Transketolase Is Identified as a Target of Herbicidal Substance α-Terthienyl by Proteomics
Abstract
:1. Introduction
2. Results
2.1. Herbicidal Activity of α-Terthienyl on Plants
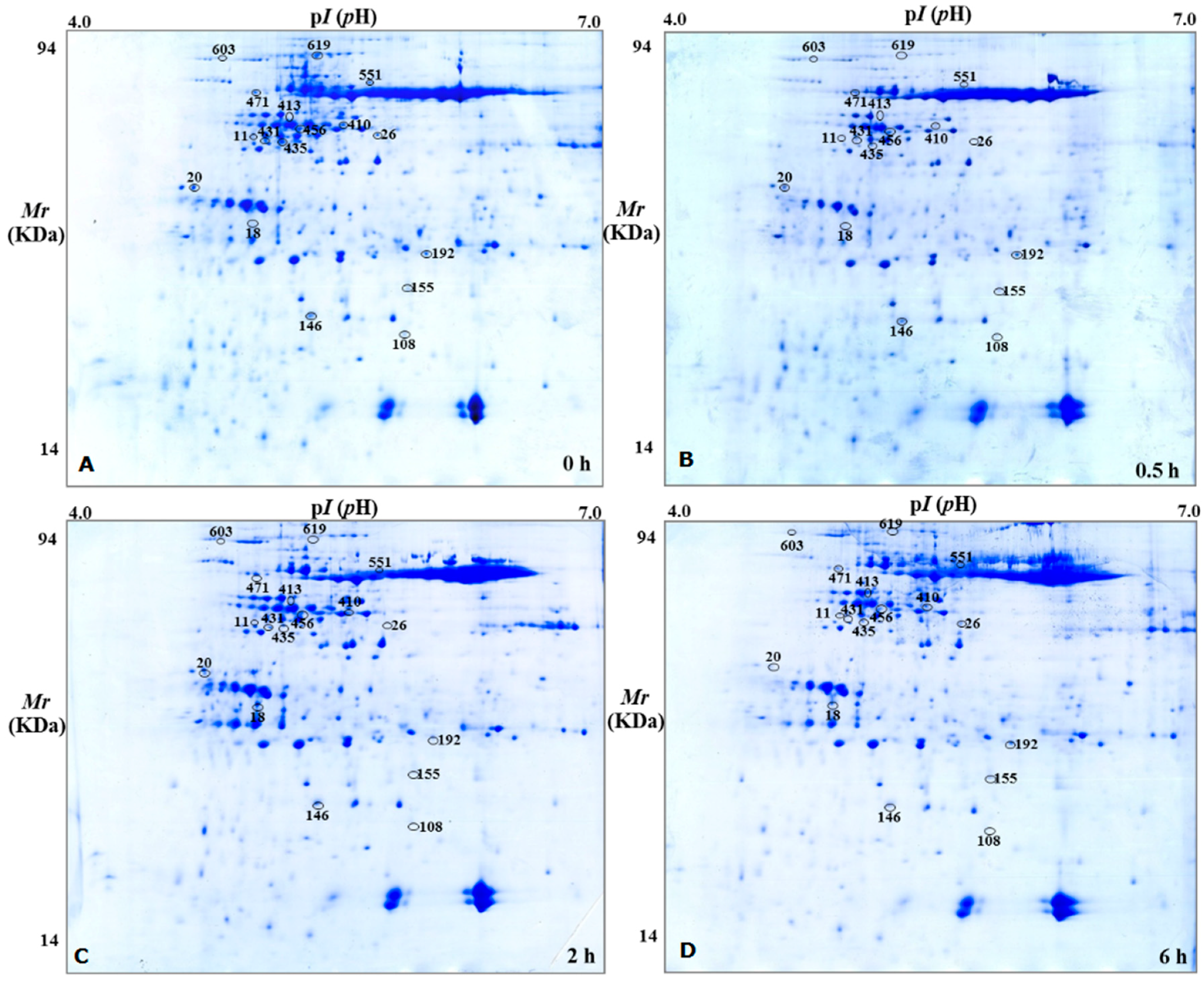
2.2. Sample Extraction and Two-Dimensional Gel Electrophoresis
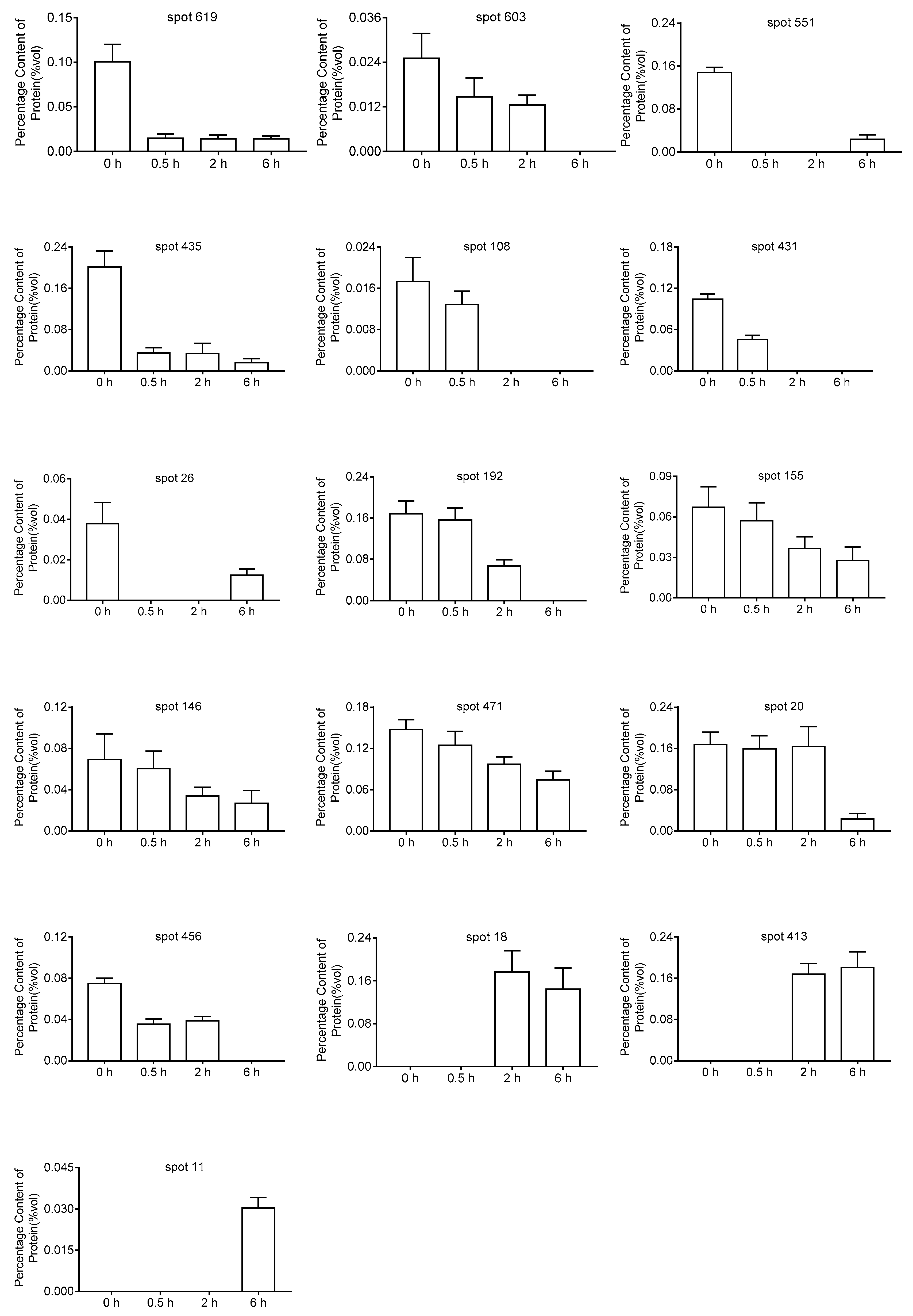
2.3. Mass Spectrometry and Analysis of DEPs
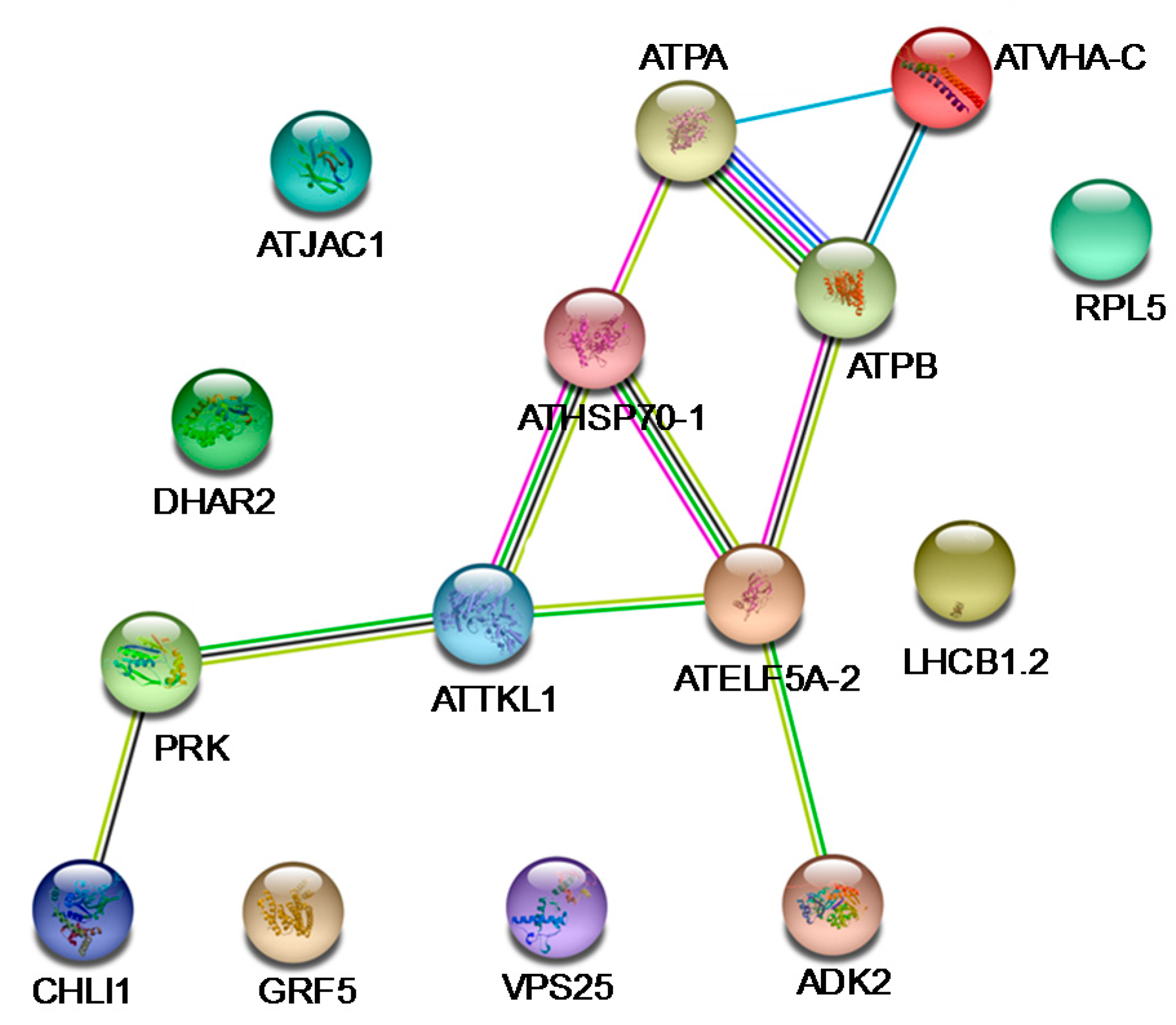
2.4. The Gene Ontology and Protein-Protein Interaction Networks Analysis of DEPs
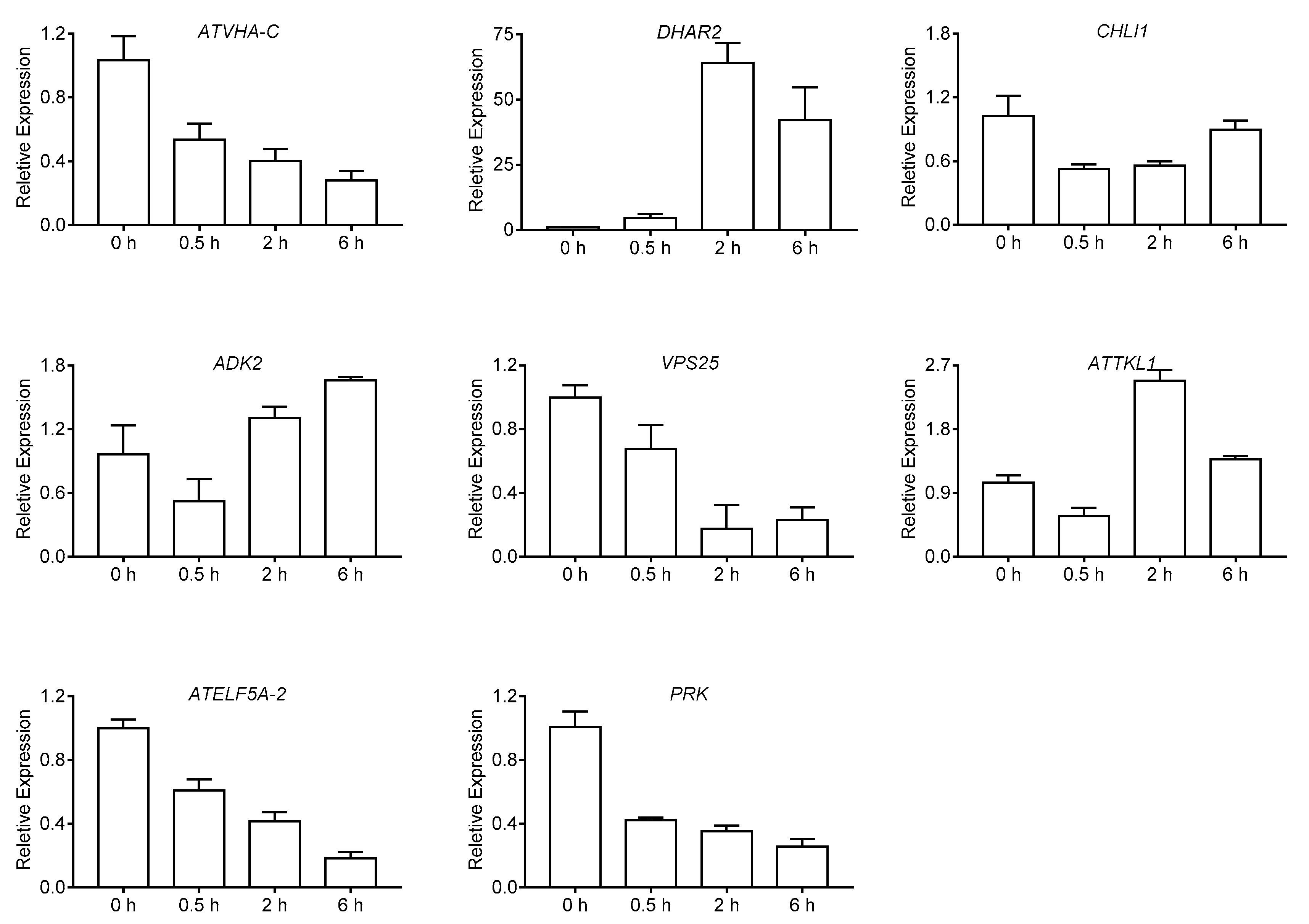
2.5. Quantitative Real-Time PCR Analysis for DEPs
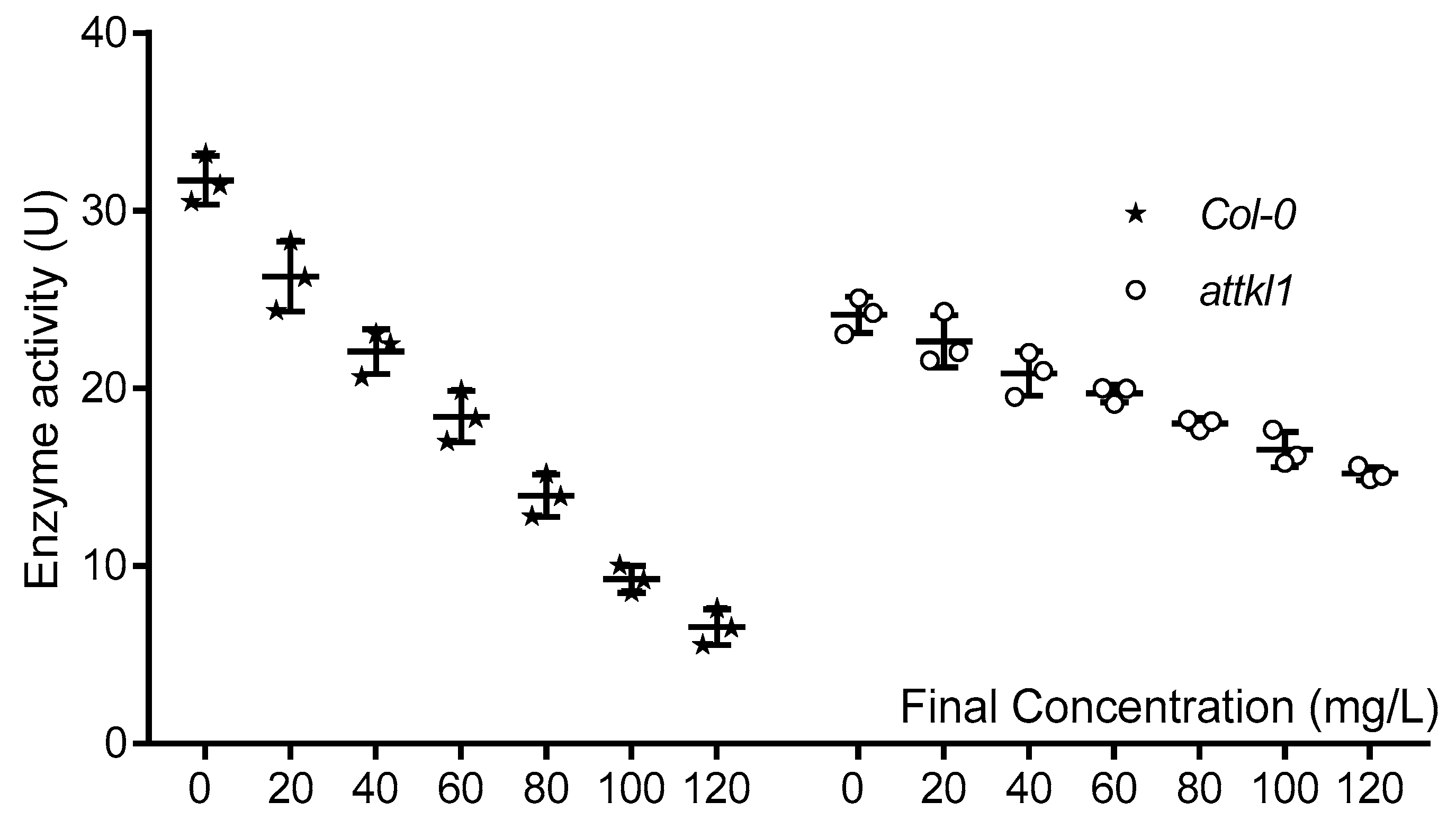
2.6. Transketolase Is One of Target Proteins Which Respond to α-Terthienyl
3. Discussion
3.1. Proteomic Approach is a Powerful Tool for Identification the Mode of Action of Herbicide
3.2. α-Terthienyl Can Inhibit Photosynthesis though Transketolase
4. Conclusions
5. Materials and Methods
5.1. The Herbicidal Bioassay of α-Terthienyl
5.2. Plant Material and Treatments
5.3. 2-DE and Data Analysis
5.4. Image Acquisition and Data Analysis
5.5. Proteins Identified by Mass Spectrometry
5.6. RNA Extraction and Q-PCR Analysis
5.7. The Sensitivity Test of α-Terthienyl in T-DNA Mutants
5.8. Enzyme Activity Assay of Transketolase
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Steinrucken, H.C.; Amrhein, N. The herbicide glyphosate is a potent inhibitor of 5-enolpyruvyl-shikimic acid-3-phosphate synthase. Biochem. Biophys. Res. Commun. 1980, 94, 1207–1212. [Google Scholar] [CrossRef]
- LaRossa, R.A.; Schloss, J.V. The sulfonylurea herbicide sulfometuron methyl is an extremely potent and selective inhibitor of acetolactate synthase in Salmonella typhimurium. J. Biol. Chem. 1984, 259, 8753–8757. [Google Scholar] [PubMed]
- Chaleff, R.S.; Mauvais, C.J. Acetolactate synthase is the site of action of two sulfonylurea herbicides in higher plants. Science 1984, 224, 1443–1445. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, E.; Zeng, L.; Baird, W.V. Alpha-tubulin missense mutations correlate with antimicrotubule drug resistance in Eleusine indica. Plant Cell 1998, 10, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.; Skerritt, J.H.; Thomas, M.; Korth, W.; Bowmer, K.H.; Larkin, K.A.; Ferguson, B.S. Quantification of the urea herbicide, diuron, in water by enzyme immunoassay. Bull. Environ. Contam. Toxicol. 1995, 55, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Riggle, B. Development of a preliminary enzyme-linked immunosorbent assay for the herbicide trifluralin. Bull. Environ. Contam. Toxicol. 1991, 46, 404–409. [Google Scholar] [CrossRef] [PubMed]
- Odell, J.T.; Caimi, P.G.; Yadav, N.S.; Mauvais, C.J. Comparison of increased expression of wild-type and herbicide-resistant acetolactate synthase genes in transgenic plants, and indication of posttranscriptional limitation on enzyme activity. Plant Physiol. 1990, 94, 1647–1654. [Google Scholar] [CrossRef] [PubMed]
- Duke, S.O.; Bajsa, J.; Pan, Z. Omics methods for probing the mode of action of natural and synthetic phytotoxins. J. Chem. Ecol. 2013, 39, 333–347. [Google Scholar] [CrossRef] [PubMed]
- Huo, J.Q.; Xing, J.H.; Zhang, L.H.; Kang, Z.H.; Zhang, J.L. Isolation and structural identification of herbicidal active substance from root of Flaveria bident (L.) kuntze. J. Integr. Agric. 2014, 13, 804–810. [Google Scholar] [CrossRef]
- Bakker, J.; Gommers, F.J.; Nieuwenhuis, I.; Wynberg, H. Photoactivation of the nematicidal compound alpha-terthienyl from roots of marigolds (Tagetes species). A possible singlet oxygen role. J. Biol. Chem. 1979, 254, 1841–1844. [Google Scholar] [PubMed]
- Chan, G.F.; Prihoda, M.; Towers, G.H.; Mitchell, J.C. Phototoxicity evoked by alpha-terthienyl. Contact Dermat. 1977, 3, 215–216. [Google Scholar]
- Yamamoto, E.; MacRae, W.D.; Garcia, F.J.; Towers, G.H. Photodynamic hemolysis caused by alpha-terthienyl. Planta Med. 1984, 50, 124–127. [Google Scholar] [CrossRef] [PubMed]
- Ji, H.F.; Shen, L. Triplet excited state characters and photosensitization mechanisms of alpha-terthienyl: A theoretical study. J. Photochem. Photobiol. B Biol. 2009, 94, 51–53. [Google Scholar] [CrossRef] [PubMed]
- Downum, K.R. Light-activated plant defence. New Phytol. 2010, 122, 401–420. [Google Scholar] [CrossRef]
- Campbell, G.; Lambert, J.D.; Arnason, T.; Towers, G.H. Allelopathic properties of alpha-terthienyl and phenylheptatriyne, naturally occurring compounds from species of asteraceae. J. Chem. Ecol. 1982, 8, 961–972. [Google Scholar] [CrossRef] [PubMed]
- Brennan, T.M. Photosensitized inhibition of photosynthetic 14CO2 fixation by α-terthienyl and Ultraviolet-A. Photochem. Photobiol. 2008, 59, 631–636. [Google Scholar]
- Harris, E.H. Chlamydomonas as a model organism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2001, 52, 363–406. [Google Scholar] [CrossRef] [PubMed]
- Fischer, B.B.; Rufenacht, K.; Dannenhauer, K.; Wiesendanger, M.; Eggen, R.I. Multiple stressor effects of high light irradiance and photosynthetic herbicides on growth and survival of the green alga Chlamydomonas reinhardtii. Environ. Toxicol. Chem. 2010, 29, 2211–2219. [Google Scholar] [CrossRef] [PubMed]
- Jamers, A.; De, C.W. Effect assessment of the herbicide paraquat on a green alga using differential gene expression and biochemical biomarkers. Environ. Toxicol. Chem. 2010, 29, 893–901. [Google Scholar] [CrossRef] [PubMed]
- Brennan, T.M. Decarboxylation of lndole-3-acetic acid and inhibition of growth in avena sativa seedlings by plant-derived photosensitizers. Photochem. Photobiol. 2010, 64, 1001–1006. [Google Scholar] [CrossRef]
- Gommers, F.J.; Bakker, J. Mode of action of α-terthienyl and related compounds may explain the suppressant effects of tagetes species on populations of free living endoparasitic plant nematodes. Bioact. Mol. 1988, 7, 61–69. [Google Scholar]
- Asirvatham, V.S.; Watson, B.S.; Sumner, L.W. Analytical and biological variances associated with proteomic studies of medicago truncatula by two-dimensional polyacrylamide gel electrophoresis. Proteomics 2002, 2, 960–968. [Google Scholar] [CrossRef]
- Henkes, S.; Sonnewald, U.; Badur, R.; Flachmann, R.; Stitt, M. A small decrease of plastid transketolase activity in antisense tobacco transformants has dramatic effects on photosynthesis and phenylpropanoid metabolism. Plant Cell 2001, 13, 535–551. [Google Scholar] [CrossRef] [PubMed]
- Rocha, A.G.; Mehlmer, N.; Stael, S.; Mair, A.; Parvin, N.; Chigri, F.; Teige, M.; Vothknecht, U.C. Phosphorylation of arabidopsis transketolase at ser428 provides a potential paradigm for the metabolic control of chloroplast carbon metabolism. Biochem. J. 2014, 458, 313–322. [Google Scholar] [CrossRef] [PubMed]
- Nestler, H.; Groh, K.J.; Schonenberger, R.; Eggen, R.I.; Suter, M.J. Linking proteome responses with physiological and biochemical effects in herbicide-exposed Chlamydomonas reinhardtii. J. Proteom. 2012, 75, 5370–5385. [Google Scholar] [CrossRef] [PubMed]
- Holmes, P.; Farquharson, R.; Hall, P.J.; Rolfe, B.G. Proteomic analysis of root meristems and the effects of acetohydroxyacid synthase-inhibiting herbicides in the root of Medicago truncatula. J. Proteome Res. 2006, 5, 2309–2316. [Google Scholar] [CrossRef] [PubMed]
- Villafranca, J.J.; Axelrod, B. Heptulose synthesis from nonphosphorylated aldoses and ketoses by spinach transketolase. J. Biol. Chem. 1971, 246, 3126–3131. [Google Scholar] [PubMed]
- Murphy, D.J.; Walker, D.A. The properties of transketolase from photosynthetic tissue. Planta 1982, 155, 316–320. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Mochizuki, N.; Yoshimura, N.; Motohashi, K.; Hisabori, T.; Masuda, T. Functional analysis of Arabidopsis thaliana isoforms of the mg-chelatase chli subunit. Photochem. Photobiol. Sci. Off. J. Eur. Photochem. Assoc. Eur. Soc. Photobiol. 2008, 7, 1188–1195. [Google Scholar] [CrossRef] [PubMed]
- Rissler, H.M.; Collakova, E.; DellaPenna, D.; Whelan, J.; Pogson, B.J. Chlorophyll biosynthesis. Expression of a second chl i gene of magnesium chelatase in arabidopsis supports only limited chlorophyll synthesis. Plant Physiol. 2002, 128, 770–779. [Google Scholar] [CrossRef] [PubMed]
- Du, S.Y.; Zhang, X.F.; Lu, Z.; Xin, Q.; Wu, Z.; Jiang, T.; Lu, Y.; Wang, X.F.; Zhang, D.P. Roles of the different components of magnesium chelatase in abscisic acid signal transduction. Plant Mol. Biol. 2012, 80, 519–537. [Google Scholar] [CrossRef] [PubMed]
- Gibson, J.L.; Chen, J.H.; Tower, P.A.; Tabita, F.R. The form ii fructose 1,6-bisphosphatase and phosphoribulokinase genes form part of a large operon in rhodobacter sphaeroides: Primary structure and insertional mutagenesis analysis. Biochemistry 1990, 29, 8085–8093. [Google Scholar] [CrossRef] [PubMed]
- Trimpin, S.; Brizzard, B. Analysis of insoluble proteins. Biotechniques 2009, 46, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Bindschedler, L.V.; Cramer, R. Quantitative plant proteomics. Proteomics 2011, 11, 756–775. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Yun, X.; Rao, W.; Xiao, C. Antioxidative cellular response of lepidopteran ovarian cells to photoactivated alpha-terthienyl. Pestic. Biochem. Physiol. 2017, 137, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Liu, Y.; Zhan, T.; Deng, Y.; He, Y. Comparable susceptibilities of human 293 cells and insect tn-5b1-4 cells to photoactivated alpha-terthienyl. J. Agric. Food Chem. 2010, 58, 2637–2642. [Google Scholar] [CrossRef] [PubMed]
- Cilento, G.; Nascimento, A.L. Generation of electronically excited triplet species at the cellular level: A potential source of genotoxicity. Toxicol. Lett. 1993, 67, 17–28. [Google Scholar] [CrossRef]
- Cilento, G.; Adam, W. Photochemistry and photobiology without light. Photochem. Photobiol. 1988, 48, 361–368. [Google Scholar] [CrossRef] [PubMed]
- Cilento, G. Photobiochemistry without Light: Intracellular Generation and Transfer of Electronic Energy; Springer: Boston, MA, USA, 1988. [Google Scholar]
- Grijalba, M.T.; Nantes, L.L.; Cilento, G.; Quina, F.H. Tris(bipyridine) ruthenium(ii): An efficient detector of excited species generated by chemiluminescent processes. Photochem. Photobiol. 1996, 63, 697–701. [Google Scholar] [CrossRef]
- Brennan, T.M.; Lee, E.; Battaglia, P.R. Participation of the photosensitizer alpha-terthienyl in the peroxidase-catalyzed oxidation of indole-3-acetic acid. Photochem. Photobiol. 2000, 71, 355–360. [Google Scholar] [CrossRef]
- Zhao, B.; Huo, J.; Xing, J.; Meng, Q.I.; Zhang, J.; Dong, J. Homologous modeling of transketolase ATTKL1 and its combination with α-terthienyl in Arabidopsis thaliana. Chem. J. Chin. Univ. 2015, 36, 682–686. [Google Scholar]
- Zhang, J.L.; Zhang, L.H.; Liu, Y.C.; Ma, J.; Li, C.; Dong, J.G. The herbicidal activity of mutant isolates from Botrytis cinerea. Agric. Sci. China 2006, 5, 622–628. [Google Scholar] [CrossRef]
- Giavalisco, P.; Nordhoff, E.; Kreitler, T.; Kloppel, K.D.; Lehrach, H.; Klose, J.; Gobom, J. Proteome analysis of arabidopsis thaliana by two-dimensional gel electrophoresis and matrix-assisted laser desorption/ionisation-time of flight mass spectrometry. Proteomics 2005, 5, 1902–1913. [Google Scholar] [CrossRef] [PubMed]
- Neuhoff, V.; Arold, N.; Taube, D.; Ehrhardt, W. Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using coomassie brilliant blue g-250 and r-250. Electrophoresis 1988, 9, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.G.; Mitsukawa, N.; Oosumi, T.; Whittier, R.F. Efficient isolation and mapping of Arabidopsis thaliana t-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. Cell Mol. Biol. 1995, 8, 457–463. [Google Scholar] [CrossRef]

| Species/Treatment | Median Inhibitory Concentration (mg/L) | Toxicity Regression Equations | R2 |
|---|---|---|---|
| D. sanguinalis/α-terthienyl | 22.03 | y = 4.30 + 0.51x | 0.9209 |
| D. sanguinalis/atrazine | 32.95 | y = −3.09 + 2.02x | 0.9162 |
| A. thaliana/α-terthienyl | 29.64 | y = 4.30 + 0.49x | 0.9199 |
| A. thaliana/atrazine | 44.57 | y = 4.01 + 0.63x | 0.9559 |
| C. reinhardti/α-terthienyl | 1.10 | y = 4.97 + 0.65x | 0.9597 |
| C. reinhardti/atrazine | 2.26 | y = 4.78 + 0.63x | 0.9778 |
| Spot ID | Biological Process | Protein Name | Gene No. | Gene Name | Coverage (%) | pI/Mr Experimental | pI/Mr Theoretical | Different Expression Protein Detected by 2-DE | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 h | 0.5 h | 2 h | 6 h | ||||||||
| 619 | Carbohydrate metabolism | Transketolase-1 | AT3G60750 | ATTKL1 | 74 | 5.42/81 | 5.94/80 | 〇 | ↓ | ↓ | ↓ |
| 603 | defense response | heat shock cognate protein 70-1 | AT5G02500 | ATHSP70-1 | 71 | 4.95/77 | 5.03/72 | 〇 | ↓ | ↓ | ↓ |
| 551 | Energy metabolism | ATP synthase subunit beta | ATCG00480 | ATPB | 88 | 5.64/60 | 5.38/54 | 〇 | ↓ | ↓ | ↓ |
| 435 | reductive pentose-phosphate cycle | Phosphoribulokinase | AT1G32060 | PRK | 71 | 5.21/41 | 5.71/44 | 〇 | ↓ | ↓ | ↓ |
| 108 | Nucleotide and amino acid translation | Eukaryotic translation initiation factor 5A-2 | AT1G26630 | ATELF5A-2 | 72 | 5.83/18 | 5.55/17 | 〇 | ≡ | ↓ | ↓ |
| 431 | Photosynthesis | Magnesium-chelatase subunit ChlI-1 | AT4G18480 | CHLI1 | 66 | 5.21/46.5 | 6.08/46.5 | 〇 | ↓ | ↓ | ↓ |
| 26 | Energy metabolism | V-type proton ATPase subunit C | AT1G12840 | ATVHA-C | 81 | 5.68/41 | 5.4/42 | 〇 | ≡ | ↓ | ↓ |
| 192 | Response to Stress | Glutathione S-transferase DHAR2 | AT1G75270 | DHAR2 | 81 | 5.97/25 | 5.79/24 | 〇 | ≡ | ↓ | ↓ |
| 155 | translation | 60S ribosomal protein L5 | ATMG00210 | RPL5 | 32 | 5.90/22 | 5.6/21 | 〇 | ≡ | ↓ | ↓ |
| 146 | Protein transport | Vacuolar protein sorting-associated protein 25 | AT4G19003 | VPS25 | 26 | 5.30/20 | 5.48/20 | 〇 | ≡ | ↓ | ↓ |
| 471 | Energy metabolism | ATP synthase subunit alpha | ATCG00120 | ATPA | 76 | 5.20/60 | 5.19/55 | 〇 | ≡ | ↓ | ↓ |
| 20 | response to cadmium ion | 14-3-3-like protein GF14 upsilon | AT5G16050 | GRF5 | 78 | 4.72/32 | 4.73/30 | 〇 | ≡ | ↓ | ↓ |
| 456 | reductive pentose-phosphate cycle | Phosphoribulokinase | AT1G32060 | PRK | 62 | 5.34/44 | 5.71/44 | 〇 | ≡ | ≡ | ↓ |
| 18 | Carbohydrate metabolism | Chlorophyll a-b binding protein 2 | AT1G29910 | LHCB1.2 | 44 | 5.02/29 | 5.29/28 | 〇 | ≡ | ≡ | ↑ |
| 413 | Response to Stress | Myrosinase-binding protein | AT3G16470 | ATJAC1 | 69 | 5.26/48 | 5.12/48 | 〇 | ≡ | ↑ | ↑ |
| 11 | AMP salvage | Adenosine kinase 2 | AT5G03300 | ADK2 | 68 | 5.01/43 | 5.14/38 | 〇 | ≡ | ≡ | ↑ |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, B.; Huo, J.; Liu, N.; Zhang, J.; Dong, J. Transketolase Is Identified as a Target of Herbicidal Substance α-Terthienyl by Proteomics. Toxins 2018, 10, 41. https://doi.org/10.3390/toxins10010041
Zhao B, Huo J, Liu N, Zhang J, Dong J. Transketolase Is Identified as a Target of Herbicidal Substance α-Terthienyl by Proteomics. Toxins. 2018; 10(1):41. https://doi.org/10.3390/toxins10010041
Chicago/Turabian StyleZhao, Bin, Jingqian Huo, Ning Liu, Jinlin Zhang, and Jingao Dong. 2018. "Transketolase Is Identified as a Target of Herbicidal Substance α-Terthienyl by Proteomics" Toxins 10, no. 1: 41. https://doi.org/10.3390/toxins10010041
APA StyleZhao, B., Huo, J., Liu, N., Zhang, J., & Dong, J. (2018). Transketolase Is Identified as a Target of Herbicidal Substance α-Terthienyl by Proteomics. Toxins, 10(1), 41. https://doi.org/10.3390/toxins10010041





