Loxosceles gaucho Spider Venom: An Untapped Source of Antimicrobial Agents
Abstract
1. Introduction
2. Results and Discussion
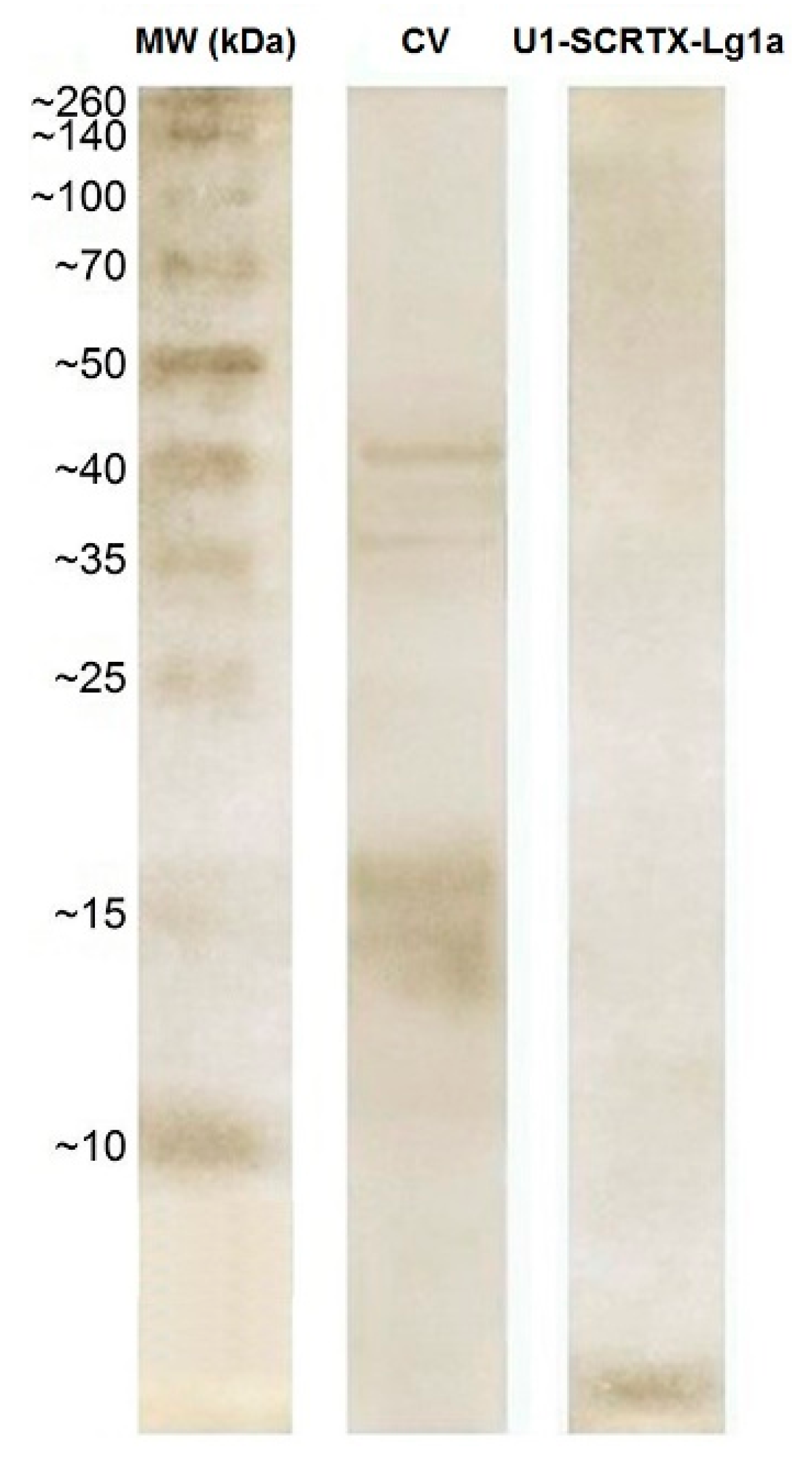
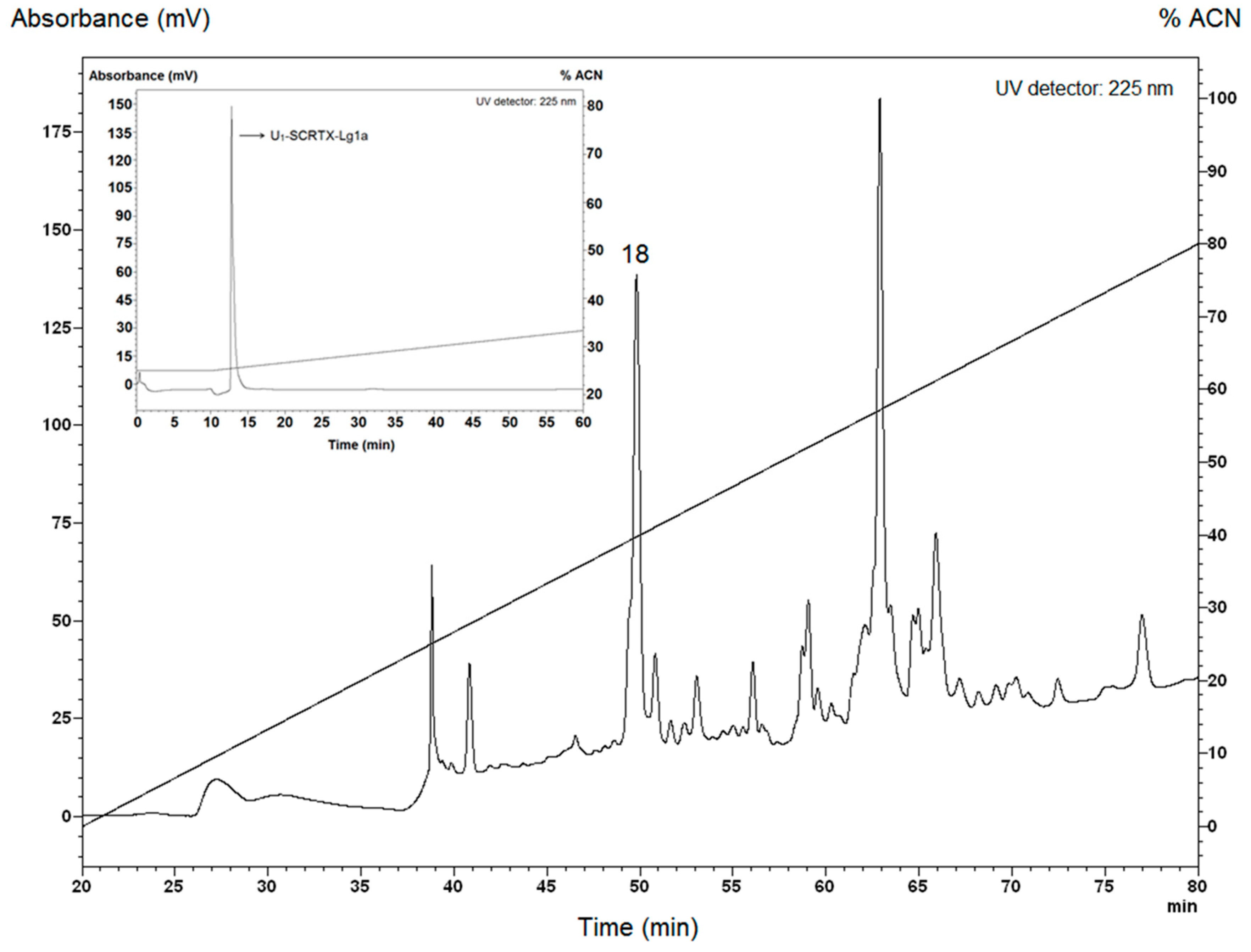
2.1. Purification of U1-SCRTX-Lg1a from the Venom of L. gaucho
2.2. Antimicrobial Activity and Minimum Inhibitory Concentrations (MICs)
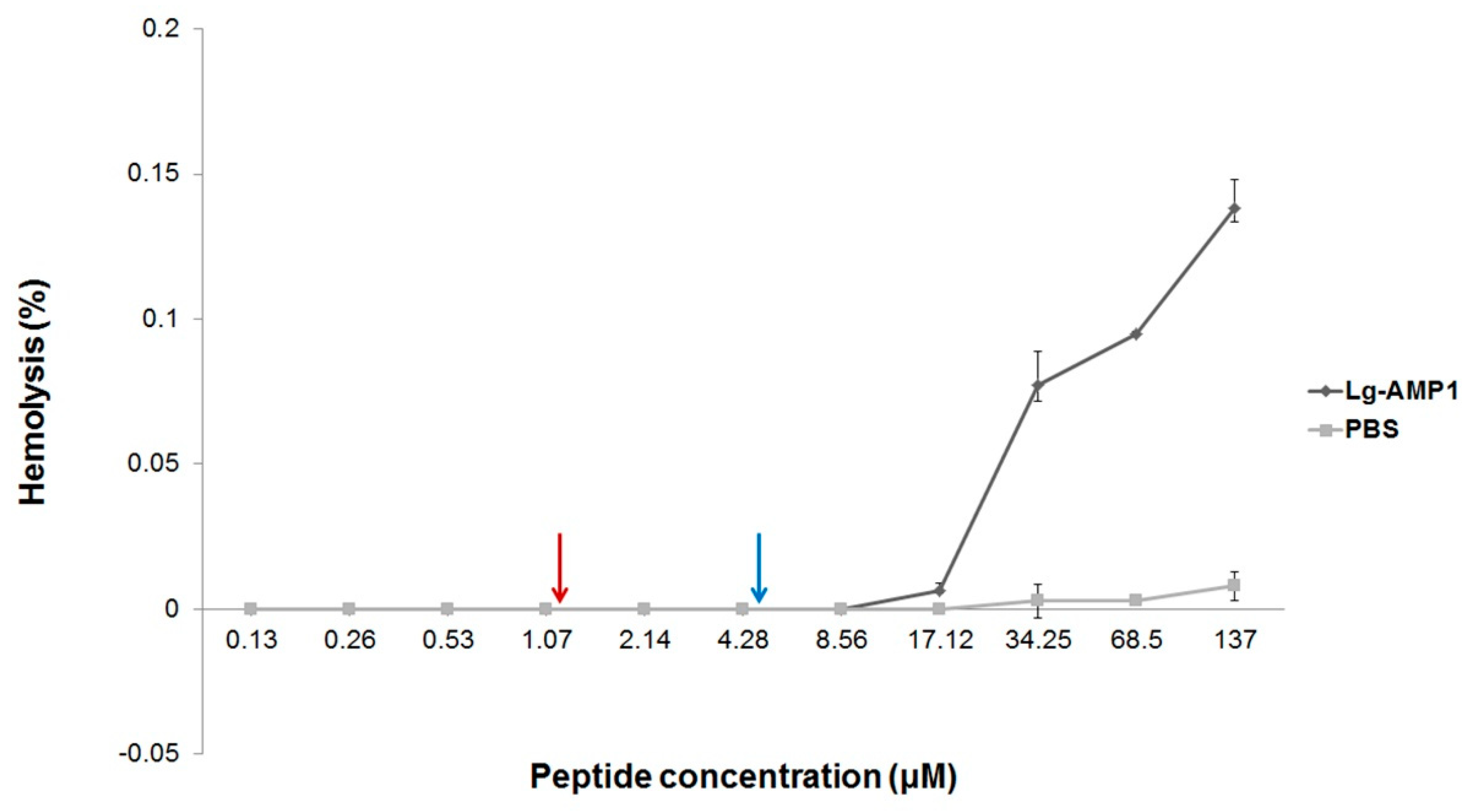
2.3. Hemolytic Activity
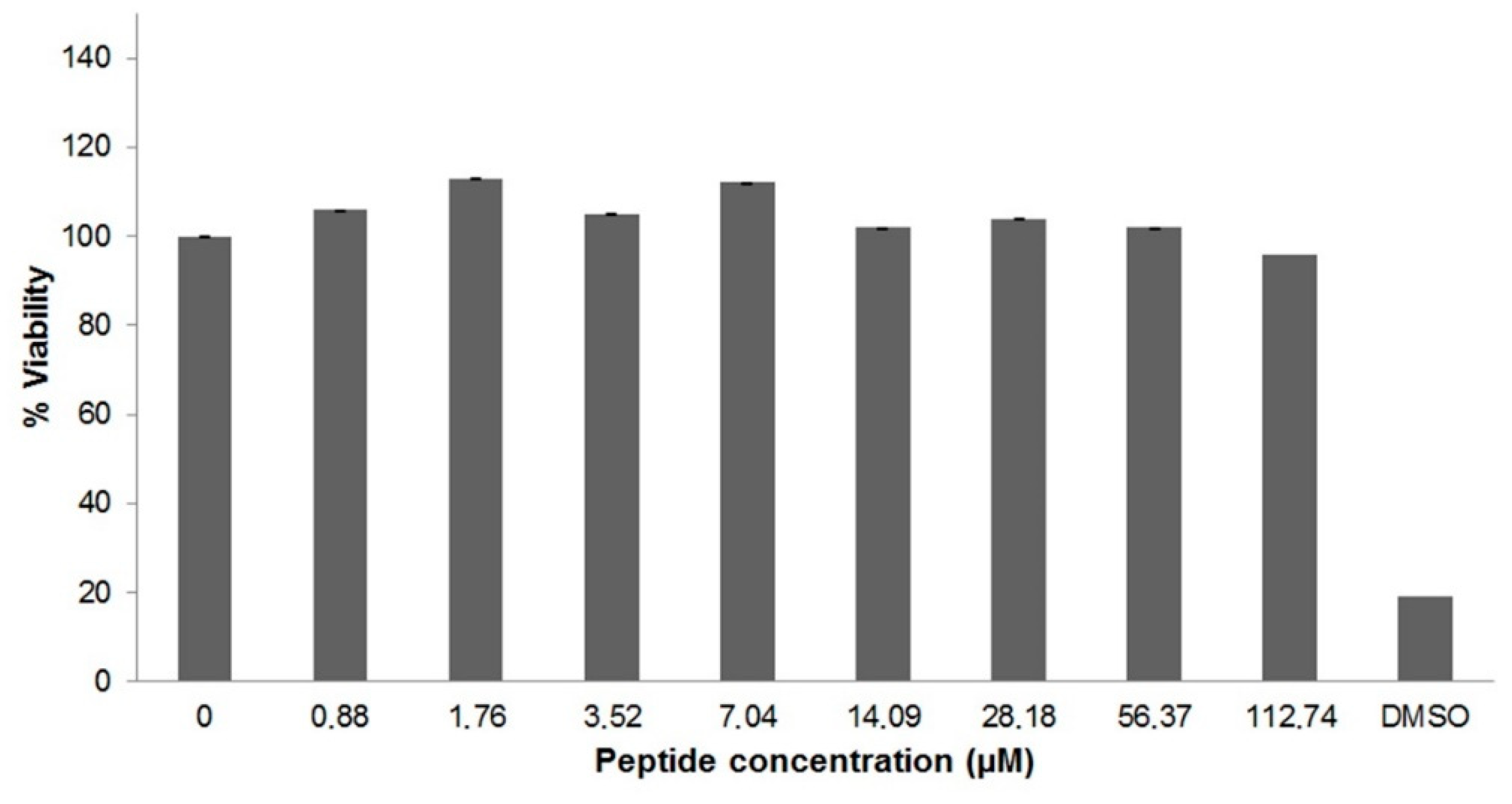
2.4. Cytotoxicity of U1-SCRTX-Lg1a
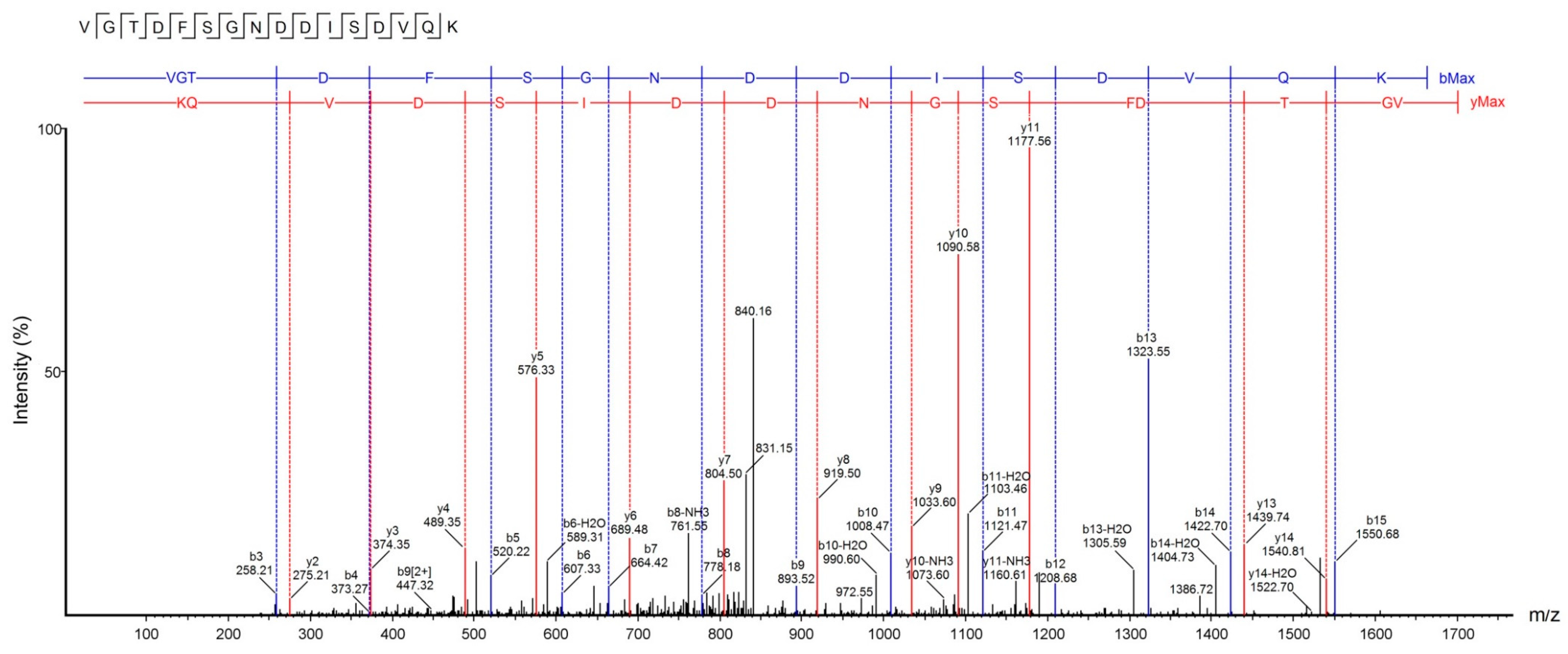
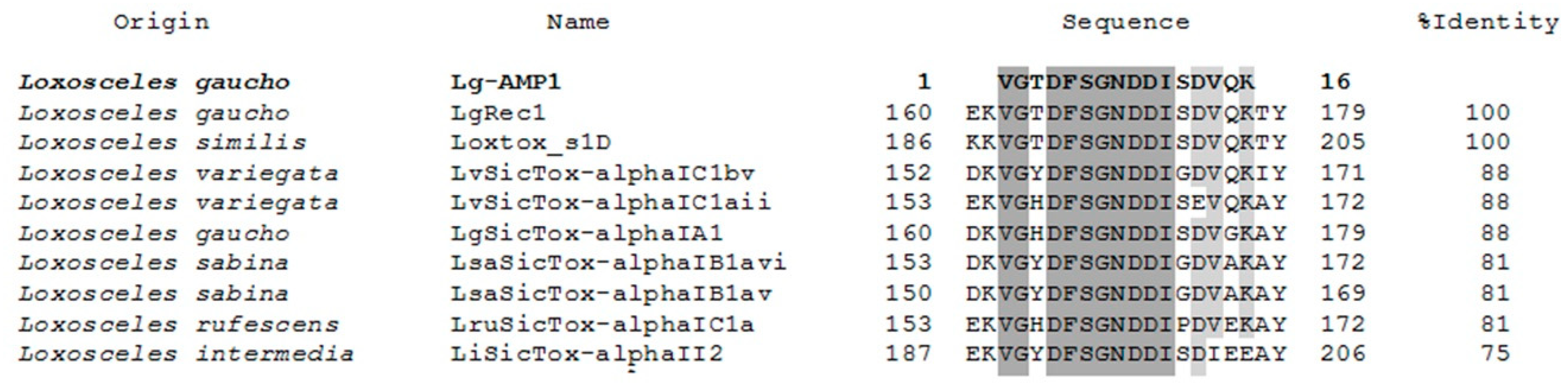
2.5. U1-SCRTX-Lg1a Identification
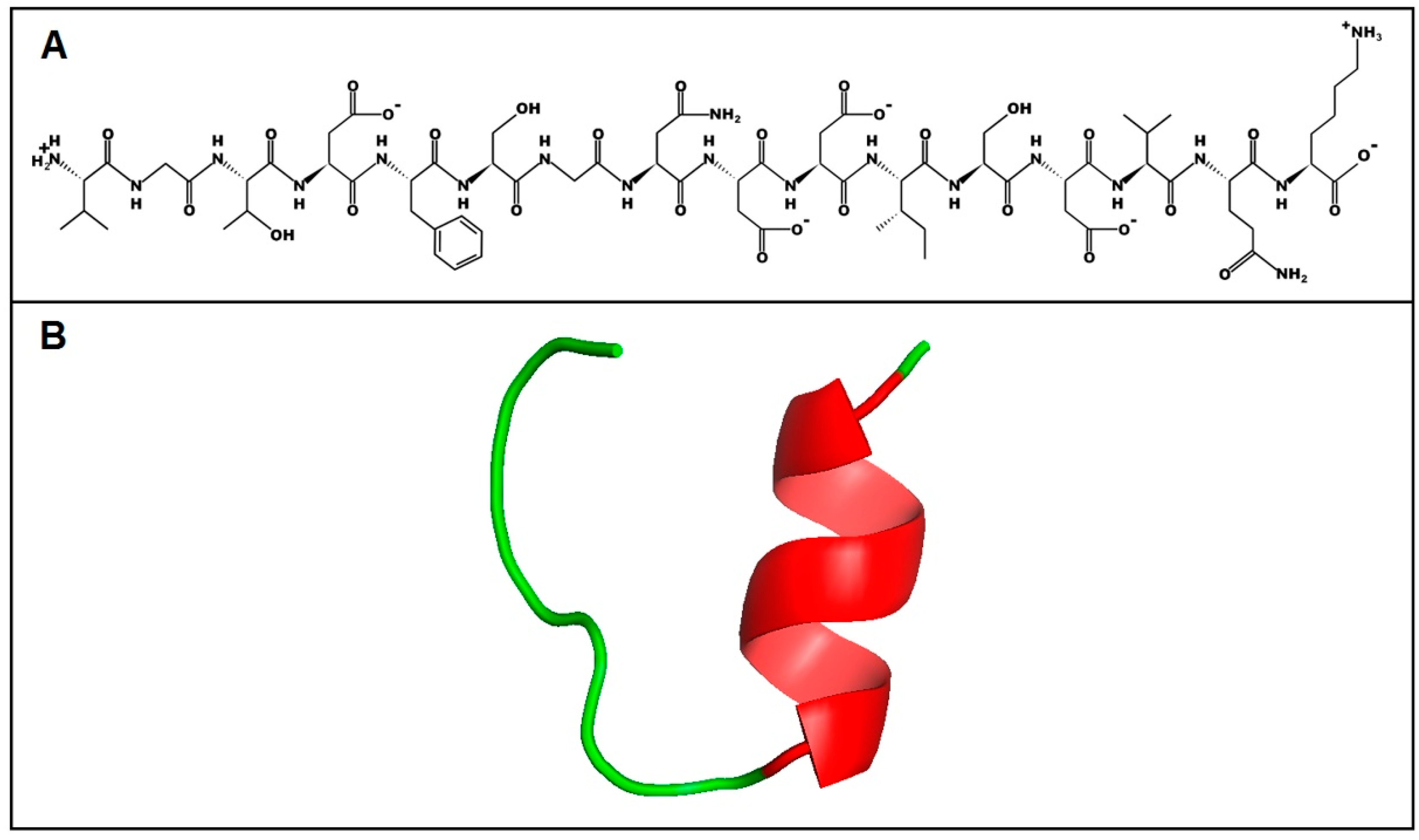
2.6. Structure and Physicochemical Characteristics of U1-SCRTX-Lg1a
3. Conclusions
4. Materials and Methods
4.1. Microbial Strains
4.2. Animals
4.3. Venom Fractionation and U1-SCRTX-Lg1a Purification
4.4. Antimicrobial Assays
4.5. Concentration of U1-SCRTX-Lg1a
4.6. MICs of U1-SCRTX-Lg1a
4.7. Determination of Hemolytic Activity
4.8. Cytotoxicity Assay against HeLa Cells
4.9. SDS-PAGE Analysis
4.10. Mass Spectrometry Analysis and U1-SCRTX-Lg1a Identification
4.11. U1-SCRTX-Lg1a Analysis with Bioinformatics Tools
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Garcia, F.; Villegas, E.; Espino-Solis, G.P.; Rodriguez, A.; Paniagua-Solis, J.F.; Sandoval-Lopez, G.; Possani, L.D.; Corzo, G. Antimicrobial peptides from arachnid venoms and their microbicidal activity in the presence of commercial antibiotics. J. Antibiot. 2013, 66, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Harrison, P.L.; Abdel-Rahman, M.A.; Miller, K.; Strong, P.N. Antimicrobial peptides from scorpion venoms. Toxicon 2014, 88, 115–137. [Google Scholar] [CrossRef]
- Peters, B.M.; Shirtliff, M.E.; Jabra-Rizk, M.A. Antimicrobial peptides: Primeval molecules or future drugs? PLoS Pathog. 2010, 6, e1001067. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, V.; Feio, M.J.; Bastos, M. Role of lipids in the interaction of antimicrobial peptides with membranes. Prog. Lipid Res. 2012, 51, 149–177. [Google Scholar] [CrossRef]
- Mechkarska, M.; Attoub, S.; Sulaiman, S.; Pantic, J.; Lukic, M.L.; Conlon, J.M. Anti-cancer, immunoregulatory, and antimicrobial activities of the frog skin host-defense peptides pseudhymenochirin-1Pb and pseudhymenochirin-2Pa. Regul. Pept. 2014, 194–195, 69–76. [Google Scholar] [CrossRef]
- Narayana, J.L.; Chen, J.Y. Antimicrobial peptides: Possible anti-infective agents. Peptides 2015, 72, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Steinstraesser, L.; Kraneburg, U.; Jacobsen, F.; Al-Benna, S. Host defense peptides and their antimicrobial-immunomodulatory duality. Immunobiology 2011, 216, 322–333. [Google Scholar] [CrossRef]
- Corzo, G.; Villegas, E.; Gómez-Lagunas, F.; Possani, L.D.; Belokoneva, O.S.; Nakajima, T. Oxyopinins, large amphipathic peptides isolated from the venom of the wolf spider Oxyopes kitabensis with cytolytic properties and positive insecticidal cooperativity with spider neurotoxins. J. Biol. Chem. 2002, 277, 23627–23637. [Google Scholar] [CrossRef]
- Kozlov, S.A.; Vassilevski, A.A.; Feofanov, A.V.; Surovoy, A.Y.; Karpunin, D.V.; Grishin, E.V. Latarcins, antimicrobial and cytolytic peptides from the venom of the spider Lachesana tarabaevi (Zodariidae) that exemplify biomolecular diversity. J. Biol. Chem. 2006, 281, 20983–20992. [Google Scholar] [CrossRef]
- Yan, L.; Adams, M.E. Lycotoxins, antimicrobial peptides from venom of the wolf spider Lycosa carolinensis. J. Biol. Chem. 1998, 273, 2059–2066. [Google Scholar] [CrossRef]
- Nicholson, G.M. Spider peptides. In Handbook of Biologically Active Peptides, 2nd ed.; Kastin, A.J., Ed.; Academic Press: Los Angeles, CA, USA, 2013; pp. 461–472. ISBN 978-0-12-385095-9. [Google Scholar]
- Abreu, T.F.; Sumitomo, B.N.; Nishiyama, M.Y., Jr.; Oliveira, U.C.; Souza, G.H.; Kitano, E.S.; Zelanis, A.; Serrano, S.M.; Junqueira-de-Azevedo, I.; Silva, P.I., Jr.; et al. Peptidomics of Acanthoscurria gomesiana spider venom reveals new toxins with potential antimicrobial activity. J. Proteomics 2017, 151, 232–242. [Google Scholar] [CrossRef] [PubMed]
- Ayroza, G.; Ferreira, I.L.; Sayegh, R.S.; Tashima, A.K.; Silva, P.I., Jr. Juruin: An antifungal peptide from the venom of the Amazonian Pink Toe spider, Avicularia juruensis, which contains the inhibitory cystine knot motif. Front Microbiol. 2012, 3, 324. [Google Scholar] [CrossRef] [PubMed]
- Lazarev, V.N.; Shkarupeta, M.M.; Polina, N.F.; Kostrjukova, E.S.; Vassilevski, A.A.; Kozlov, S.A.; Grishin, E.V.; Govorun, V.M. Antimicrobial peptide from spider venom inhibits Chlamydia trachomatis infection at an early stage. Arch. Microbiol. 2013, 195, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Lei, Q.; Yu, H.; Peng, X.; Yan, S.; Wang, J.; Yan, Y.; Wang, X. Isolation and preliminary characterization of proteinaceous toxins with insecticidal and antibacterial activities from black widow spider (L. tredecimguttatus) eggs. Toxins 2015, 7, 886–889. [Google Scholar] [CrossRef] [PubMed]
- Santos, D.M.; Verly, R.M.; Piló-Veloso, D.; de Maria, M.; de Carvalho, M.A.; Cisalpino, P.S.; Soares, B.M.; Diniz, C.G.; Farias, L.M.; Moreira, D.F.; et al. LyeTx I, a potent antimicrobial peptide from the venom of the spider Lycosa erythrognatha. Amino Acids 2010, 39, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.; Ding, X.; Meng, S.; Liu, C.; Wang, H.; Xia, L.; Liu, Z.; Liang, S. Antimicrobial potential of lycosin-I, a cationic and amphiphilic peptide from the venom of the spider Lycosa singorensis. Curr. Mol. Med. 2013, 13, 900–910. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, L.; Yang, H.; Xiao, H.; Farooq, A.; Liu, Z.; Hu, M.; Shi, X. The spider venom peptide Lycosin-II has potent antimicrobial activity against clinically isolated bacteria. Toxins 2016, 8, 119. [Google Scholar] [CrossRef]
- De Oliveira, K.C.; Gonçalves de Andrade, R.M.; Piazza, R.M.; Ferreira, J.M., Jr.; van den Berg, C.W.; Tambourgi, D.V. Variations in Loxosceles spider venom composition and toxicity contribute to the severity of envenomation. Toxicon 2005, 45, 421–429. [Google Scholar] [CrossRef]
- Chaim, O.M.; Trevisan-Silva, D.; Chaves-Moreira, D.; Wille, A.C.; Ferrer, V.P.; Matsubara, F.H.; Mangili, O.C.; da Silveira, R.B.; Gremski, L.H.; Gremski, W.; et al. Brown spider (Loxosceles genus) venom toxins: Tools for biological purposes. Toxins 2011, 3, 309–344. [Google Scholar] [CrossRef]
- Gremski, L.H.; Trevisan-Silva, D.; Ferrer, V.P.; Matsubara, F.H.; Meissner, G.O.; Wille, A.C.; Vuitika, L.; Dias-Lopes, C.; Ullah, A.; de Moraes, F.R.; et al. Recent advances in the understanding of brown spider venoms: From the biology of spiders to the molecular mechanisms of toxins. Toxicon 2014, 83, 91–120. [Google Scholar] [CrossRef]
- Manríquez, J.J.; Silva, S. Cutaneous and visceral loxoscelism: A systematic review. Rev. Chilena Infectol. 2009, 26, 420–432. [Google Scholar] [CrossRef] [PubMed]
- Buch, D.R.; Souza, F.N.; Meissner, G.O.; Morgon, A.M.; Gremski, L.H.; Ferrer, V.P.; Trevisan-Silva, D.; Matsubara, F.H.; Bóia-Ferreira, M.; Sade, Y.B.; et al. Brown spider (Loxosceles genus) venom toxins: Evaluation of biological conservation by immune cross-reactivity. Toxicon 2015, 108, 154–166. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, F.H.; Gremski, L.H.; Meissner, G.O.; Soares, E.C.; Gremski, W.; Senff-Ribeiro, A.; Chaim, O.M.; Veiga, S.S. A novel ICK peptide from the Loxosceles intermedia (brown spider) venom gland: Cloning, heterologous expression and immunological cross-reactivity approaches. Toxicon 2013, 71, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Senff-Ribeiro, A.; da Silva, P.H.; Chaim, O.M.; Gremski, L.H.; Paludo, K.S.; da Silveira, R.B.; Gremski, W.; Mangili, O.C.; Veiga, S.S. Biotechnological applications of brown spider (Loxosceles genus) venom toxins. Biotechnol. Adv. 2008, 26, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Appel, M.H.; da Silveira, R.B.; Chaim, O.M.; Paludo, K.S.; Trevisan-Silva, D.; Chaves-Moreira, D.; da Silva, P.H.; Mangili, O.C.; Senff-Ribeiro, A.; Gremski, W.; et al. Identification, cloning and functional characterization of a novel dermonecrotic toxin (phospholipase D) from brown spider (Loxosceles intermedia) venom. Biochim. Biophys. Acta 2008, 1780, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Binford, G.J.; Cordes, M.H.; Wells, M.A. Sphingomyelinase D from venoms of Loxosceles spiders: Evolutionary insights from cDNA sequences and gene structure. Toxicon 2005, 45, 547–560. [Google Scholar] [CrossRef] [PubMed]
- Da Silveira, R.B.; Pigozzo, R.B.; Chaim, O.M.; Appel, M.H.; Dreyfuss, J.L.; Toma, L.; Mangili, O.C.; Gremski, W.; Dietrich, C.P.; Nader, H.B.; et al. Molecular cloning and functional characterization of two isoforms of dermonecrotic toxin from Loxosceles intermedia (Brown spider) venom gland. Biochimie 2006, 88, 1241–1253. [Google Scholar] [CrossRef]
- Da Silveira, R.B.; Pigozzo, R.B.; Chaim, O.M.; Appel, M.H.; Trevisan-Silva, D.; Dreyfuss, J.L.; Toma, L.; Dietrich, C.P.; Nader, H.B.; Veiga, S.S.; et al. Two novel dermonecrotic toxins LiRecDT4 and LiRecDT5 from Brown spider (Loxosceles intermedia) venom: From cloning to functional characterization. Biochimie 2007, 89, 289–300. [Google Scholar] [CrossRef]
- Dantas, A.E.; Carmo, A.O.; Horta, C.C.; Leal, H.G.; Oliveira-Mendes, B.B.; Martins, A.P.; Chávez-Olórtegui, C.; Kalapothakis, E. Description of Loxtox protein family and identification of a new group of Phospholipases D from Loxosceles similis venom gland. Toxicon 2016, 120, 97–106. [Google Scholar] [CrossRef]
- De Giuseppe, P.O.; Ullah, A.; Trevisan-Silva, D.; Gremski, L.H.; Wille, A.C.; Chaves-Moreira, D.; Senff-Ribeiro, A.; Chaim, O.M.; Murakami, M.T.; Veiga, S.S.; et al. Structure of a novel class II phospholipase D: Catalytic cleft is modified by a disulphide bridge. Biochem. Biophys. Res. Commun. 2011, 409, 622–627. [Google Scholar] [CrossRef]
- Kalapothakis, E.; Chatzaki, M.; Gonçalves-Dornelas, H.; de Castro, C.S.; Silvestre, F.G.; Laborne, F.V.; de Moura, J.F.; Veiga, S.S.; Chávez-Olórtegui, C.; Granier, C.; et al. The loxtox protein family in Loxosceles intermedia (Mello-Leitão) venom. Toxicon 2007, 50, 938–946. [Google Scholar] [CrossRef] [PubMed]
- Magalhães, G.S.; Caporrino, M.C.; Della-Casa, M.S.; Kimura, L.F.; Prezotto-Neto, J.P.; Fukuda, D.A.; Portes-Junior, J.A.; Neves-Ferreira, A.G.; Santoro, M.L.; Barbaro, K.C. Cloning, expression and characterization of a phospholipase D from Loxosceles gaucho venom gland. Biochimie 2013, 95, 1773–1783. [Google Scholar] [CrossRef] [PubMed]
- Wille, A.C.; Chaves-Moreira, D.; Trevisan-Silva, D.; Magnoni, M.G.; Boia-Ferreira, M.; Gremski, L.H.; Gremski, W.; Chaim, O.M.; Senff-Ribeiro, A.; Veiga, S.S. Modulation of membrane phospholipids, the cytosolic calcium influx and cell proliferation following treatment of B16-F10 cells with recombinant phospholipase-D from Loxosceles intermedia (brown spider) venom. Toxicon 2013, 67, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Zobel-Thropp, P.A.; Kerins, A.E.; Binford, G.J. Sphingomyelinase D in sicariid spider venom is a potent insecticidal toxin. Toxicon 2012, 60, 265–271. [Google Scholar] [CrossRef] [PubMed]
- Da Silveira, R.B.; dos Santos Filho, J.F.; Mangili, O.C.; Veiga, S.S.; Gremski, W.; Nader, H.B.; von Dietrich, C.P. Identification of proteases in the extract of venom glands from brown spiders. Toxicon 2002, 40, 815–822. [Google Scholar] [CrossRef]
- Da Silveira, R.B.; Wille, A.C.; Chaim, O.M.; Appel, M.H.; Silva, D.T.; Franco, C.R.; Toma, L.; Mangili, O.C.; Gremski, W.; Dietrich, C.P.; et al. Identification, cloning, expression and functional characterization of an astacin-like metalloprotease toxin from Loxosceles intermedia (brown spider) venom. Biochem. J. 2007, 406, 355–363. [Google Scholar] [CrossRef] [PubMed]
- Feitosa, L.; Gremski, W.; Veiga, S.S.; Elias, M.C.; Graner, E.; Mangili, O.C.; Brentani, R.R. Detection and characterization of metalloproteinases with gelatinolytic, fibronectinolytic and fibrinogenolytic activities in brown spider (Loxosceles intermedia) venom. Toxicon 1998, 36, 1039–1051. [Google Scholar] [CrossRef]
- Morgon, A.M.; Belisario-Ferrari, M.R.; Trevisan-Silva, D.; Meissner, G.O.; Vuitika, L.; Marin, B.; Tashima, A.K.; Gremski, L.H.; Gremski, W.; Senff-Ribeiro, A.; et al. Expression and immunological cross-reactivity of LALP3, a novel astacin-like metalloprotease from brown spider (Loxosceles intermedia) venom. Biochimie 2016, 128–129, 8–19. [Google Scholar] [CrossRef]
- Trevisan-Silva, D.; Gremski, L.H.; Chaim, O.M.; da Silveira, R.B.; Meissner, G.O.; Mangili, O.C.; Barbaro, K.C.; Gremski, W.; Veiga, S.S.; Senff-Ribeiro, A. Astacin-like metalloproteases are a gene family of toxins present in the venom of different species of the brown spider (genus Loxosceles). Biochimie 2010, 92, 21–32. [Google Scholar] [CrossRef]
- Trevisan-Silva, D.; Bednaski, A.V.; Gremski, L.H.; Chaim, O.M.; Veiga, S.S.; Senff-Ribeiro, A. Differential metalloprotease content and activity of three Loxosceles spider venoms revealed using two-dimensional electrophoresis approaches. Toxicon 2013, 76, 11–22. [Google Scholar] [CrossRef]
- Barbaro, K.C.; Knysak, I.; Martins, R.; Hogan, C.; Winkel, K. Enzymatic characterization, antigenic cross-reactivity and neutralization of dermonecrotic activity of five Loxosceles spider venoms of medical importance in the Americas. Toxicon 2005, 45, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Da Silveira, R.B.; Chaim, O.M.; Mangili, O.C.; Gremski, W.; Dietrich, C.P.; Nader, H.B.; Veiga, S.S. Hyaluronidases in Loxosceles intermedia (Brown spider) venom are endo-beta-N-acetyl-d-hexosaminidases hydrolases. Toxicon 2007, 49, 758–768. [Google Scholar] [CrossRef]
- Dos Santos, L.D.; Dias, N.B.; Roberto, J.; Pinto, A.S.; Palma, M.S. Brown recluse spider venom: Proteomic analysis and proposal of a putative mechanism of action. Protein Pept. Lett. 2009, 16, 933–943. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, V.P.; de Mari, T.L.; Gremski, L.H.; Trevisan-Silva, D.; da Silveira, R.B.; Gremski, W.; Chaim, O.M.; Senff-Ribeiro, A.; Nader, H.B.; Veiga, S.S. A novel hyaluronidase from brown spider (Loxosceles intermedia) venom (Dietrich’s Hyaluronidase): From cloning to functional characterization. PLoS Negl. Trop. Dis. 2013, 7, e2206. [Google Scholar] [CrossRef] [PubMed]
- Young, A.R.; Pincus, S.J. Comparison of enzymatic activity from three species of necrotising arachnids in Australia: Loxosceles rufescens, Badumna insignis and Lampona cylindrata. Toxicon 2001, 39, 391–400. [Google Scholar] [CrossRef]
- Fernandes-Pedrosa, M.F.; Junqueira-de-Azevedo, I.L.; Gonçalves-de-Andrade, R.M.; Kobashi, L.S.; Almeida, D.D.; Ho, P.L.; Tambourgi, D.V. Transcriptome analysis of Loxosceles laeta (Araneae, Sicariidae) spider venomous gland using expressed sequence tags. BMC Genomics 2008, 9, 279. [Google Scholar] [CrossRef] [PubMed]
- Gremski, L.H.; da Silveira, R.B.; Chaim, O.M.; Probst, C.M.; Ferrer, V.P.; Nowatzki, J.; Weinschutz, H.C.; Madeira, H.M.; Gremski, W.; Nader, H.B.; et al. A novel expression profile of the Loxosceles intermedia spider venomous gland revealed by transcriptome analysis. Mol. Biosyst. 2010, 6, 2403–2416. [Google Scholar] [CrossRef] [PubMed]
- Machado, L.F.; Laugesen, S.; Botelho, E.D.; Ricart, C.A.; Fontes, W.; Barbaro, K.C.; Roepstorff, P.; Sousa, M.V. Proteome analysis of brown spider venom: Identification of loxnecrogin isoforms in Loxosceles gaucho venom. Proteomics 2005, 5, 2167–2176. [Google Scholar] [CrossRef]
- Veiga, S.S.; da Silveira, R.B.; Dreyfus, J.L.; Haoach, J.; Pereira, A.M.; Mangili, O.C.; Gremski, W. Identification of high molecular weight serine-proteases in Loxosceles intermedia (brown spider) venom. Toxicon 2000, 38, 825–839. [Google Scholar] [CrossRef]
- Sade, Y.B.; Bóia-Ferreira, M.; Gremski, L.H.; da Silveira, R.B.; Gremski, W.; Senff-Ribeiro, A.; Chaim, O.M.; Veiga, S.S. Molecular cloning, heterologous expression and functional characterization of a novel translationally-controlled tumor protein (TCTP) family member from Loxosceles intermedia (brown spider) venom. Int. J. Biochem. Cell Biol. 2012, 44, 170–177. [Google Scholar] [CrossRef]
- De Castro, C.S.; Silvestre, F.G.; Araújo, S.C.; Yazbeck, G.M.; Mangili, O.C.; Cruz, I.; Chávez-Olórtegui, C.; Kalapothakis, E. Identification and molecular cloning of insecticidal toxins from the venom of the brown spider Loxosceles intermedia. Toxicon 2004, 44, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Meissner, G.O.; de Resende Lara, P.T.; Scott, L.P.; Braz, A.S.; Chaves-Moreira, D.; Matsubara, F.H.; Soares, E.M.; Trevisan-Silva, D.; Gremski, L.H.; Veiga, S.S.; et al. Molecular cloning and in silico characterization of knottin peptide, U2-SCRTX-Lit2, from brown spider (Loxosceles intermedia) venom glands. J. Mol. Model. 2016, 22, 196. [Google Scholar] [CrossRef] [PubMed]
- King, G.F.; Gentz, M.C.; Escoubas, P.; Nicholson, G.M. A rational nomenclature for naming peptide toxins from spiders and other venomous animals. Toxicon 2008, 52, 264–276. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira Domingos, M.; Neves, I.V.; Vigerelli, H.; Pimenta, D.C.; de Carvalho Lins Fernandes Távora, B.; Lemos, T.J.; Franzolin, M.R.; Marques, V.D.; Barbaro, K.C. The potential of Loxosceles gaucho spider venom to regulate Pseudomonas aeruginosa mechanisms of virulence. Toxicon 2018, 152, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Chaparro, E.; da Silva, P.I., Jr. Lacrain: The first antimicrobial peptide from the body extract of the Brazilian centipede Scolopendra viridicornis. Int. J. Antimicrob. Agents 2016, 48, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.I., Jr.; Daffre, S.; Bulet, P. Isolation and characterization of Gomesin, an 18-residue cysteine-rich defense peptide from the spider Acanthoscurria gomesiana hemocytes with sequence similarities to horseshoe crab antimicrobial peptides of the tachyplesin family. J. Biol. Chem. 2000, 275, 33464–33470. [Google Scholar] [CrossRef] [PubMed]
- Gruenheid, S.; Le Moual, H. Resistance to antimicrobial peptides in Gram-negative bacteria. FEMS Microbiol. Lett. 2012, 330, 81–89. [Google Scholar] [CrossRef]
- Sardar, A.H.; Das, S.; Agnihorti, S.; Kumar, M.; Ghosh, A.K.; Abhishek, K.; Kumar, A.; Purkait, B.; Ansari, M.Y.; Das, P. Spinigerin induces apoptotic like cell death in a caspase independent manner in Leishmania donovani. Exp. Parasitol. 2013, 135, 715–725. [Google Scholar] [CrossRef]
- Oddo, A.; Hansen, P.R. Hemolytic activity of antimicrobial peptides. Methods Mol. Biol. 2017, 1548, 427–435. [Google Scholar] [CrossRef]
- Betts, M.J.; Russell, R.B. Amino acid properties and consequences of substitutions. In Bioinformatics for Geneticists; Barnes, M.R., Gray, I.C., Eds.; John Wiley & Sons: Hoboken, NJ, USA, 2003; pp. 289–316. ISBN 9780470867303. [Google Scholar]
- Santamaría, C.; Larios, S.; Quirós, S.; Pizarro-Cerda, J.; Gorvel, J.P.; Lomonte, B.; Moreno, E. Bactericidal and antiendotoxic properties of short cationic peptides derived from a snake venom Lys49 phospholipase A2. Antimicrob. Agents Chemother. 2005, 49, 1340–1345. [Google Scholar] [CrossRef]
- Harris, F.; Dennison, S.R.; Phoenix, D.A. Anionic antimicrobial peptides from eukaryotic organisms. Curr. Protein Pept. Sci. 2009, 10, 585–606. [Google Scholar] [CrossRef] [PubMed]
- Torrent, M.; Andreu, D.; Nogués, V.M.; Boix, E. Connecting peptide physicochemical and antimicrobial properties by a rational prediction model. PLoS ONE 2011, 6, e16968. [Google Scholar] [CrossRef] [PubMed]
- Dennison, S.R.; Howe, J.; Morton, L.H.; Brandenburg, K.; Harris, F.; Phoenix, D.A. Interactions of an anionic antimicrobial peptide with Staphylococcus aureus membranes. Biochem. Biophys. Res. Commun. 2006, 347, 1006–1010. [Google Scholar] [CrossRef] [PubMed]
- Dennison, S.R.; Harris, F.; Mura, M.; Phoenix, D.A. An atlas of anionic antimicrobial peptides from amphibians. Curr. Protein Pept. Sci. 2018, 19, 823–838. [Google Scholar] [CrossRef] [PubMed]
- Bulet, P. Strategies for the discovery, isolation, and characterization of natural bioactive peptides from the immune system of invertebrates. Methods Mol. Biol. 2008, 494, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Hetru, C.; Bulet, P. Strategies for the isolation and characterization of antimicrobial peptides of invertebrates. Methods Mol. Biol. 1997, 78, 35–49. [Google Scholar] [CrossRef]
- Anthis, N.J.; Clore, G.M. Sequence-specific determination of protein and peptide concentrations by absorbance at 205 nm. Protein Sci. 2013, 22, 851–858. [Google Scholar] [CrossRef]
- Ehret-Sabatier, L.; Loew, D.; Goyffon, M.; Fehlbaum, P.; Hoffmann, J.A.; van Dorsselaer, A.; Bulet, P. Characterization of novel cysteine-rich antimicrobial peptides from scorpion blood. J. Biol. Chem. 1996, 271, 29537–29544. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Morrissey, J.H. Silver stain for proteins in polyacrylamide gels: A modified procedure with enhanced uniform sensitivity. Anal. Biochem. 1981, 117, 307–310. [Google Scholar] [CrossRef]
- Ma, B.; Zhang, K.; Hendrie, C.; Liang, C.; Li, M.; Doherty-Kirby, A.; Lajoie, G. PEAKS: Powerful software for peptide de novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2003, 17, 2337–2342. [Google Scholar] [CrossRef] [PubMed]



| Microorganism | MIC (μM (μg/mL)) 1 |
|---|---|
| Gram-negative bacteria | |
| Escherichia coli SBS363 | 4.6 (7.6) |
| E. coli D31 | 4.6 (7.6) |
| Pseudomonas aeruginosa ATCC 27853 P. aeruginosa PA14 Enterobacter cloacae β-12 | 1.15 (1.9) 4.6 (7.6) 2.3 (3.8) |
| Gram-positive bacteria | |
| Micrococcus luteus A270 | ND |
| Staphylococcus aureus ATCC 29213 Bacillus subtilis ATCC 6633 | ND ND |
| Fungus | |
| Aspergillus niger | ND |
| Yeasts | |
| Candida albicans MDM8 Candida krusei IOC 4559 | ND ND |
| Net charge | −3 |
| Theoretical isoelectric point (pI) | 3.77 |
| Molar extinction coefficient (ε) | 51,100 M−1 cm−1 |
| Aliphatic index | 60.62 |
| GRAVY (grand average of hydropathicity) | −0.769 1 |
| Instability index | −11.26 2 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Segura-Ramírez, P.J.; Silva Júnior, P.I. Loxosceles gaucho Spider Venom: An Untapped Source of Antimicrobial Agents. Toxins 2018, 10, 522. https://doi.org/10.3390/toxins10120522
Segura-Ramírez PJ, Silva Júnior PI. Loxosceles gaucho Spider Venom: An Untapped Source of Antimicrobial Agents. Toxins. 2018; 10(12):522. https://doi.org/10.3390/toxins10120522
Chicago/Turabian StyleSegura-Ramírez, Paula J, and Pedro I Silva Júnior. 2018. "Loxosceles gaucho Spider Venom: An Untapped Source of Antimicrobial Agents" Toxins 10, no. 12: 522. https://doi.org/10.3390/toxins10120522
APA StyleSegura-Ramírez, P. J., & Silva Júnior, P. I. (2018). Loxosceles gaucho Spider Venom: An Untapped Source of Antimicrobial Agents. Toxins, 10(12), 522. https://doi.org/10.3390/toxins10120522




