Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins
Abstract
:1. Introduction
2. Materials and Methods
2.1. Milk Samples
2.2. Removal of Fat and Denatured Protein
2.3. Filter Aided Sample Preparation (FASP)
2.4. LC-MS/MS Analysis
2.5. Data Analysis
2.6. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Loss, G.; Apprich, S.; Waser, M.; Kneifel, W.; Genuneit, J.; Buchele, G.; Weber, J.; Sozanska, B.; Danielewicz, H.; Horak, E.; et al. The protective effect of farm milk consumption on childhood asthma and atopy: The gabriela study. J. Allergy Clin. Immunol. 2011, 128, 766–773. [Google Scholar] [CrossRef] [PubMed]
- Brick, T.; Schober, Y.; Bocking, C.; Pekkanen, J.; Genuneit, J.; Loss, G.; Dalphin, J.C.; Riedler, J.; Lauener, R.; Nockher, W.A.; et al. Omega-3 fatty acids contribute to the asthma-protective effect of unprocessed cow’s milk. J. Allergy Clin. Immunol. 2016, 137, 1699–1706. [Google Scholar] [CrossRef] [PubMed]
- Loss, G.; Depner, M.; Ulfman, L.H.; van Neerven, R.J.; Hose, A.J.; Genuneit, J.; Karvonen, A.M.; Hyvarinen, A.; Kaulek, V.; Roduit, C.; et al. Consumption of unprocessed cow’s milk protects infants from common respiratory infections. J. Allergy Clin. Immunol. 2015, 135, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Van Neerven, R.J.; Knol, E.F.; Heck, J.M.; Savelkoul, H.F. Which factors in raw cow’s milk contribute to protection against allergies? J. Allergy Clin. Immunol. 2012, 130, 853–858. [Google Scholar] [CrossRef] [PubMed]
- Li-Chan, E.; Kummer, A.; Losso, J.N.; Kitts, D.D.; Nakai, S. Stability of bovine immunoglobulins to thermal treatment and processing. Food Res. Int. 1995, 28, 9–16. [Google Scholar] [CrossRef]
- Van Gysel, M.; Cossey, V.; Fieuws, S.; Schuermans, A. Impact of pasteurization on the antibacterial properties of human milk. Eur. J. Pediatr. 2012, 171, 1231–1237. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Boeren, S.; Smits, M.; Hooijdonk, T.V.; Vervoort, J.; Hettinga, K. Proteomic study on the stability of proteins in bovine, camel, and caprine milk sera after processing. Food Res. Int. 2016, 82, 104–111. [Google Scholar] [CrossRef]
- Anema, S.G. Chapter 8—The whey proteins in milk: Thermal denaturation, physical interactions and effects on the functional properties of milk a2—Thompson, abby. In Milk Proteins; Boland, M., Singh, H., Eds.; Academic Press: San Diego, CA, USA, 2008; pp. 239–281. [Google Scholar]
- Navarro, F.; Harouna, S.; Calvo, M.; Perez, M.D.; Sanchez, L. Kinetic and thermodynamic parameters for thermal denaturation of ovine milk lactoferrin determined by its loss of immunoreactivity. J. Dairy Sci. 2015, 98, 4328–4337. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, T. Whey protein aggregation under shear conditions—Effects of lactose and heating temperature on aggregate size and structure. Int. J. Food Sci. Technol. 1999, 34, 523–531. [Google Scholar] [CrossRef]
- Van Boekel, M.A.J.S. Effect of heating on maillard reactions in milk. Food Chem. 1998, 62, 403–414. [Google Scholar] [CrossRef]
- Milkovska-Stamenova, S.; Hoffmann, R. Identification and quantification of bovine protein lactosylation sites in different milk products. J. Proteom. 2016, 134, 112–126. [Google Scholar] [CrossRef] [PubMed]
- Kirchner, B.; Pfaffl, M.W.; Dumpler, J.; von Mutius, E.; Ege, M.J. Microrna in native and processed cow’s milk and its implication for the farm milk effect on asthma. J. Allergy Clin. Immunol. 2016, 137, 1893–1895. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Boeren, S.; de Vries, S.C.; van Valenberg, H.J.; Vervoort, J.; Hettinga, K. Filter-aided sample preparation with dimethyl labeling to identify and quantify milk fat globule membrane proteins. J. Proteom. 2011, 75, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Wisniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. Maxquant enables high peptide identification rates, individualized p.P.B.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- UniProt, C. Universal Protein Resource (Uniprot). Avaliable online: http://www.uniprot.org/ (accessed on 31 August 2017).
- Schwanhausser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Corrigendum: Global quantification of mammalian gene expression control. Nature 2013, 495, 126–127. [Google Scholar] [CrossRef] [PubMed]
- The R Development Core Team. R: A Language and Environment for Statistical Computing. Avaliable online: https://www.R-project.org/ (accessed on 31 August 2017).
- Laleye, L.C.; Jobe, B.; Wasesa, A.A. Comparative study on heat stability and functionality of camel and bovine milk whey proteins. J. Dairy Sci. 2008, 91, 4527–4534. [Google Scholar] [CrossRef] [PubMed]
- Riedler, J.; Eder, W.; Oberfeld, G.; Schreuer, M. Austrian children living on a farm have less hay fever, asthma and allergic sensitization. Clin. Exp. Allergy 2000, 30, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.K.; Dalgleish, D.G. Interactions between milk serum proteins and synthetic fat globule membrane during heating of homogenized whole milk. J. Agric. Food Chem. 1993, 41, 1407–1412. [Google Scholar] [CrossRef]
- Waser, M.; Michels, K.B.; Bieli, C.; Floistrup, H.; Pershagen, G.; von Mutius, E.; Ege, M.; Riedler, J.; Schram-Bijkerk, D.; Brunekreef, B.; et al. Inverse association of farm milk consumption with asthma and allergy in rural and suburban populations across europe. Clin. Exp. Allergy 2007, 37, 661–670. [Google Scholar] [CrossRef] [PubMed]
- Corredig, M.; Dalgleish, D.G. Effect of temperature and ph on the interactions of whey proteins with casein micelles in skim milk. Food Res. Int. 1996, 29, 49–55. [Google Scholar] [CrossRef]
- Lee, S.J.; Sherbon, J.W. Chemical changes in bovine milk fat globule membrane caused by heat treatment and homogenization of whole milk. J. Dairy Res. 2002, 69, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Dewit, J.N.; Klarenbeek, G. Effects of various heat treatments on structure and solubility of whey proteins. J. Dairy Sci. 1984, 67, 2701–2710. [Google Scholar] [CrossRef]
- Troost, F.J.; Steijns, J.; Saris, W.H.; Brummer, R.J. Gastric digestion of bovine lactoferrin in vivo in adults. J. Nutr. 2001, 131, 2101–2104. [Google Scholar] [PubMed]
- Davidson, L.A.; Lonnerdal, B. Fecal alpha 1-antitrypsin in breast-fed infants is derived from human milk and is not indicative of enteric protein loss. Acta Paediatr. Scand. 1990, 79, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Jasion, V.S.; Burnett, B.P. Survival and digestibility of orally-administered immunoglobulin preparations containing igg through the gastrointestinal tract in humans. Nutr. J. 2015, 14, 22. [Google Scholar] [CrossRef] [PubMed]
- Giansanti, F.; Panella, G.; Leboffe, L.; Antonini, G. Lactoferrin from milk: Nutraceutical and pharmacological properties. Pharmaceuticals (Basel) 2016, 9, 61. [Google Scholar] [CrossRef] [PubMed]
- Drago-Serrano, M.E.; Campos-Rodriguez, R.; Carrero, J.C.; de la Garza, M. Lactoferrin: Balancing ups and downs of inflammation due to microbial infections. Int. J. Mol. Sci. 2017, 18, 501. [Google Scholar] [CrossRef] [PubMed]
- Kussendrager, K.D.; van Hooijdonk, A.C. Lactoperoxidase: Physico-chemical properties, occurrence, mechanism of action and applications. Br. J. Nutr. 2000, 84 (Suppl. 1), S19–S25. [Google Scholar] [CrossRef] [PubMed]
- Al Obaidi, A.H. Role of airway lactoperoxidase in scavenging of hydrogen peroxide damage in asthma. Ann. Thorac. Med. 2007, 2, 107–110. [Google Scholar] [CrossRef] [PubMed]
- Setiawan, H.; Nagaoka, K.; Kubo, M.; Fujikura, Y.; Ogino, K. Involvement of xanthine oxidoreductase-related oxidative stress in a dermatophagoides farinae-induced asthma model of nc/nga mice. Acta Med. Okayama 2016, 70, 175–182. [Google Scholar] [PubMed]
- Varga, L.; Farkas, H.; Fust, G. Role of complement in allergy. In The Complement System: Novel Roles in Health and Disease; Szebeni, J., Ed.; Springer: Boston, MA, USA, 2004; pp. 345–360. [Google Scholar]
- Zhang, X.; Kohl, J. A complex role for complement in allergic asthma. Exp. Rev. Clin. Immunol. 2010, 6, 269–277. [Google Scholar] [CrossRef]
- Drouin, S.M.; Corry, D.B.; Kildsgaard, J.; Wetsel, R.A. Cutting edge: The absence of c3 demonstrates a role for complement in th2 effector functions in a murine model of pulmonary allergy. J. Immunol. 2001, 167, 4141–4145. [Google Scholar] [CrossRef] [PubMed]
- Sklyarova, T.; van Hengel, J.; Van Wonterghem, E.; Libert, C.; van Roy, F.; Vandenbroucke, R.E. Hematopoietic plakophilin-3 regulates acute tissue-specific and systemic inflammation in mice. Eur. J. Immunol. 2015, 45, 2898–2910. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; He, H.; Day, A.J.; Tseng, S.C. Constitutive expression of inter-alpha-inhibitor (ialphai) family proteins and tumor necrosis factor-stimulated gene-6 (tsg-6) by human amniotic membrane epithelial and stromal cells supporting formation of the heavy chain-hyaluronan (hc-ha) complex. J. Biol. Chem. 2012, 287, 12433–12444. [Google Scholar] [CrossRef] [PubMed]
- Hettinga, K.A.; Reina, F.M.; Boeren, S.; Zhang, L.; Koppelman, G.H.; Postma, D.S.; Vervoort, J.J.; Wijga, A.H. Difference in the breast milk proteome between allergic and non-allergic mothers. PLoS ONE 2015, 10, e0122234. [Google Scholar] [CrossRef] [PubMed]
- Gregory, L.G.; Lloyd, C.M. Orchestrating house dust mite-associated allergy in the lung. Trends Immunol. 2011, 32, 402–411. [Google Scholar] [CrossRef] [PubMed]
- Kudo, M.; Khalifeh Soltani, S.M.; Sakuma, S.A.; McKleroy, W.; Lee, T.H.; Woodruff, P.G.; Lee, J.W.; Huang, K.; Chen, C.; Arjomandi, M.; et al. Mfge8 suppresses airway hyperresponsiveness in asthma by regulating smooth muscle contraction. Proc. Natl. Acad. Sci. USA 2013, 110, 660–665. [Google Scholar] [CrossRef] [PubMed]
- Hupin, C.; Rombaux, P.; Bowen, H.; Gould, H.; Lecocq, M.; Pilette, C. Downregulation of polymeric immunoglobulin receptor and secretory iga antibodies in eosinophilic upper airway diseases. Allergy 2013, 68, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Kaetzel, C.S. The polymeric immunoglobulin receptor: Bridging innate and adaptive immune responses at mucosal surfaces. Immunol. Rev. 2005, 206, 83–99. [Google Scholar] [CrossRef] [PubMed]
- Ege, M.J.; Strachan, D.P.; Cookson, W.O.; Moffatt, M.F.; Gut, I.; Lathrop, M.; Kabesch, M.; Genuneit, J.; Buchele, G.; Sozanska, B.; et al. Gene-environment interaction for childhood asthma and exposure to farming in central europe. J. Allergy Clin. Immunol. 2011, 127, 138–144. [Google Scholar] [CrossRef] [PubMed]
- Bieli, C.; Eder, W.; Frei, R.; Braun-Fahrlander, C.; Klimecki, W.; Waser, M.; Riedler, J.; von Mutius, E.; Scheynius, A.; Pershagen, G.; et al. A polymorphism in cd14 modifies the effect of farm milk consumption on allergic diseases and cd14 gene expression. J. Allergy Clin. Immunol. 2007, 120, 1308–1315. [Google Scholar] [CrossRef] [PubMed]
- Paulsson, M.A.; Svensson, U.; Kishore, A.R.; Naidu, A.S. Thermal behavior of bovine lactoferrin in water and its relation to bacterial interaction and antibacterial activity. J. Dairy Sci. 1993, 76, 3711–3720. [Google Scholar] [CrossRef]
- Marín, E.; Sánchez, L.; Pérez, M.D.; Puyol, P.; Calvo, M. Effect of heat treatment on bovine lactoperoxidase activity in skim milk: Kinetic and thermodynamic analysis. J. Food Sci. 2003, 68, 89–93. [Google Scholar] [CrossRef]
- Hettinga, K.; van Valenberg, H.; de Vries, S.; Boeren, S.; van Hooijdonk, T.; van Arendonk, J.; Vervoort, J. The host defense proteome of human and bovine milk. PLoS ONE 2011, 6, e19433. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Lara, N.R.; Escuder-Vieco, D.; Garcia-Algar, O.; De la Cruz, J.; Lora, D.; Pallas-Alonso, C. Effect of freezing time on macronutrients and energy content of breastmilk. Breastfeed. Med. 2012, 7, 295–301. [Google Scholar] [CrossRef] [PubMed]
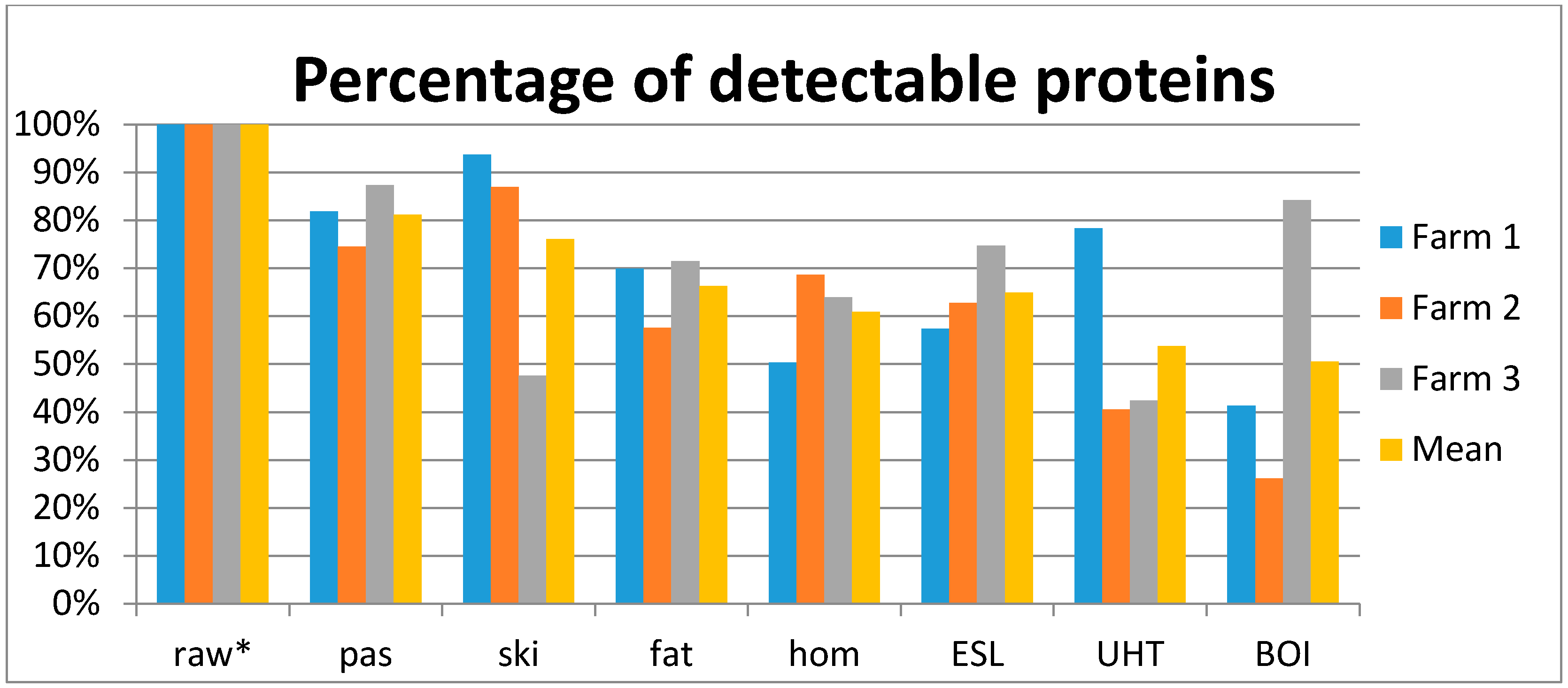
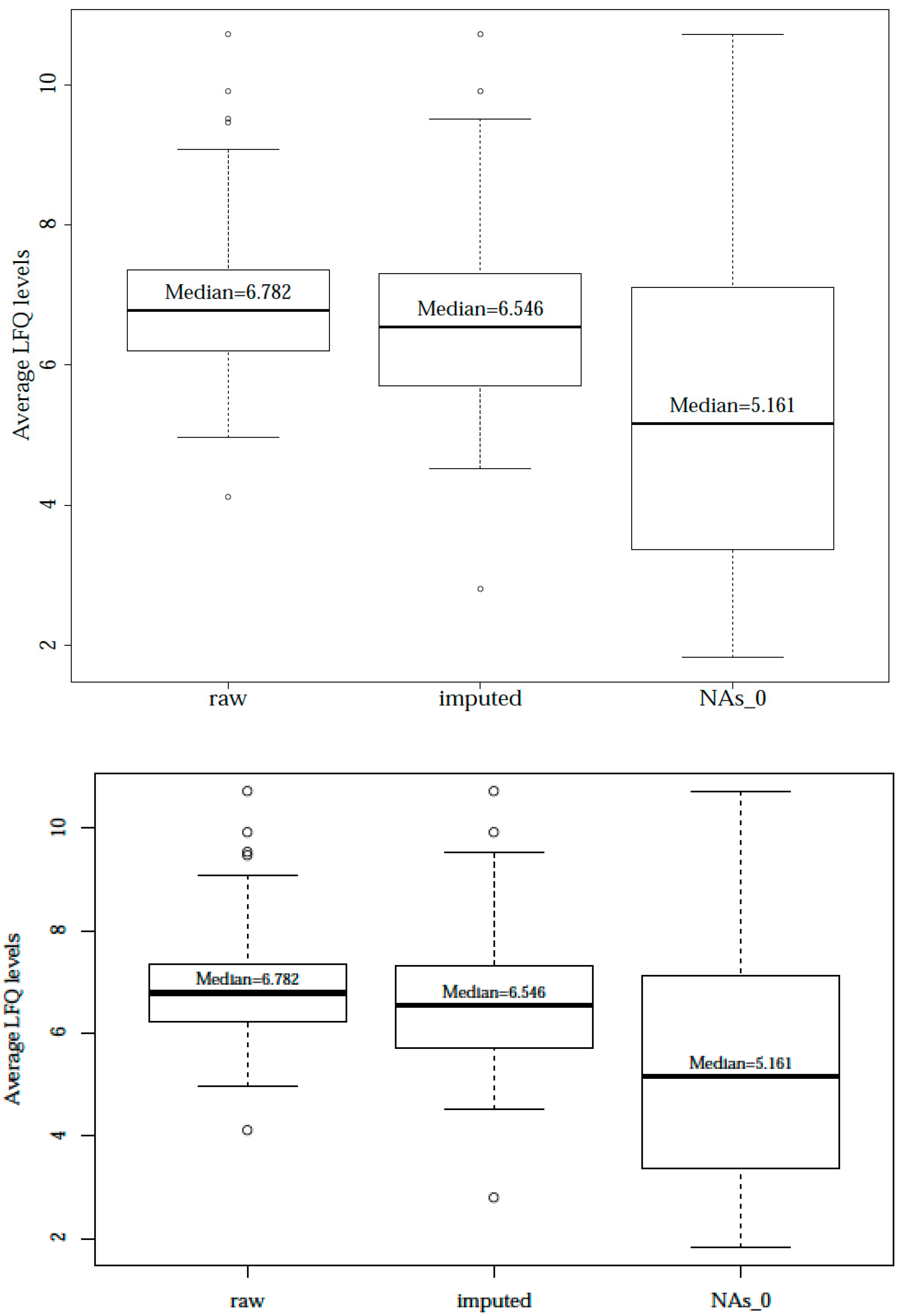
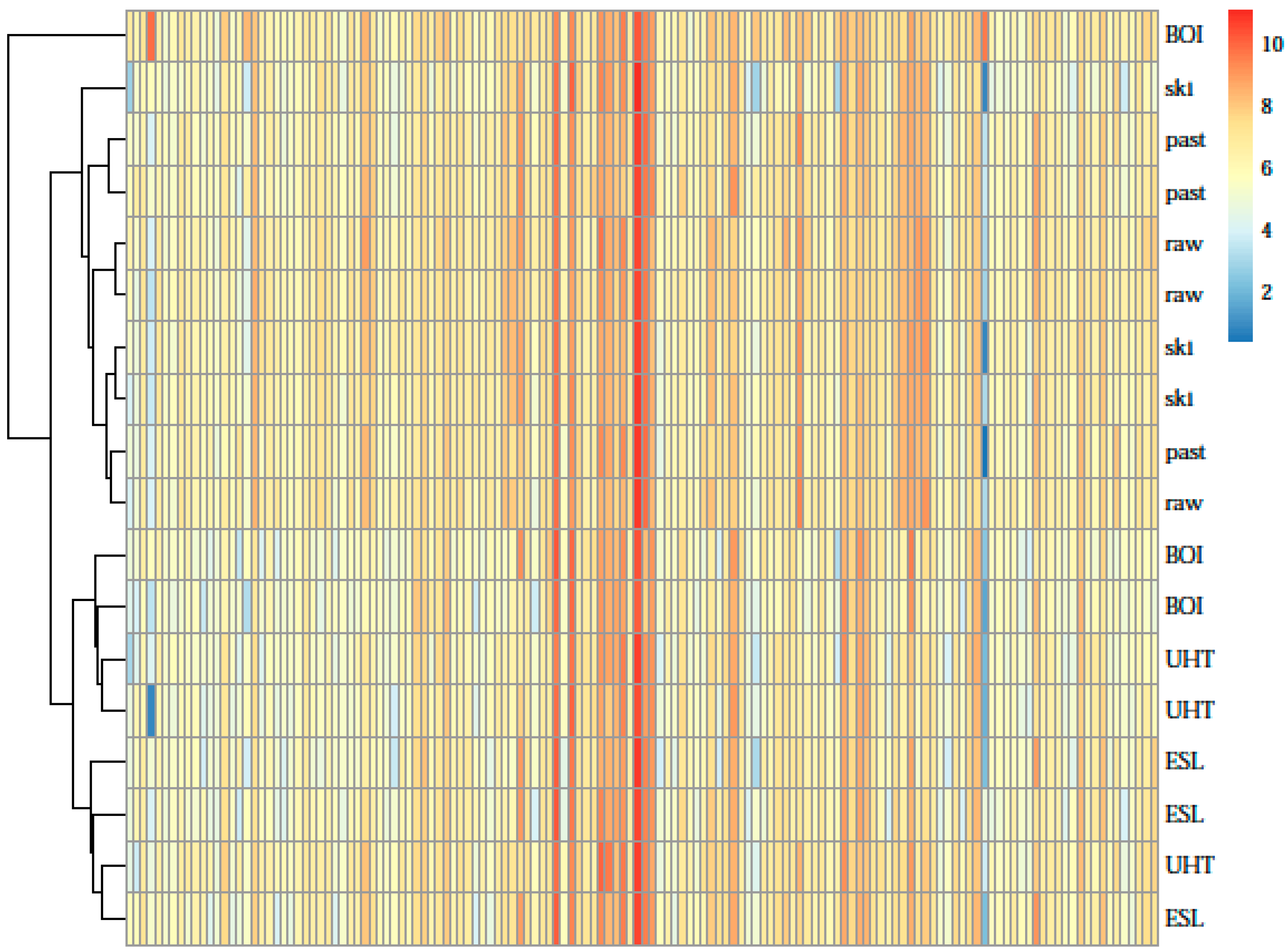
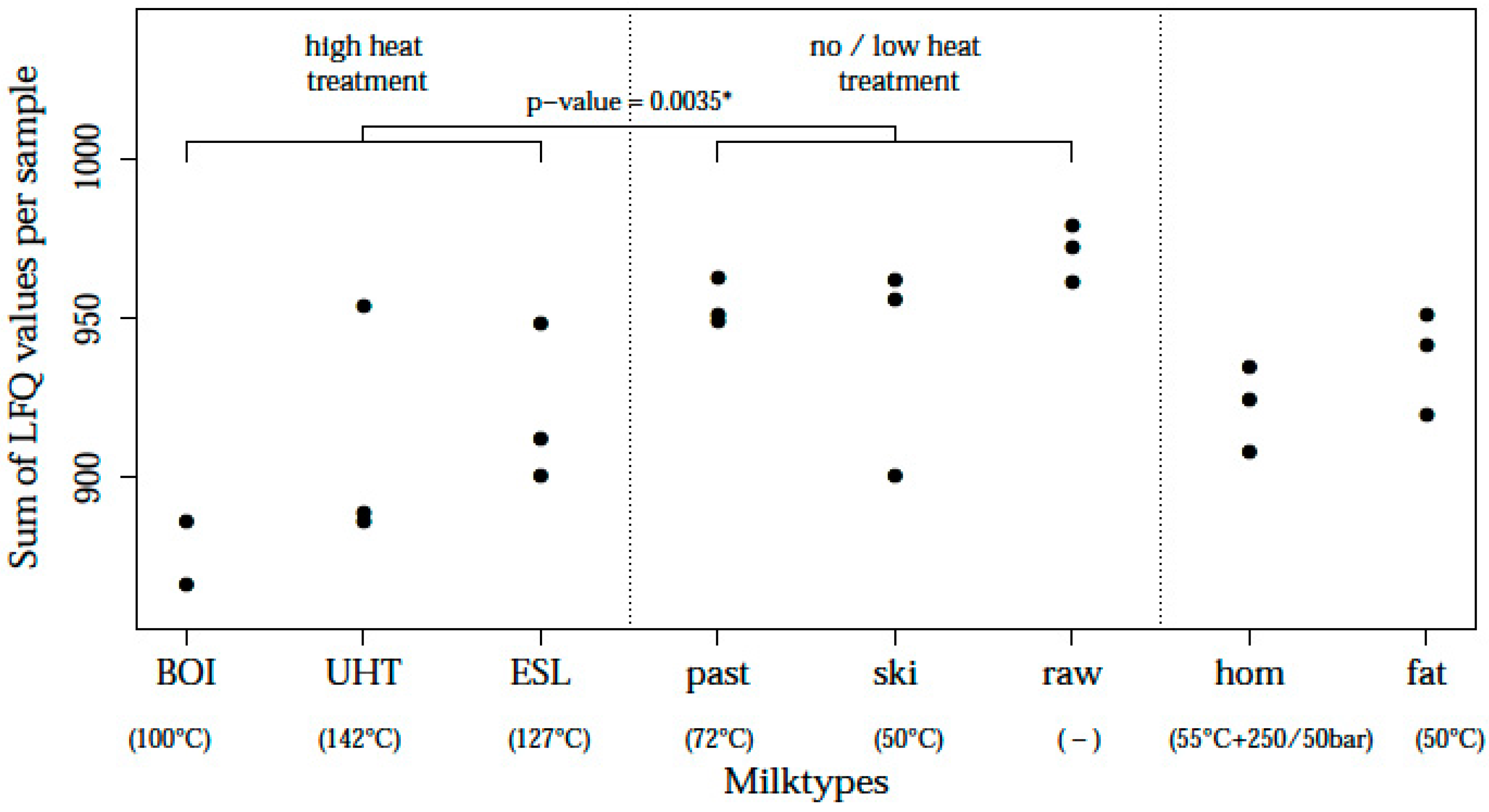

| Sample Origin | Farms (in Bavaria) | ||
|---|---|---|---|
| Traunstein | Freising | Starnberg | |
| No. of cows | 13 | 60 | 30 |
| Time point of milking for pooled samples | Morning and evening | Morning and evening | Morning and evening |
| No. of detectable milk serum proteins in raw milk samples | 143 | 153 | 158 |
| Code | Milk Fraction | Processing Conditions | Day of Processing * | Grouping of Milk Types ** |
|---|---|---|---|---|
| RAW | Native raw milk | - | Wednesday | No-low heat |
| PAS | Pasteurized | 72 °C for 20 s | Wednesday | No-low heat |
| Total processing time *** 60 s | ||||
| SKI | Skim milk | Separation at 50 °C | Tuesday | No-low heat |
| FAT | Fat fraction/cream | Separation at 50 °C | Tuesday | - |
| HOM | Homogenized milk | Preheating to 55 °C, 2-stage homogenization at 250/50 bar | Tuesday | - |
| ESL | Extended shelf life milk | Preheating at 95 °C for 20 s, direct steam injection at 127 °C for 5 s | Monday | High heat |
| Total processing time *** 60 s | ||||
| UHT | Ultra-high heat treated | Preheating at 93 °C for 23 s, direct steam injection at 142 °C for 5 s | Monday | High heat |
| Total processing time *** 85 s | ||||
| BOI | Boiled milk | Preheating at >80 °C for >300 s, boiling at 100 °C for 30 s | Tuesday | High heat |
| Total processing time *** 2000 s |
| Protein Code | Number of Peptides | p-Value * | Log2 Fold Change (95% CI) | Protein Name | Protein Function |
|---|---|---|---|---|---|
| P80457 | 67 | 0.001 | −0.44 (−0.56; −0.31) | Xanthine dehydrogenase/oxidase | immunity |
| P24627 | 71 | 0.004 | −0.37 (−0.51; −0.22) | Lactoferrin | immunity |
| G3X6N3 | 57 | 0.006 | −0.35 (−0.50; −0.20) | Serotransferrin | transport |
| F1MR22 | 42 | 0.004 | −0.34 (−0.47; −0.21) | Polymeric immunoglobulin receptor | immunity |
| P80025 | 37 | 0.001 | −0.33 (−0.43; −0.23) | Lactoperoxidase | immunity |
| G3N1R1 | 4 | 0.002 | −0.32 (−0.44; −0.21) | Uncharacterized protein | unknown |
| F1MGU7 | 7 | 0.04 | −0.30 (−0.52; −0.07) | Fibrinogen gamma-B chain | Blood coagulation |
| G3X7A5 | 80 | 0.002 | −0.29 (−0.41; −0.18) | Complement C3 | immunity |
| F1MZ96 | 10 | 0.002 | −0.27 (−0.36; −0.18) | Uncharacterized protein | unknown |
| F1MX50 | 4 | 0.01 | −0.27 (−0.40; −0.13) | Uncharacterized protein | cell |
| F1MM32 | 8 | 0.026 | −0.26 (−0.43; −0.08) | Sulfhydryl oxidase | enzyme |
| P81265 | 42 | 0.006 | −0.24 (−0.35; −0.14) | Polymeric immunoglobulin receptor | immunity |
| F1N076 | 12 | 0.001 | −0.23 (−0.30; −0.15) | Ceruloplasmin | cell |
| F1MXX6 | 26 | 0.02 | −0.22 (−0.35; −0.08) | Lactadherin | cell |
| Q08DQ0 | 6 | 0.017 | −0.21 (−0.34; −0.08) | Plakophilin-3 | cell |
| P07589 | 6 | 0.004 | −0.20 (−0.30; −0.11) | Fibronectin | immunity |
| A6QNL0 | 6 | 0.01 | −0.20 (−0.32; −0.09) | Monocyte differentiation antigen CD 14 | immunity |
| P10152 | 11 | 0.048 | −0.20 (−0.37; −0.04) | Angiogenin-1 (ribonuclease 5) | cell |
| F1MMD7 | 5 | 0.031 | −0.20 (−0.34; −0.06) | Inter-alpha-trypsin inhibitor heavy chain H4 | Protease inhibitor |
| Q3MHN2 | 6 | 0.043 | −0.20 (−0.35; −0.04) | Complement component C9 | immunity |
| P00735 | 7 | 0.028 | −0.18 (−0.30; −0.05) | Prothrombin | immunity |
| F1MCF8 | 9 | 0.001 | −0.17 (−0.22; −0.12) | Uncharacterized protein | immunity |
| P17690 | 9 | 0.005 | −0.16 (−0.23; −0.09) | Beta-2-glycoprotein 1 | Blood coagulation |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brick, T.; Ege, M.; Boeren, S.; Böck, A.; Von Mutius, E.; Vervoort, J.; Hettinga, K. Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins. Nutrients 2017, 9, 963. https://doi.org/10.3390/nu9090963
Brick T, Ege M, Boeren S, Böck A, Von Mutius E, Vervoort J, Hettinga K. Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins. Nutrients. 2017; 9(9):963. https://doi.org/10.3390/nu9090963
Chicago/Turabian StyleBrick, Tabea, Markus Ege, Sjef Boeren, Andreas Böck, Erika Von Mutius, Jacques Vervoort, and Kasper Hettinga. 2017. "Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins" Nutrients 9, no. 9: 963. https://doi.org/10.3390/nu9090963
APA StyleBrick, T., Ege, M., Boeren, S., Böck, A., Von Mutius, E., Vervoort, J., & Hettinga, K. (2017). Effect of Processing Intensity on Immunologically Active Bovine Milk Serum Proteins. Nutrients, 9(9), 963. https://doi.org/10.3390/nu9090963





