Maternal Prenatal Folic Acid Supplementation Programs Offspring Lipid Metabolism by Aberrant DNA Methylation in Hepatic ATGL and Adipose LPL in Rats
Abstract
1. Introduction
2. Materials and Methods
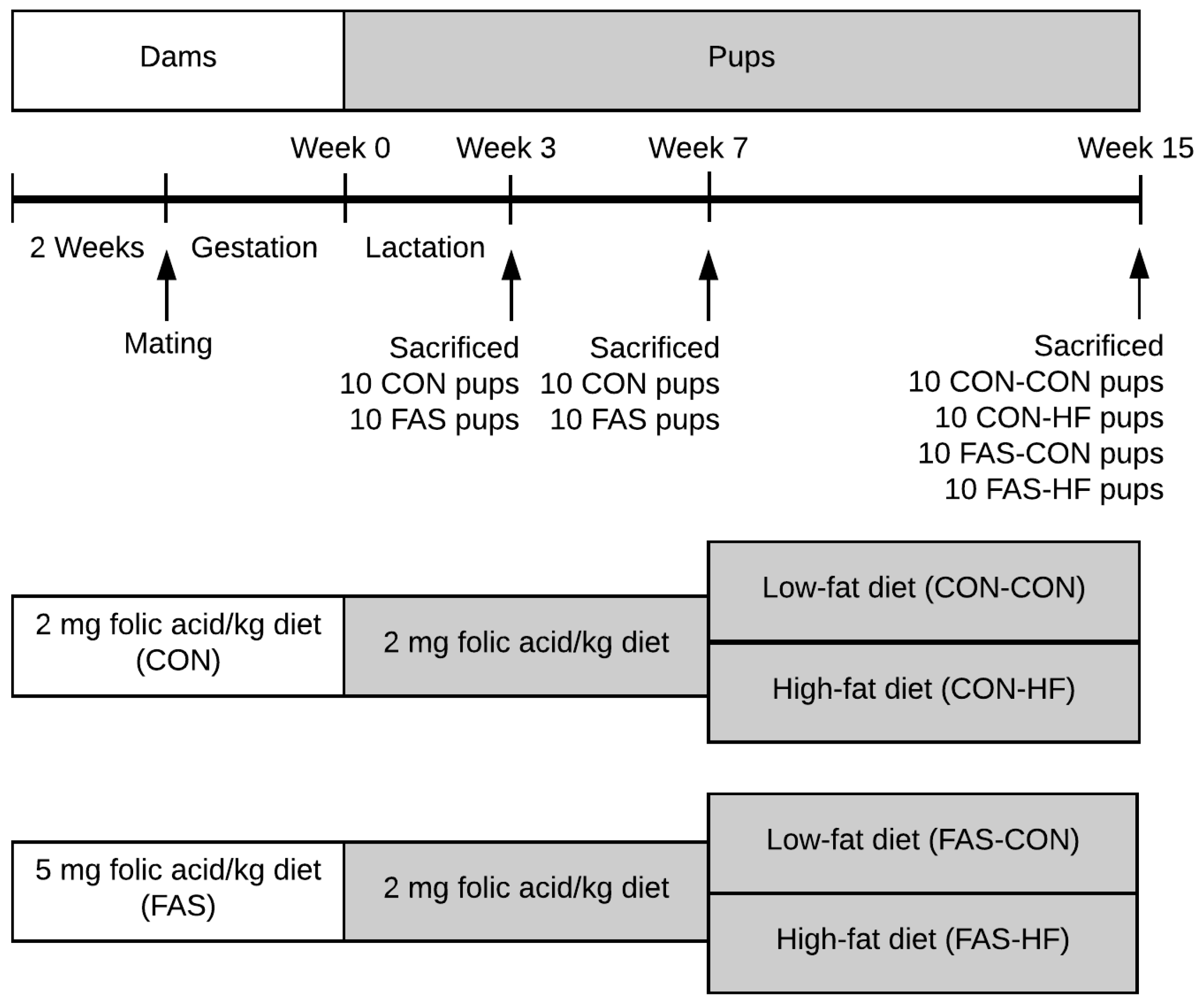
2.1. Folic Acid Doses Selection and Diets
2.2. Animal Experimentation
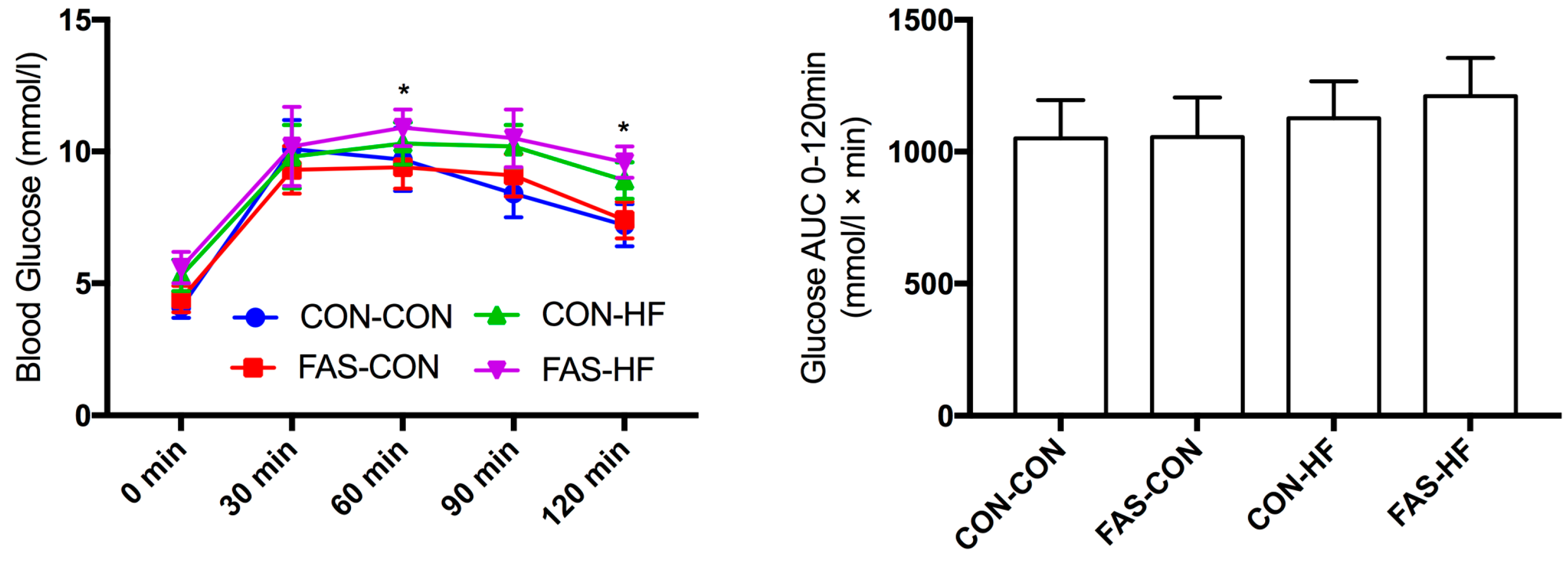
2.3. Oral Glucose Tolerance Test (OGTT)
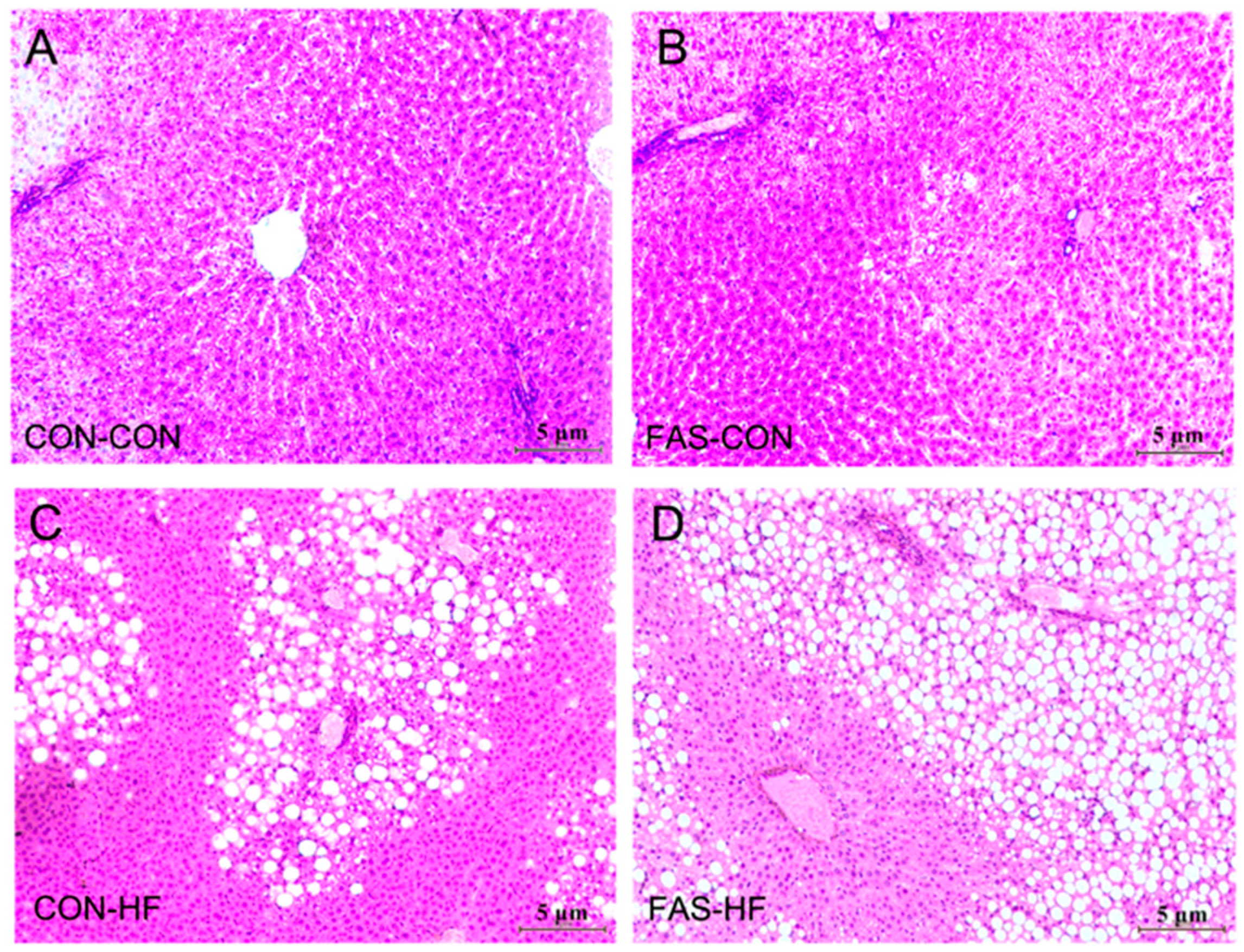
2.4. Liver Tissue Triglycerides and Histology and Blood Biochemical Measurements
2.5. Quantitative Real-Time PCR and Western Blotting
2.6. DNA Methylation Determination
2.7. Statistical Analysis
3. Results
3.1. Maternal and Offspring Food Intake and Serum Folic Acid Levels
3.2. Offspring Body Weight, Glucose and Lipid Metabolism at 15 Weeks
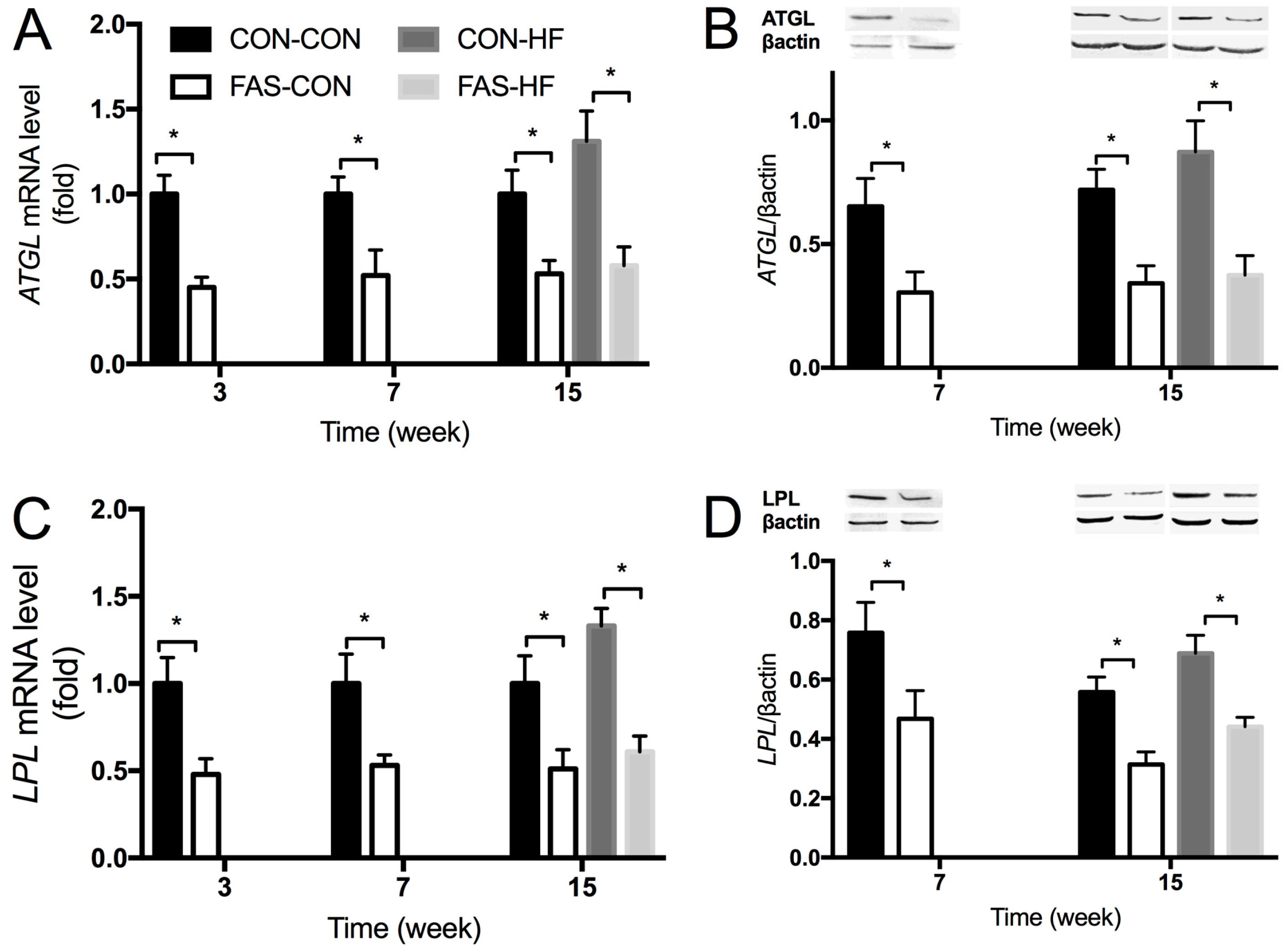
3.3. Expression of Lipid Metabolic Related Genes in Liver and White Adipose Tissue
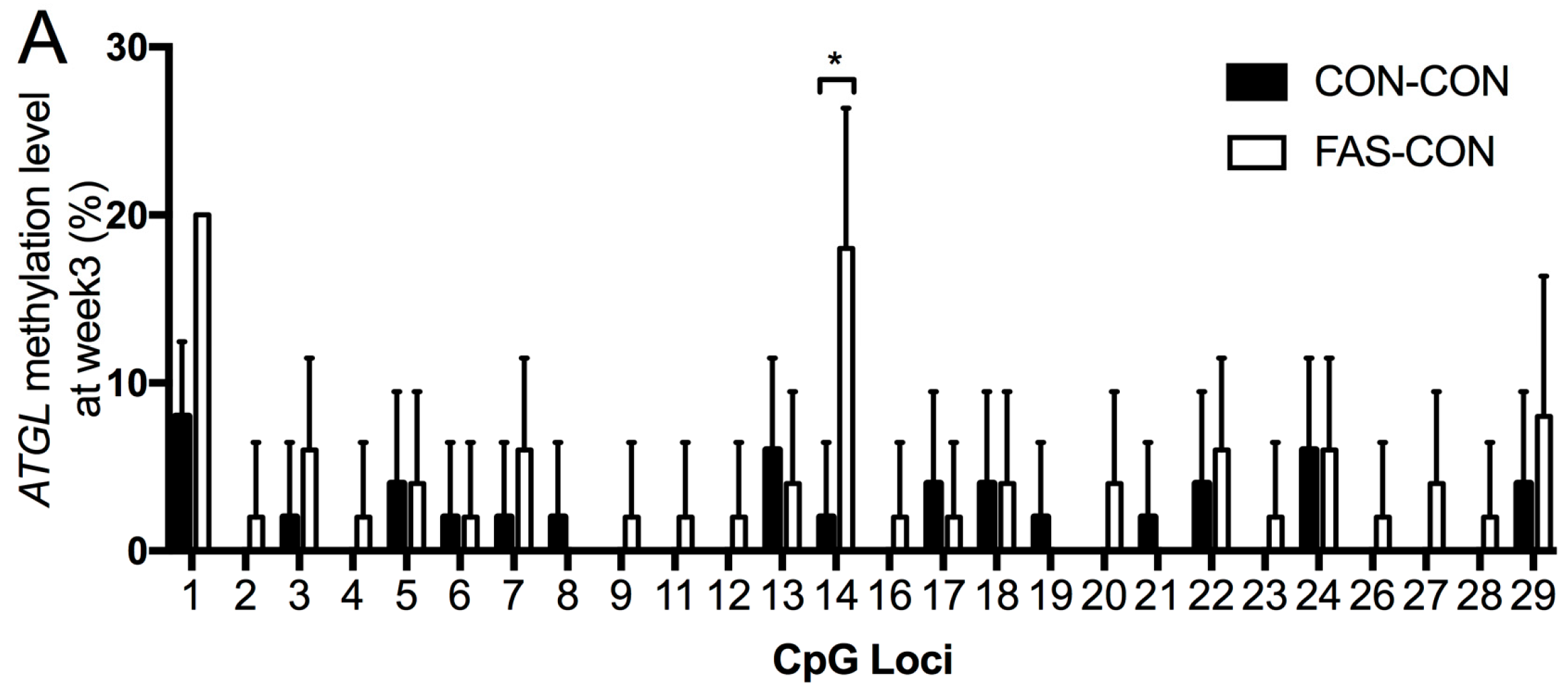
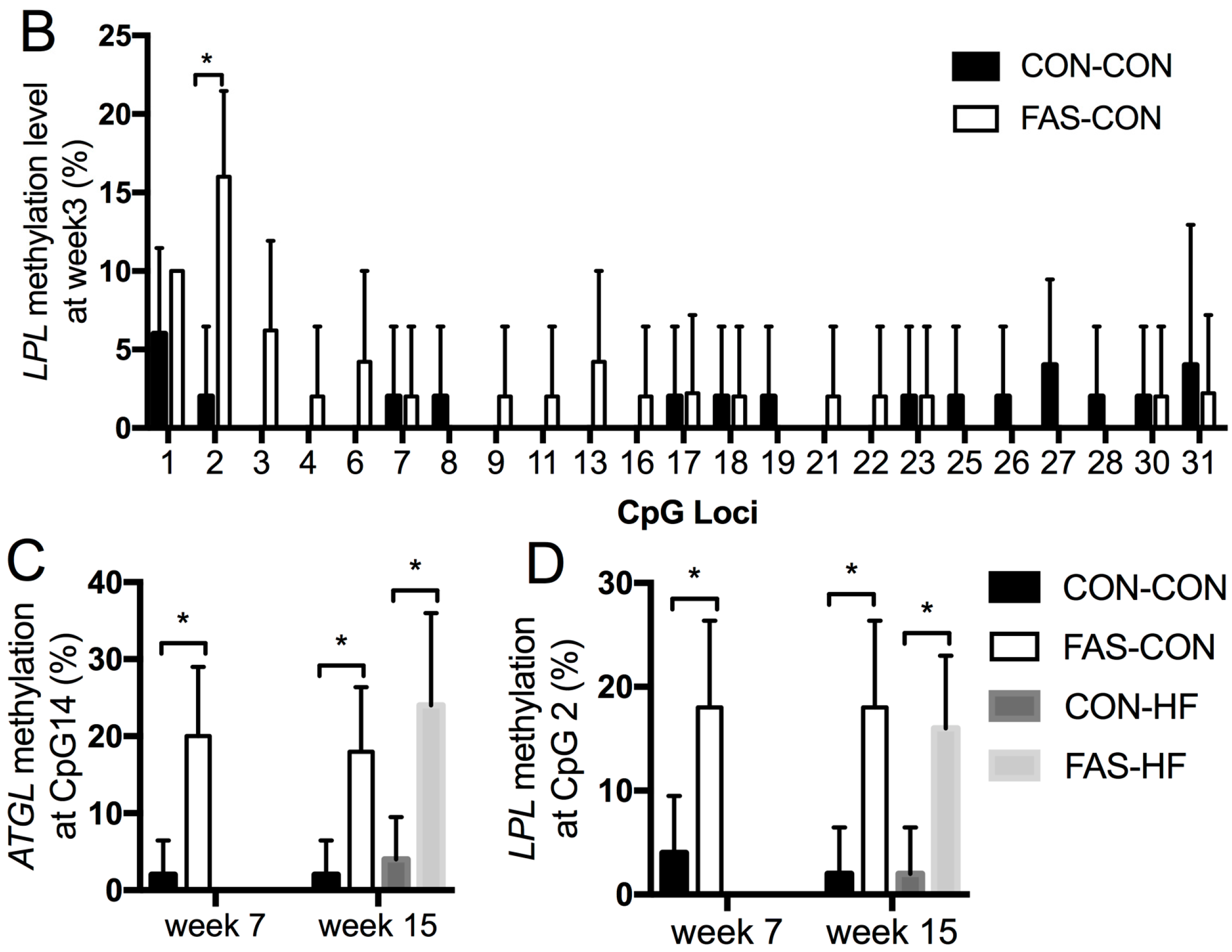
3.4. DNA Methylation of ATGL and LPL
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gluckman, P.D.; Hanson, M.A.; Cooper, C.; Thornburg, K.L. Effect of in utero and early-life conditions on adult health and disease. N. Engl. J. Med. 2008, 359, 61–73. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Jaddoe, V.W.; Qi, L.; He, Y.; Wang, D.; Lai, J.; Zhang, J.; Fu, P.; Yang, X.; Hu, F.B. Exposure to the chinese famine in early life and the risk of metabolic syndrome in adulthood. Diabetes Care 2011, 34, 1014–1018. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, S.; Li, S.; Feng, R.; Na, L.; Chu, X.; Wu, X.; Niu, Y.; Sun, Z.; Han, T.; et al. Prenatal exposure to famine and the development of hyperglycemia and type 2 diabetes in adulthood across consecutive generations: A population-based cohort study of families in Suihua, China. Am. J. Clin. Nutr. 2017, 105, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Erhuma, A.; Bellinger, L.; Langley-Evans, S.C.; Bennett, A.J. Prenatal exposure to undernutrition and programming of responses to high-fat feeding in the rat. Br. J. Nutr. 2007, 98, 517–524. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Crider, K.S.; Yang, T.P.; Berry, R.J.; Bailey, L.B. Folate and DNA methylation: A review of molecular mechanisms and the evidence for folate’s role. Adv. Nutr. 2012, 3, 21–38. [Google Scholar] [CrossRef] [PubMed]
- Waterland, R.A.; Jirtle, R.L. Transposable elements: Targets for early nutritional effects on epigenetic gene regulation. Mol. Cell. Biol. 2003, 23, 5293–5300. [Google Scholar] [CrossRef] [PubMed]
- Waterland, R.A.; Dolinoy, D.C.; Lin, J.R.; Smith, C.A.; Shi, X.; Tahiliani, K.G. Maternal methyl supplements increase offspring DNA methylation at axin fused. Genesis 2006, 44, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Berry, R.J.; Li, Z.; Erickson, J.D.; Li, S.; Moore, C.A.; Wang, H.; Mulinare, J.; Zhao, P.; Wong, L.Y.; Gindler, J.; et al. Prevention of neural-tube defects with folic acid in china. N. Engl. J. Med. 1999, 341, 1485–1490. [Google Scholar] [CrossRef] [PubMed]
- Lumley, J.; Watson, L.; Watson, M.; Bower, C. Periconceptional supplementation with folate and/or multivitamins for preventing neural tube defects. Cochrane Database Syst. Rev. 2001, CD001056. [Google Scholar] [CrossRef]
- Krishnaveni, G.V.; Veena, S.R.; Karat, S.C.; Yajnik, C.S.; Fall, C.H. Association between maternal folate concentrations during pregnancy and insulin resistance in indian children. Diabetologia 2014, 57, 110–121. [Google Scholar] [CrossRef] [PubMed]
- Yajnik, C.S.; Deshpande, S.S.; Jackson, A.A.; Refsum, H.; Rao, S.; Fisher, D.J.; Bhat, D.S.; Naik, S.S.; Coyaji, K.J.; Joglekar, C.V.; et al. Vitamin B12 and folate concentrations during pregnancy and insulin resistance in the offspring: The pune maternal nutrition study. Diabetologia 2008, 51, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Schaible, T.D.; Harris, R.A.; Dowd, S.E.; Smith, C.W.; Kellermayer, R. Maternal methyl-donor supplementation induces prolonged murine offspring colitis susceptibility in association with mucosal epigenetic and microbiomic changes. Hum. Mol. Genet. 2011, 20, 1687–1696. [Google Scholar] [CrossRef] [PubMed]
- Hollingsworth, J.W.; Maruoka, S.; Boon, K.; Garantziotis, S.; Li, Z.; Tomfohr, J.; Bailey, N.; Potts, E.N.; Whitehead, G.; Brass, D.M.; et al. In utero supplementation with methyl donors enhances allergic airway disease in mice. J. Clin. Investig. 2008, 118, 3462–3469. [Google Scholar] [CrossRef] [PubMed]
- Yates, A.A.; Schlicker, S.A.; Suitor, C.W. Dietary reference intakes: The new basis for recommendations for calcium and related nutrients, B vitamins, and choline. J. Am. Diet. Assoc. 1998, 98, 699–706. [Google Scholar] [CrossRef]
- Reeves, P.G. Components of the AIN-93 diets as improvements in the AIN-76A diet. J. Nutr. 1997, 127, 838S–841S. [Google Scholar] [PubMed]
- Folch, J.; Lees, M.; Sloane Stanley, G.H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−delta delta c(t)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Lu, N.; Li, Y.; Qin, H.; Zhang, Y.L.; Sun, C.H. Gossypin up-regulates ldl receptor through activation of erk pathway: A signaling mechanism for the hypocholesterolemic effect. J. Agric. Food Chem. 2008, 56, 11526–11532. [Google Scholar] [CrossRef] [PubMed]
- Chmurzynska, A.; Stachowiak, M.; Gawecki, J.; Pruszynska-Oszmalek, E.; Tubacka, M. Protein and folic acid content in the maternal diet determine lipid metabolism and response to high-fat feeding in rat progeny in an age-dependent manner. Genes Nutr. 2012, 7, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Cho, C.E.; Sanchez-Hernandez, D.; Reza-Lopez, S.A.; Huot, P.S.; Kim, Y.I.; Anderson, G.H. High folate gestational and post-weaning diets alter hypothalamic feeding pathways by DNA methylation in wistar rat offspring. Epigenetics 2013, 8, 710–719. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; He, Y.; Sun, X.; He, Y.; Li, Y.; Sun, C. Maternal high folic acid supplement promotes glucose intolerance and insulin resistance in male mouse offspring fed a high-fat diet. Int. J. Mol. Sci 2014, 15, 6298–6313. [Google Scholar] [CrossRef] [PubMed]
- Mierzecki, A.; Kloda, K.; Bukowska, H.; Chelstowski, K.; Makarewicz-Wujec, M.; Kozlowska-Wojciechowska, M. Association between low-dose folic acid supplementation and blood lipids concentrations in male and female subjects with atherosclerosis risk factors. Med. Sci. Monit. 2013, 19, 733–739. [Google Scholar] [PubMed]
- Lewis, S.J.; Leary, S.; Davey Smith, G.; Ness, A. Body composition at age 9 years, maternal folate intake during pregnancy and methyltetrahydrofolate reductase (MTHFR) C677T genotype. Br. J. Nutr. 2009, 102, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Burdge, G.C.; Lillycrop, K.A.; Phillips, E.S.; Slater-Jefferies, J.L.; Jackson, A.A.; Hanson, M.A. Folic acid supplementation during the juvenile-pubertal period in rats modifies the phenotype and epigenotype induced by prenatal nutrition. J. Nutr. 2009, 139, 1054–1060. [Google Scholar] [CrossRef] [PubMed]
- Hoile, S.P.; Lillycrop, K.A.; Grenfell, L.R.; Hanson, M.A.; Burdge, G.C. Increasing the folic acid content of maternal or post-weaning diets induces differential changes in phosphoenolpyruvate carboxykinase mrna expression and promoter methylation in rats. Br. J. Nutr. 2012, 108, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, R.; Strauss, J.G.; Haemmerle, G.; Schoiswohl, G.; Birner-Gruenberger, R.; Riederer, M.; Lass, A.; Neuberger, G.; Eisenhaber, F.; Hermetter, A.; et al. Fat mobilization in adipose tissue is promoted by adipose triglyceride lipase. Science 2004, 306, 1383–1386. [Google Scholar] [CrossRef] [PubMed]
- Ong, K.T.; Mashek, M.T.; Bu, S.Y.; Greenberg, A.S.; Mashek, D.G. Adipose triglyceride lipase is a major hepatic lipase that regulates triacylglycerol turnover and fatty acid signaling and partitioning. Hepatology 2011, 53, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Zechner, R.; Strauss, J.; Frank, S.; Wagner, E.; Hofmann, W.; Kratky, D.; Hiden, M.; Levak-Frank, S. The role of lipoprotein lipase in adipose tissue development and metabolism. Int. J. Obes. Relat. Metab. Disord. 2000, 24 (Suppl. 4), S53–S56. [Google Scholar] [CrossRef] [PubMed]
- Pugni, L.; Riva, E.; Pietrasanta, C.; Rabacchi, C.; Bertolini, S.; Pederiva, C.; Mosca, F.; Calandra, S. Severe hypertriglyceridemia in a newborn with monogenic lipoprotein lipase deficiency: An unconventional therapeutic approach with exchange transfusion. JIMD Rep. 2014, 13, 59–64. [Google Scholar] [PubMed]
- Salbaum, J.M.; Kappen, C. Genetic and epigenomic footprints of folate. Prog. Mol. Biol. Transl. Sci. 2012, 108, 129–158. [Google Scholar] [PubMed]
- Ly, A.; Ishiguro, L.; Kim, D.; Im, D.; Kim, S.E.; Sohn, K.J.; Croxford, R.; Kim, Y.I. Maternal folic acid supplementation modulates DNA methylation and gene expression in the rat offspring in a gestation period-dependent and organ-specific manner. J. Nutr. Biochem. 2016, 33, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Joubert, B.R.; den Dekker, H.T.; Felix, J.F.; Bohlin, J.; Ligthart, S.; Beckett, E.; Tiemeier, H.; van Meurs, J.B.; Uitterlinden, A.G.; Hofman, A.; et al. Maternal plasma folate impacts differential DNA methylation in an epigenome-wide meta-analysis of newborns. Nat. Commun. 2016, 7, 10577. [Google Scholar] [CrossRef] [PubMed]
- Gueant, J.L.; Namour, F.; Gueant-Rodriguez, R.M.; Daval, J.L. Folate and fetal programming: A play in epigenomics? Trends Endocrinol. Metab. 2013, 24, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Jirtle, R.L.; Skinner, M.K. Environmental epigenomics and disease susceptibility. Nat. Rev. Genet. 2007, 8, 253–262. [Google Scholar] [CrossRef] [PubMed]
- McKay, J.A.; Xie, L.; Adriaens, M.; Evelo, C.T.; Ford, D.; Mathers, J.C. Maternal folate depletion during early development and high fat feeding from weaning elicit similar changes in gene expression, but not in DNA methylation, in adult offspring. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
| CON-CON | FAS-CON | CON-HF | FAS-HF | p1 | p2 | p3 | |
|---|---|---|---|---|---|---|---|
| Body Weight, g | 460.3 ± 23.8 | 468.2 ± 23.6 | 565.4 ± 24.3 | 583.7 ± 24.9 | 0.48 | <0.001 | 0.5 |
| Fasting glucose, mmol/L | 4.62 ± 0.81 | 4.52 ± 0.54 | 4.93 ± 0.79 | 5.62 ± 0.76 | 0.17 | 0.002 | 0.07 |
| Total cholesterol, mmol/L | 1.33 ± 0.66 | 1.41 ± 0.48 | 1.92 ± 0.74 | 2.20 ± 0.38 | 0.29 | <0.001 | 0.55 |
| Triglyceride, mmol/L | 0.71 ± 0.09 | 0.84 ± 0.12 | 1.24 ± 0.37 | 1.92 ± 0.27 | <0.001 | <0.001 | 0.001 |
| High density lipoprotein cholesterol, mmol/L | 0.82 ± 0.07 | 0.82 ± 0.06 | 0.92 ± 0.13 | 0.96 ± 0.08 | 0.59 | <0.001 | 0.45 |
| Low density lipoprotein cholesterol, mmol/L | 0.31 ± 0.05 | 0.32 ± 0.06 | 0.47 ± 0.10 | 0.53 ± 0.05 | 0.11 | <0.001 | 0.26 |
| Alanine aminotransferase, IU/L | 34.74 ± 4.23 | 34.61 ± 3.55 | 70.49 ± 7.83 | 77.84 ± 7.87 | 0.06 | <0.001 | 0.06 |
| Aspartate aminotransferase, IU/L | 128.02 ± 15.26 | 127.34 ± 13.34 | 148.03 ± 19.29 | 155.67 ± 15.43 | 0.46 | <0.001 | 0.38 |
| Liver body weight ratio, % | 2.36 ± 0.15 | 2.39 ± 0.15 | 3.17 ± 0.25 | 3.47 ± 0.27 | 0.02 | <0.001 | 0.04 |
| Liver triglyceride, mmol/g | 39.07 ± 6.11 | 52.60 ± 7.36 | 85.51 ± 15.81 | 137.98 ± 16.63 | <0.001 | <0.001 | <0.001 |
| Genes | Fold Changes in Liver | Fold Changes in White Adipose Tissues |
|---|---|---|
| Acetyl-CoA carboxylase | 1.06 ± 0.09 | 1.35 ± 0.15 |
| Acyl-CoA oxidase 1 | 1.29 ± 0.15 | 0.93 ± 0.09 |
| Adiponectin | 1.32 ± 0.12 | 1.07 ± 0.09 |
| Adiponectin receptor | 0.82 ± 0.07 | 1.15 ± 0.1 |
| AMP-activated protein kinase 1 | 0.93 ± 0.09 | 0.86 ± 0.09 |
| AMP-activated protein kinase 2 | 1.13 ± 0.12 | 1.45 ± 0.12 |
| Adipose triglyceride lipase | 0.45 ± 0.06 * | 0.91 ± 0.10 |
| Cluster of Differentiation 36 | 1.09 ± 0.1 | 1.27 ± 0.15 |
| Carnitine palmitoyltransferase 1 | 1.04 ± 0.12 | 1.12 ± 0.12 |
| Diglyceride acyltransferase 1 | 0.83 ± 0.1 | 1.09 ± 0.1 |
| Diglyceride acyltransferase 2 | 1.46 ± 0.16 | 1.16 ± 0.14 |
| Fatty acid-binding protein | 0.81 ± 0.09 | 1.28 ± 0.15 |
| Fatty acid synthase | 1.14 ± 0.12 | 1.31 ± 0.15 |
| Hormone-sensitive lipase | 1.32 ± 0.11 | 1.35 ± 0.13 |
| Leptin | 0.88 ± 0.1 | 1.16 ± 0.11 |
| Lipoprotein lipase | 0.82 ± 0.09 | 0.53 ± 0.06 * |
| Perilipin | 1.05 ± 0.09 | 0.93 ± 0.09 |
| Peroxisome proliferator-activated receptor α | 1.11 ± 0.11 | 0.88 ± 0.08 |
| Peroxisome proliferator-activated receptor γ | 1.21 ± 0.11 | 1.31 ± 0.18 |
| Resistin | 0.84 ± 0.09 | 0.82 ± 0.07 |
| Stearoyl-CoA desaturase-1 | 1.11 ± 0.11 | 0.88 ± 0.08 |
| NAD-dependent deacetylase sirtuin-3 | 0.82 ± 0.09 | 0.78 ± 0.09 |
| Sterol regulatory element-binding protein 1c | 1.18 ± 0.18 | 0.85 ± 0.08 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, X.; Huang, Y.; Sun, C.; Li, J. Maternal Prenatal Folic Acid Supplementation Programs Offspring Lipid Metabolism by Aberrant DNA Methylation in Hepatic ATGL and Adipose LPL in Rats. Nutrients 2017, 9, 935. https://doi.org/10.3390/nu9090935
Yang X, Huang Y, Sun C, Li J. Maternal Prenatal Folic Acid Supplementation Programs Offspring Lipid Metabolism by Aberrant DNA Methylation in Hepatic ATGL and Adipose LPL in Rats. Nutrients. 2017; 9(9):935. https://doi.org/10.3390/nu9090935
Chicago/Turabian StyleYang, Xue, Yifan Huang, Changhao Sun, and Jie Li. 2017. "Maternal Prenatal Folic Acid Supplementation Programs Offspring Lipid Metabolism by Aberrant DNA Methylation in Hepatic ATGL and Adipose LPL in Rats" Nutrients 9, no. 9: 935. https://doi.org/10.3390/nu9090935
APA StyleYang, X., Huang, Y., Sun, C., & Li, J. (2017). Maternal Prenatal Folic Acid Supplementation Programs Offspring Lipid Metabolism by Aberrant DNA Methylation in Hepatic ATGL and Adipose LPL in Rats. Nutrients, 9(9), 935. https://doi.org/10.3390/nu9090935




