Cyclopiazonic Acid Biosynthesis of Aspergillus flavus and Aspergillus oryzae
Abstract
:1. Introduction
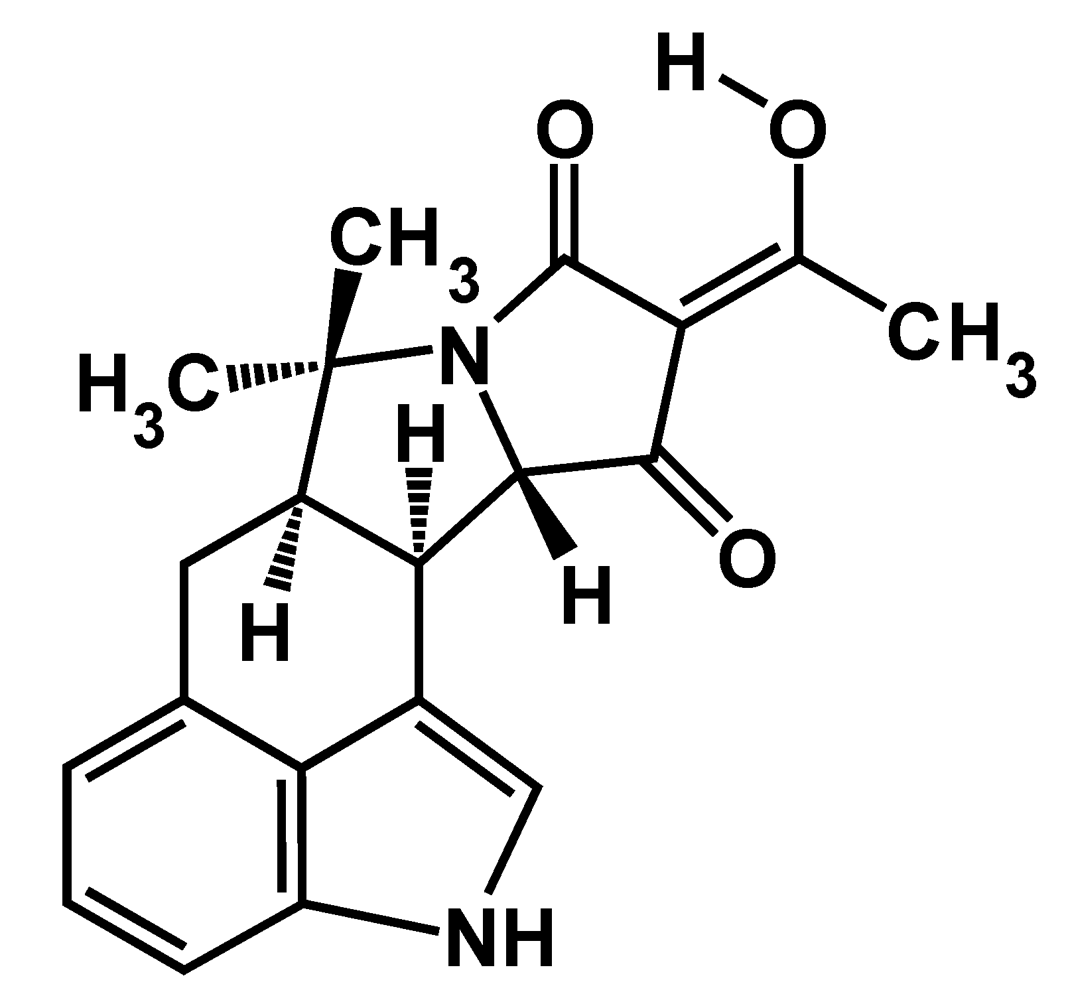
2. CPA-Producing Fungi
| Sources | No.a | BGb | AF/CPA | AF/− | −/CPA | −/− | References |
|---|---|---|---|---|---|---|---|
| Peanuts, soybean, wheat, Argentina | 87 | 5 | 27 | 2 | 57 | 14 | [27] |
| Peanuts, Argentina | 38 | 2 | 79 | 0 | 21 | 0 | [28] |
| Peanuts, Argentina | 29 | 3 | 49 | 3 | 24 | 24 | [29] |
| Soils, Argentina | 218L | 8 | 77 | 11 | 11 | 1 | [30] |
| 73S | 88 | 4 | 8 | 0 | |||
| 70N | 78 | 9 | 10 | 3 | |||
| Dried vine berries, Argentina | 5 | 0 | 0 | 100 | 0 | [31] | |
| Grain, smoked dried meat products, Croatia | 96 | 0 | 10 | 5 | 85 | [32] | |
| Corn, wheat, feeds, Hungary | 32 | 0 | 0 | 59 | 41 | [33] | |
| Sour lime, India | 25 | 20 | 40 | ? | ? | [34] | |
| Soils, Iran | 58 | 21 | 7 | 22 | 50 | [35] | |
| Peanuts, Israel | 200 | 19 | 73 | 4 | 4 | [36] | |
| Maize, Italy | 62 | 8 | 45 | 21 | 13 | 21 | [37] |
| Almonds, Portugal | 15 | 1 | 20 | 0 | 0 | 80 | [38] |
| Feeds, Queensland | 31 | 7 | 65 | 3 | 22 | 10 | [39] |
| Cocoa beans, Spain | 100 | 20 | 15 | 32 | 17 | 36 | [40] |
| Maize, US | 19 | 58 | 5 | 16 | 21 | [41] | |
| Soils, US | 774L | 71 | <1 | 12 | 16 | [42] | |
| 309S | 99 | <1 | <1 | 0 | |||
| Corn, nuts, animals and humans, Brazil, Uganda, US | 54 | 26 | 7 | 26 | 41 | [43] |
3. Toxicological and Pathological Effects of CPA
4. Biosynthesis of CPA
4.1. The mevalonate pathway key to CPA formation in A. flavus
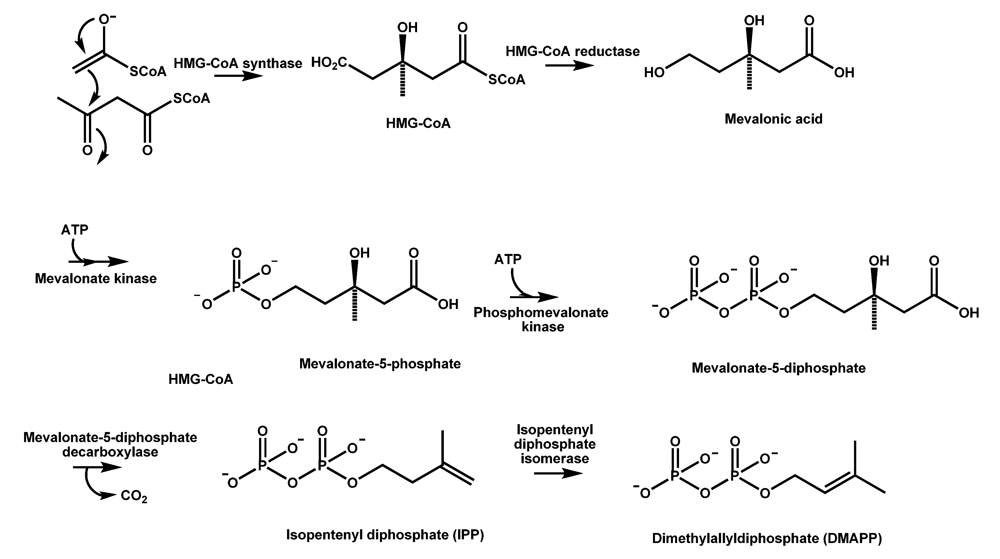
| Enzyme | Gene Locusa | Location |
|---|---|---|
| `3-Hydroxy-3-methylglutaryl-CoA synthase | AFL2G_02388.2 | Contig 2, 2508858~2510466 − |
| AFL2G_11649.2 | Contig 14, 1056212~1057748 − | |
| AO090003000611 | Chromosome 2, SC003,1640889~1642497 + | |
| AO090010000487 | Chromosome 8, SC010, 1307684~1309220 − | |
| HMG-CoA reductase | AFL2G_08234.2 | Contig 9, 779011~783044 - |
| AFL2G_12082.2 | Contig 15, 462348~465804 - | |
| AO090120000217 | Chromosome 5, SC113, 541346~544055 - | |
| AO090103000311 | Chromosome 8, SC103, 805908~807609 + | |
| Mevalonate kinase | AFL2G_04604.2 | Contig 4, 2076279~2077883 − |
| AO090023000793 | Chromosome 3, SC023, 2058446~2060050 - | |
| Phosphomevalonate kinase | AFL2G_11633.2 | Contig 14, 1009373~1011005 - |
| AO090010000471 | Chromosome 8, SC010, 1260354~1261854 - | |
| Pyrophosphomevalonate decarboxylase | AFL2G_04673.2 | Contig 4: 2266676~2268145 - |
| AO090023000862 | Chromosome 3, SC023, 2250974~2252258 − | |
| Isopentenyl pyrophosphate isomerase | AFL2G_04341.2 | Contig 4, 1322573~1323720 + |
| AFL2G_04245.2 | Contig 4: 1026352~1027394 + | |
| AO090023000500 | Chromosome 3, SC023, 1306204~1307211 + | |
| AO090023000391 | Chromosome 3, SC023, 1004129~1005046 + |
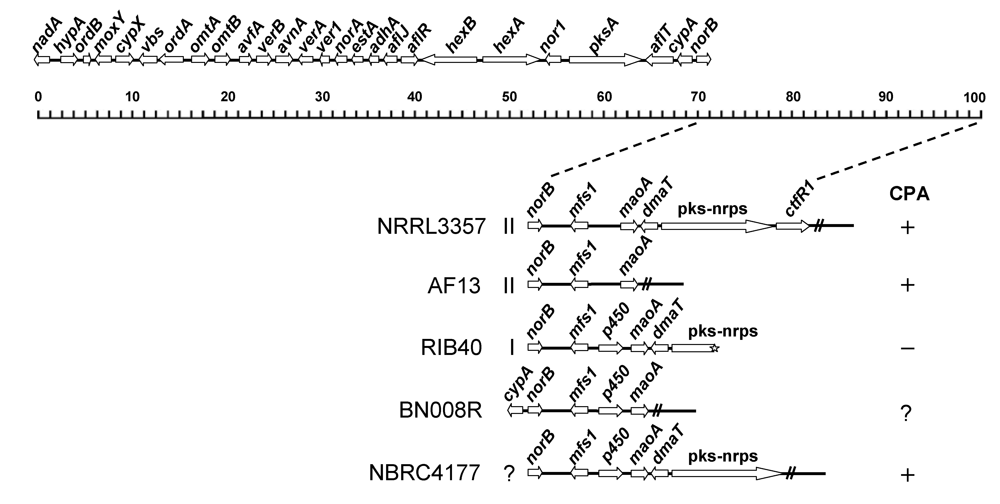
4.2. The gene cluster involved in CPA biosynthesis is adjacent to the aflatoxin gene cluster in A. flavus/oryzae
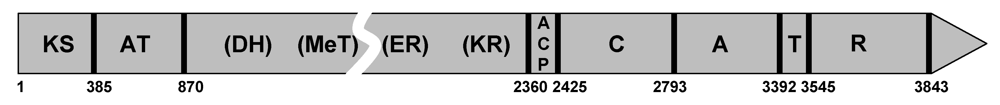
5. Characterization of the Enzymes Involved in CPA Biosynthesis
5.1. Formation of cAATrp by CpaS, a hybrid PKS-NRPS
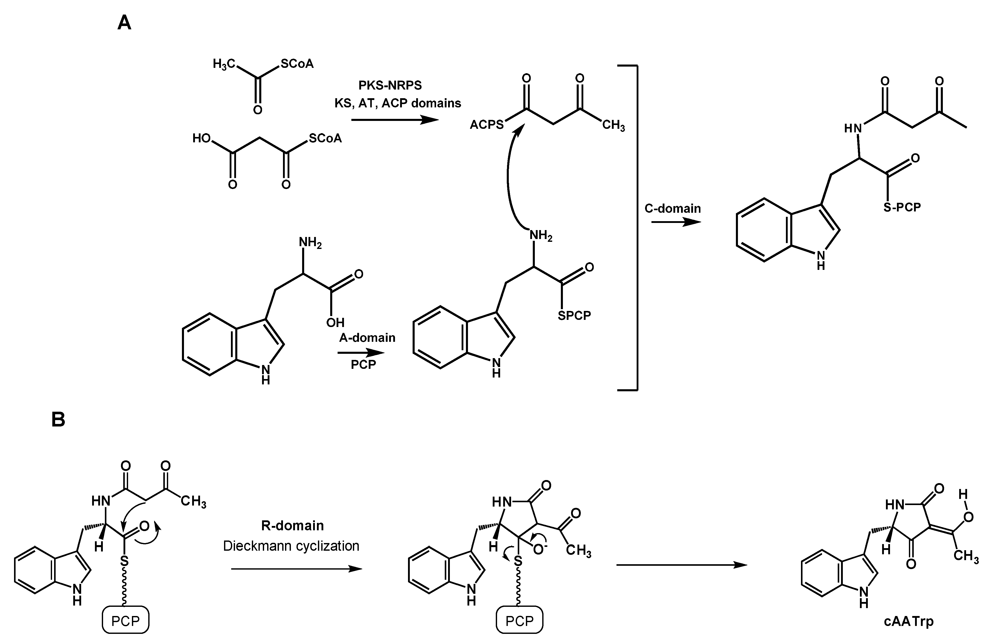
5.2. Conversion of cAATrp to β-CPA
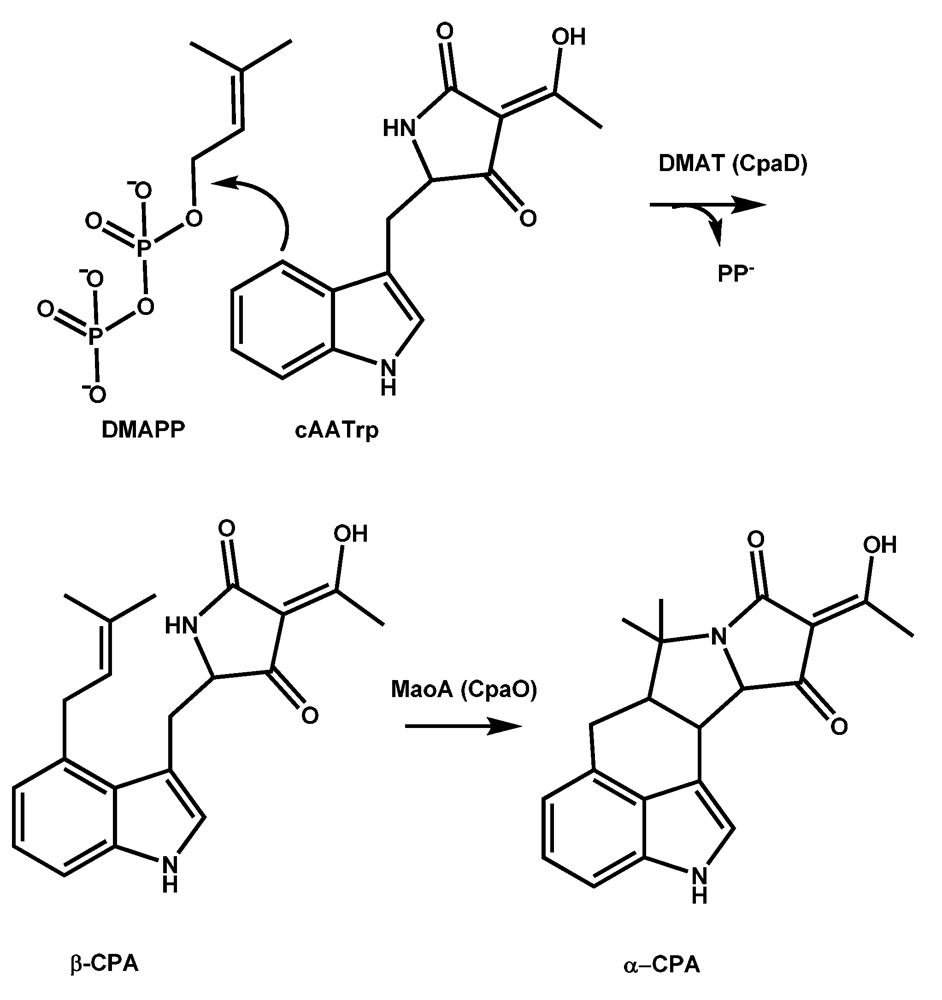
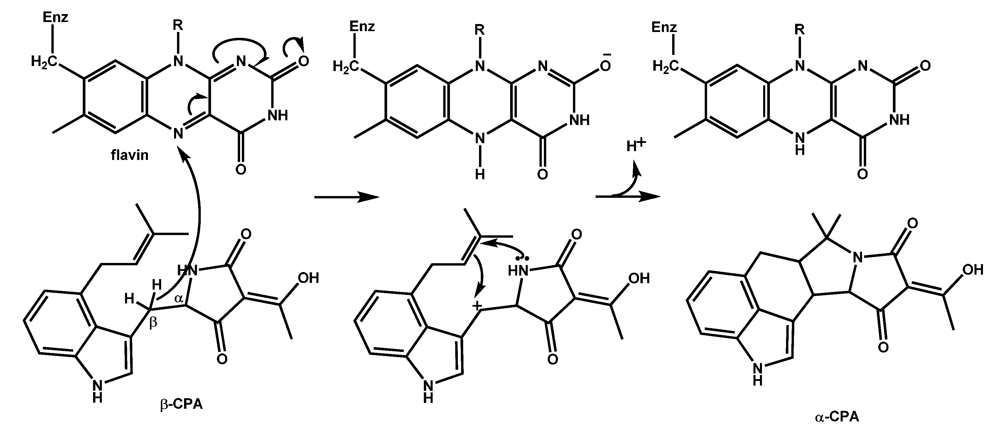
5.3. Conversion of β-CPA to α-CPA is catalyzed by the monoamine oxidase MaoA (CpaO)
6. Genetic Diversity in CPA Biosynthesis of A. flavus and A. oryzae
| Species | Strain | Sclerotial Morphotypea | AF/CPA Productionb | norB-cypA Patternc | p450d |
|---|---|---|---|---|---|
| A. flavus | CA28 | S | +/+ | I | N |
| CA42 | S | +/+ | I | N | |
| CA43 | S | +/+ | I | N | |
| CA44 | S | +/+ | I | N | |
| AF12 | S | +/+ | I | N | |
| AF70 | S | +/+ | I | N | |
| GA10-18 | S | +/+ | I | N | |
| VA4-36 | S | +/+ | I | N | |
| AF13 | L | +/+ | II | N | |
| CA14 | L | +/+ | II | N | |
| CA19 | L | +/+ | II | N | |
| GA9-9 | L | +/+ | II | N | |
| GA13-9 | L | +/+ | II | N | |
| NRRL3357 | L | +/+ | II | N | |
| VA2-9 | L | +/+ | II | N | |
| LA4-5 | L | −/+ | I | N | |
| SC6-9 | L | −/+ | I | N | |
| TX9-8 | L | −/+ | I | N | |
| GA4-4 | L | −/+ | II | N | |
| LA10-4 | L | −/+ | II | N | |
| MS1-1 | L | −/+ | I | Y | |
| NC3-6 | L | −/+ | I | Y | |
| SC3-5 | L | −/+ | I | Y | |
| TX21-9 | L | −/+ | I | Y | |
| A. oryzae | NBRC 4177 | ? | −/+ | ? | Y |
| RIB40 | N | −/− | I | Y | |
| SRRC304 | N | −/? | I | Y | |
| SRRC493 | N | −/? | I | Y | |
| SRRC2044 | N | −/? | I | Y | |
| A. parasiticus | BN009 | L | +/− | intact | N |
| SRRC2043 | L | +*/− | intact | N | |
| SRRC2999 | L | +/− | intact | N | |
| A. sojae | SRRC299 | N | −/? | intact | N |
| SRRC1123 | N | −/? | intact | N | |
| SRRC1126 | N | −/? | intact | N |
7. Possible Advantage of CPA to Fungi
8. Conclusions
References
- Abbas, H.K.; Accinelli, C.; Zablotowicz, R.M.; Abel, C.A.; Bruns, H.A.; Dong, Y.; Shier, W.T. Dynamics of mycotoxin and Aspergillus flavus levels in aging Bt and non-Bt corn residues under Mississippi no-till conditions. J. Agric. Food Chem. 2008, 56, 7578–7585. [Google Scholar] [PubMed]
- Njobeh, P.B.; Dutton, M.F.; Koch, S.H.; Chuturgoon, A.; Stoev, S.; Seifert, K. Contamination with storage fungi of human food from Cameroon. Int. J. Food Microbiol. 2009, 135, 193–198. [Google Scholar] [PubMed]
- Finoli, C.; Vecchio, A.; Galli, A.; Franzetti, L. Production of cyclopiazonic acid by molds isolated from Taleggio cheese. J. Food Prot. 1999, 62, 1198–1202. [Google Scholar] [PubMed]
- Sosa, M.J.; Cordoba, J.J.; Diaz, C.; Rodriguez, M.; Bermudez, E.; Asensio, M.A.; Nunez, F. Production of cyclopiazonic acid by Penicillium commune isolated from dry-cured ham on a meat extract-based substrate. J. Food. Prot. 2002, 65, 988–992. [Google Scholar] [PubMed]
- Lopez-Diaz, T.M.; Santos, J.A.; Garcia-Lopez, M.L.; Otero, A. Surface mycoflora of a Spanish fermented meat sausage and toxigenicity of Penicillium isolates. Int. J. Food Microbiol. 2001, 68, 69–74. [Google Scholar] [PubMed]
- Dorner, J.W.; Cole, R.J.; Erlington, D.J.; Suksupath, S.; McDowell, G.H.; Bryden, W.L. Cyclopiazonic acid residues in milk and eggs. J. Agric. Food Chem. 1994, 42, 1516–1518. [Google Scholar]
- Norred, W.P.; Porter, J.K.; Dorner, J.W.; Cole, R.J. Occurrence of the mycotoxin cyclopiazonic acid in meat after oral administration to chickens. J. Agri. Food Chem. 1988, 36, 113–116. [Google Scholar]
- Oliveira, C.A.F.; Sebastio, L.S.; Fagundes, H.; Rosim, R.E.; Fernandes, A.M. Aflatoxins and cyclopiazonic acid in feed and milk from dairy farms in Sao Paulo, Brazil. Food Add. Contam. 2008, 1, 147–152. [Google Scholar] [CrossRef]
- Holzapfel, C.W. The biosynthesis of cyclopizonic acid and related tetramic acids. In The Biosynthesis of Mycotoxins; Steyn, P.S., Ed.; Academic Press: New York, NY, USA, 1980; pp. 327–355. [Google Scholar]
- Spatz, J.H.; Welsch, S.J.; Duhaut, D.; Jäger, N.; Boursier, T.; Fredrich, M.; Allmendinger, L.; Ross, G.; Kolb, J.; Burdack, C.; Umkehrer, M. Tetramic acid derivatives via Ugi-Dieckmann-reaction. Tetrahedron Lett. 2009, 50, 1705–1707. [Google Scholar]
- Yang, Y.L.; Lu, C.P.; Chen, M.Y.; Chen, K.Y.; Wu, Y.C.; Wu, S.H. Cytotoxic polyketides containing tetramic acid moieties isolated from the fungus Myceliophthora thermophila: Elucidation of the relationship between cytotoxicity and stereoconfiguration. Chemistry 2007, 13, 6985–6991. [Google Scholar] [PubMed]
- Iwata, Y.; Maekawara, N.; Tanino, K.; Miyashita, M. Tetramic acid antibiotics: Stereoselective synthesis of streptolic acid and tirandalydigin. Angew. Chem. Int. Ed. 2005, 44, 1532–1536. [Google Scholar]
- Chang, P.-K.; Horn, B.W.; Dorner, J.W. Clustered genes involved in cyclopiazonic acid production are next to the aflatoxin biosynthesis gene cluster in Aspergillus flavus. Fungal Genet. Biol. 2009, 46, 176–182. [Google Scholar] [PubMed]
- Seshime, Y.; Juvvadi, P.R.; Tokuoka, M.; Koyama, Y.; Kitamoto, K.; Ebizuka, Y.; Fujii, I. Functional expression of the Aspergillus flavus PKS-NRPS hybrid CpaA involved in the biosynthesis of cyclopiazonic acid. Bioorg. Med. Chem. Lett. 2009, 19, 3288–3292. [Google Scholar] [PubMed]
- Tokuoka, M.; Seshime, Y.; Fujii, I.; Kitamoto, K.; Takahashi, T.; Koyama, Y. Identification of a novel polyketide synthase-nonribosomal peptide synthetase (PKS-NRPS) gene required for the biosynthesis of cyclopiazonic acid in Aspergillus oryzae. Fungal Genet. Biol. 2008, 45, 1608–1615. [Google Scholar] [PubMed]
- Holzapfel, C.W. The isolation and structure of cyclopiazonic acid, a toxic metabolite of Penicillium cyclopium Westling. Tetrahedron 1968, 24, 2101–2119. [Google Scholar] [CrossRef] [PubMed]
- Hermansen, K.; Frisvad, J.C.; Emborg, C.; Hansen, J. Cyclopiazonic acid production by submerged cultures of Penicillium and Aspergillus strains. FEMS Microbiol. Lett. 1984, 21, 253–261. [Google Scholar]
- Frisvad, J.C. The connection between the Penicillia and Aspergilli and mycotoxins with special emphasis on misidentified isolates. Arch. Environ. Contam. Toxicol. 1989, 18, 452–467. [Google Scholar] [PubMed]
- Burdock, G.A.; Flamm, W.G. Review Article: Safety assessment of the mycotoxin cyclopiazonic acid. Int. J. Toxicol. 2000, 19, 195–218. [Google Scholar]
- El-Banna, A.A.; Pitt, J.I.; Leistner, L. Production of mycotoxins by Penicillium species. Sys. Appl. Microbiol. 1987, 1, 42–46. [Google Scholar]
- Dorner, J.W. Production of cyclopiazonic acid by Aspergillus tamarii Kita. Appl. Environ. Microbiol. 1983, 46, 1435–1437. [Google Scholar] [PubMed]
- Dorner, J.W.; Cole, R.J.; Lomax, L.G.; Gosser, H.S.; Diener, U.L. Cyclopiazonic acid production by Aspergillus flavus and its effects on broiler chickens. Appl. Environ. Microbiol. 1983, 46, 698–703. [Google Scholar] [PubMed]
- Dorner, J.W.; Cole, R.J.; Diener, U.L. The relationship of Aspergillus flavus and Aspergillus parasiticus with reference to production of aflatoxins and cyclopiazonic acid. Mycopathologia 1984, 87, 13–15. [Google Scholar] [PubMed]
- Vinokurova, N.G.; Ivanushkina, N.E.; Khmel'nitskaia, II; Arinbasarov, M.U. Synthesis of alpha-cyclopiazonic acid by fungi of the genus Aspergillus. Prikl. Biokhim. Mikrobiol. 2007, 43, 486–489. [Google Scholar] [PubMed]
- Ohmomo, S.; Sugita, M.; Matazo, A. Isolation of cyclopiazonic acid, cyclopiazonic acid imine and bissecodehydrocyclopiazonic acid from the cultures of Aspergillus versicolor (Vuill.) Tiraboschi. J. Agr. Chem. Soc. Jpn. 1973, 47, 57–63. [Google Scholar]
- Domsch, K.H.; Gams, W.; Anderson, T.-H. Compendium of Soil Fungi; Academic Press: London, UK, 1980. [Google Scholar]
- Vaamonde, G.; Patriarca, A.; Fernandez Pinto, V.; Comerio, R.; Degrossi, C. Variability of aflatoxin and cyclopiazonic acid production by Aspergillus section Flavi from different substrates in Argentina. Int. J. Food Microbiol. 2003, 88, 79–84. [Google Scholar] [PubMed]
- Pildain, M.B.; Vaamonde, G.; Cabral, D. Analysis of population structure of Aspergillus flavus from peanut based on vegetative compatibility, geographic origin, mycotoxin and sclerotia production. Int. J. Food Microbiol. 2004, 93, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Novas, M.V.; Cabral, D. Association of mycotoxin and sclerotia production with compatibility groups in Aspergillus flavus from peanut in Argentina. Plant Dis. 2002, 86, 215–219. [Google Scholar]
- Barros, G.; Torres, A.; Chulze, S. Aspergillus flavus population isolated from soil of Argentina's peanut growing region. Sclerotia production and toxigenic profile. J. Sci. Food Agric. 2005, 85, 2349–2353. [Google Scholar] [CrossRef]
- Romero, S.M.; Comerio, R.M.; Larumbe, G.; Ritieni, A.; Vaamonde, G.; Fernandez Pinto, V. Toxigenic fungi isolated from dried vine fruits in Argentina. Int. J. Food Microbiol. 2005, 104, 43–49. [Google Scholar] [PubMed]
- Cvetnic, Z.; Pepeljnjak, S. Production of cyclopiazonic acid by aflatoxigenic and non-aflatoxigenic strains of Aspergillus flavus. Nahrung 1998, 42, 321–323. [Google Scholar] [PubMed]
- Richard, J.L.; Bhatnagar, D.; Peterson, S.; Sandor, G. Assessment of aflatoxin and cyclopiazonic acid production by Aspergillus flavus isolates from Hungary. Mycopathologia 1992, 120, 183–188. [Google Scholar]
- Bamba, R.; Sumbali, G. Co-occurrence of aflatoxin B1 and cyclopiazonic acid in sour lime (Citrus aurantifolia Swingle) during post-harvest pathogenesis by Aspergillus flavus. Mycopathologia 2005, 159, 407–411. [Google Scholar] [PubMed]
- Razzaghi-Abyaneh, M.; Shams-Ghahfarokhi, M.; Allameh, A.; Kazeroon-Shiri, A.; Ranjbar-Bahadori, S.; Mirzahoseini, H.; Rezaee, M.B. A survey on distribution of Aspergillus section Flavi in corn field soils in Iran: Population patterns based on aflatoxins, cyclopiazonic acid and sclerotia production. Mycopathologia 2006, 161, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Lisker, N.; Michaeli, R.; Frank, Z.R. Mycotoxigenic potential of Aspergillus flavus strains isolated from groundnuts growing in Israel. Mycopathologia 1993, 122, 177–183. [Google Scholar] [PubMed]
- Giorni, P.; Magan, N.; Pietri, A.; Bertuzzi, T.; Battilani, P. Studies on Aspergillus section Flavi isolated from maize in northern Italy. Int. J. Food Microbiol. 2007, 113, 330–338. [Google Scholar] [PubMed]
- Rodrigues, P.; Venancio, A.; Kozakiewicz, Z.; Lima, N. A polyphasic approach to the identification of aflatoxigenic and non-aflatoxigenic strains of Aspergillus section Flavi isolated from Portuguese almonds. Int. J. Food Microbiol. 2009, 129, 187–193. [Google Scholar] [PubMed]
- Blaney, B.J.; Kelly, M.A.; Tyler, A.L.; Connole, M.D. Aflatoxin and cyclopiazonic acid production by Queensland isolates of Aspergillus flavus and Aspergillus parasiticus. Aust. J. Agric. Res. 1989, 40, 395–400. [Google Scholar]
- Sanchez-Hervas, M.; Gil, J.V.; Bisbal, F.; Ramon, D.; Martinez-Culebras, P.V. Mycobiota and mycotoxin producing fungi from cocoa beans. Int. J. Food Microbiol. 2008, 125, 336–340. [Google Scholar] [PubMed]
- Lee, Y.J.; Hagler, W.M.J. Aflatoxin and cyclopiazonic acid production by Aspergillus flavus isolated from contaminated maize. J. Food Sci. 1991, 56, 871–872. [Google Scholar]
- Horn, B.W.; Dorner, J.W. Regional differences in production of aflatoxin B1 and cyclopiazonic acid by soil isolates of Aspergillus flavus along a transect within the United States. Appl. Environ. Microbiol. 1999, 65, 1444–1449. [Google Scholar] [PubMed]
- Gallagher, R.T.; Richard, J.L.; Stahr, H.M.; Cole, R.J. Cyclopiazonic acid production by aflatoxigenic and non-aflatoxigenic strains of Aspergillus flavus. Mycopathologia 1978, 66, 31–36. [Google Scholar] [PubMed]
- Urano, T.; Trucksess, M.W.; Beaver, R.W.; Wilson, D.M.; Dorner, J.W.; Dowell, F.E. Co-occurrence of cyclopiazonic acid and aflatoxins in corn and peanuts. J. Off. Anal. Chem. Int. 1992, 75, 838–841. [Google Scholar]
- Widiastuti, R.; Maryam, R.; Blaney, B.J.; Salfina; Stoltz, D.R. Cyclopiazonic acid in combination with aflatoxins, zearalenone and ochratoxin A in Indonesian corn. Mycopathologia 1988, 104, 153–156. [Google Scholar] [CrossRef] [PubMed]
- Mphande, F.A.; Siame, B.A.; Taylor, J.E. Fungi, aflatoxins and cyclopiazonic acid associated with peanut retailing in Botswana. J. Food Prot. 2004, 67, 96–102. [Google Scholar] [PubMed]
- Hayashi, Y.; Yoshizawa, T. Survey of cyclopiazonic acid contamination in corn from China and Southeast Asian countries. Mycotoxins 2005, 55, 3–8. [Google Scholar]
- Lansden, J.A.; Davidson, J.I. Occurrence of cyclopiazonic acid in peanuts. Appl. Environ. Microbiol. 1983, 45, 766–769. [Google Scholar] [PubMed]
- Castro, M.; Soares, L.; Furlani, R. Mycoflora, aflatoxigenic species and mycotoxins in freshly harvested corn (Zea mays L.): A preliminary study. Rev. Microbiol. 1995, 26, 289–295. [Google Scholar]
- Adebajo, L.O.; Idowu, A.A.; Adesanya, O.O. Mycoflora, and mycotoxins production in Nigerian corn and corn-based snacks. Mycopathologia 1994, 126, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Goncalez, E.; Nogueira, J.H.; Fonseca, H.; Felicio, J.D.; Pino, F.A.; Correa, B. Mycobiota and mycotoxins in Brazilian peanut kernels from sowing to harvest. Int. J. Food Microbiol. 2008, 123, 184–190. [Google Scholar] [PubMed]
- Bhattacharya, K.; Raha, S. Deteriorative changes of maize, groundnut and soybean seeds by fungi in storage. Mycopathologia 2002, 155, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Khosravi, A.R.; Mansouri, M.; Bahonar, A.R.; Shokri, H. Mycoflora of maize harvested from Iran and imported maize. Pak. J. Biol. Sci. 2007, 10, 4432–4437. [Google Scholar] [PubMed][Green Version]
- Ito, Y.; Peterson, S.W.; Wicklo, D.T.; Goto, T. Aspergillus pseudotamarii, a new aflatoxin producing species in Aspergillus section Flavi. Mycological Res. 2001, 105, 233–239. [Google Scholar] [CrossRef]
- Peterson, S.W.; Ito, Y.; Horn, B.W.; Goto, T. Aspergillus bombycis, a new aflatoxigenic species and genetic variation in its sibling species, A. nomius. Mycologia 2001, 93, 689–703. [Google Scholar] [CrossRef]
- Hesseltine, C.W.; Shotwell, O.D.; Smith, M.; Ellis, J.J.; Vandergraft, E.E.; Shannon, G.M. Production of various aflatoxins bystrains of the Aspergillus flavus series. In Toxic Microorganisms: Mycotoxin, Botulism, Proceedings of the First U.S.-Japan Conference on Toxic Microorganisms; Herzberg, M., Ed.; US Government Printing Office: Washington D.C.,USA, 1970; pp. 202–210. [Google Scholar]
- Cotty, P.J. Virulence and cultural characteristics of two Aspergillus flavus strains pathogenic on cotton. Phytopathology 1989, 79, 808–814. [Google Scholar]
- Geiser, D.M.; Dorner, J.W.; Horn, B.W.; Taylor, J.W. The phylogenetics of mycotoxin and sclerotium production in Aspergillus flavus and Aspergillus oryzae. Fungal Genet. Biol. 2000, 31, 169–179. [Google Scholar] [PubMed]
- Saito, M.; Tsuruta, O. A new variety of Aspergillus flavus from tropical soil in Thailand and its aflatoxin productivity. Proc. Jpn. Assoc. Mycotoxicol. 1993, 37, 31–36. [Google Scholar]
- Ehrlich, K.C.; Kobbeman, K.; Montalbano, B.G.; Cotty, P.J. Aflatoxin-producing Aspergillus species from Thailand. Int. J. Food Microbiol. 2007, 114, 153–159. [Google Scholar] [PubMed]
- Cotty, P.J.; Cardwell, K.F. Divergence of West African and North American communities of Aspergillus section Flavi. Appl. Environ. Microbiol. 1999, 65, 2264–2266. [Google Scholar] [PubMed]
- Pildain, M.B.; Frisvad, J.C.; Vaamonde, G.; Cabral, D.; Varga, J.; Samson, R.A. Two novel aflatoxin-producing Aspergillus species from Argentinean peanuts. Int. J. Syst. Evol. Microbiol. 2008, 58, 725–735. [Google Scholar] [PubMed]
- Luk, K.C.; Kobbe, B.; Townsend, J.M. Production of cyclopiazonic acid by Aspergillus flavus Link. Appl. Environ. Microbiol. 1977, 33, 211–212. [Google Scholar] [PubMed]
- Riley, R.T.; Goeger, D.E.; Yoo, H.; Showker, J.L. Comparison of three tetramic acids and their ability to alter membrane function in cultured skeletal muscle cells and sarcoplasmic reticulum vesicles. Toxicol. Appl. Pharmacol. 1992, 114, 261–267. [Google Scholar] [PubMed]
- Goeger, D.E.; Riley, R.T.; Dorner, J.W.; Cole, R.J. Cyclopiazonic acid inhibition of the Ca2+-transport ATPase in rat skeletal muscle sarcoplasmic reticulum vesicles. Biochem. Pharmacol. 1988, 37, 978–981. [Google Scholar] [PubMed]
- Moncoq, K.; Trieber, C.A.; Young, H.S. The molecular basis for cyclopiazonic acid inhibition of the sarcoplasmic reticulum calcium pump. J. Biol. Chem. 2007, 282, 9748–9757. [Google Scholar] [PubMed]
- Purchase, I.F. The acute toxicity of the mycotoxin cyclopiazonic acid to rats. Toxicol. Appl. Pharmacol. 1971, 18, 114–123. [Google Scholar] [PubMed]
- Nishie, K.; Cole, R.J.; Dorner, J.W. Toxic effects of cyclopiazonic acid in the early phase of pregnancy in mice. Res. Commun. Chem. Pathol. Pharmacol. 1987, 55, 303–315. [Google Scholar] [PubMed]
- Nuehring, L.P.; Rowland, G.N.; Harrison, L.R.; Cole, R.J.; Dorner, J.W. Cyclopiazonic acid mycotoxicosis in the dog. Am. J. Vet. Res. 1985, 46, 1670–1676. [Google Scholar] [PubMed]
- Lomax, L.G.; Cole, R.J.; Dorner, J.W. The toxicity of cyclopiazonic acid in weaned pigs. Vet. Pathol. 1984, 21, 418–424. [Google Scholar] [PubMed]
- Smith, E.E.; Kubena, L.F.; Braithwaite, C.E.; Harvey, R.B.; Phillips, T.D.; Reine, A.H. Toxicological evaluation of aflatoxin and cyclopiazonic acid in broiler chickens. Poult. Sci. 1992, 71, 1136–1144. [Google Scholar] [PubMed]
- Keblys, M.; Bernhoft, A.; Hofer, C.C.; Morrison, E.; Larsen, H.J.; Flaoyen, A. The effects of the Penicillium mycotoxins citrinin, cyclopiazonic acid, ochratoxin A, patulin, penicillic acid, and roquefortine C on in vitro proliferation of porcine lymphocytes. Mycopathologia 2004, 158, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Bernhoft, A.; Keblys, M.; Morrison, E.; Larsen, H.J.; Flaoyen, A. Combined effects of selected Penicillium mycotoxins on in vitro proliferation of porcine lymphocytes. Mycopathologia 2004, 158, 441–450. [Google Scholar] [PubMed]
- Gentles, A.; Smith, E.E.; Kubena, L.F.; Duffus, E.; Johnson, P.; Thompson, J.; Harvey, R.B.; Edrington, T.S. Toxicological evaluations of cyclopiazonic acid and ochratoxin A in broilers. Poult. Sci. 1999, 78, 1380–1384. [Google Scholar] [PubMed]
- Kubena, L.F.; Smith, E.E.; Gentles, A.; Harvey, R.B.; Edrington, T.S.; Phillips, T.D.; Rottinghaus, G.E. Individual and combined toxicity of T-2 toxin and cyclopiazonic acid in broiler chicks. Poult. Sci. 1994, 73, 1390–1397. [Google Scholar] [PubMed]
- Venkatesh, P.K.; Vairamuthu, S.; Balachandran, C.; Manohar, B.M.; Raj, G.D. Induction of apoptosis by fungal culture materials containing cyclopiazonic acid and T-2 toxin in primary lymphoid organs of broiler chickens. Mycopathologia 2005, 159, 393–400. [Google Scholar] [PubMed]
- Kamalavenkatesh, P.; Vairamuthu, S.; Balachandran, C.; Manohar, B.M.; Raj, G.D. Immunopathological effect of the mycotoxins cyclopiazonic acid and T-2 toxin on broiler chicken. Mycopathologia 2005, 159, 273–279. [Google Scholar] [PubMed]
- Antony, M.; Shukla, Y.; Janardhanan, K.K. Potential risk of acute hepatotoxicity of kodo poisoning due to exposure to cyclopiazonic acid. J. Ethnopharmacol. 2003, 87, 211–214. [Google Scholar] [PubMed]
- Bradburn, N.; Coker, R.D.; Blunden, G. The aetiology of turkey 'X' disease. Phytochemistry 1994, 35, 817. [Google Scholar]
- Spensley, P.C. Aflatoxin, the active principle in turkey 'X' disease. Endeavour 1963, 22, 75–79. [Google Scholar] [PubMed]
- Cole, R.J. Etiology of turkey "X" disease in retrospect: A case for the involvement of cyclopiazonic acid. Mycotoxin Res. 1986, 2, 3–7. [Google Scholar]
- Rao, L.B.; Husain, A. Presence of cyclopiazonic acid in kodo millet (Paspalum scrobiculatum) causing 'kodua poisoning' in man and its production by associated fungi. Mycopathologia 1985, 89, 177–180. [Google Scholar] [PubMed]
- Holzapfel, C.W.; Wilkins, D.C. On the biosynthesis of cyclopiazonic acid. Phytochemistry 1971, 10, 351–358. [Google Scholar]
- McGrath, R.M.; Steyn, P.S.; Ferreira, N.P.; Neethling, D.C. Biosynthesis of cyclopiazonic acids in Penicillium cyclopium: The Isolation of dimethylallylpyrophosphate: Cyclo-acetoacetyltryptophanyl dimethylallyltransferase. Biorg. Chem. 1976, 4, 11–23. [Google Scholar]
- Agurell, S.; Lindgren, J.E. Natural occurrence of 4-dimethylallyltryptophan-an ergot alkloid precursor. Tetrahedron Lett. 1968, 49, 5127–5128. [Google Scholar] [PubMed]
- Gebler, J.C.; Poulter, C.D. Purification and characterization of dimethylallyl tryptophan synthase from Clavicepsa purpurea. Arch. Biochem. Biophys. 1992, 296, 308–313. [Google Scholar] [PubMed]
- Plieninger, H.; Immel, H.; Volkl, A. Synthesis and incorporation of a 14C-and 3H-labeled 4-dimethylallyl-tryptophan and a 14C-labeled 4-dimethylallyl-tryptamine as well as incorporation of a 14C-labelled dimethylallyl-pyrophosphate. Justus Liebigs Ann. Chem. 1967, 706, 223–229. [Google Scholar] [PubMed]
- Robbers, J.E.; Floss, H.G. Biosynthesis of ergot alkaloids: Formation of 4-dimethylallyltryptophan by the ergot fungus. Arch. Biochem. Biophys. 1968, 126, 967–969. [Google Scholar] [PubMed]
- McGrath, R.M.; Steyn, P.S.; Ferreira, N.P. α-Acetyl-γ-(β-indolyl)methyltetramic acid. A biosynthetic intermediate of cyclopiazonic acid and of bis-secodehydrocyclopiazonic acid. J. Chem. Soc.,Chem. Commun. 1973, 21, 812–813. [Google Scholar]
- Heinstein, P.F.; Lee, S.I.; Floss, H.G. Isolation of dimethylallylpyrophosphate: Tryptophan dimethylallyl transferase from the ergot fungus (Claviceps spec.). Biochem. Biophys. Res. Commun. 1971, 44, 1244–1251. [Google Scholar] [CrossRef] [PubMed]
- McGrath, R.M.; Steyn, P.S.; Nourse, P.N.; Neethling, D.C.; Ferreira, N.P. The diversion of dimethylallyl pyrophosphate from polyisoprenoid to cyclopiazonic acid in Penicillium cyclopium Westling. Biorg. Chem. 1977, 6, 53–69. [Google Scholar]
- Steffan, N.; Grundmann, A.; Yin, W.B.; Kremer, A.; Li, S.M. Indole prenyltransferases from fungi: A new enzyme group with high potential for the production of prenylated indole derivatives. Curr. Med. Chem. 2009, 16, 218–231. [Google Scholar] [PubMed]
- Schabort, J.C.; Wilkens, D.C.; Holzapfel, C.W.; Potgieter, D.J.; Neitz, A.W. β-cyclopiazonate oxidocyclase from Penicillium cyclopium. I. Assay methods,isolation and purification. Biochim. Biophys. Acta 1971, 250, 311–328. [Google Scholar] [PubMed]
- Steenkamp, D.J.; Schabort, J.C.; Holzapfel, C.W.; Ferreira, N.P. The role of essential histidines in the mechanism of catalysis of the flavoenzyme, β-cyclopiazonate oxidocyclase. Biochim. Biophys. Acta 1974, 358, 126–143. [Google Scholar] [PubMed]
- Steenkamp, D.J.; Schabort, J.C.; Ferreira, N.P. β-cyclopiazonate oxidocyclase from Penicillium cyclopium. 3. Preliminary studies on the mechanism of action. Biochim. Biophys. Acta 1973, 309, 440–456. [Google Scholar] [PubMed]
- Wilding, E.I.; Brown, J.R.; Bryant, A.P.; Chalker, A.F.; Holmes, D.J.; Ingraham, K.A.; Iordanescu, S.; So, C.Y.; Rosenberg, M.; Gwynn, M.N. Identification, evolution, and essentiality of the mevalonate pathway for isopentenyl diphosphate biosynthesis in gram-positive cocci. J. Bacteriol. 2000, 182, 4319–4327. [Google Scholar] [PubMed]
- Andreassi, J.L., 2nd; Vetting, M.W.; Bilder, P.W.; Roderick, S.L.; Leyh, T.S. Structure of the ternary complex of phosphomevalonate kinase: The enzyme and its family. Biochemistry 2009, 48, 6461–6468. [Google Scholar] [PubMed]
- Durbecq, V.; Sainz, G.; Oudjama, Y.; Clantin, B.; Bompard-Gilles, C.; Tricot, C.; Caillet, J.; Stalon, V.; Droogmans, L.; Villeret, V. Crystal structure of isopentenyl diphosphate:dimethylallyl diphosphate isomerase. EMBO J. 2001, 20, 1530–1537. [Google Scholar] [PubMed]
- Machida, M.; Yamada, O.; Gomi, K. Genomics of Aspergillus oryzae: Learning from the history of koji mold and exploration of its future. DNA Res. 2008, 15, 173–183. [Google Scholar] [PubMed]
- Payne, G.A.; Nierman, W.C.; Wortman, J.R.; Pritchard, B.L.; Brown, D.; Dean, R.A.; Bhatnagar, D.; Cleveland, T.E.; Machida, M.; Yu, J. Whole genome comparison of Aspergillus flavus and A. oryzae. Med. Mycol. 2006, 44 Suppl, 9–11. [Google Scholar]
- Chang, P.-K.; Horn, B.W.; Dorner, J.W. Sequence breakpoints in the aflatoxin biosynthesis gene cluster and flanking regions in nonaflatoxigenic Aspergillus flavus isolates. Fungal Genet. Biol. 2005, 42, 914–923. [Google Scholar] [PubMed]
- Christensen, B.E.; Mollgaard, H.; Kaasgaard, S.; Lehmbeck, J. Methods for producing polypeptides in Aspergillus mutant cells. US Patent No. 6383781, 2002. [Google Scholar]
- Christensen, B.E.; Mollgaard, H.; Kaasgaard, S.; Lehmbeck, J. Methods for producing polypeptides in Aspergillus mutant cells. US Patent No. 7241614, 2007. [Google Scholar]
- Ehrlich, K.C.; Montalbano, B.G.; Cary, J.W. Binding of the C6-zinc cluster protein, AFLR, to the promoters of aflatoxin pathway biosynthesis genes in Aspergillus parasiticus. Gene 1999, 230, 249–257. [Google Scholar] [PubMed]
- Gulick, A.M. Conformational dynamics in the acyl-CoA synthetases, adenylation domains of non-ribosomal peptide synthetases, and firefly luciferase. ACS Chem. Biol. 2009, 4, 811–827. [Google Scholar] [CrossRef] [PubMed]
- Rausch, C.; Weber, T.; Kohlbacher, O.; Wohlleben, W.; Huson, D.H. Specificity prediction of adenylation domains in nonribosomal peptide synthetases (NRPS) using transductive support vector machines (TSVMs). Nucleic Acids Res. 2005, 33, 5799–5808. [Google Scholar] [PubMed]
- Stachelhaus, T.; Marahiel, M.A. Modular structure of peptide synthetases revealed by dissection of the multifunctional enzyme GrsA. J. Biol. Chem. 1995, 270, 6163–6169. [Google Scholar] [PubMed]
- Finking, R.; Marahiel, M.A. Biosynthesis of nonribosomal peptides1. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [PubMed]
- Fischbach, M.A.; Walsh, C.T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: Logic, machinery, and mechanisms. Chem. Rev. 2006, 106, 3468–3496. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Van Lanen, S.G.; Shen, B. A free-standing condensation enzyme catalyzing ester bond formation in C-1027 biosynthesis. Proc. Natl. Acad. Sci. U S A 2009, 106, 4183–4188. [Google Scholar] [PubMed]
- Liu, X.; Walsh, C.T. Cyclopiazonic acid biosynthesis in Aspergillus sp.: Characterization of a reductase-like R* domain in cyclopiazonate synthetase that forms and releases cyclo-acetoacetyl-L-tryptophan. Biochemistry 2009, 48, 8746–8757. [Google Scholar] [PubMed]
- Metzger, U.; Schall, C.; Zocher, G.; Unsold, I.; Stec, E.; Li, S.M.; Heide, L.; Stehle, T. The structure of dimethylallyl tryptophan synthase reveals a common architecture of aromatic prenyltransferases in fungi and bacteria. Proc. Natl. Acad. Sci. USA 2009, 106, 14309–14314. [Google Scholar]
- Kenney, W.C.; Edmondson, D.E.; Singer, T.P.; Steenkamp, D.J.; Schabort, J.C. The covalently bound flavin prosthetic group of β-cyclopiazonate oxidocyclase. FEBS Lett. 1974, 41, 111–114. [Google Scholar] [PubMed]
- Kenney, W.C.; Edmondson, D.E.; Singer, T.P.; Steenkamp, D.J.; Schabort, J.C. Identification and properties of the covalently bound flavin of β-cyclopiazonate oxidocyclase. Biochemistry 1976, 15, 4931–4935. [Google Scholar] [PubMed]
- Edmondson, D.E.; Binda, C.; Mattevi, A. Structural insights into the mechanism of amine oxidation by monoamine oxidases A and B. Arch. Biochem. Biophys. 2007, 464, 269–276. [Google Scholar] [PubMed]
- Edmondson, D.E.; Binda, C.; Wang, J.; Upadhyay, A.K.; Mattevi, A. Molecular and mechanistic properties of the membrane-bound mitochondrial monoamine oxidases. Biochemistry 2009, 48, 4220–4230. [Google Scholar] [PubMed]
- Fitzpatrick, P.F. Oxidation of amines by flavoproteins. Arch. Biochem. Biophys. 2009. [Google Scholar]
- Winkler, A.; Lyskowski, A.; Riedl, S.; Puhl, M.; Kutchan, T.M.; Macheroux, P.; Gruber, K. A concerted mechanism for berberine bridge enzyme. Nat. Chem. Biol. 2008, 4, 739–741. [Google Scholar] [PubMed]
- Winkler, A.; Motz, K.; Riedl, S.; Puhl, M.; Macheroux, P.; Gruber, K. Structural and mechanistic studies reveal the functional role of bicovalent flavinylation in berberine bridge enzyme. J. Biol. Chem. 2009, 284, 19993–20001. [Google Scholar] [PubMed]
- Winkler, A.; Puhl, M.; Weber, H.; Kutchan, T.M.; Gruber, K.; Macheroux, P. Berberine bridge enzyme catalyzes the six electron oxidation of (S)-reticuline to dehydroscoulerine. Phytochemistry 2009, 70, 1092–1097. [Google Scholar] [PubMed]
- Ehrlich, K.C.; Yu, J.; Cotty, P.J. Aflatoxin biosynthesis gene clusters and flanking regions. J. Appl. Microbiol. 2005, 99, 518–527. [Google Scholar] [PubMed]
- Tokuoka, M.; Takahashi, T.; Seshime, Y.; Fujii, I.; Kitamoto, K.; Koyama, Y. Identification of the biosynthetic pathway of cyclopiazonic acid in Aspergillus oryzae. 9th European Conference on Fungal Genetics; 2008. [Google Scholar]
- Chang, P.K.; Ehrlich, K.C.; Hua, S.S. Cladal relatedness among Aspergillus oryzae isolates and Aspergillus flavus S and L morphotype isolates. Int. J. Food Microbiol. 2006, 108, 172–177. [Google Scholar] [PubMed]
- Kusumoto, K.; Nogata, Y.; Ohta, H. Directed deletions in the aflatoxin biosynthesis gene homolog cluster of Aspergillus oryzae. Curr. Genet. 2000, 37, 104–111. [Google Scholar] [PubMed]
- Tominaga, M.; Lee, Y.H.; Hayashi, R.; Suzuki, Y.; Yamada, O.; Sakamoto, K.; Gotoh, K.; Akita, O. Molecular analysis of an inactive aflatoxin biosynthesis gene cluster in Aspergillus oryzae RIB strains. Appl. Environ. Microbiol. 2006, 72, 484–490. [Google Scholar] [PubMed]
- Riley, R.T.; Goeger, D.E. Cyclopiazonic acid: Speculation on its function on fungi. In Handbook of Applied Mycology: Mycotoxins in Ecological Systems; Deepak Bhatnagar, D., Lillehoj, E.B., Arora, D.K., Eds.; Marcel Dekker Inc.: New York, NY, USA, 1992; pp. 385–402. [Google Scholar]
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chang, P.-K.; Ehrlich, K.C.; Fujii, I. Cyclopiazonic Acid Biosynthesis of Aspergillus flavus and Aspergillus oryzae. Toxins 2009, 1, 74-99. https://doi.org/10.3390/toxins1020074
Chang P-K, Ehrlich KC, Fujii I. Cyclopiazonic Acid Biosynthesis of Aspergillus flavus and Aspergillus oryzae. Toxins. 2009; 1(2):74-99. https://doi.org/10.3390/toxins1020074
Chicago/Turabian StyleChang, Perng-Kuang, Kenneth C. Ehrlich, and Isao Fujii. 2009. "Cyclopiazonic Acid Biosynthesis of Aspergillus flavus and Aspergillus oryzae" Toxins 1, no. 2: 74-99. https://doi.org/10.3390/toxins1020074
APA StyleChang, P.-K., Ehrlich, K. C., & Fujii, I. (2009). Cyclopiazonic Acid Biosynthesis of Aspergillus flavus and Aspergillus oryzae. Toxins, 1(2), 74-99. https://doi.org/10.3390/toxins1020074




