High Fat Diets Sex-Specifically Affect the Renal Transcriptome and Program Obesity, Kidney Injury, and Hypertension in the Offspring
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Design
2.2. Biochemical Analysis
2.3. Histology and Morphometric Study
2.4. Detection of l-arginine, l-citrulline, ADMA, and SDMA by HPLC
2.5. Next-Generation Sequencing and Analysis
2.6. Quantitative Real-time Polymerase Chain Reaction (PCR)
2.7. Western Blot
2.8. Immunohistochemistry Staining for 8-OHdG
2.9. Statistical Analysis
3. Results
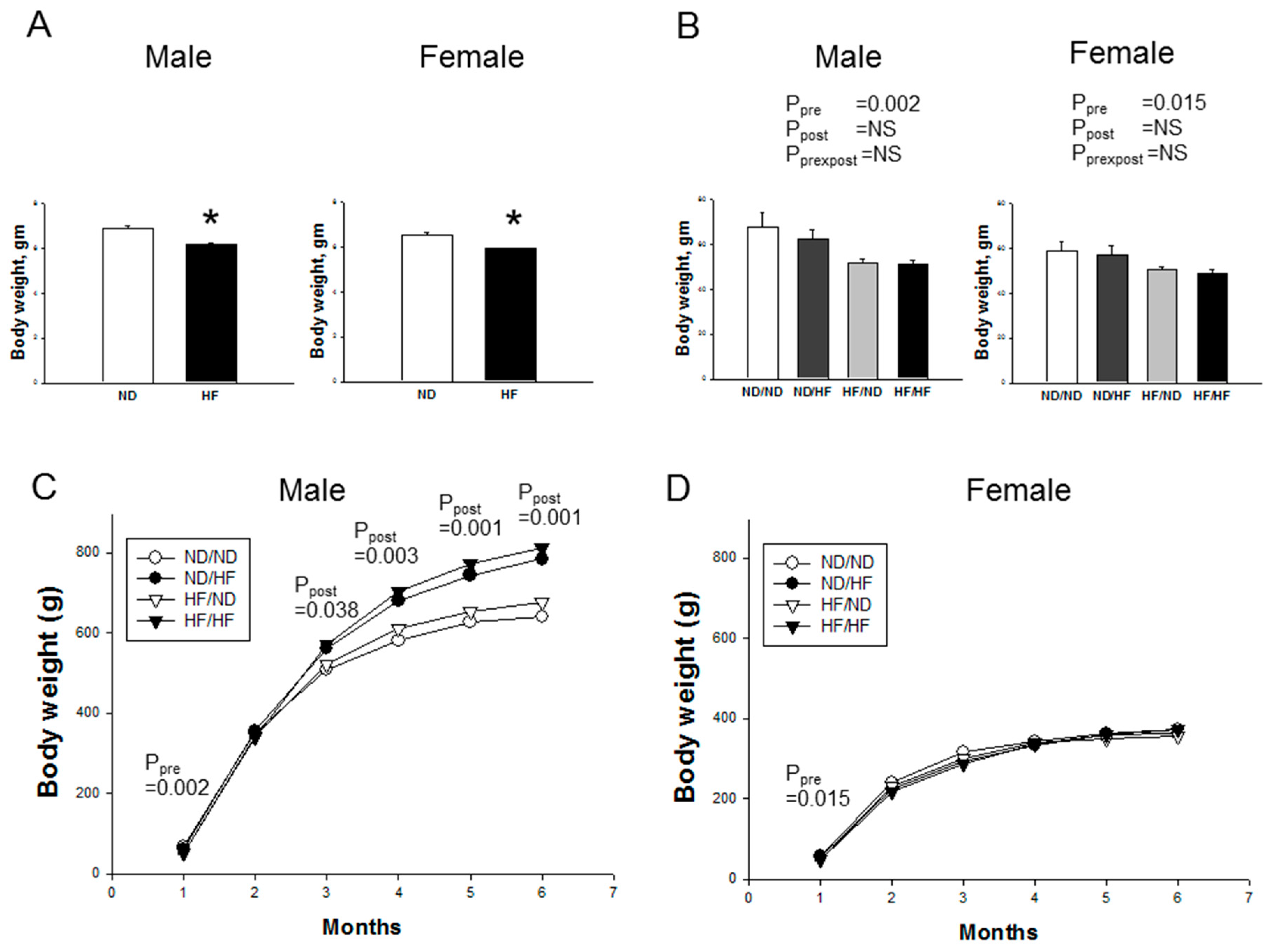
3.1. Morphological Features and Biochemistry
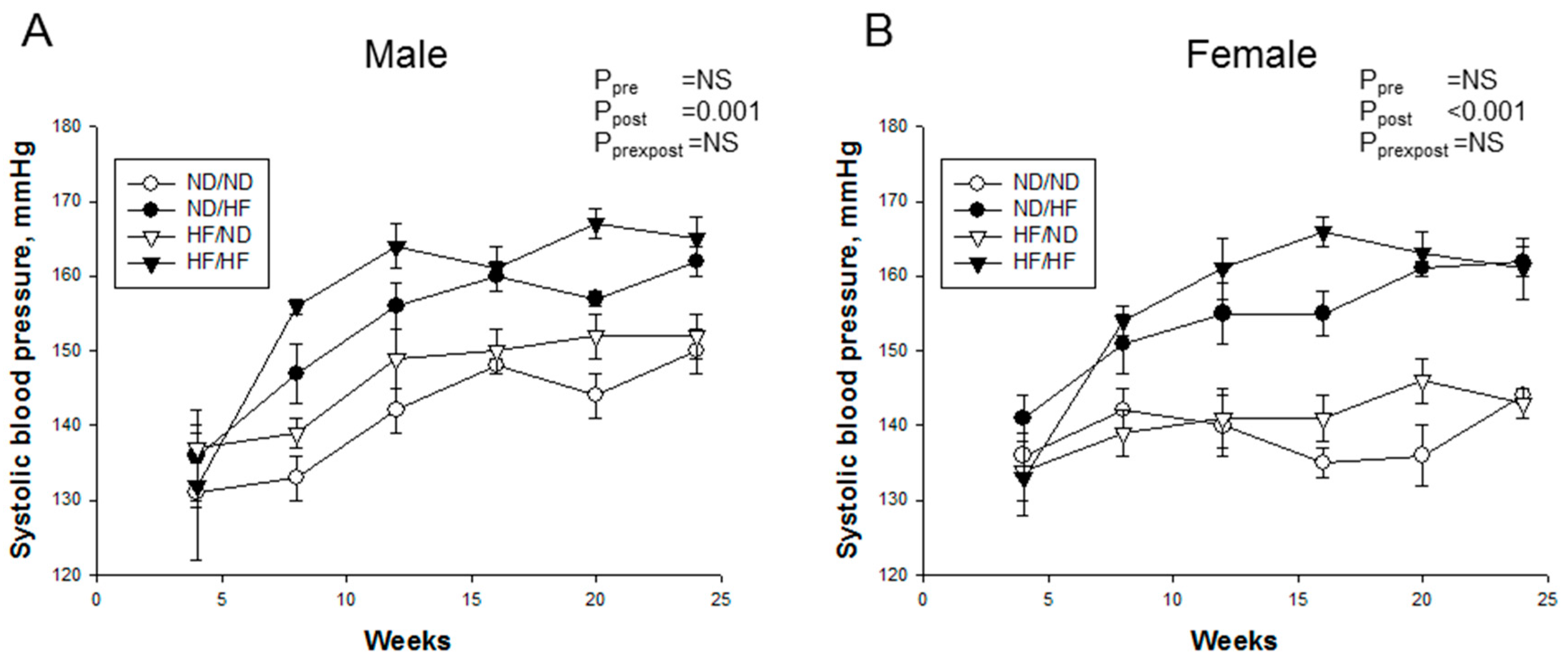
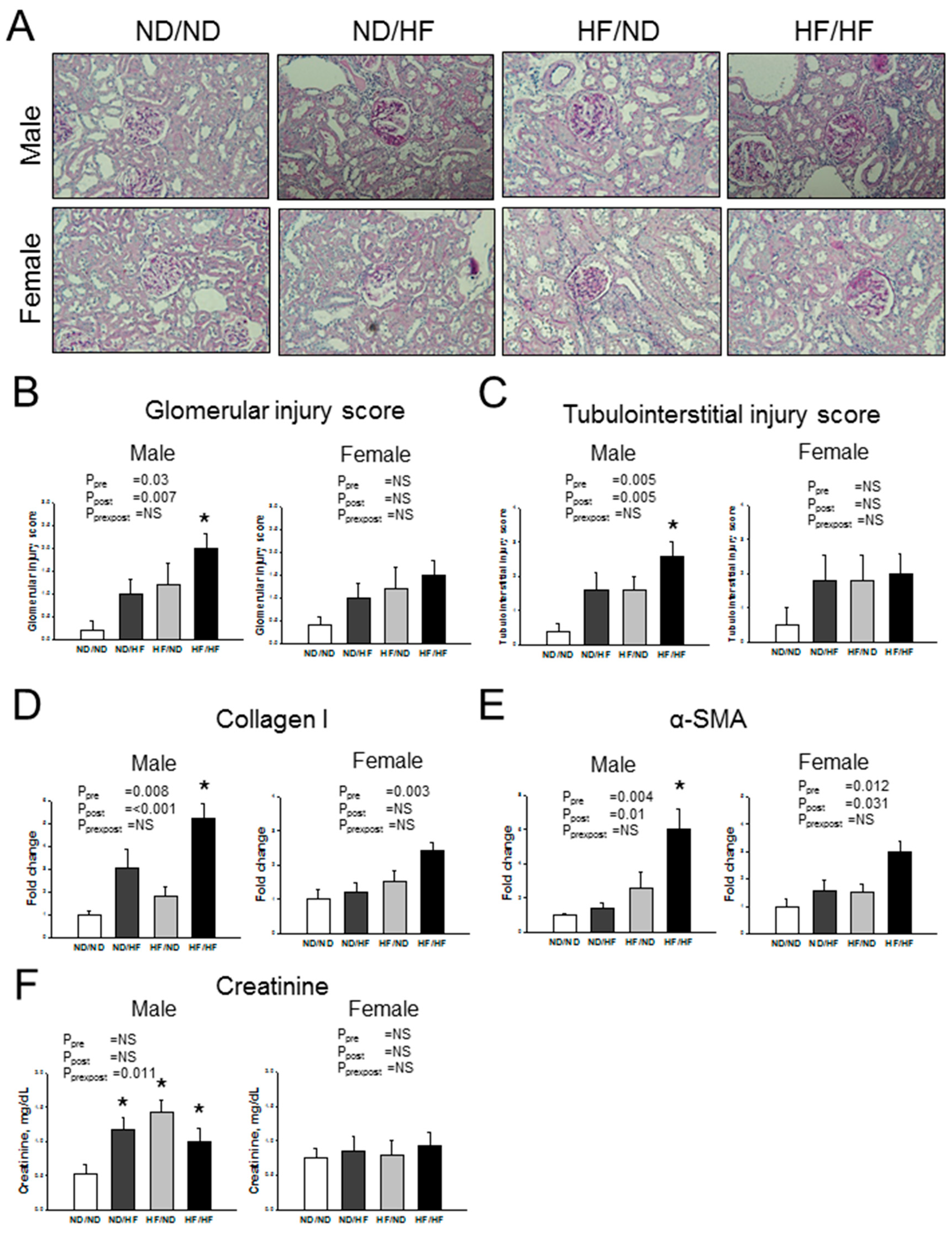
3.2. Blood Pressure and Renal Outcome
3.3. Renal Transcriptome
3.4. Oxidative Stress and Nitric Oxide Pathway
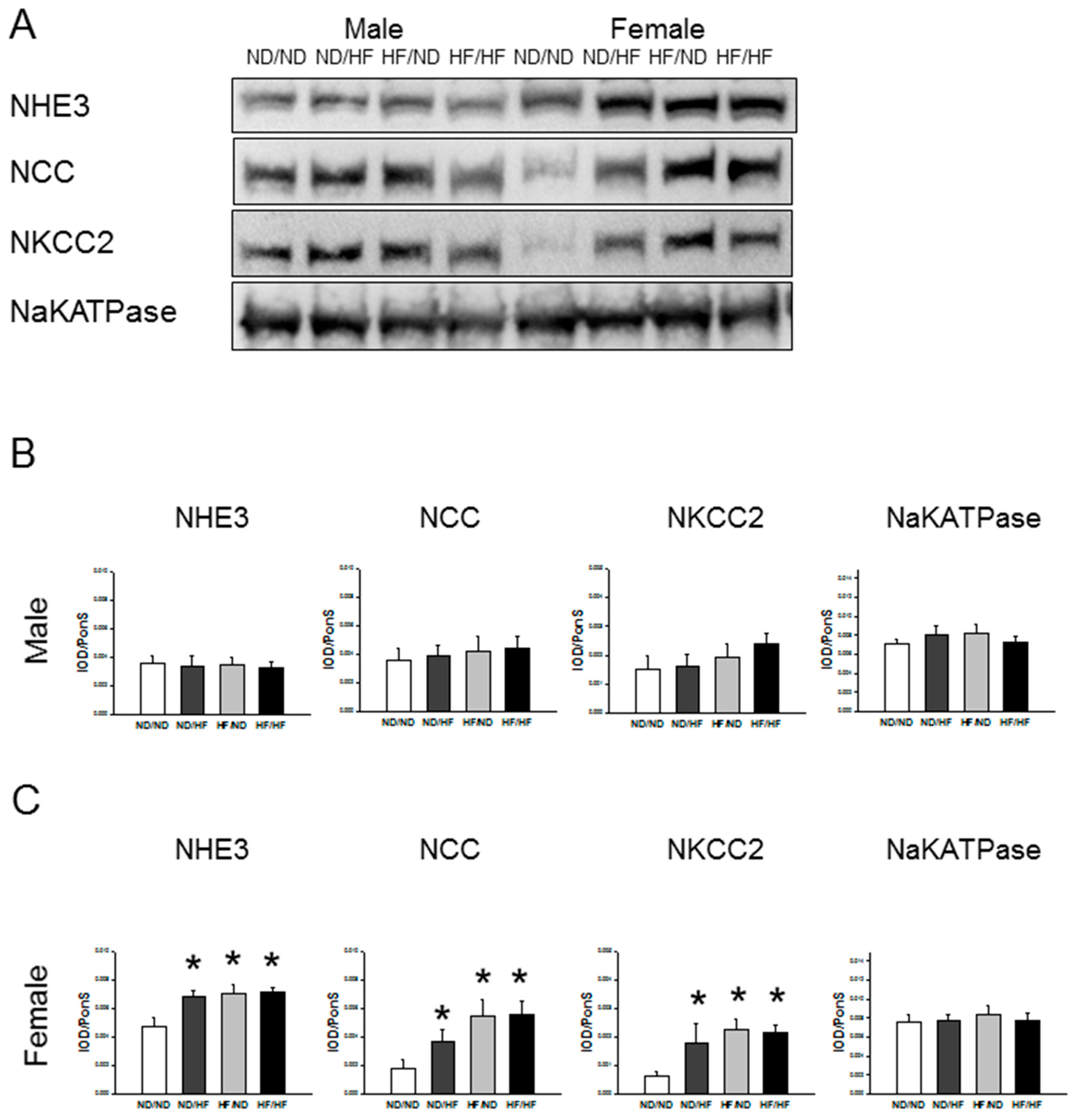
3.5. RAS and Sodium Transporters
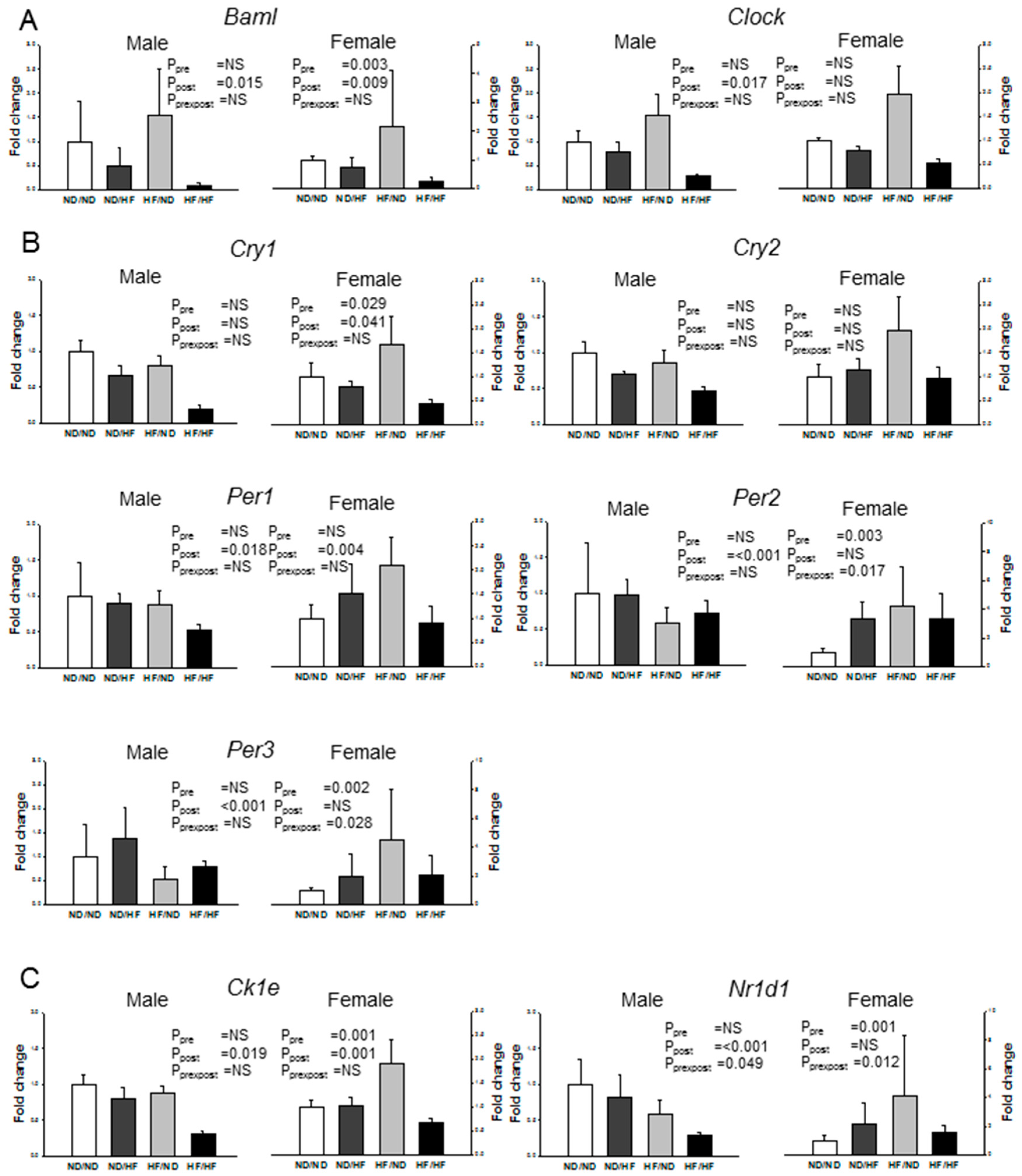
3.6. Clock and Clock-Controlled Genes
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Vickers, M.H. Early life nutrition, epigenetics and programming of later life disease. Nutrients 2014, 6, 2165–2178. [Google Scholar] [CrossRef] [PubMed]
- Misra, A.; Singhal, N.; Khurana, L. Obesity, the metabolic syndrome, and type 2 diabetes in developing countries: Role of dietary fats and oils. J. Am. Coll. Nutr. 2010, 29, 289S–301S. [Google Scholar] [CrossRef] [PubMed]
- Buettner, R.; Parhofer, K.G.; Woenckhaus, M.; Wrede, C.E.; Kunz-Schughart, L.A.; Schölmerich, J.; Bolheimer, L.C. Defining high-fat-diet rat models: Metabolic and molecular effects of different fat types. J. Mol. Endocrinol. 2006, 36, 485–501. [Google Scholar] [CrossRef] [PubMed]
- Buettner, R.; Schölmerich, J.; Bollheimer, L.C. High-fat diets: Modeling the metabolic disorders of human obesity in rodents. Obesity 2007, 15, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Kakimoto, P.A.; Kowaltowski, A.J. Effects of high fat diets on rodent liver bioenergetics and oxidative imbalance. Redox Biol. 2016, 8, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Williams, L.; Seki, Y.; Vuguin, P.M.; Charron, M.J. Animal models of in utero exposure to a high fat diet: A review. Biochim. Biophys. Acta 2014, 1842, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Pan, Y.X. Pathophysiological basis for compromised health beyond generations: Role of maternal high-fat diet and low-grade chronic inflammation. J. Nutr. Biochem. 2015, 26, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Paixão, A.D.; Alexander, B.T. How the kidney is impacted by the perinatal maternal environment to develop hypertension. Biol. Reprod. 2013, 89, 144. [Google Scholar] [CrossRef] [PubMed]
- Luyckx, V.A.; Bertram, J.F.; Brenner, B.M.; Fall, C.; Hoy, W.E.; Ozanne, S.E.; Vikse, B.E. Effect of fetal and child health on kidney development and long-term risk of hypertension and kidney disease. Lancet 2013, 382, 273–283. [Google Scholar] [CrossRef]
- Tain, Y.L.; Joles, J.A. Reprogramming: A preventive strategy in hypertension focusing on the kidney. Int. J. Mol. Sci. 2015, 17, E23. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Hsu, C.N. Developmental origins of chronic kidney disease: Should we focus on early life? Int. J. Mol. Sci. 2017, 18, 381. [Google Scholar] [CrossRef] [PubMed]
- Tokonami, N.; Mordasini, D.; Pradervand, S.; Centeno, G.; Jouffe, C.; Maillard, M.; Bonny, O.; Gachon, F.; Gomez, R.A.; Sequeira-Lopez, M.L.; et al. Local renal circadian clocks control fluid-electrolyte homeostasis and BP. J. Am. Soc. Nephrol. 2014, 25, 1430–1439. [Google Scholar] [CrossRef] [PubMed]
- Feng, D.; Lazar, M.A. Clocks, metabolism, and the epigenome. Mol. Cell 2012, 47, 158–167. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, M.C.; Yang, S.C.; Tseng, H.L.; Hwang, L.L.; Chen, C.T.; Shieh, K.R. Abnormal expressions of circadian-clock and circadian clock-controlled genes in the livers and kidneys of long-term, high-fat-diet-treated mice. Int. J. Obes. 2010, 34, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Roberts, C.K.; Barnard, R.J.; Sindhu, R.K.; Jurczak, M.; Ehdaie, A.; Vaziri, N.D. Oxidative stress and dysregulation of NAD(P)H oxidase and antioxidant enzymes in diet-induced metabolic syndrome. Metabolism 2006, 55, 928–934. [Google Scholar] [CrossRef] [PubMed]
- Riazi, S.; Tiwari, S.; Sharma, N.; Rash, A.; Ecelbarger, C.M. Abundance of the Na-K-2Cl cotransporter NKCC2 is increased by high-fat feeding in Fischer 344 X Brown Norway (F1) rats. Am. J. Physiol. Renal Physiol. 2009, 296, F762–F770. [Google Scholar] [CrossRef] [PubMed]
- Perucca, J.; Bouby, N.; Valeix, P.; Bankir, L. Sex difference in urine concentration across differing ages, sodium intake, and level of kidney disease. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 292, R700–R705. [Google Scholar] [CrossRef] [PubMed]
- Sandberg, K.; Ji, H. Sex differences in primary hypertension. Biol. Sex Differ. 2012, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Mauvais-Jarvis, F. Sex differences in metabolic homeostasis, diabetes, and obesity. Biol. Sex Differ. 2015, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Sheen, J.M.; Yu, H.R.; Chen, C.C.; Tiao, M.M.; Hsu, C.N.; Lin, Y.J.; Kuo, K.C.; Huang, L.T. Maternal melatonin therapy rescues prenatal dexamethasone and postnatal high-fat diet induced programmed hypertension in male rat offspring. Front. Physiol. 2015, 6, 377. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Wu, M.S.; Lin, Y.J. Sex differences in renal transcriptome and programmed hypertension in offspring exposed to prenatal dexamethasone. Steroids 2016, 115, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Ordovas, J.M. Nutrigenomics in cardiovascular medicine. Circ. Cardiovasc. Genet. 2009, 2, 637–651. [Google Scholar] [CrossRef] [PubMed]
- Juma, S.; Imrhan, V.; Vijayagopal, P.; Prasad, C. Prescribing personalized nutrition for cardiovascular health: Are we ready? J. Nutrigenet. Nutrigenomics 2014, 7, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Hsu, C.N.; Chan, J.Y.; Huang, L.T. Renal transcriptome analysis of programmed hypertension induced by maternal nutritional insults. Int. J. Mol. Sci. 2015, 16, 17826–17837. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Chan, J.Y.; Hsu, C.N. Maternal fructose intake affects transcriptome changes and programmed hypertension in offspring in later life. Nutrients 2016, 8, 757. [Google Scholar] [CrossRef] [PubMed]
- Sheen, J.M.; Hsieh, C.S.; Tain, Y.L.; Li, S.W.; Yu, H.R.; Chen, C.C.; Tiao, M.M.; Chen, Y.C.; Huang, L.T. Programming effects of prenatal glucocorticoid exposure with a postnatal high-fat diet in diabetes mellitus. Int. J. Mol. Sci. 2016, 17, E533. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Hsieh, C.S.; Lin, I.C.; Chen, C.C.; Sheen, J.M.; Huang, L.T. Effects of maternal l-citrulline supplementation on renal function and blood pressure in offspring exposed to maternal caloric restriction: The impact of nitric oxide pathway. Nitric Oxide 2010, 23, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Lee, W.C.; Leu, S.; Wu, K.; Chan, J. High salt exacerbates programmed hypertension in maternal fructose-fed male offspring. Nutr. Metab. Cardiovasc. Dis. 2015, 25, 1146–1151. [Google Scholar] [CrossRef] [PubMed]
- NIH DAVID Bioinformatics Resources 6.8. Available online: Https://david.ncifcrf.gov/ (accessed on 9 January 2017).
- Jackson, C.M.; Alexander, B.T.; Roach, L.; Haggerty, D.; Marbury, D.C.; Hutchens, Z.M.; Flynn, E.R.; Maric-Bilkan, C. Exposure to maternal overnutrition and a high-fat diet during early postnatal development increases susceptibility to renal and metabolic injury later in life. Am. J. Physiol. Renal Physiol. 2012, 302, F774–F783. [Google Scholar] [CrossRef] [PubMed]
- Aliou, Y.; Liao, M.C.; Zhao, X.P.; Chang, S.Y.; Chenier, I.; Ingelfinger, J.R.; Zhang, S.L. Post-weaning high-fat diet accelerates kidney injury, but not hypertension programmed by maternal diabetes. Pediatr. Res. 2016, 79, 416–424. [Google Scholar] [CrossRef] [PubMed]
- Férézou-Viala, J.; Roy, A.F.; Sérougne, C.; Gripois, D.; Parquet, M.; Bailleux, V.; Gertler, A.; Delplanque, B.; Djiane, J.; Riottot, M.; Taouis, M. Long-term consequences of maternal high-fat feeding on hypothalamic leptin sensitivity and diet-induced obesity in the offspring. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 293, R1056–R1062. [Google Scholar] [CrossRef] [PubMed]
- Howie, G.J.; Sloboda, D.M.; Kamal, T.; Vickers, M.H. Maternal nutritional history predicts obesity in adult offspring independent of postnatal diet. J. Physiol. 2009, 587, 905–915. [Google Scholar] [CrossRef] [PubMed]
- Khan, I.Y.; Taylor, P.D.; Dekou, V.; Seed, P.T.; Lakasing, L.; Graham, D.; Dominiczak, A.F.; Hanson, M.A.; Poston, L. Gender-linked hypertension in offspring of lard-fed pregnant rats. Hypertension 2003, 41, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Armitage, J.A.; Lakasing, L.; Taylor, P.D.; Balachandran, A.A.; Jensen, R.I.; Dekou, V.; Ashton, N.; Nyengaard, J.R.; Poston, L. Developmental programming of aortic and renal structure in offspring of rats fed fat-rich diets in pregnancy. J. Physiol. 2005, 565, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.; Alvers, K.M.; Crump, E.M.; Rowland, N.E. Effect of high-fat diet during gestation, lactation, or postweaning on physiological and behavioral indexes in borderline hypertensive rats. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2009, 296, R20–R28. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, S.; Arai, T.; Iwata, K.; Oka, M.; Nagase, M. A high-fat diet aggravates tubulointerstitial but not glomerular lesions in obese Zucker rats. Kidney Int. 1999, 71, S150–S152. [Google Scholar] [CrossRef]
- Tomat, A.L.; Salazar, F.J. Mechanisms involved in developmental programming of hypertension and renal diseases. Gender differences. Horm. Mol. Biol. Clin. Investig. 2014, 18, 63–77. [Google Scholar] [CrossRef] [PubMed]
- Elliot, S.J.; Berho, M.; Korach, K.; Doublier, S.; Lupia, E.; Striker, G.E.; Karl, M. Gender-specific effects of endogenous testosterone: Female alpha-estrogen receptor-deficient C57Bl/6J mice develop glomerulosclerosis. Kidney Int. 2007, 72, 464–472. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Wu, K.L.; Lee, W.C.; Leu, S.; Chan, J.Y. Maternal fructose-intake-induced renal programming in adult male offspring. J. Nutr. Biochem. 2015, 26, 642–650. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Lee, C.T.; Huang, L.T. Long-term effects of maternal citrulline supplementation on renal transcriptome prevention of nitric oxide depletion-related programmed hypertension: The impact of gene-nutrient interactions. Int. J. Mol. Sci. 2014, 15, 23255–23268. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.; Zhang, X.; Sieli, P.T.; Falduto, M.T.; Torres, K.E.; Rosenfeld, C.S. Contrasting effects of different maternal diets on sexually dimorphic gene expression in the murine placenta. Proc. Natl. Acad. Sci. USA 2010, 107, 5557–5562. [Google Scholar] [CrossRef] [PubMed]
- Cox, L.A.; Li, C.; Glenn, J.P.; Lange, K.; Spradling, K.D.; Nathanielsz, P.W.; Jansson, T. Expression of the placental transcriptome in maternal nutrient reduction in baboons is dependent on fetal sex. J. Nutr. 2013, 143, 1698–1708. [Google Scholar] [CrossRef] [PubMed]
- Pinto, V.; Pinho, M.J.; Soares-da-Silva, P. Renal amino acid transport systems and essential hypertension. FASEBJ. 2013, 27, 2927–2938. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Hsu, C.N. Targeting on asymmetric dimethylarginine-related nitric oxide-reactive oxygen species imbalance to reprogram the development of hypertension. Int. J. Mol. Sci. 2016, 17, E2020. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Ristow, M. Mitochondria and metabolic homeostasis. Antioxid. Redox Signal. 2013, 19, 240–242. [Google Scholar] [CrossRef] [PubMed]
- Bonny, O.; Vinciguerra, M.; Gumz, M.L.; Mazzoccoli, G. Molecular bases of circadian rhythmicity in renal physiology and pathology. Nephrol. Dial. Transplant. 2013, 28, 2421–2431. [Google Scholar] [CrossRef] [PubMed]
- Paauw, N.D.; van Rijn, B.B.; Lely, A.T.; Joles, J.A. Pregnancy as a critical window for blood pressure regulation in mother and child: Programming and reprogramming. Acta Physiol. 2017, 219, 241–259. [Google Scholar] [CrossRef] [PubMed]
- Oosterman, J.E.; Kalsbeek, A.; la Fleur, S.E.; Belsham, D.D. Impact of nutrients on circadian rhythmicity. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2015, 308, R337–R350. [Google Scholar] [CrossRef] [PubMed]
- Firsov, D.; Bonny, O. Circadian regulation of renal function. Kidney Int. 2010, 78, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.; Diaz, A.N.; Gumz, M.L. Clock genes in hypertension: Novel insights from rodent models. Blood Press. Monit. 2014, 19, 249–254. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Fu, O.; Yao, L.; Sun, L.; Zhuge, F.; Fu, Z. Differential responses of peripheral circadian clocks to a short-term feeding stimulus. Mol. Biol. Rep. 2012, 39, 9783–9789. [Google Scholar] [CrossRef] [PubMed]
- Kume, S.; Uzu, T.; Araki, S.; Sugimoto, T.; Isshiki, K.; Chin-Kanasaki, M.; Sakaguchi, M.; Kubota, N.; Terauchi, Y.; Kadowaki, T.; Haneda, M.; Kashiwagi, A.; Koya, D. Role of altered renal lipid metabolism in the development of renal injury induced by a high-fat diet. J. Am. Soc. Nephrol. 2007, 18, 2715–2723. [Google Scholar] [CrossRef] [PubMed]


| Gene | Forward | Reverse |
|---|---|---|
| Collagen I | 5 aggcataaagggtcatcgtg 3 | 5 accgttgagtccatctttgc 3 |
| α-SMA | 5 gaccctgaagtatccgatagaaca 3 | 5 cacgcgaagctcgttatagaag 3 |
| Ren | 5 aacattaccagggcaactttcact 3 | 5 acccccttcatggtgatctg 3 |
| Atp6ap2 | 5 gaggcagtgaccctcaacat 3 | 5 ccctcctcacacaacaaggt 3 |
| Agt | 5 gcccaggtcgcgatgat 3 | 5 tgtacaagatgctgagtgaggcaa 3 |
| Ace | 5 caccggcaaggtctgctt 3 | 5 cttggcatagtttcgtgaggaa 3 |
| Ace2 | 5 acccttcttacatcagccctactg 3 | 5 tgtccaaaacctaccccacatat 3 |
| Agtr1a | 5 gctgggcaacgagtttgtct 3 | 5 cagtccttcagctggatcttca 3 |
| Agtr1b | 5 caatctggctgtggctgactt 3 | 5 tgcacatcacaggtccaaaga 3 |
| Mas1 | 5 catctctcctctcggctttgtg 3 | 5 cctcatccggaagcaaagg 3 |
| Clock | 5 ccactgtacaatacgatggtgatctc 3 | 5 tgcggcatactggatggaat3 |
| Bmal1 | 5 attccagggggaaccaga 3 | 5 gaaggtgatgaccctcttatcct 3 |
| Per1 | 5 gcttgtgtggactgtggtagca 3 | 5 gccccaatccatccagttgt 3 |
| Per2 | 5 catctgccacctcagactca 3 | 5 ctggtgtgacttgtatcactgct 3 |
| Per3 | 5 tggccacagcatcagtaca 3 | 5 tacactgctggcactgcttc 3 |
| Cry1 | 5 atcgtgcgcatttcacatac 3 | 5 tccgccattgagttctatgat 3 |
| Cry2 | 5 gggagcatcagcaacacag 3 | 5 gcttccagcttgcgtttg 3 |
| Ck1e | 5 gcctctatcaacacccacct 3 | 5 ggagcccaggttgaagtaca 3 |
| Nr1d1 | 5 ctactggctccctcacccagga 3 | 5 gacactcggctgctgtcttcca 3 |
| Rn18s | 5 gccgcggtaattccagctcca 3 | 5 cccgcccgctcccaagatc 3 |
| Groups | ND/ND | ND/HF | HF/ND | HF/HF | p Value | |||
|---|---|---|---|---|---|---|---|---|
| Number | M = 6; F = 6 | M = 6; F = 6 | M = 6; F = 6 | M = 7; F = 6 | Pre | Post | Pre × Post | |
| Body weight (g) | Male | 641 ± 39 | 785 ± 39 * | 677 ± 21 | 813 ± 41*,$ | NS | 0.001 | NS |
| Female | 372 ± 17 | 362 ± 19 | 355 ± 14 | 372 ± 10 | NS | NS | NS | |
| Left kidney (LK) weight (g) | Male | 2.16 ± 0.14 | 2.12 ± 0.08 | 2.26 ± 0.08 | 2.08 ± 0.07 | NS | NS | NS |
| Female | 1.28 ± 0.04 | 1.45 ± 0.1 | 1.37 ± 0.05 | 1.38 ± 0.01 | NS | NS | NS | |
| LK weight/100 g BW | Male | 0.34 ± 0.02 | 0.27 ± 0.01 * | 0.33 ± 0.01 # | 0.26 ± 0.01*,$ | NS | <0.001 | NS |
| Female | 0.35 ± 0.02 | 0.4 ± 0.02 | 0.39 ± 0.01 | 0.37 ± 0.01 | NS | NS | 0.021 | |
| AST (U/L) | Male | 88 ± 11 | 308 ± 58 * | 82 ± 10 # | 145 ± 19 # | 0.019 | <0.001 | 0.028 |
| Female | 73 ± 3 | 160 ± 24 | 83 ± 12 | 82 ± 8 | 0.026 | 0.007 | 0.006 | |
| ALT (U/L) | Male | 27 ± 3 | 196 ± 52 * | 23 ± 2 # | 66 ± 17 # | 0.031 | 0.002 | 0.041 |
| Female | 19 ± 2 | 67 ± 13 | 22 ± 3 | 30 ± 4 | 0.026 | 0.001 | 0.009 | |
| Total cholesterol (mg/dL) | Male | 71 ± 8 | 82 ± 7 | 58 ± 5 | 65 ± 5 | 0.027 | NS | NS |
| Female | 81 ± 9 | 95 ± 4 | 104 ± 7 | 95 ± 16 | NS | NS | NS | |
| Triglyceride (mg/dL) | Male | 101 ± 26 | 61 ± 13 | 105 ± 9 | 87 ± 12 | NS | NS | NS |
| Female | 97 ± 23 | 58 ± 9 | 120 ± 23 # | 60 ± 8 | NS | NS | NS | |
| HDL (mg/dL) | Male | 43 ± 4 | 49 ± 6 | 34 ± 4 | 42 ± 4 | NS | NS | NS |
| Female | 39 ± 4 | 52 ± 2 | 59 ± 4 | 58 ± 10 | NS | NS | NS | |
| Glucose (mg/dL) | Male | 81 ± 2 | 91 ± 3 | 93 ± 4 | 81 ± 3 | NS | NS | NS |
| Female | 75 ± 4 | 76 ± 1 | 73 ± 2 | 62 ± 3 # | NS | NS | NS | |
| IPGTT (AUC, mg/dL·120 min) | Male | 22,071 ± 1354 | 23,923 ± 2345 | 23,498 ± 2286 | 25,922 ± 1973 | - | - | - |
| Female | 26,420 ± 1406 | 31,389 ± 1773 * | 26,890 ± 1820 | 26,949 ± 2416 | - | - | - | |
| KEGG Pathway | Count | Gene Symbol | p-Value | Benjamini |
|---|---|---|---|---|
| Male | ||||
| Protein digestion and absorption | 2 | Slc15a1, Slc6a19 | 5.6 × 10−2 | 5.6 × 10−2 |
| Female | ||||
| Oxidative phosphorylation | 5 | Atp5j2, Atp6v0d2, Ndufa5, Cox6c, Cox7c | 1.6 × 10−2 | 1.6 × 10−2 |
| Protein digestion and absorption | 4 | Slc15a1, Slc6a19, Slc7a7, Slc7a8 | 2.2 × 10−2 | 2.2 × 10−2 |
| Metabolic pathways | 16 | Dhcr24, Abat, Atp5j2, Atp6v0d2, C1qalt1c1, Mgat4c, Ndufa5, Alox15, Cyp24a1, Cox6c, Cox7c, Dse, Dqkq, Gatm, Hykk, Polr2k | 2.3 × 10−2 | 2.3 × 10−2 |
| Ribosome | 5 | Mrpl33, Mrps18c, Rpl22l1, Rpl30, LOC100362027 | 2.9 × 10−2 | 2.9 × 10−2 |
| Cardiac muscle contraction | 3 | Cacna2d2, Cox6c, Cox7c | 9.9 × 10−2 | 9.9 × 10−2 |
| Groups | ND/ND | ND/HF | HF/ND | HF/HF | P Value | |||
|---|---|---|---|---|---|---|---|---|
| Number | M = 5; F = 6 | M = 6; F = 6 | M = 6; F = 6 | M = 7; F = 6 | Pre | Post | Pre × Post | |
| l-Citrulline (μM) | Male | 42.4 ± 2.2 | 42.0 ± 2.0 | 41.6 ± 1.6 | 46.2 ± 1.9 | NS | NS | NS |
| Female | 48.7 ± 4.1 | 61.7 ± 9.5 | 44.5 ± 4.8 | 62.9 ± 4.7 | NS | 0.019 | NS | |
| l-Arginine (μM) | Male | 168.0 ± 15.9 | 46.1 ± 12.8 * | 172.0 ± 9.5 # | 101.8 ± 15.5 | 0.041 | <0.001 | NS |
| Female | 152.4 ± 27.7 | 120.8 ± 23.4 | 179.0 ± 10.2 | 119.6 ± 5.2 | NS | 0.027 | NS | |
| ADMA (μM) | Male | 1.02 ± 0.03 | 0.92 ± 0.03 | 1.23 ± 0.03 # | 1.03 ± 0.07 | 0.001 | 0.003 | NS |
| Female | 1.45 ± 0.09 | 1.25 ± 0.07 | 1.38 ± 0.03 | 1.1 ± 0.04 *,$ | NS | 0.001 | NS | |
| SDMA(μM) | Male | 0.43 ± 0.03 | 0.55 ± 0.02 | 0.67 ± 0.03 * | 0.53 ± 0.02 | 0.001 | NS | <0.001 |
| Female | 0.72 ± 0.08 | 0.65 ± 0.03 | 0.68 ± 0.05 | 0.5 ± 0.03* | NS | 0.028 | NS | |
| l-Arginine-to-ADMA ratio (μM/μM) | Male | 165.7 ± 15.1 | 49.7 ± 13.9 * | 138.6 ± 7.7 # | 101.7 ± 17.6* | NS | <0.001 | 0.011 |
| Female | 107.2 ± 20.8 | 93.3 ± 16.0 | 129.1 ± 9.6 | 107.9 ± 3.0 | NS | NS | NS | |
| NOx (NO2− + NO−) (μM) | Male | 218.6 ± 15.1 | 167.5 ± 5.1 | 195.7 ± 6.6 | 178 ± 7.4 | NS | 0.001 | NS |
| Female | 172.8 ± 16.5 | 161.5 ± 17.6 | 176.6 ± 16.5 | 159.1 ± 24.5 | NS | NS | NS | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tain, Y.-L.; Lin, Y.-J.; Sheen, J.-M.; Yu, H.-R.; Tiao, M.-M.; Chen, C.-C.; Tsai, C.-C.; Huang, L.-T.; Hsu, C.-N. High Fat Diets Sex-Specifically Affect the Renal Transcriptome and Program Obesity, Kidney Injury, and Hypertension in the Offspring. Nutrients 2017, 9, 357. https://doi.org/10.3390/nu9040357
Tain Y-L, Lin Y-J, Sheen J-M, Yu H-R, Tiao M-M, Chen C-C, Tsai C-C, Huang L-T, Hsu C-N. High Fat Diets Sex-Specifically Affect the Renal Transcriptome and Program Obesity, Kidney Injury, and Hypertension in the Offspring. Nutrients. 2017; 9(4):357. https://doi.org/10.3390/nu9040357
Chicago/Turabian StyleTain, You-Lin, Yu-Ju Lin, Jiunn-Ming Sheen, Hong-Ren Yu, Mao-Meng Tiao, Chih-Cheng Chen, Ching-Chou Tsai, Li-Tung Huang, and Chien-Ning Hsu. 2017. "High Fat Diets Sex-Specifically Affect the Renal Transcriptome and Program Obesity, Kidney Injury, and Hypertension in the Offspring" Nutrients 9, no. 4: 357. https://doi.org/10.3390/nu9040357
APA StyleTain, Y.-L., Lin, Y.-J., Sheen, J.-M., Yu, H.-R., Tiao, M.-M., Chen, C.-C., Tsai, C.-C., Huang, L.-T., & Hsu, C.-N. (2017). High Fat Diets Sex-Specifically Affect the Renal Transcriptome and Program Obesity, Kidney Injury, and Hypertension in the Offspring. Nutrients, 9(4), 357. https://doi.org/10.3390/nu9040357








