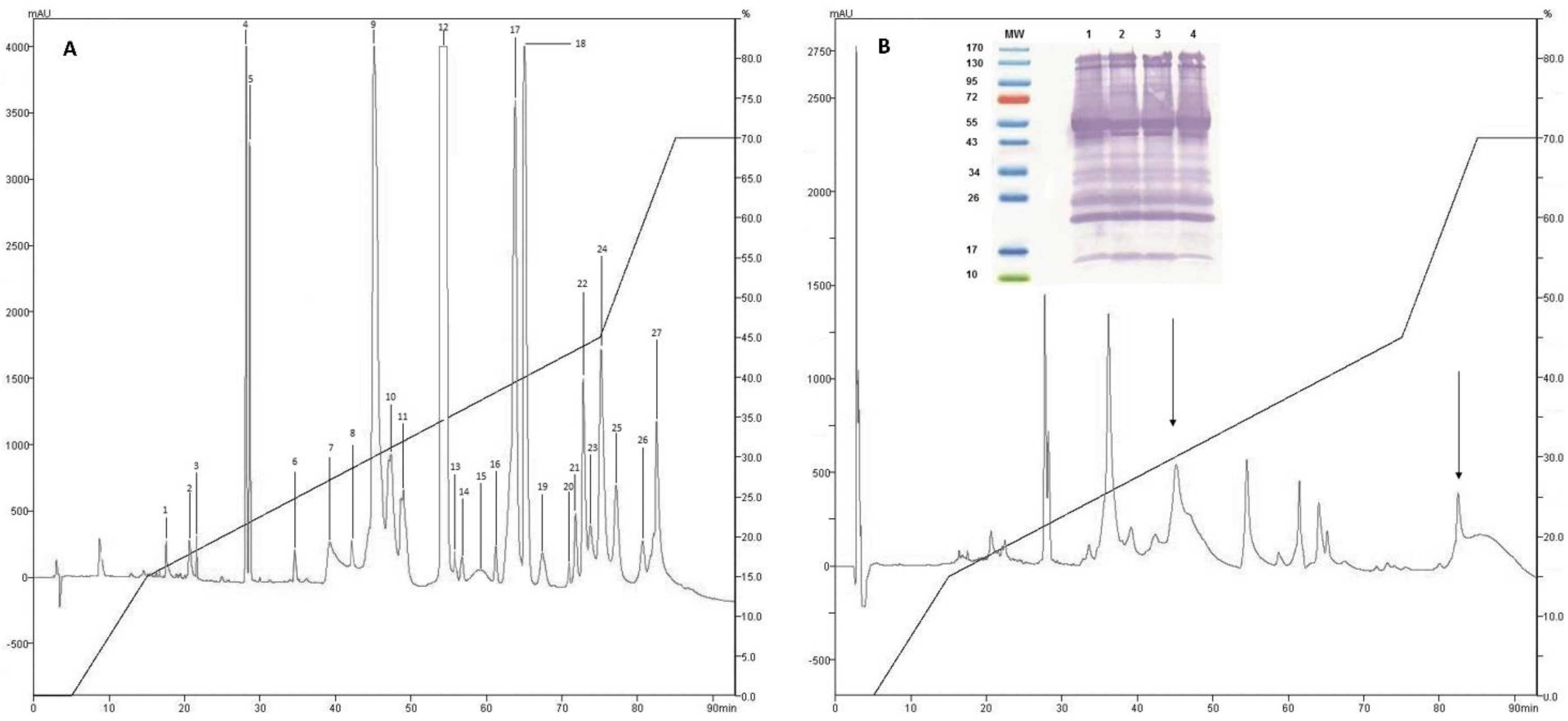
2.1. Isolation of Fractions of C. d. cumanensis
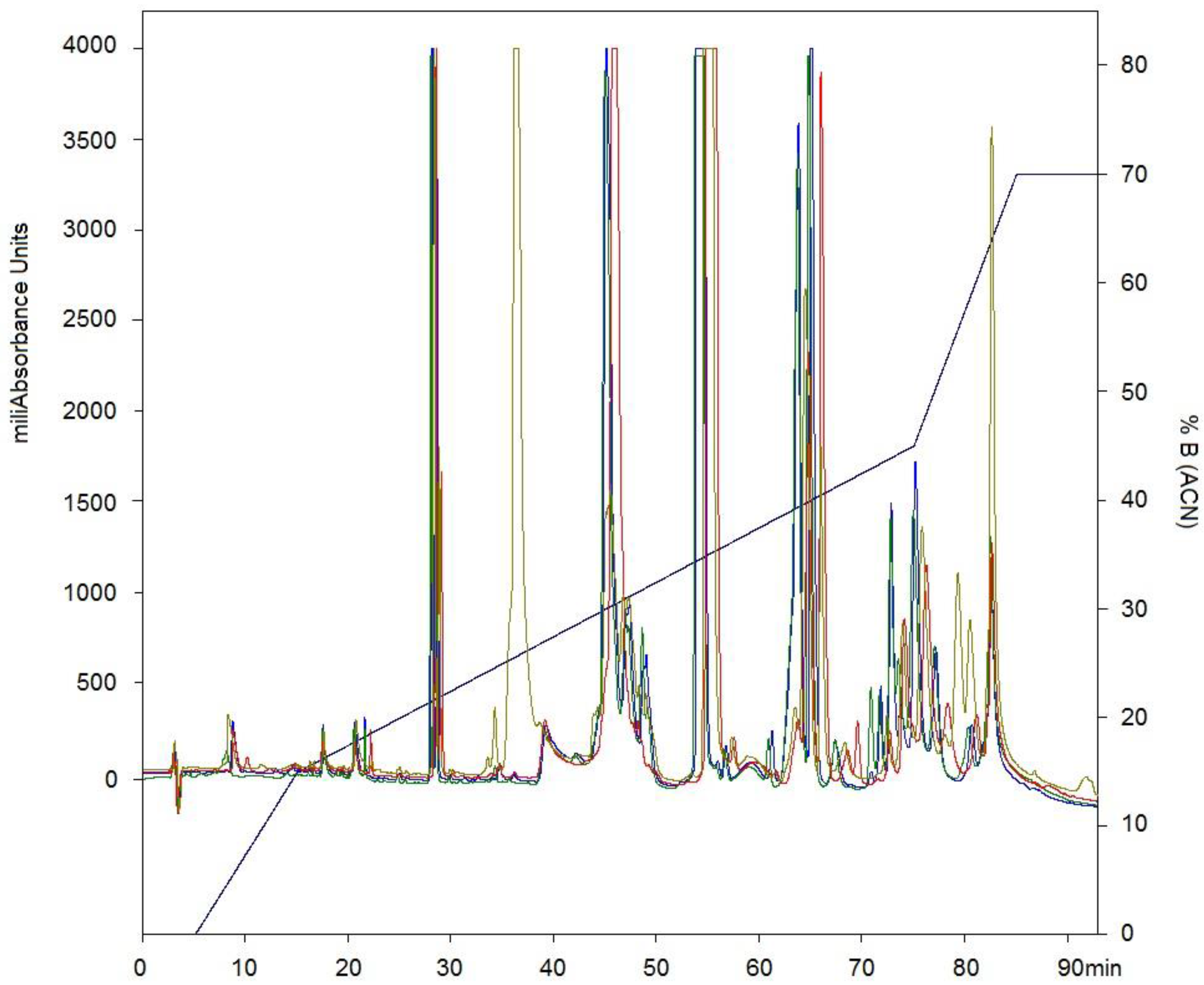
2.2. Western Blotting and Immunodepletion
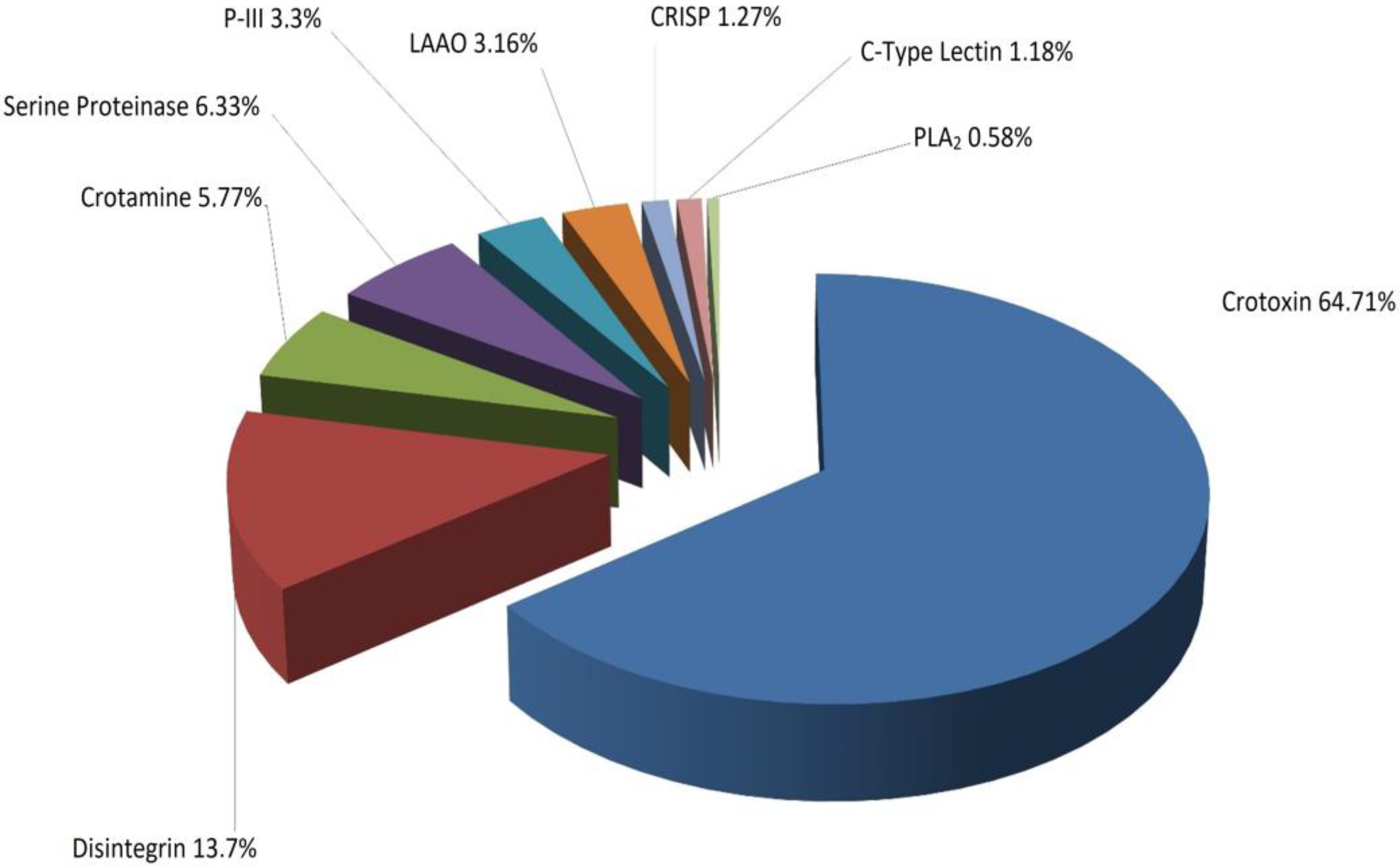
2.3. Identification of Proteins
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Venoms, Chemical Products, and Reagents
5.2. Isolation and Characterization of Venom Proteins
5.3. Electrophoresis and Determination of Molecular Mass
5.4. Identification of Proteins by HPLC-nESI-MS/MS
5.5. Immunodepletion of Venom Proteins by a Mexican Polyvalent Antivenom
5.6. Search Database
5.7. BLAST Search of the Identified Peptides
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Campbell, J.A.; Lamar, W. The Venomous Reptiles of the Western Hemisphere; Cornell University Press: New York, NY, USA, 2004; Volume II, p. 870. [Google Scholar]
- Klauber, L.M. Rattlesnakes: Their Habits, Life Histories, and Influence on Mankind; Zoological Society of San Diego: San Diego, CA, USA, 1972. [Google Scholar]
- Bucaretchi, F.; Herrera, S.R.; Hyslop, S.; Baracat, E.C.; Vieira, R.J. Snakebites by Crotalus durissus ssp. in children in Campinas, Sao Paulo, Brazil. Rev. Inst. Med. Trop. São Paulo 2002, 44, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Oshima-Franco, Y.; Hyslop, S.; Prado-Franceschi, J.; Cruz-Hofling, M.A.; Rodrigues-Simioni, L. Neutralizing capacity of antisera raised in horses and rabbits against Crotalus durissus terrificus (South American rattlesnake) venom and its main toxin, crotoxin. Toxicon 1999, 37, 1341–1357. [Google Scholar] [CrossRef]
- Bon, C. Multicomponent neurotoxic phospholipases A2. In Venom Phospholipase A2 Enzymes: Structure, Function and Mechanism; Wiley: Chichester, UK, 1997; pp. 269–285.A2. In Venom Phospholipase A2 Enzymes: Structure, Function and Mechanism; Wiley: Chichester, UK, 1997; pp. 269–285. [Google Scholar]
- Sampaio, S.C.; Hyslop, S.; Fontes, M.R.; Prado-Franceschi, J.; Zambelli, V.O.; Magro, A.J.; Brigatte, P.; Gutierrez, V.P.; Cury, Y. Crotoxin: Novel activities for a classic beta-neurotoxin. Toxicon 2010, 55, 1045–1060. [Google Scholar] [CrossRef] [PubMed]
- Azevedo-Marques, M.M.; Cupo, P.; Coimbra, T.M.; Hering, S.E.; Rossi, M.A.; Laure, C.J. Myonecrosis, myoglobinuria and acute renal failure induced by South American rattlesnake (Crotalus durissus terrificus) envenomation in Brazil. Toxicon 1985, 23, 631–636. [Google Scholar] [CrossRef]
- Martins, A.M.; Toyama, M.H.; Havt, A.; Novello, J.C.; Marangoni, S.; Fonteles, M.C.; Monteiro, H.S. Determination of Crotalus durissus cascavella venom components that induce renal toxicity in isolated rat kidneys. Toxicon 2002, 40, 1165–1171. [Google Scholar] [CrossRef]
- Cupo, P.; Azevedo-Marques, M.M.; Hering, S.E. Clinical and laboratory features of South American rattlesnake (Crotalus durissus terrificus) envenomation in children. Trans. R. Soc. Trop. Med. Hyg. 1988, 82, 924–929. [Google Scholar] [CrossRef]
- De Rezende, N.A.; Torres, F.M.; Dias, M.B.; Campolina, D.; Chavez-Olortegui, C.; Amaral, C.F. South American rattlesnake bite (Crotalus durissus sp.) without envenoming: Insights on diagnosis and treatment. Toxicon 1998, 36, 2029–2032. [Google Scholar] [CrossRef]
- Jorge, M.T.; Ribeiro, L.A. [the epidemiology and clinical picture of an accidental bite by the South American rattlesnake (Crotalus durissus)]. Rev. Inst. Med. Trop. São Paulo 1992, 34, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Pinho, F.M.; Zanetta, D.M.; Burdmann, E.A. Acute renal failure after Crotalus durissus snakebite: A prospective survey on 100 patients. Kidney Int. 2005, 67, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Pinho, F.O.; Zanetta, D.M.T.; Burdmann, E.A. Risk factors and prevalence of acute renal failure (ARF) after Crotalus snakebite (CSB): A prospective survey on 100 consecutive patients. J. Am. Soc. Nephrol. 2003, 14, 511a–512a. [Google Scholar]
- Sano-Martins, I.S.; Tomy, S.C.; Campolina, D.; Dias, M.B.; de Castro, S.C.; de Sousa-e-Silva, M.C.; Amaral, C.F.; Rezende, N.A.; Kamiguti, A.S.; Warrell, D.A.; et al. Coagulopathy following lethal and non-lethal envenoming of humans by the South American rattlesnake (Crotalus durissus) in Brazil. QJM 2001, 94, 551–559. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, J.M.; Leon, G.; Rojas, G.; Lomonte, B.; Rucavado, A.; Chaves, F. Neutralization of local tissue damage induced by Bothrops asper (terciopelo) snake venom. Toxicon 1998, 36, 1529–1538. [Google Scholar] [CrossRef]
- Leon, G.; Valverde, J.M.; Rojas, G.; Lomonte, B.; Gutierrez, J.M. Comparative study on the ability of IgG and Fab sheep antivenoms to neutralize local hemorrhage, edema and myonecrosis induced by Bothrops asper (terciopelo) snake venom. Toxicon 2000, 38, 233–244. [Google Scholar] [CrossRef]
- Theakston, R.D.G. The Kinetics of snakebite envenoming and therapy. In Proceedings of the 1st International Congress on Envenomations and their Treatments, Paris, France, 7–9 June 1995; p. 69. [Google Scholar]
- Sasa, M. Diet and snake venom evolution: Can local selection alone explain intraspecific venom variation? Toxicon 1999, 37, 249–252, author reply 253–260. [Google Scholar] [PubMed]
- Saldarriaga, M.M.; Otero, R.; Nunez, V.; Toro, M.F.; Diaz, A.; Gutierrez, J.M. Ontogenetic variability of Bothrops atrox and Bothrops asper snake venoms from Colombia. Toxicon 2003, 42, 405–411. [Google Scholar] [CrossRef]
- Menezes, M.C.; Furtado, M.F.; Travaglia-Cardoso, S.R.; Camargo, A.C.; Serrano, S.M. Sex-based individual variation of snake venom proteome among eighteen Bothrops jararaca siblings. Toxicon 2006, 47, 304–312. [Google Scholar] [CrossRef] [PubMed]
- Antunez, J.; Fernandez, J.; Lomonte, B.; Angulo, Y.; Sanz, L.; Perez, A.; Calvete, J.J.; Gutierrez, J.M. Antivenomics of Atropoides mexicanus and Atropoides picadoi snake venoms: Relationship to the neutralization of toxic and enzymatic activities. J. Venom. Res. 2010, 1, 8–17. [Google Scholar] [PubMed]
- Gutierrez, J.M.; dos Santos, M.C.; Furtado Mde, F.; Rojas, G. Biochemical and pharmacological similarities between the venoms of newborn Crotalus durissus durissus and adult Crotalus durissus terrificus rattlesnakes. Toxicon 1991, 29, 1273–1277. [Google Scholar] [CrossRef]
- Currier, R.B.; Harrison, R.A.; Rowley, P.D.; Laing, G.D.; Wagstaff, S.C. Intra-specific variation in venom of the african puff adder (bitis arietans): Differential expression and activity of snake venom metalloproteinases (SVMPS). Toxicon 2010, 55, 864–873. [Google Scholar] [CrossRef] [PubMed]
- Tun, P.; Nu Nu, L.; Aye Aye, M.; Kyi May, H.; Khin Aung, C. Biochemical and biological properties of the venom from Russell’s viper (Daboia russelli siamensis) of varying ages. Toxicon 1995, 33, 817–821. [Google Scholar] [CrossRef]
- Rael, E.D.; Rivas, J.Z.; Chen, T.; Maddux, N.; Huizar, E.; Lieb, C.S. Differences in fibrinolysis and complement inactivation by venom from different northern blacktailed rattlesnakes (Crotalus molossus molossus). Toxicon 1997, 35, 505–513. [Google Scholar] [CrossRef]
- Ogawa, T.; Nakashima, K.; Nobuhisa, I.; Deshimaru, M.; Shimohigashi, Y.; Fukumaki, Y.; Sakaki, Y.; Hattori, S.; Ohno, M. Accelerated evolution of snake venom phospholipase A2 isozymes for acquisition of diverse physiological functions. Toxicon 1996, 34, 1229–1236. [Google Scholar] [CrossRef]
- Monteiro, R.Q.; Carlini, C.R.; Guimaraes, J.A.; Bon, C.; Zingali, R.B. Distinct bothrojaracin isoforms produced by individual jararaca (Bothrops jararaca) snakes. Toxicon 1997, 35, 649–657. [Google Scholar] [CrossRef]
- Chippaux, J.P.; Williams, V.; White, J. Snake venom variability: Methods of study, results and interpretation. Toxicon 1991, 29, 1279–1303. [Google Scholar] [CrossRef]
- Boldrini-Franca, J.; Correa-Netto, C.; Silva, M.M.; Rodrigues, R.S.; De La Torre, P.; Perez, A.; Soares, A.M.; Zingali, R.B.; Nogueira, R.A.; Rodrigues, V.M.; et al. Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management. J. Proteom. 2010, 73, 1758–1776. [Google Scholar] [CrossRef] [PubMed]
- Markland, F.S. Snake venoms and the hemostatic system. Toxicon 1998, 36, 1749–1800. [Google Scholar] [CrossRef]
- Calvete, J.J.; Sanz, L.; Cid, P.; de la Torre, P.; Flores-Diaz, M.; Dos Santos, M.C.; Borges, A.; Bremo, A.; Angulo, Y.; Lomonte, B.; et al. Snake venomics of the central american rattlesnake crotalus simus and the South American Crotalus durissus complex points to neurotoxicity as an adaptive paedomorphic trend along Crotalus dispersal in South America. J. Proteom. Res. 2010, 9, 528–544. [Google Scholar] [CrossRef] [PubMed]
- Saravia, P.; Rojas, E.; Arce, V.; Guevara, C.; López, J.C.; Chaves, E.; Velásquez, R.; Rojas, G.; Gutiérrez, J.M. Geographic and ontogenic variability in the venom of the neotropical rattlesnake Crotalus durissus: Pathophysiological and therapeutic implications. Rev. Biol. Trop. 2002, 50, 337–346. [Google Scholar] [PubMed]
- Céspedes, N.; Castro, F.; Jiménez, E.; Montealegre, L.; Castellanos, A.; Cañas, C.; Arévalo-Herrera, M.; Herrera, S. Biochemical comparison of venoms from young colombian Crotalus durissus cumanensis and their parents. J. Venom. Anim. Toxins Incl. Trop. Dis. 2010, 16, 268–284. [Google Scholar] [CrossRef]
- Otero-Patino, R.; Segura, A.; Herrera, M.; Angulo, Y.; Leon, G.; Gutierrez, J.M.; Barona, J.; Estrada, S.; Pereanez, A.; Quintana, J.C.; et al. Comparative study of the efficacy and safety of two polyvalent, caprylic acid fractionated [IgG and F(ab’)2] antivenoms, in Bothrops asper bites in Colombia. Toxicon 2012, 59, 344–355. [Google Scholar] [CrossRef] [PubMed]
- Patino, A.C.; Lopez, J.; Aristizabal, M.; Quintana, J.C.; Benjumea, D. [evaluation of the inhibitory effect of extracts from leaves of Renealmia alpinia rottb. Maas (Zingiberaceae) on the venom of Bothrops asper (mapana)]. Biomedica 2012, 32, 365–374. [Google Scholar] [PubMed]
- Angulo, Y.; Escolano, J.; Lomonte, B.; Gutierrez, J.M.; Sanz, L.; Calvete, J.J. Snake venomics of central american pitvipers: Clues for rationalizing the distinct envenomation profiles of Atropoides nummifer and Atropoides picadoi. J. Proteom. Res. 2008, 7, 708–719. [Google Scholar] [CrossRef] [PubMed]
- Valente, R.H.; Guimaraes, P.R.; Junqueira, M.; Neves-Ferreira, A.G.; Soares, M.R.; Chapeaurouge, A.; Trugilho, M.R.; Leon, I.R.; Rocha, S.L.; Oliveira-Carvalho, A.L.; et al. Bothrops insularis venomics: A proteomic analysis supported by transcriptomic-generated sequence data. J. Proteom. 2009, 72, 241–255. [Google Scholar] [CrossRef] [PubMed]
- Tashima, A.K.; Sanz, L.; Camargo, A.C.; Serrano, S.M.; Calvete, J.J. Snake venomics of the brazilian pitvipers Bothrops cotiara and Bothrops fonsecai. Identification of taxonomy markers. J. Proteom. 2008, 71, 473–485. [Google Scholar] [CrossRef] [PubMed]
- Alape-Giron, A.; Sanz, L.; Escolano, J.; Flores-Diaz, M.; Madrigal, M.; Sasa, M.; Calvete, J.J. Snake venomics of the lancehead pitviper Bothrops asper: Geographic, individual, and ontogenetic variations. J. Proteom. Res. 2008, 7, 3556–3571. [Google Scholar] [CrossRef] [PubMed]
- Angulo, Y.; Lomonte, B. Biochemistry and toxicology of toxins purified from the venom of the snake Bothrops asper. Toxicon 2009, 54, 949–957. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Escolano, J.; Fernandez, J.; Sanz, L.; Angulo, Y.; Gutierrez, J.M.; Calvete, J.J. Snake venomics and antivenomics of the arboreal neotropical pitvipers Bothriechis lateralis and Bothriechis schlegelii. J. Proteom. Res. 2008, 7, 2445–2457. [Google Scholar] [CrossRef] [PubMed]
- Sanz, L.; Escolano, J.; Ferretti, M.; Biscoglio, M.J.; Rivera, E.; Crescenti, E.J.; Angulo, Y.; Lomonte, B.; Gutierrez, J.M.; Calvete, J.J. Snake venomics of the south and central american bushmasters. Comparison of the toxin composition of Lachesis muta gathered from proteomic versus transcriptomic analysis. J. Proteom. 2008, 71, 46–60. [Google Scholar] [CrossRef] [PubMed]
- Bogarin, G.; Romero, M.; Rojas, G.; Lutsch, C.; Casadamont, M.; Lang, J.; Otero, R.; Gutierrez, J.M. Neutralization, by a monospecific bothrops lanceolatus antivenom, of toxic activities induced by homologous and heterologous Bothirops snake venoms. Toxicon 1999, 37, 551–557. [Google Scholar] [CrossRef]
- Otero, R.; Gutierrez, J.; Beatriz Mesa, M.; Duque, E.; Rodriguez, O.; Luis Arango, J.; Gomez, F.; Toro, A.; Cano, F.; Maria Rodriguez, L.; et al. Complications of Bothrops, Porthidium, and Bothriechis snakebites in Colombia. A clinical and epidemiological study of 39 cases attended in a university hospital. Toxicon 2002, 40, 1107–1114. [Google Scholar] [CrossRef]
- Otero, R.; Leon, G.; Gutierrez, J.M.; Rojas, G.; Toro, M.F.; Barona, J.; Rodriguez, V.; Diaz, A.; Nunez, V.; Quintana, J.C.; et al. Efficacy and safety of two whole IgG polyvalent antivenoms, refined by caprylic acid fractionation with or without beta-propiolactone, in the treatment of Bothrops asper bites in Colombia. Trans. R. Soc. Trop. Med. Hyg. 2006, 100, 1173–1182. [Google Scholar] [CrossRef] [PubMed]
- Otero-Patino, R. Epidemiological, clinical and therapeutic aspects of Bothrops asper bites. Toxicon 2009, 54, 998–1011. [Google Scholar] [CrossRef] [PubMed]
- Otero-Patino, R.; Cardoso, J.L.; Higashi, H.G.; Nunez, V.; Diaz, A.; Toro, M.F.; Garcia, M.E.; Sierra, A.; Garcia, L.F.; Moreno, A.M.; et al. A randomized, blinded, comparative trial of one pepsin-digested and two whole IgG antivenoms for Bothrops snake bites in Uraba, Colombia. The Regional Group on Antivenom Therapy Research (REGATHER). Am. J. Trop. Med. Hyg. 1998, 58, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Perez, A.; Lomonte, B.; Sanchez, E.E.; Sanz, L. Snake venomics of crotalus tigris: The minimalist toxin arsenal of the deadliest nearctic rattlesnake venom. Evolutionary clues for generating a pan-specific antivenom against crotalid type ii venoms [corrected]. J. Proteom. Res. 2012, 11, 1382–1390. [Google Scholar] [CrossRef] [PubMed]
- Quintana, J.C.; Chacon, A.M.; Vargas, L.; Segura, C.; Gutierrez, J.M.; Alarcon, J.C. Antiplasmodial effect of the venom of Crotalus durissus cumanensis, crotoxin complex and crotoxin B. Acta Trop. 2012, 124, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- BLAST Search. Available online: http://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins (accessed on 12 May 2017).
- Mascot. Matrix Science Mascot Ms/Ms Ion Search. Available online: http://www.matrixscience.com/cgi/search_form.pl?FORMVER=2&SEARCH=MIS (accessed on 12 May 2017).
| HPLC Fraction | Molecular Mass | Peptide Ion | MS/MS Sequence | Score | Protein Family | ||
|---|---|---|---|---|---|---|---|
| Monoisotopic Mass | Z | Spectrum Mill | Mascot | ||||
| 1 | 7591.4 | 2.036.852 | 3 | LRPGAQCADGLCCDQCR | 15.75 | 42 | Disintegrin accession number SP|A2CJE7 |
| 2 | 7431.3 | 2.036.852 | 3 | LRPGAQCADGLCCDQCR | 20 | Disintegrin accession number SP|P21858.1 | |
| 2.242.927 | 3 | IARGDDMDDYCNGISAGCPR | 13.31 | ||||
| 3 | 7874.2 | 2.036.852 | 3 | LRPGAQCADGLCCDQCR | 19.9 | Disintegrin accession number SP|A2CJE7 | |
| 1.902.705 | 3 | GDDMDDYCNGISAGCPR | 12.7 | ||||
| 4 | 7545.6 | 2.050.867 | 2 | LRPGAQCAEGLCCDQCR | 18.89 | 27 | Disintegrin accession number SP|P21858.1 |
| 5 | 7762.9 | 2.050.867 | 3 | LRPGAQCAEGLCCDQCR | 17.25 | Disintegrin accession number SP|P21858.1 | |
| 6 | 7703.7 | 2.036.852 | 3 | LRPGAQCADGLCCDQCR | 19.45 | 42 | Disintegrin accession number SP|A2CJE7 |
| 1.954.729 | 3 | GDWNDDTCTGQSADCPR | 14.33 | ||||
| 1.230.730 | 2 | EAGEECDCGTPGNPCCDAATCK | 53 | ||||
| 7 | 10,730.4 | 1.549.533 | 2 | CCFEHDCCYAK | 15.7 | 35 | Crotoxin acid chain accession number SP|P08878 |
| 1.661.748 | 3 | LTGCDPTTDVYTYR | 22.08 | ||||
| 8 | 4809.9 | 864.349 | 2 | MDCPWR | 6.61 | 30 | Crotamine accession number SP|P24331 |
| 1220.105 | 2 | ICIPPSSDFGK | 7.13 | 41 | |||
| 9 | 4810.2 | 1477.736 | 3 | EKICIPPSSDFGK | 11.5 | 61 | Crotamine accession number SP|P24331 |
| 1220.105 | 2 | ICIPPSSDFGK | 6.43 | 69 | |||
| 13,856.2 | 1137.419 | 2 | NCLEESEPC | 12.29 | PLA2-01 accession number GB|BAA08383.1 | ||
| 10 | 10,319.1 | 1549.533 | 3 | CCFEHDCCYAK | 15.7 | 20 | Crotoxin acid chain accession number SP|P08878 |
| 808.413 | 2 | AAAICFR | 12.22 | 35 | |||
| 1294.425 | 2 | GGHGRPQDASDR | 19 | ||||
| 1539.185 | 2 | FSPENCQGESQPC | 54 | ||||
| 11 | 10,400.6 | 1549.533 | 3 | CCFEHDCCYAK | 12.56 | 14 | Crotoxin acid chain accession number SP|P08878 |
| 808.413 | 2 | AAAICFR | 9.54 | 25 | |||
| 1539.185 | 2 | FSPENCQGESQPC | 48 | ||||
| 12 | 7502.4 | 2036.852 | 2 | LRPGAQCADGLCCDQCR | 19.9 | Disintegrin accession number SP|A2CJE7 | |
| 10040.7 | 1549.533 | 3 | CCFEHDCCYAK | 14.43 | 59 | Crotoxin acid chain accession number SP|P08878 | |
| 13 | 10,043.2 | 1549.533 | 3 | CCFEHDCCYAK | 11.31 | Crotoxin acid chain accession number SP|P08878 | |
| 2278.725 | 2 | SSYGCYCGAGGQGWPQDASDR | 57 | ||||
| 808.413 | 2 | AAAICFR | 31 | ||||
| 14 | 10,538.0 | 1661.748 | 2 | LTGCDPTTDVYTYR | 19.39 | 44 | Crotoxin acid chain accession number SP|P08878 |
| 808.413 | 2 | AAAICFR | 33 | ||||
| 2278.725 | 2 | SSYGCYCGAGGQGWPQDASDR | 61 | ||||
| 15 | 14,538.0 | 1298.551 | 2 | CCFVHDCCYGK | 15.89 | 40 | Crotoxin basic chain accession number SP|P62022 |
| 1297.825 | 2 | YGYMFYPDSR | 6.35 | 45 | |||
| 16 | 13,970.6 | 2287.021 | 3 | KNAIPFYAFYGCYCGWGGR | 21.42 | 44 | Crotoxin basic chain accession number SP|P62022 |
| 1687.815 | 3 | CNTKWDIYPYSLK | 21.08 | ||||
| 1297.825 | 2 | YGYMFYPDSR | 12.53 | 54 | |||
| 1977.905 | 3 | SLSTYKYGYMFYPDSR | 20.01 | 49 | |||
| 966.377 | 2 | CRGPSETC | 7.5 | ||||
| 17 | 14,464.4 | 1687.815 | 3 | CNTKWDIYPYSLK | 15.76 | 43 | Crotoxin basic chain accession number SP|P62022 |
| 1505.543 | 3 | CCFVHDCCYGK | 14.39 | 47 | |||
| 10,925.4 | 1549.533 | 2 | CCFEHDCCYAK | 8.69 | Crotoxin acid chain accession number SP|P08878 | ||
| 25,435.6 | 2202.999 | 3 | YSYFYVCQYCPAGNIIGK | 12.13 | Cysteine-rich secretory protein accession number GB|ACE73575.1 | ||
| 18 | 13,993.8 | 1505.543 | 3 | CCFVHDCCYGK | 11.31 | 26 | Crotoxin basic chain accession number SP|P62022 |
| 1297.825 | 2 | YGYMFYPDSR | 54 | ||||
| 29,842.0 | 2890.425 | 3 | LDSPVSDSEHIAPLSLPSSPPSVGSVCR | 18.46 | Serine proteinase accession number SP|Q5W959 | ||
| 27,298.5 | 2291.145 | 3 | NSAHIEPLSLPSSPPSVGSVCR | 13.59 | Serine proteinase accession number SP|Q71QJ2 | ||
| 19 | 29,420.1 | 2142.042 | 2 | LLDDAVCQPPYPELPATSR | 13.76 | 33 | Kallikrein-like accession number SP|Q8QHK2 |
| 1068.825 | 2 | EKFFCPNK | 30 | ||||
| 1390.704 | 3 | TLPDVPYCANIK | 6.51 | 34 | |||
| 20 | 14,123.2 | 1505.543 | 2 | CCFVHDCCYGK | 15.57 | 26 | Crotoxin basic chain accession number SP|P62022 |
| 21 | 14,735.6 | 1505.543 | 3 | CCFVHDCCYGK | 11.41 | Crotoxin basic chain accession number SP|P62022 | |
| 22 | 17,789.3 | 1278.578 | 2 | LWNDQVCESK | 10 | 62 | C-type lectin accession number SP|P84987 |
| 1916.891 | 3 | YGESLEIAEYISDYHK | 9 | 56 | |||
| 23 | 14,319.1 | 1505.543 | 3 | CCFVHDCCYGK | 13.43 | Crotoxin basic chain accession number SP|P62022 | |
| 27,292.3 | 1136.57 | 2 | SVQFDKEQR | 5.61 | Serine proteinase gyroxin-like accession number GB|ABY65930 | ||
| 24 | 56,513.5 | 1236.658 | 2 | SAAQLYVESLR | 16.19 | 72 | LAAO accession number SP|P56742 |
| 1165.698 | 2 | IKFEPPLPPK | 8.82 | 32 | |||
| 970.522 | 2 | VQVHFNAR | 8.87 | 51 | |||
| 2065.063 | 3 | DCADIVINDLSLIHELPK | 10.22 | ||||
| 25 | 54,609.0 | 1236.658 | 2 | SAAQLYVESLR | 16.33 | 76 | LAAO accession number SP|P56742 |
| 2065.063 | 3 | DCADIVINDLSLIHELPK | 8.94 | ||||
| 1222.567 | 2 | DWYANLGPMR | 6.97 | ||||
| 26 | 51,346.6 | 1851.008 | 3 | KKHDNAQLLTAIDLDR | 15.17 | P-III metalloproteinase accession number GB|ACV83931 | |
| 1160.693 | 2 | FVELVLVVDK | 15.94 | 71 | |||
| 2052.055 | 3 | ITVKPEAGYTLNAFGEWR | 12.2 | ||||
| 27 | 49,476.6 | 1160.693 | 2 | FVELVLVVDK | 16.51 | 68 | P-III metalloproteinase accession number GB|ACV83931 |
| 1570.753 | 3 | ENGNKIPCAPEDVK | 9.12 | ||||
| 1052.425 | 2 | GNYYGYCR | 9.05 | ||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Quintana-Castillo, J.C.; Vargas, L.J.; Segura, C.; Estrada-Gómez, S.; Bueno-Sánchez, J.C.; Alarcón, J.C. Characterization of the Venom of C. d. cumanesis of Colombia: Proteomic Analysis and Antivenomic Study. Toxins 2018, 10, 85. https://doi.org/10.3390/toxins10020085
Quintana-Castillo JC, Vargas LJ, Segura C, Estrada-Gómez S, Bueno-Sánchez JC, Alarcón JC. Characterization of the Venom of C. d. cumanesis of Colombia: Proteomic Analysis and Antivenomic Study. Toxins. 2018; 10(2):85. https://doi.org/10.3390/toxins10020085
Chicago/Turabian StyleQuintana-Castillo, Juan Carlos, Leidy Johana Vargas, Cesar Segura, Sebastián Estrada-Gómez, Julio César Bueno-Sánchez, and Juan Carlos Alarcón. 2018. "Characterization of the Venom of C. d. cumanesis of Colombia: Proteomic Analysis and Antivenomic Study" Toxins 10, no. 2: 85. https://doi.org/10.3390/toxins10020085
APA StyleQuintana-Castillo, J. C., Vargas, L. J., Segura, C., Estrada-Gómez, S., Bueno-Sánchez, J. C., & Alarcón, J. C. (2018). Characterization of the Venom of C. d. cumanesis of Colombia: Proteomic Analysis and Antivenomic Study. Toxins, 10(2), 85. https://doi.org/10.3390/toxins10020085





