Lactobacillus reuteri I5007 Modulates Intestinal Host Defense Peptide Expression in the Model of IPEC-J2 Cells and Neonatal Piglets
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Bacterial Strain, Growth and Storage Conditions
2.3. Cell Culture and Treatment
2.4. Animals and Treatments
2.5. Analysis of Porcine Gene Expression by Real Time PCR
2.6. Colonic Short Chain Fatty Acid Concentrations
2.7. Fecal Microbiota Analysis
2.8. Statistical Analysis
3. Results
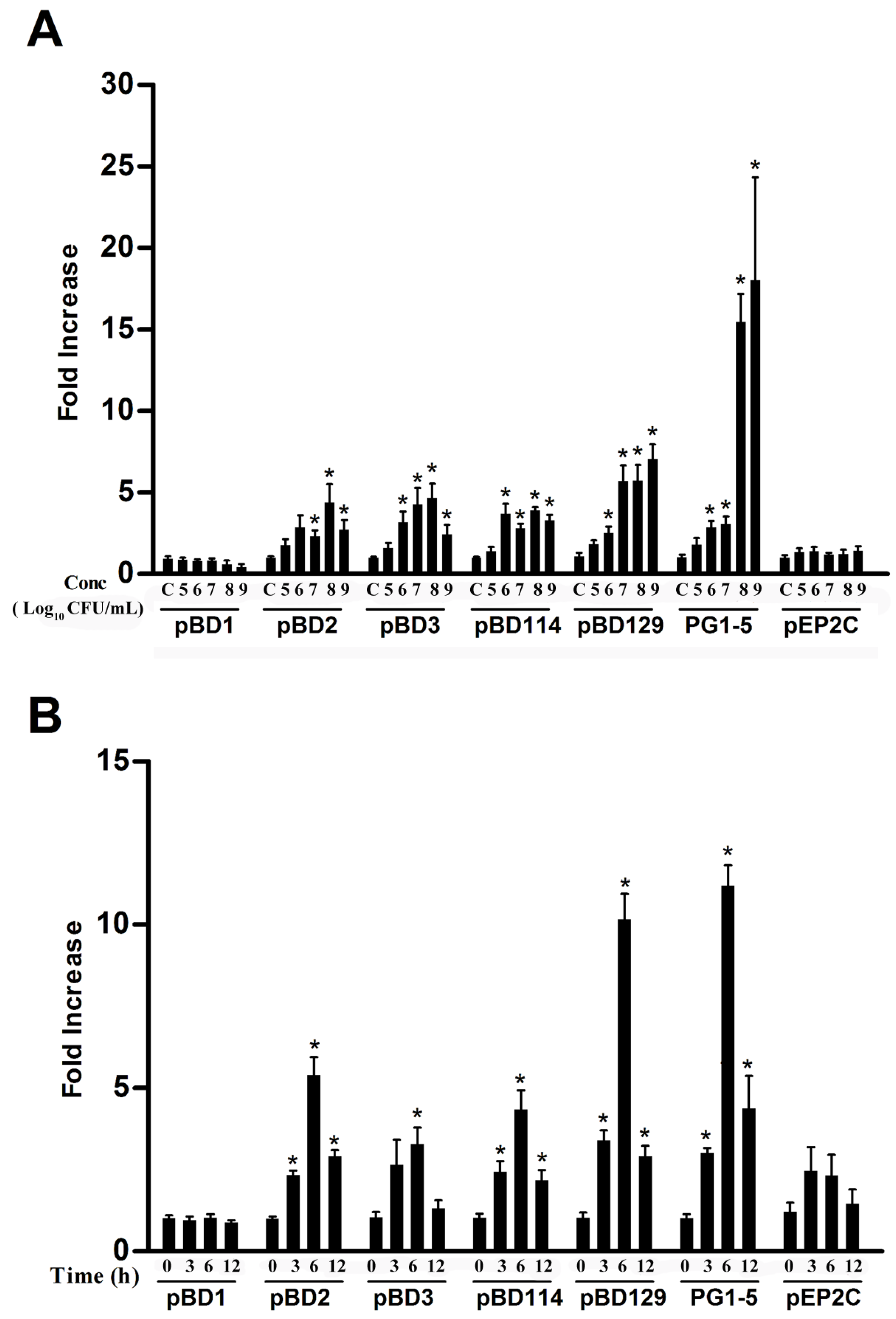
3.1. Effects of L. reuteri I5007 on Host Defense Peptide Expression in IPEC-J2 cells
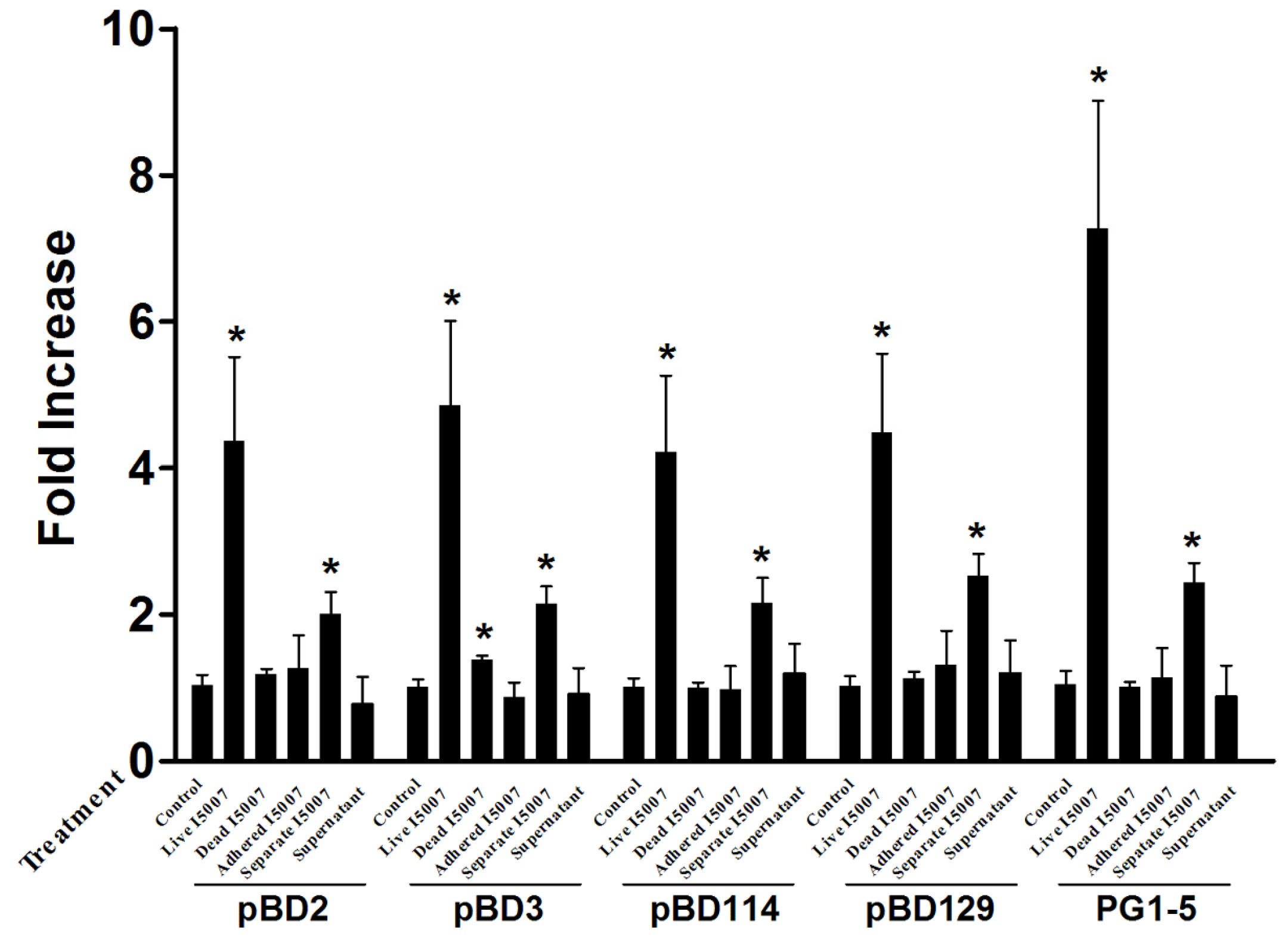
3.2. Effects of Different Processing Conditions on L. reuteri I5007 Induced Host Defense Peptide Expression in IPEC-J2 Cells
3.3. Effects of L. reuteri I5007 on Neonatal Piglet Performance
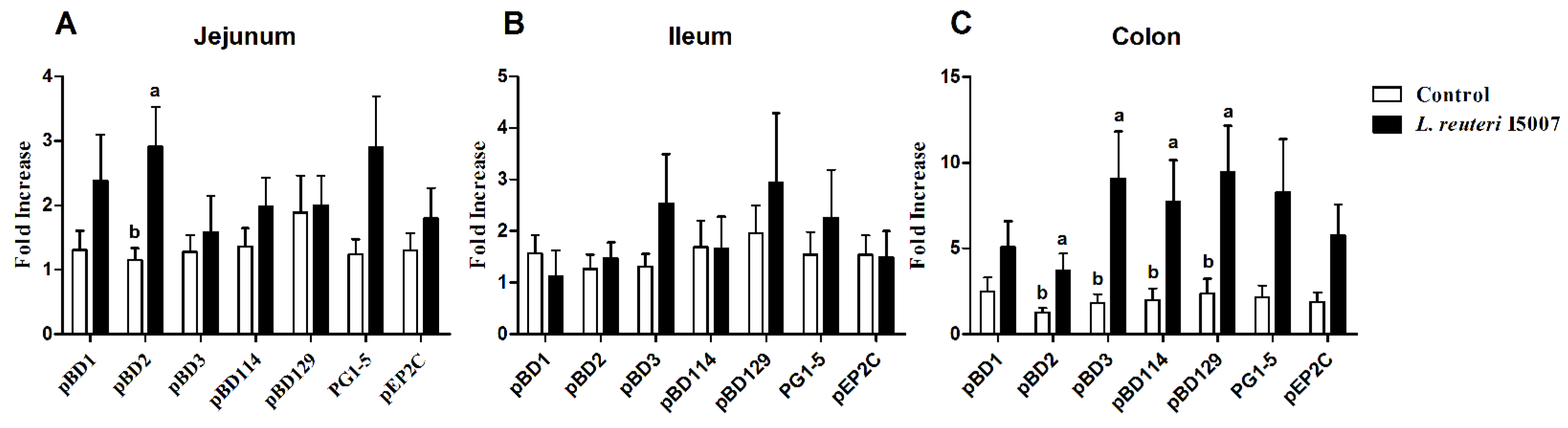
3.4. Effects of L. reuteri I5007 on Host Defense Peptide Expression in Neonatal Piglets
3.5. Effects of L. reuteri I5007 on Short Chain Fatty Acid Concentrations in Colonic Digesta
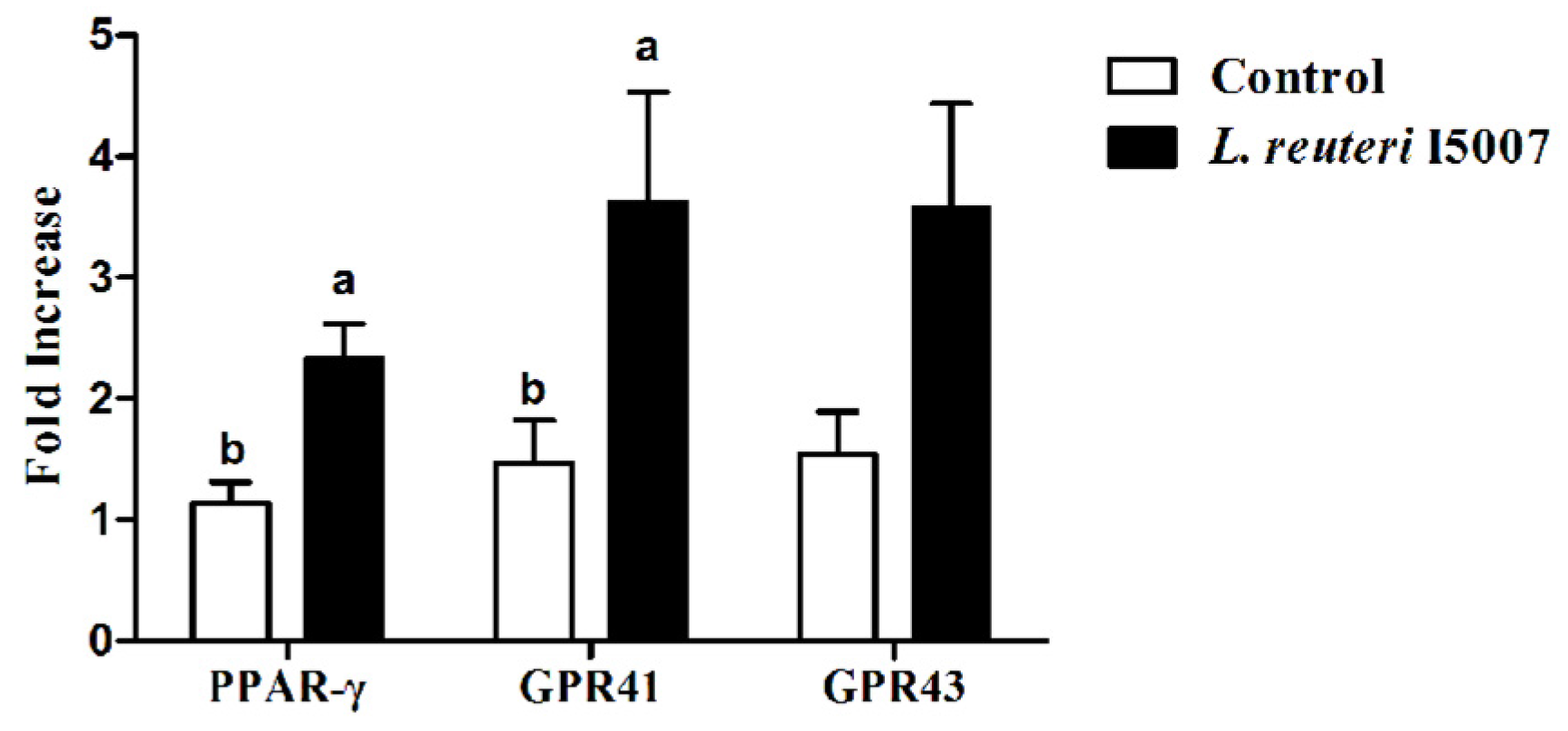
3.6. Effects of L. reuteri I5007 on PPAR-γ, GPR41 and GPR43 in Colonic Tissue
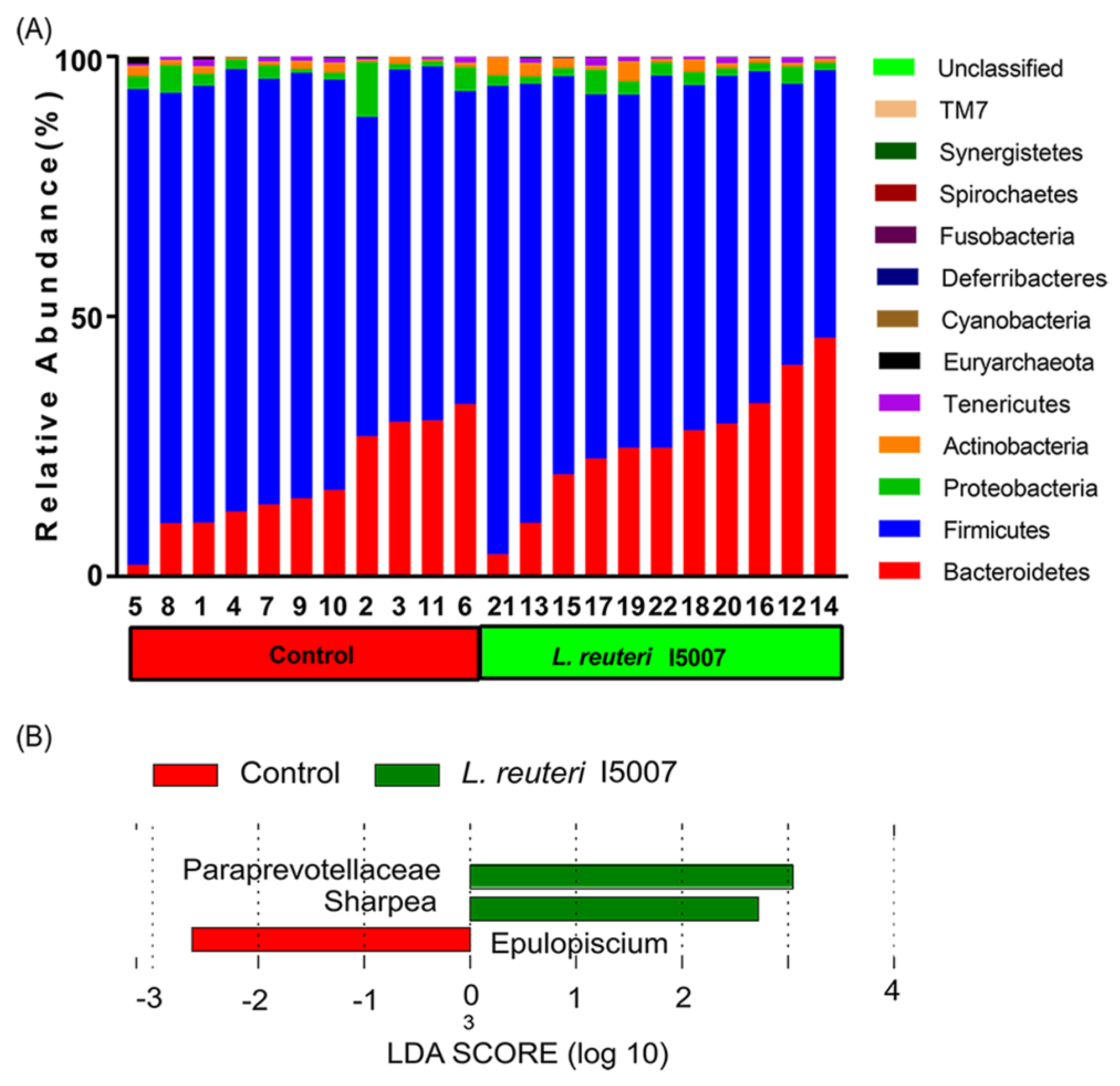
3.7. Effects of L. reuteri I5007 on Bacterial Community Structure in Colonic Digesta
3.8. Effects of L. reuteri I5007 on Pattern-Recognition Receptor (PRRs) Expression in Intestinal Tissue
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
References
- Laxminarayan, R.; Matsoso, P.; Pant, S.; Brower, C.; Rottingen, J.A.; Klugman, K.; Davies, S. Access to effective antimicrobials: A worldwide challenge. Lancet 2016, 387, 168–175. [Google Scholar] [CrossRef]
- Nijnik, A.; Hancock, R. Host defence peptides: Antimicrobial and immunomodulatory activity and potential applications for tackling antibiotic-resistant infections. Emerg. Health Threats J. 2009, 2, e1. [Google Scholar] [CrossRef] [PubMed]
- Hancock, R.E.; Sahl, H.G. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat. Biotechnol. 2006, 24, 1551–1557. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Sunkara, L.T.; Jiang, W.; Bible, M.; Carter, S.; Ma, X.; Qiao, S.; Zhang, G. Induction of porcine host defense peptide gene expression by short-chain fatty acids and their analogs. PLoS ONE 2013, 8, e72922. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, L.M.; Williamson, I.; Piedrahita, J.A.; Blikslager, A.T.; Magness, S.T. Cell lineage identification and stem cell culture in a porcine model for the study of intestinal epithelial regeneration. PLoS ONE 2013, 8, e66465. [Google Scholar] [CrossRef] [PubMed]
- Meurens, F.; Summerfield, A.; Nauwynck, H.; Saif, L.; Gerdts, V. The pig: A model for human infectious diseases. Trends Microbiol. 2012, 20, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Selsted, M.E.; Ouellette, A.J. Mammalian defensins in the antimicrobial immune response. Nat. Immunol. 2005, 6, 551–557. [Google Scholar] [CrossRef] [PubMed]
- Zanetti, M. Cathelicidins, multifunctional peptides of the innate immunity. J. Leukoc. Biol. 2004, 75, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Talukder, P.; Satho, T.; Irie, K.; Sharmin, T.; Hamady, D.; Nakashima, Y.; Kashige, N.; Miake, F. Trace metal zinc stimulates secretion of antimicrobial peptide LL-37 from Caco-2 cells through ERK and p38 MAP kinase. Int. Immunopharmacol. 2011, 11, 141–144. [Google Scholar] [CrossRef] [PubMed]
- Fehlbaum, P.; Rao, M.; Zasloff, M.; Anderson, G.M. An essential amino acid induces epithelial beta -defensin expression. Proc. Natl. Acad. Sci. USA 2000, 97, 12723–12728. [Google Scholar] [CrossRef] [PubMed]
- Cederlund, A.; Kai-Larsen, Y.; Printz, G.; Yoshio, H.; Alvelius, G.; Lagercrantz, H.; Stromberg, R.; Jornvall, H.; Gudmundsson, G.H.; Agerberth, B.; et al. Lactose in human breast milk an inducer of innate immunity with implications for a role in intestinal homeostasis. PLoS ONE 2013, 8, e53876. [Google Scholar] [CrossRef] [PubMed]
- Gombart, A.F.; Borregaard, N.; Koeffler, H.P. Human cathelicidin antimicrobial peptide (CAMP) gene is a direct target of the vitamin D receptor and is strongly up-regulated in myeloid cells by 1,25-dihydroxyvitamin D3. FASEB J. 2005, 19, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Schlee, M.; Harder, J.; Koten, B.; Stange, E.F.; Wehkamp, J.; Fellermann, K. Probiotic lactobacilli and VSL#3 induce enterocyte beta-defensin 2. Clin. Exp. Immunol. 2008, 151, 528–535. [Google Scholar] [PubMed]
- Zhang, J.; Deng, J.; Li, Y.; Yang, Q. The effect of Lactobacillus on the expression of porcine β-defensin-2 in the digestive tract of piglets. Livest. Sci. 2011, 138, 259–265. [Google Scholar] [CrossRef]
- Valeur, N.; Engel, P.; Carbajal, N.; Connolly, E.; Ladefoged, K. Colonization and immunomodulation by Lactobacillus reuteri ATCC 55730 in the human gastrointestinal tract. Appl. Environ. Microbiol. 2004, 70, 1176–1181. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.; Zeng, X.; Yang, F.; Liu, H.; Qiao, S. Study and use of the probiotic Lactobacillus reuteri in pigs: A review. J. Anim. Sci. Biotechnol. 2015, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.; Wang, Q.; Zeng, X.; Yang, F.; Zhang, J.; Liu, H.; Ma, X.; Qiao, S. Complete genome sequence of Lactobacillus reuteri I5007, a probiotic strain isolated from healthy piglet. J. Biotechnol. 2014, 179, 63–64. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Qiao, S.; Li, D.; Piao, X.; Ren, J. Effects of lactobacilli on the performance, diarrhea incidence, VFA concentration and gastrointestinal microbial flora of weaning pigs. Asian Australas. J. Anim. Sci. 2004, 17, 401–409. [Google Scholar] [CrossRef]
- Li, X.J.; Yue, L.Y.; Guan, X.F.; Qiao, S.Y. The adhesion of putative probiotic lactobacilli to cultured epithelial cells and porcine intestinal mucus. J. Appl. Microbiol. 2008, 104, 1082–1091. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yang, F.; Liu, C.; Zhou, H.; Wu, G.; Qiao, S.; Li, D.; Wang, J. Dietary Supplementation with the Probiotic Lactobacillus fermentum I5007 and the Antibiotic Aureomycin Differentially Affects the Small Intestinal Proteomes of Weanling Piglets. J. Nutr. 2011, 142, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.F.; Wang, A.N.; Li, X.J.; Qiao, S.Y. Effect of viable Lactobacillus fermentum on the growth performance, nutrient digestibility and immunity of weaned pigs. J. Anim. Feed Sci. 2008, 17, 61–69. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, J.; Zhang, S.; Yang, F.; Thacker, P.A.; Zhang, G.; Qiao, S.; Ma, X. Oral administration of Lactobacillus fermentum I5007 favors intestinal development and alters the intestinal microbiota in formula-fed piglets. J. Agric. Food Chem. 2014, 62, 860–866. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.N.; Cai, C.J.; Zeng, X.F.; Zhang, F.R.; Zhang, G.L.; Thacker, P.A.; Wang, J.J.; Qiao, S.Y. Dietary supplementation with. J. Appl. Microbiol. 2013, 114, 1582–1591. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.; Liu, H.; Zhang, J.; Zhang, S.; Yang, F.; Zeng, X.; Thacker, P.; Zhang, G.; Qiao, S. Intestinal Microbiota Succession and Immunomodulatory Consequences after Introduction of Lactobacillus reuteri I5007 in Neonatal Piglets. PLoS ONE 2015, 10, e0119505. [Google Scholar] [CrossRef] [PubMed]
- Marquardt, R.R.; Jin, L.Z.; Kim, J.W.; Fang, L.; Frohlich, A.A.; Baidoo, S.K. Passive protective effect of egg-yolk antibodies against enterotoxigenic Escherichia coli K88+ infection in neonatal and early-weaned piglets. FEMS Immunol. Med. Microbiol. 1999, 23, 283–288. [Google Scholar] [CrossRef] [PubMed]
- Ou, D.; Li, D.; Cao, Y.; Li, X.; Yin, J.; Qiao, S.; Wu, G. Dietary supplementation with zinc oxide decreases expression of the stem cell factor in the small intestine of weanling pigs. J. Nutr. Biochem. 2007, 18, 820–826. [Google Scholar] [CrossRef] [PubMed]
- Aranda, P.S.; LaJoie, D.M.; Jorcyk, C.L. Bleach gel: A simple agarose gel for analyzing RNA quality. Electrophoresis 2012, 33, 366–369. [Google Scholar] [CrossRef] [PubMed]
- Arce, C.; Ramirez-Boo, M.; Lucena, C.; Garrido, J.J. Innate immune activation of swine intestinal epithelial cell lines (IPEC-J2 and IPI-2I) in response to LPS from Salmonella typhimurium. Comp. Immunol. Microbiol. Infect. Dis. 2010, 33, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zerby, H.N.; Lee, K. Heart fatty acid binding protein is upregulated during porcine adipocyte development. J. Anim. Sci. 2007, 85, 1651–1659. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Su, H.; Zhou, Z.; Yao, W. Identification of the porcine G protein-coupled receptor 41 and 43 genes and their expression pattern in different tissues and development stages. PloS ONE 2014, 9, e97342. [Google Scholar] [CrossRef] [PubMed]
- Mariani, V.; Palermo, S.; Fiorentini, S.; Lanubile, A.; Giuffra, E. Gene expression study of two widely used pig intestinal epithelial cell lines: IPEC-J2 and IPI-2I. Vet. Immunol. Immunopathol. 2009, 131, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Jin, X. Development and optimization of organic acid analysis in tobacco with ion chromatography and suppressed conductivity detection. J. Chromatogr. A 2002, 950, 81–88. [Google Scholar] [CrossRef]
- Wehkamp, J.; Harder, J.; Wehkamp, K.; Wehkamp-von Meissner, B.; Schlee, M.; Enders, C.; Sonnenborn, U.; Nuding, S.; Bengmark, S.; Fellermann, K.; et al. NF-κB-and AP-1-mediated induction of human beta defensin-2 in intestinal epithelial cells by Escherichia coli Nissle 1917: A novel effect of a probiotic bacterium. Infect. Immun. 2004, 72, 5750–5758. [Google Scholar] [CrossRef] [PubMed]
- Topping, D.L.; Clifton, P.M. Short-chain fatty acids and human colonic function: Roles of resistant starch and nonstarch polysaccharides. Physiol. Rev. 2001, 81, 1031–1064. [Google Scholar] [PubMed]
- Weber, N.; Nielsen, J.P.; Jakobsen, A.S.; Pedersen, L.L.; Hansen, C.F.; Pedersen, K.S. Occurrence of diarrhoea and intestinal pathogens in non-medicated nursery pigs. Acta Vet. Scand. 2015, 57, 64. [Google Scholar] [CrossRef] [PubMed]
- Sang, Y.; Blecha, F. Porcine host defense peptides: Expanding repertoire and functions. Dev. Comp. Immunol. 2009, 33, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Dou, X.; Han, J.; Song, W.; Dong, N.; Xu, X.; Zhang, W.; Shan, A. Sodium butyrate improves porcine host defense peptide expression and relieves the inflammatory response upon Toll-like receptor 2 activation and histone deacetylase inhibition in porcine kidney cells. Oncotarget 2017, 8, 26532–26551. [Google Scholar] [CrossRef] [PubMed]
- Gallo, R.L.; Hooper, L.V. Epithelial antimicrobial defence of the skin and intestine. Nat. Rev. Immunol. 2012, 12, 503–516. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.H.; Ingale, S.L.; Kim, J.S.; Kim, K.H.; Lohakare, J.; Park, Y.K.; Kwon, I.K.; Chae, B.J. Effects of dietary supplementation with antimicrobial peptide-P5 on growth performance, apparent total tract digestibility, faecal and intestinal microflora and intestinal morphology of weanling pigs. J. Sci. Food Agric. 2013, 93, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Wen, L.F.; He, J.G. Dose-response effects of an antimicrobial peptide, a cecropin hybrid, on growth performance, nutrient utilisation, bacterial counts in the digesta and intestinal morphology in broilers. Br. J. Nutr. 2012, 108, 1756–1763. [Google Scholar] [CrossRef] [PubMed]
- Cullen, T.W.; Schofield, W.B.; Barry, N.A.; Putnam, E.E.; Rundell, E.A.; Trent, M.S.; Degnan, P.H.; Booth, C.J.; Yu, H.; Goodman, A.L. Gut microbiota. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 2015, 347, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Goyette-Desjardins, G.; Auger, J.P.; Xu, J.; Segura, M.; Gottschalk, M. Streptococcus suis, an important pig pathogen and emerging zoonotic agent-an update on the worldwide distribution based on serotyping and sequence typing. Emerg. Microbes. Infect. 2014, 3, e45. [Google Scholar] [CrossRef] [PubMed]
- Basson, M.D.; Liu, Y.; Hanly, A.M.; Emenaker, N.J.; Shenoy, S.G.; Rothberg, B.E.G. Identification and comparative analysis of human colonocyte short-chain fatty acid response genes. J. Gastrointest. Surg. 2000, 4, 501–512. [Google Scholar] [CrossRef]
- Sanderson, I.R. Short chain fatty acid regulation of signaling genes expressed by the intestinal epithelium. J. Nutr. 2004, 134, 2450S–2454S. [Google Scholar] [PubMed]
- Alex, S.; Lange, K.; Amolo, T.; Grinstead, J.S.; Haakonsson, A.K.; Szalowska, E.; Koppen, A.; Mudde, K.; Haenen, D.; Roelofsen, H.; et al. Short-chain fatty acids stimulate angiopoietin-like 4 synthesis in human colon adenocarcinoma cells by activating peroxisome proliferator-activated receptor γ. Mol. Cell. Biol. 2013, 33, 1303–1316. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.; Goldsworthy, S.M.; Barnes, A.A.; Eilert, M.M.; Tcheang, L.; Daniels, D.; Muir, A.I.; Wigglesworth, M.J.; Kinghorn, I.; Fraser, N.J. The Orphan G protein-coupled receptors GPR41 and GPR43 are activated by propionate and other short chain carboxylic acids. J. Biol. Chem. 2003, 278, 11312–11319. [Google Scholar] [CrossRef] [PubMed]
- Tazoe, H.; Otomo, Y.; Kaji, I.; Tanaka, R.; Karaki, S.I.; Kuwahara, A. Roles of short-chain fatty acids receptors, GPR41 and GPR43 on colonic functions. J. Physiol. Pharmacol. 2008, 59 (Suppl. 2), 251–262. [Google Scholar] [PubMed]
- Leser, T.D.; Mølbak, L. Better living through microbial action: The benefits of the mammalian gastrointestinal microbiota on the host. Environ. Microbiol. 2009, 11, 2194–2206. [Google Scholar] [CrossRef] [PubMed]
- Lopez, C.A.; Kingsbury, D.D.; Velazquez, E.M.; Baumler, A.J. Collateral damage: Microbiota-derived metabolites and immune function in the antibiotic era. Cell Host Microbe 2014, 16, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Savard, P.; Lamarche, B.; Paradis, M.E.; Thiboutot, H.; Laurin, E.; Roy, D. Impact of Bifidobacterium animalis subsp. lactis BB-12 and, Lactobacillus acidophilus LA-5-containing yoghurt, on fecal bacterial counts of healthy adults. Int. J. Food Microbiol. 2011, 149, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Yamano, T.; Iino, H.; Takada, M.; Blum, S.; Rochat, F.; Fukushima, Y. Improvement of the human intestinal flora by ingestion of the probiotic strain Lactobacillus johnsonii La1. Br. J. Nutr. 2006, 95, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Lahti, L.; Salonen, A.; Kekkonen, R.A.; Salojärvi, J.; Jalanka-Tuovinen, J.; Palva, A.; Orešič, M.; de Vos, W.M. Associations between the human intestinal microbiota, Lactobacillus rhamnosus GG and serum lipids indicated by integrated analysis of high-throughput profiling data. PeerJ 2013, 1, e32. [Google Scholar] [CrossRef] [PubMed]
- Rajkumar, H.; Kumar, M.; Das, N.; Kumar, S.N.; Challa, H.R.; Nagpal, R. Effect of Probiotic Lactobacillus salivarius UBL S22 and Prebiotic Fructo-oligosaccharide on Serum Lipids, Inflammatory Markers, Insulin Sensitivity, and Gut Bacteria in Healthy Young Volunteers: A Randomized Controlled Single-Blind Pilot Study. J. Cardiovasc. Pharmacol. Ther. 2015, 20, 289–298. [Google Scholar] [CrossRef] [PubMed]
- Berni, C.R.; Sangwan, N.; Stefka, A.T.; Nocerino, R.; Paparo, L.; Aitoro, R.; Calignano, A.; Khan, A.A.; Gilbert, J.A.; Nagler, C.R. Lactobacillus rhamnosus GG-supplemented formula expands butyrate-producing bacterial strains in food allergic infants. ISME J. 2015, 10, 742–750. [Google Scholar]
- Kristensen, N.B.; Bryrup, T.; Allin, K.H.; Nielsen, T.; Hansen, T.H.; Pedersen, O. Alterations in fecal microbiota composition by probiotic supplementation in healthy adults: A systematic review of randomized controlled trials. Genome. Med. 2016, 8, 52. [Google Scholar] [CrossRef] [PubMed]
- Morita, H.; Shiratori, C.; Murakami, M.; Takami, H.; Toh, H.; Kato, Y.; Nakajima, F.; Takagi, M.; Akita, H.; Masaoka, T.; et al. Sharpea azabuensis gen. nov., sp. nov., a Gram-positive, strictly anaerobic bacterium isolated from the faeces of thoroughbred horses. Int. J. Syst. Evol. Microbiol. 2008, 58, 2682–2686. [Google Scholar] [CrossRef] [PubMed]
- Eloe-Fadrosh, E.A.; Brady, A.; Crabtree, J.; Drabek, E.F.; Ma, B.; Mahurkar, A.; Ravel, J.; Haverkamp, M.; Fiorino, A.M.; Botelho, C.; et al. Functional dynamics of the gut microbiome in elderly people during probiotic consumption. MBio 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- McNulty, N.P.; Yatsunenko, T.; Hsiao, A.; Faith, J.J.; Muegge, B.D.; Goodman, A.L.; Henrissat, B.; Oozeer, R.; Cools-Portier, S.; Gobert, G.; et al. The Impact of a Consortium of Fermented Milk Strains on the Gut Microbiome of Gnotobiotic Mice and Monozygotic Twins. Sci. Transl. Med. 2011, 3, 106ra106. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Mazmanian, S.K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 2013, 14, 668–675. [Google Scholar] [CrossRef] [PubMed]
- Turroni, F.; Milani, C.; Duranti, S.; Mancabelli, L.; Mangifesta, M.; Viappiani, A.; Lugli, G.A.; Ferrario, C.; Gioiosa, L.; Ferrarini, A.; et al. Deciphering bifidobacterial-mediated metabolic interactions and their impact on gut microbiota by a multi-omics approach. ISME J. 2016, 10, 1656–1668. [Google Scholar] [CrossRef] [PubMed]
| Items | Content |
|---|---|
| Crude protein | 24.78 |
| Gross energy (MJ/kg) | 20.46 |
| Lactose | 35.10 |
| Calcium | 0.92 |
| Total phosphorus | 0.73 |
| The analyzed contents of amino acids in diets | |
| Asparate | 2.58 |
| Threoline | 1.72 |
| Serine | 1.35 |
| Glutamate | 4.52 |
| Proline | 1.54 |
| Glycine | 0.55 |
| Alanine | 1.30 |
| Valine | 1.48 |
| Isoleucine | 1.42 |
| Leucine | 2.54 |
| Tyrosine | 0.81 |
| Phenylalanine | 0.92 |
| Histidine | 0.52 |
| Lysine | 2.03 |
| Methonine | 0.65 |
| Argine | 0.76 |
| Items | Control | L. reuteri I5007 | SEM | p Value |
|---|---|---|---|---|
| Body weight at day 4 (kg) | 1.81 | 1.81 | 0.07 | 0.99 |
| Body weight at day 14 (kg) | 2.45 | 2.50 | 0.07 | 0.72 |
| Body weight at day 24 (kg) | 3.72 | 4.00 | 0.10 | 0.16 |
| Average daily gain (g) | ||||
| 4–14 days | 64 | 69 | 3.91 | 0.52 |
| 14–24 days | 127 b | 151 a | 4.86 | 0.01 |
| 4–24 days | 96 b | 110 a | 3.31 | 0.03 |
| Average feed intake (g/day) | ||||
| 4–14 days | 53 | 58 | 3.43 | 0.08 |
| 14–24 days | 140 | 152 | 5.25 | 0.82 |
| 4–24 days | 97 | 105 | 3.24 | 0.18 |
| Diarrhea score 2 | 0.20 | 0.12 | ||
| Diarrhea incidence 3 (%) | 5.91 | 3.64 | 0.32 | |
| Fatty acid | Control | L. reuteri I5007 | SEM | p Value |
|---|---|---|---|---|
| Formic acid | 0.03 | 0.03 | 0.01 | 0.66 |
| Acetic acid | 32.49 | 37.74 | 2.03 | 0.20 |
| Propionic acid | 12.99 | 15.90 | 0.83 | 0.08 |
| Butyric acid | 7.10 b | 9.51 a | 0.58 | 0.04 |
| Lactic acid | 3.67 | 3.66 | 0.35 | 1.00 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, H.; Hou, C.; Wang, G.; Jia, H.; Yu, H.; Zeng, X.; Thacker, P.A.; Zhang, G.; Qiao, S. Lactobacillus reuteri I5007 Modulates Intestinal Host Defense Peptide Expression in the Model of IPEC-J2 Cells and Neonatal Piglets. Nutrients 2017, 9, 559. https://doi.org/10.3390/nu9060559
Liu H, Hou C, Wang G, Jia H, Yu H, Zeng X, Thacker PA, Zhang G, Qiao S. Lactobacillus reuteri I5007 Modulates Intestinal Host Defense Peptide Expression in the Model of IPEC-J2 Cells and Neonatal Piglets. Nutrients. 2017; 9(6):559. https://doi.org/10.3390/nu9060559
Chicago/Turabian StyleLiu, Hongbin, Chengli Hou, Gang Wang, Hongmin Jia, Haitao Yu, Xiangfang Zeng, Philip A Thacker, Guolong Zhang, and Shiyan Qiao. 2017. "Lactobacillus reuteri I5007 Modulates Intestinal Host Defense Peptide Expression in the Model of IPEC-J2 Cells and Neonatal Piglets" Nutrients 9, no. 6: 559. https://doi.org/10.3390/nu9060559
APA StyleLiu, H., Hou, C., Wang, G., Jia, H., Yu, H., Zeng, X., Thacker, P. A., Zhang, G., & Qiao, S. (2017). Lactobacillus reuteri I5007 Modulates Intestinal Host Defense Peptide Expression in the Model of IPEC-J2 Cells and Neonatal Piglets. Nutrients, 9(6), 559. https://doi.org/10.3390/nu9060559







