PhcrTx2, a New Crab-Paralyzing Peptide Toxin from the Sea Anemone Phymanthus crucifer
Abstract
:1. Introduction
2. Results
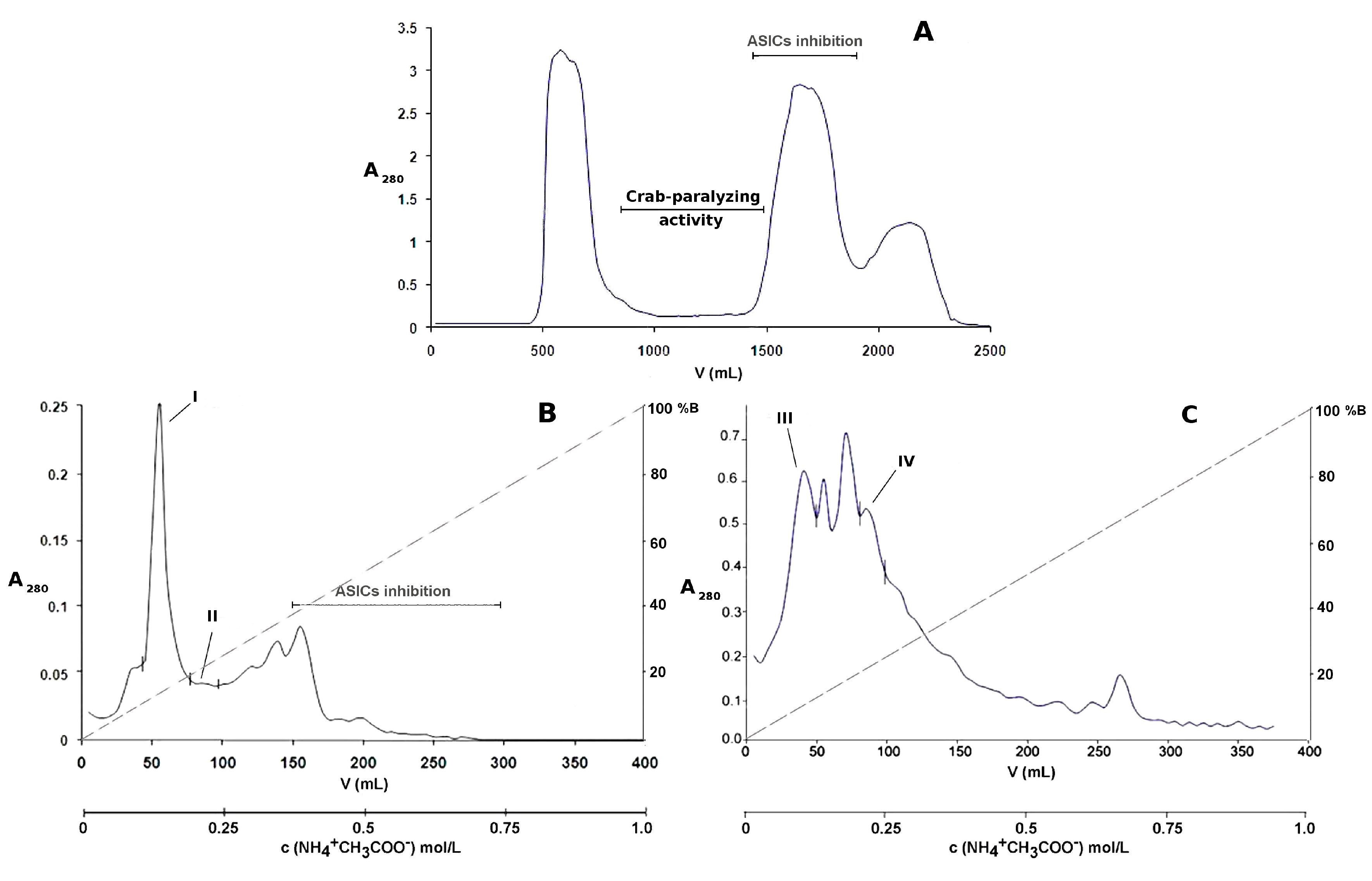
2.1. Bioassay-Guided Purification of P. crucifer Toxins
2.2. Biological Evaluation of PhcrTx2
2.2.1. Crab Toxicity Assay
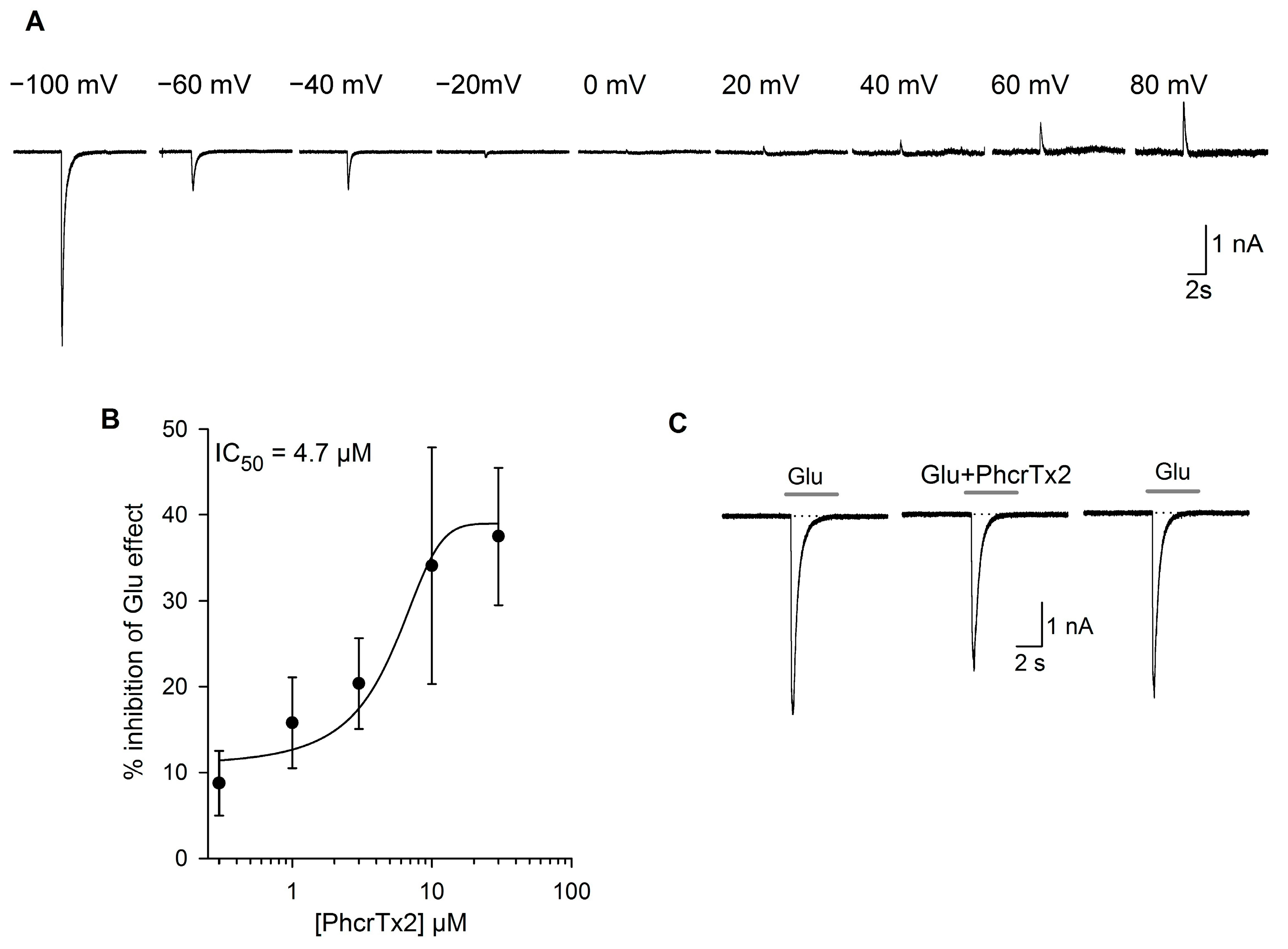
2.2.2. Effects of PhcrTx2 on Native Nav, Kv and Glutamate-Gated Currents
PhcrTx2 Evaluation on Snail Neurons
PhcrTx2 Evaluation on Rat DRG Neurons
2.2.3. Effects of PhcrTx2 on Cloned Voltage-Gated Ion Channels
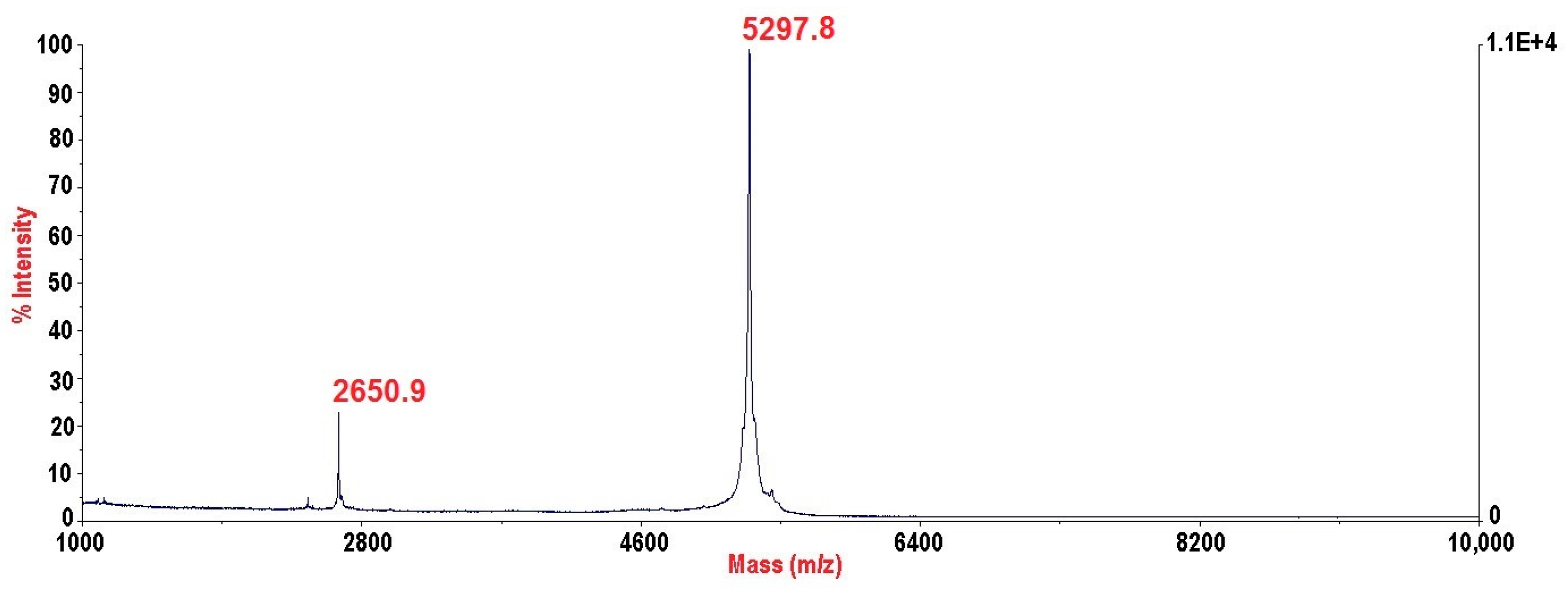
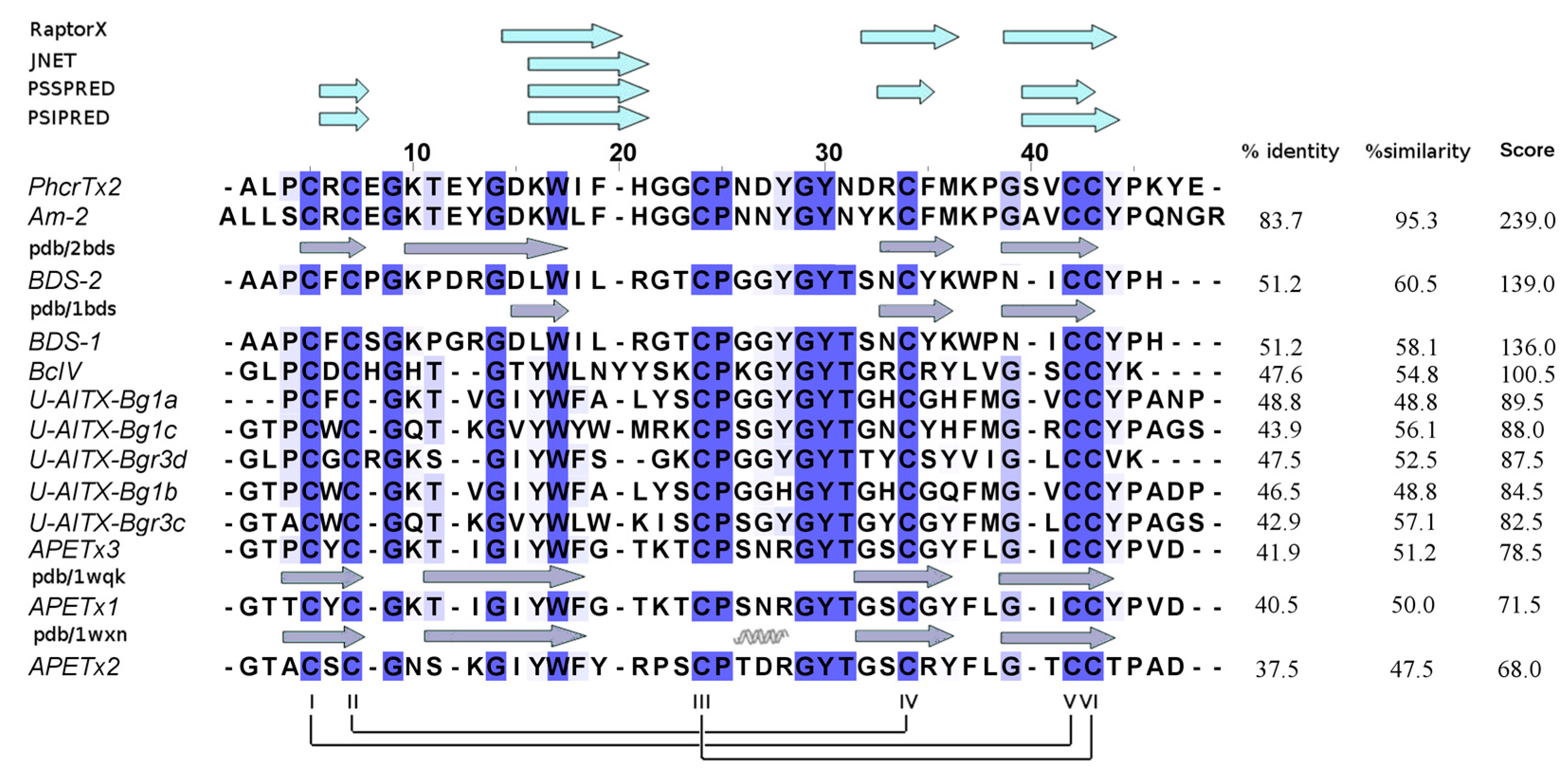
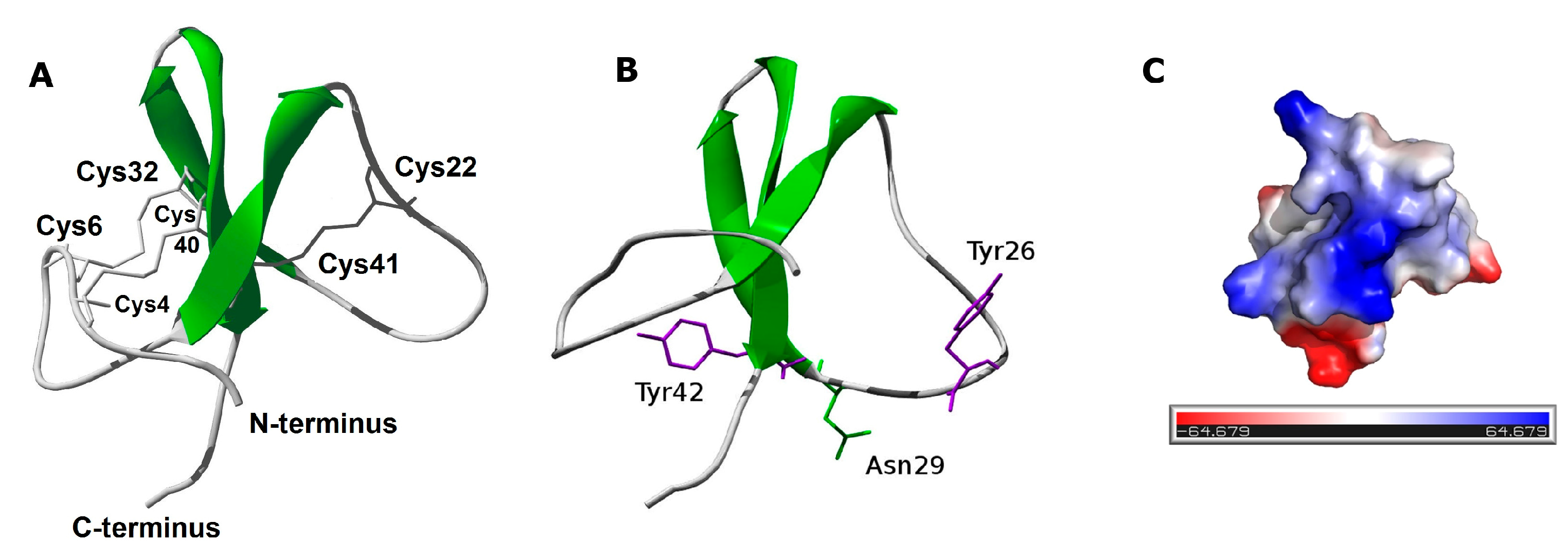
2.3. PhcrTx2 Sequence and Computational Analysis
3. Discussion
3.1. P. crucifer Produces a Diversity of Crab-Paralyzing Toxins
3.2. PhcrTx2 Is a New Defensin-Like Toxin
3.3. PhcrTx2 Is a Crab-Paralyzing Toxin Exhibiting a Low Potency Activity on Ion Channels
4. Conclusions
5. Materials and Methods
5.1. Chromatographic Separation of the Toxic Compounds
5.2. Biological Assays
5.2.1. Crab Toxicity Assay
5.2.2. Patch Clamp Experiments on Neuronal Ion Channels
Cell Culture of Rat Dorsal Root Ganglion (DRG) Neurons and Electrophysiological Recordings
Voltage Clamp Experiments on Isolated Snail Neurons
5.2.3. Expression of Voltage-Gated Ion Channels in Xenopus laevis Oocytes and Electrophysiological Recordings
5.3. Mass Spectrometry and N-Terminal Sequencing
5.4. Computational Analyses
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Oliveira, J.S.; Fuentes-Silva, D.; King, G.F. Development of a rational nomenclature for naming peptide and protein toxins from sea anemones. Toxicon 2012, 60, 539–550. [Google Scholar] [CrossRef] [PubMed]
- Honma, T.; Shiomi, K. Peptide toxins in sea anemones: Structural and functional aspects. Mar. Biotechnol. (N. Y.) 2006, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.C. Physiologic principles underlying ion channelopathies. Neurotherapeutics 2007, 4, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Osmakov, D.I.; Kozlov, S.A.; Andreev, Y.A.; Koshelev, S.G.; Sanamyan, N.P.; Sanamyan, K.E.; Dyachenko, I.A.; Bondarenko, D.A.; Murashev, A.N.; Mineev, K.S.; et al. Sea anemone peptide with uncommon beta-hairpin structure inhibits acid-sensing ion channel 3 (asic3) and reveals analgesic activity. J. Biol. Chem. 2013, 288, 23116–23127. [Google Scholar] [CrossRef] [PubMed]
- Peigneur, S.; Beress, L.; Moller, C.; Mari, F.; Forssmann, W.G.; Tytgat, J. A natural point mutation changes both target selectivity and mechanism of action of sea anemone toxins. FASEB J. 2012, 26, 5141–5151. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, A.A.; Salceda, E.; Garateix, A.G.; Zaharenko, A.J.; Peigneur, S.; Lopez, O.; Pons, T.; Richardson, M.; Diaz, M.; Hernandez, Y.; et al. A novel sea anemone peptide that inhibits acid-sensing ion channels. Peptides 2014, 53, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Putnam, N.H.; Srivastava, M.; Hellsten, U.; Dirks, B.; Chapman, J.; Salamov, A.; Terry, A.; Shapiro, H.; Lindquist, E.; Kapitonov, V.V.; et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 2007, 317, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Sabourault, C.; Ganot, P.; Deleury, E.; Allemand, D.; Furla, P. Comprehensive est analysis of the symbiotic sea anemone, Anemonia viridis. BMC Genom. 2009, 10, 333. [Google Scholar] [CrossRef] [PubMed]
- Cassoli, J.S.; Verano-Braga, T.; Oliveira, J.S.; Montandon, G.G.; Cologna, C.T.; Peigneur, S.; Pimenta, A.M.; Kjeldsen, F.; Roepstorff, P.; Tytgat, J.; et al. The proteomic profile of Stichodactyla duerdeni secretion reveals the presence of a novel o-linked glycopeptide. J. Proteom. 2013, 87, 89–102. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, S.; Grishin, E. The mining of toxin-like polypeptides from est database by single residue distribution analysis. BMC Genom. 2011, 12, 88. [Google Scholar] [CrossRef] [PubMed]
- Moran, Y.; Praher, D.; Schlesinger, A.; Ayalon, A.; Tal, Y.; Technau, U. Analysis of soluble protein contents from the nematocysts of a model sea anemone sheds light on venom evolution. Mar. Biotechnol. (N. Y.) 2013, 15, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Zaharenko, A.J.; Ferreira, W.A., Jr.; Oliveira, J.S.; Richardson, M.; Pimenta, D.C.; Konno, K.; Portaro, F.C.; de Freitas, J.C. Proteomics of the neurotoxic fraction from the sea anemone Bunodosoma cangicum venom: Novel peptides belonging to new classes of toxins. Comp. Biochem. Physiol. Part D Genom. Proteom. 2008, 3, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, A.A.; Cassoli, J.S.; Sa, F.; Dong, Z.Q.; de Freitas, J.C.; Pimenta, A.M.; de Lima, M.E.; Konno, K.; Lee, S.M.; Garateix, A.; et al. Peptide fingerprinting of the neurotoxic fractions isolated from the secretions of sea anemones Stichodactyla helianthus and Bunodosoma granulifera. New members of the apetx-like family identified by a 454 pyrosequencing approach. Peptides 2012, 34, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Madio, B.; Undheim, E.A.B.; King, G.F. Revisiting venom of the sea anemone Stichodactyla haddoni: Omics techniques reveal the complete toxin arsenal of a well-studied sea anemone genus. J. Proteom. 2017, 166, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Orts, D.J.; Moran, Y.; Cologna, C.T.; Peigneur, S.; Madio, B.; Praher, D.; Quinton, L.; De Pauw, E.; Bicudo, J.E.; Tytgat, J.; et al. Bcstx3 is a founder of a novel sea anemone toxin family of potassium channel blocker. FEBS J. 2013, 280, 4839–4852. [Google Scholar] [CrossRef] [PubMed]
- Shiomi, K.; Qian, W.H.; Lin, X.Y.; Shimakura, K.; Nagashima, Y.; Ishida, M. Novel polypeptide toxins with crab lethality from the sea anemone anemonia erythraea. Biochim. Biophys. Acta 1997, 1335, 191–198. [Google Scholar] [CrossRef]
- Béress, L.; Béress, R. Reinigung zweier krabbenlähmender toxine aus der seeanemone Anemonia sulcata. Kiel. Meeresforsch 1971, 27, 117–127. [Google Scholar]
- Shapiro, B.I. Purification of a toxin from tentacles of the anemone Condylactis gigantea. Toxicon 1968, 5, 253–259. [Google Scholar] [CrossRef]
- Maeda, M.; Honma, T.; Shiomi, K. Isolation and cdna cloning of type 2 sodium channel peptide toxins from three species of sea anemones (Cryptodendrum adhaesivum, Heterodactyla hemprichii and Thalassianthus aster) belonging to the family thalassianthidae. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2010, 157, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Honma, T.; Kawahata, S.; Ishida, M.; Nagai, H.; Nagashima, Y.; Shiomi, K. Novel peptide toxins from the sea anemone Stichodactyla haddoni. Peptides 2008, 29, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Honma, T.; Hasegawa, Y.; Ishida, M.; Nagai, H.; Nagashima, Y.; Shiomi, K. Isolation and molecular cloning of novel peptide toxins from the sea anemone Antheopsis maculata. Toxicon 2005, 45, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Shiomi, K.; Honma, T.; Ide, M.; Nagashima, Y.; Ishida, M.; Chino, M. An epidermal growth factor-like toxin and two sodium channel toxins from the sea anemone Stichodactyla gigantea. Toxicon 2003, 41, 229–236. [Google Scholar] [CrossRef]
- Bruhn, T.; Schaller, C.; Schulze, C.; Sanchez-Rodriguez, J.; Dannmeier, C.; Ravens, U.; Heubach, J.F.; Eckhardt, K.; Schmidtmayer, J.; Schmidt, H.; et al. Isolation and characterisation of five neurotoxic and cardiotoxic polypeptides from the sea anemone Anthopleura elegantissima. Toxicon 2001, 39, 693–702. [Google Scholar] [CrossRef]
- Ishida, M.; Yokoyama, A.; Shimakura, K.; Nagashima, Y.; Shiomi, K. Halcurin, a polypeptide toxin from the sea anemone Halcurias sp., with a structural resemblance to type 1 and 2 toxins. Toxicon 1997, 35, 537–544. [Google Scholar] [CrossRef]
- Lin, X.Y.; Ishida, M.; Nagashima, Y.; Shiomi, K. A polypeptide toxin in the sea anemone Actinia equina homologous with other sea anemone sodium channel toxins: Isolation and amino acid sequence. Toxicon 1996, 34, 57–65. [Google Scholar] [CrossRef]
- Schweitz, H.; Bidard, J.N.; Frelin, C.; Pauron, D.; Vijverberg, H.P.; Mahasneh, D.M.; Lazdunski, M.; Vilbois, F.; Tsugita, A. Purification, sequence, and pharmacological properties of sea anemone toxins from radianthus paumotensis. A new class of sea anemone toxins acting on the sodium channel. Biochemistry 1985, 24, 3554–3561. [Google Scholar] [CrossRef] [PubMed]
- Ständker, L.; Beress, L.; Garateix, A.; Christ, T.; Ravens, U.; Salceda, E.; Soto, E.; John, H.; Forssmann, W.G.; Aneiros, A. A new toxin from the sea anemone Condylactis gigantea with effect on sodium channel inactivation. Toxicon 2006, 48, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Shiomi, K. Novel peptide toxins recently isolated from sea anemones. Toxicon 2009, 54, 1112–1118. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, A.; Takeuchi, N. The effect on crayfish muscle of iontophoretically applied glutamate. J. Physiol. 1964, 170, 296–317. [Google Scholar] [CrossRef] [PubMed]
- Garateix, A.; Flores, A.; Garcia-Andrade, J.M.; Palmero, A.; Aneiros, A.; Vega, R.; Soto, E. Antagonism of glutamate receptors by a chromatographic fraction from the exudate of the sea anemone Phyllactis flosculifera. Toxicon 1996, 34, 443–450. [Google Scholar] [CrossRef]
- Lewis, R.J.; Dutertre, S.; Vetter, I.; Christie, M.J. Conus venom peptide pharmacology. Pharmacol. Rev. 2012, 64, 259–298. [Google Scholar] [CrossRef] [PubMed]
- De Figueiredo, S.G.; de Lima, M.E.; Nascimento Cordeiro, M.; Diniz, C.R.; Patten, D.; Halliwell, R.F.; Gilroy, J.; Richardson, M. Purification and amino acid sequence of a highly insecticidal toxin from the venom of the brazilian spider Phoneutria nigriventer which inhibits nmda-evoked currents in rat hippocampal neurones. Toxicon 2001, 39, 309–317. [Google Scholar] [CrossRef]
- Rodriguez, A.A.; Standker, L.; Zaharenko, A.J.; Garateix, A.G.; Forssmann, W.G.; Beress, L.; Valdes, O.; Hernandez, Y.; Laguna, A. Combining multidimensional liquid chromatography and maldi-tof-ms for the fingerprint analysis of secreted peptides from the unexplored sea anemone species Phymanthus crucifer. J. Chromatogr. B Anal. Technol. Biomed. Life. Sci. 2012, 903, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Yeung, S.Y.; Thompson, D.; Wang, Z.; Fedida, D.; Robertson, B. Modulation of kv3 subfamily potassium currents by the sea anemone toxin bds: Significance for cns and biophysical studies. J. Neurosci. 2005, 25, 8735–8745. [Google Scholar] [CrossRef] [PubMed]
- Diochot, S.; Schweitz, H.; Beress, L.; Lazdunski, M. Sea anemone peptides with a specific blocking activity against the fast inactivating potassium channel kv3.4. J. Biol. Chem. 1998, 273, 6744–6749. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Jo, S.; Bean, B.P. Modulation of neuronal sodium channels by the sea anemone peptide bds-i. J. Neurophysiol. 2012, 107, 3155–3167. [Google Scholar] [CrossRef] [PubMed]
- Llewellyn, L.E.; Norton, R.S. Binding of the sea anemone polypeptide bds ii to the voltage-gated sodium channel. Biochem. Int. 1991, 24, 937–946. [Google Scholar] [PubMed]
- Diochot, S.; Baron, A.; Rash, L.D.; Deval, E.; Escoubas, P.; Scarzello, S.; Salinas, M.; Lazdunski, M. A new sea anemone peptide, apetx2, inhibits asic3, a major acid-sensitive channel in sensory neurons. EMBO J. 2004, 23, 1516–1525. [Google Scholar] [CrossRef] [PubMed]
- Driscoll, P.C.; Gronenborn, A.M.; Beress, L.; Clore, G.M. Determination of the three-dimensional solution structure of the antihypertensive and antiviral protein bds-i from the sea anemone Anemonia sulcata: A study using nuclear magnetic resonance and hybrid distance geometry-dynamical simulated annealing. Biochemistry 1989, 28, 2188–2198. [Google Scholar] [CrossRef] [PubMed]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J.; et al. The pfam protein families database. Nucleic Acids Res. 2012, 40, D290–D301. [Google Scholar] [CrossRef] [PubMed]
- Chagot, B.; Diochot, S.; Pimentel, C.; Lazdunski, M.; Darbon, H. Solution structure of apetx1 from the sea anemone Anthopleura elegantissima: A new fold for an herg toxin. Proteins 2005, 59, 380–386. [Google Scholar] [CrossRef] [PubMed]
- Diochot, S.; Loret, E.; Bruhn, T.; Beress, L.; Lazdunski, M. Apetx1, a new toxin from the sea anemone Anthopleura elegantissima, blocks voltage-gated human ether-a-go-go-related gene potassium channels. Mol. Pharmacol. 2003, 64, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Chagot, B.; Escoubas, P.; Diochot, S.; Bernard, C.; Lazdunski, M.; Darbon, H. Solution structure of apetx2, a specific peptide inhibitor of asic3 proton-gated channels. Protein Sci. 2005, 14, 2003–2010. [Google Scholar] [CrossRef] [PubMed]
- Torres, A.M.; Kuchel, P.W. The beta-defensin-fold family of polypeptides. Toxicon 2004, 44, 581–588. [Google Scholar] [CrossRef] [PubMed]
- Janda, J.O.; Meier, A.; Merkl, R. Clips-4d: A classifier that distinguishes structurally and functionally important residue-positions based on sequence and 3d data. Bioinformatics 2013, 29, 3029–3035. [Google Scholar] [CrossRef] [PubMed]
- Anangi, R.; Rash, L.D.; Mobli, M.; King, G.F. Functional expression in Escherichia coli of the disulfide-rich sea anemone peptide apetx2, a potent blocker of acid-sensing ion channel 3. Mar. Drugs 2012, 10, 1605–1618. [Google Scholar] [CrossRef] [PubMed]
- Anangi, R.; Chen, C.C.; Lin, Y.W.; Cheng, Y.R.; Cheng, C.H.; Chen, Y.C.; Chu, Y.P.; Chuang, W.J. Expression in Pichia pastoris and characterization of apetx2, a specific inhibitor of acid sensing ion channel 3. Toxicon 2010, 56, 1388–1397. [Google Scholar] [CrossRef] [PubMed]
- Jensen, J.E.; Mobli, M.; Brust, A.; Alewood, P.F.; King, G.F.; Rash, L.D. Cyclisation increases the stability of the sea anemone peptide apetx2 but decreases its activity at acid-sensing ion channel 3. Mar. Drugs 2012, 10, 1511–1527. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, J.S.; Fuentes-Silva, D.; Zaharenko, A.J. Sea anemone peptides. Biological activities, structure-function relationships and phylogenetic aspects. In Animal Toxins: State of the Art. Perspective in Health and Biotechnology, 1st ed.; de Lima, M.E., Pimenta, A.M., Martin-Eauclaire, M.F., Zingali, R.B., Rochat, H., Eds.; Editora UFMG: Belo, Horizonte, 2009. [Google Scholar]
- Bosmans, F.; Tytgat, J. Sea anemone venom as a source of insecticidal peptides acting on voltage-gated Na+ channels. Toxicon 2007, 49, 550–560. [Google Scholar] [CrossRef] [PubMed]
- Walker, R.J.; Roberts, C.J. The pharmacology of Limulus central neurons. Comp. Biochem. Physiol. C 1982, 72, 391–401. [Google Scholar] [CrossRef]
- Salceda, E.; Perez-Castells, J.; Lopez-Mendez, B.; Garateix, A.; Salazar, H.; Lopez, O.; Aneiros, A.; Standker, L.; Beress, L.; Forssmann, W.G.; et al. Cgna, a type i toxin from the giant caribbean sea anemone Condylactis gigantea shows structural similarities to both type i and ii toxins, as well as distinctive structural and functional properties(1). Biochem. J. 2007, 406, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Salceda, E.; Garateix, A.; Aneiros, A.; Salazar, H.; Lopez, O.; Soto, E. Effects of apc, a sea anemone toxin, on sodium currents of mammalian neurons. Brain Res. 2006, 1110, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Garateix, A.; Salceda, E.; Menendez, R.; Regalado, E.L.; Lopez, O.; Garcia, T.; Morales, R.A.; Laguna, A.; Thomas, O.P.; Soto, E. Antinociception produced by Thalassia testudinum extract bm-21 is mediated by the inhibition of acid sensing ionic channels by the phenolic compound thalassiolin b. Mol. Pain 2011, 7, 10. [Google Scholar] [CrossRef] [PubMed]
- Moran, Y.; Gordon, D.; Gurevitz, M. Sea anemone toxins affecting voltage-gated sodium channels—Molecular and evolutionary features. Toxicon 2009, 54, 1089–1101. [Google Scholar] [CrossRef] [PubMed]
- Gondran, M.; Eckeli, A.L.; Migues, P.V.; Gabilan, N.H.; Rodrigues, A.L. The crude extract from the sea anemone, Bunodosoma caissarum elicits convulsions in mice: Possible involvement of the glutamatergic system. Toxicon 2002, 40, 1667–1674. [Google Scholar] [CrossRef]
- Garateix, A.; Menéndez, R.; Díaz, M.; Martínez, J.R.; Más, R. Bunodosoma granulifera: Una especie de interés para el estudio de las neurotoxinas de anémonas. Biología 1987, 1, 15–26. [Google Scholar]
- Blanchard, M.G.; Rash, L.D.; Kellenberger, S. Inhibition of voltage-gated Na+ currents in sensory neurones by the sea anemone toxin apetx2. Br. J. Pharmacol. 2012, 165, 2167–2177. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.P.; Saez, N.J.; Cristofori-Armstrong, B.; Anangi, R.; King, G.F.; Smith, M.T.; Rash, L.D. Inhibition of acid-sensing ion channels by diminazene and apetx2 evoke partial and highly variable antihyperalgesia in a rat model of inflammatory pain. Br. J. Pharmacol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.K.; Krohn, R.I.; Hermanson, G.T.; Mallia, A.K.; Gartner, F.H.; Provenzano, M.D.; Fujimoto, E.K.; Goeke, N.M.; Olson, B.J.; Kenk, D.C. Measurement of protein using bicinchoninic acid. Anal. Biochem. 1985, 150, 76–85. [Google Scholar] [CrossRef]
- Garza, A.; Lopez-Ramirez, O.; Vega, R.; Soto, E. The aminoglycosides modulate the acid-sensing ionic channel currents in dorsal root ganglion neurons from the rat. J. Pharmacol. Exp. Ther. 2010, 332, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Salceda, E.; Lopez, O.; Zaharenko, A.J.; Garateix, A.; Soto, E. The sea anemone Bunodosoma caissarum toxin bciii modulates the sodium current kinetics of rat dorsal root ganglia neurons and is displaced in a voltage-dependent manner. Peptides 2010, 31, 412–418. [Google Scholar] [CrossRef] [PubMed]
- Liman, E.R.; Tytgat, J.; Hess, P. Subunit stoichiometry of a mammalian K+ channel determined by construction of multimeric cdnas. Neuron 1992, 9, 861–871. [Google Scholar] [CrossRef]
- Peri, S.; Steen, H.; Pandey, A. Gpmaw—A software tool for analyzing proteins and peptides. Trends Biochem. Sci. 2001, 26, 687–689. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped blast and psi-blast: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- McWilliam, H.; Li, W.; Uludag, M.; Squizzato, S.; Park, Y.M.; Buso, N.; Cowley, A.P.; Lopez, R. Analysis tool web services from the embl-ebi. Nucleic Acids Res. 2013, 41, W597–W600. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. Mafft multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.; Clamp, M.; Barton, G.J. Jalview version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 1999, 292, 195–502. [Google Scholar] [CrossRef] [PubMed]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The swiss-model workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Kelley, L.A.; Sternberg, M.J. Protein structure prediction on the web: A case study using the phyre server. Nat. Protoc. 2009, 4, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. I-tasser server for protein 3d structure prediction. BMC Bioinform. 2008, 9, 40. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zhang, Y. Lomets: A local meta-threading-server for protein structure prediction. Nucleic Acids Res. 2007, 35, 3375–3382. [Google Scholar] [CrossRef] [PubMed]
- Kallberg, M.; Wang, H.; Wang, S.; Peng, J.; Wang, Z.; Lu, H.; Xu, J. Template-based protein structure modeling using the raptorx web server. Nat. Protoc. 2012, 7, 1511–1522. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Rullmannn, J.A.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. Aqua and procheck-nmr: Programs for checking the quality of protein structures solved by nmr. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Hooft, R.W.; Vriend, G.; Sander, C.; Abola, E.E. Errors in protein structures. Nature 1996, 381, 272. [Google Scholar] [CrossRef] [PubMed]
- Bowie, J.U.; Luthy, R.; Eisenberg, D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 1991, 253, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Guex, N.; Peitsch, M.C. Swiss-model and the swiss-pdbviewer: An environment for comparative protein modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef] [PubMed]
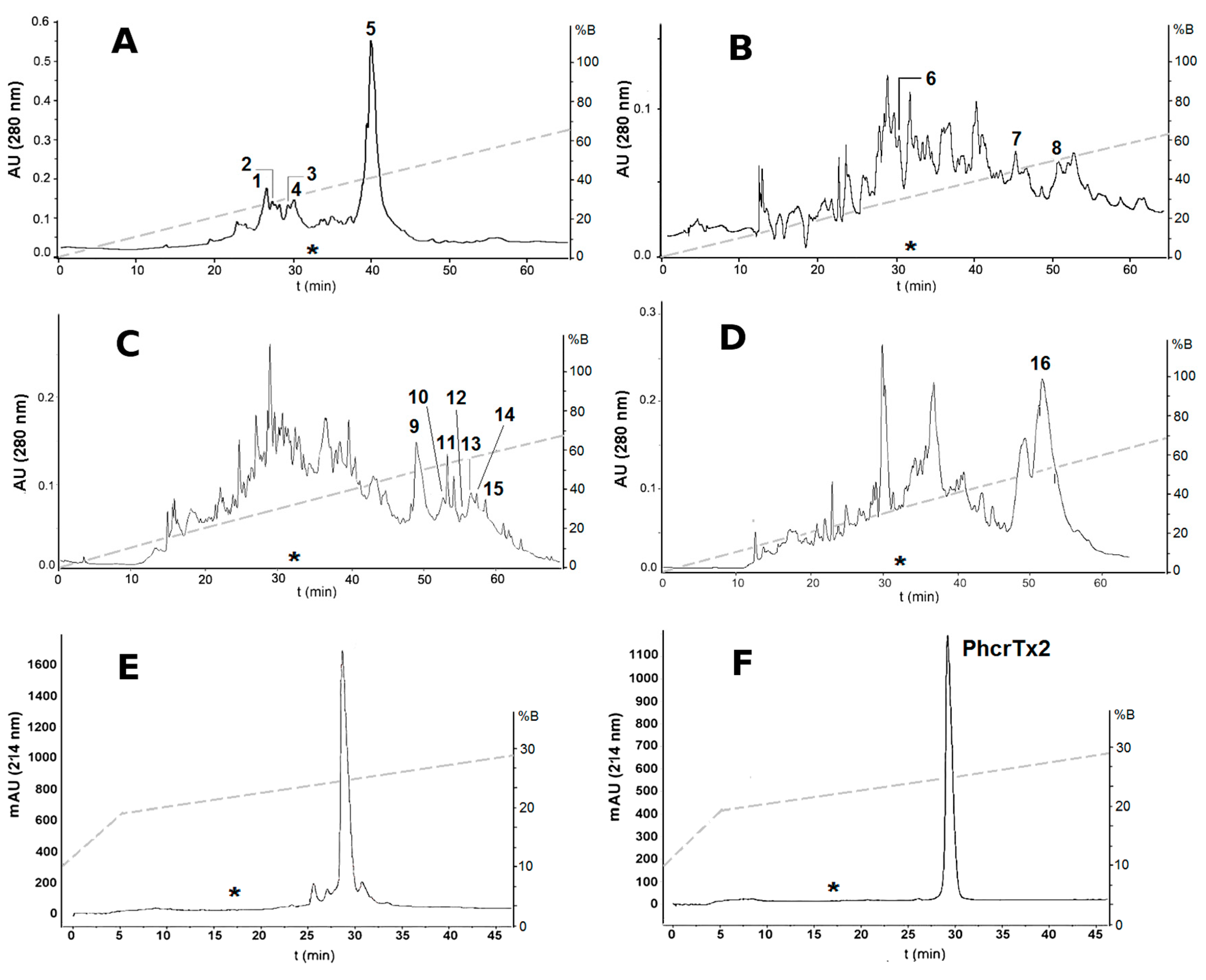
| RP-HPLC Fraction Number | Features | Paralyzing Effects on Crab Uca thayeri |
|---|---|---|
| 1–4 and 6 | tR = 25–30 min * 1986–5671 Da ** basic peptides *** | Progressive slowing down of legs movements until remaining motionless approximately after 30 min. Lethal. |
| 5 and 7 | tR = 40–45 min 5294–5296 Da basic peptides | Moderate tetanic paralysis starting after 30 min from toxin administration. Partial recovery from paralysis followed by uncoordinated leg movements. Not lethal. |
| 8 | tR = 51 min 3082 Da basic peptide | Severe tetanic paralysis after a few seconds, resembling the effects provoked by Nav toxins [27]. Lethal. |
| 9–15 | tR = 49–59 min 4573–9847 Da acidic peptides | Severe tetanic paralysis after a few seconds, resembling the effects provoked by Nav toxins [27]. Lethal. |
| 16 | tR = 50–55 min 19,758 Da acidic protein | Moderate tetanic paralysis 5 min after toxin administration. Not lethal. |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodríguez, A.A.; Garateix, A.; Salceda, E.; Peigneur, S.; Zaharenko, A.J.; Pons, T.; Santos, Y.; Arreguín, R.; Ständker, L.; Forssmann, W.-G.; et al. PhcrTx2, a New Crab-Paralyzing Peptide Toxin from the Sea Anemone Phymanthus crucifer. Toxins 2018, 10, 72. https://doi.org/10.3390/toxins10020072
Rodríguez AA, Garateix A, Salceda E, Peigneur S, Zaharenko AJ, Pons T, Santos Y, Arreguín R, Ständker L, Forssmann W-G, et al. PhcrTx2, a New Crab-Paralyzing Peptide Toxin from the Sea Anemone Phymanthus crucifer. Toxins. 2018; 10(2):72. https://doi.org/10.3390/toxins10020072
Chicago/Turabian StyleRodríguez, Armando Alexei, Anoland Garateix, Emilio Salceda, Steve Peigneur, André Junqueira Zaharenko, Tirso Pons, Yúlica Santos, Roberto Arreguín, Ludger Ständker, Wolf-Georg Forssmann, and et al. 2018. "PhcrTx2, a New Crab-Paralyzing Peptide Toxin from the Sea Anemone Phymanthus crucifer" Toxins 10, no. 2: 72. https://doi.org/10.3390/toxins10020072
APA StyleRodríguez, A. A., Garateix, A., Salceda, E., Peigneur, S., Zaharenko, A. J., Pons, T., Santos, Y., Arreguín, R., Ständker, L., Forssmann, W.-G., Tytgat, J., Vega, R., & Soto, E. (2018). PhcrTx2, a New Crab-Paralyzing Peptide Toxin from the Sea Anemone Phymanthus crucifer. Toxins, 10(2), 72. https://doi.org/10.3390/toxins10020072







