The Role of Dietary Histone Deacetylases (HDACs) Inhibitors in Health and Disease
Abstract
:1. Introduction
2. Histones
2.1. A Brief History
2.2. Modifications
3. Histone Modifying Enzymes
| Modification Type | Amino Acid Modified | Abbreviation | Examples of Modifying Enzymes Identified in Humans | Role |
|---|---|---|---|---|
| Acetylation | Lysine | K-ac | Histone Acetyltransferases (HATs): e.g., HAT1 Histone Deacetylases (HDACs): e.g., HDAC1 | Transcription, Repair, Replication, Condensation |
| Methylation | Lysine Arginine | K-me1, -me2, -me3 R-me1, -me2 | Lysine Methyltransferases: e.g., SUV39H1 Lysine Demethylases: e.g., LSD1/BHC110 Arginine Methyltransferases: e.g., PRMT4 Arginine Demethylases: e.g., JMJD6 | Transcription, Repair Transcription |
| Phosphorylation | Serine Threonine | S-ph T-ph | Serine/Threonine Kinases: e,g. WEE1 Dephosphorylated by Phosphatases: e.g., PP4 | Transcription, Repair, Condensation |
| Ubiquitination | Lysine | K-ub | Ubiquinases (Ubiquitin Ligases): e.g., RING1B Deubiquinating Enzymes: e.g., USP22 | Transcription, Repair |
| SUMOylation | Lysine | K-su | Small Ubiquitin-like Modifier (SUMO) proteins: e.g., SUMO-1 De-SUMOylating Enzymes: Sentrin-Specific Proteases: e.g., SENP1 | Transcription |
| ADP ribosylation | Glutamate | E-ar | ADP-Ribosyltransferases: e.g., ARTD1 (PARP1) | Transcription |
| Deimination | Arginine (to Citrulline) | K to Cit | Peptidylarginine Deiminases: e.g., PADI4 | Transcription |
| Proline Isomerisation | Proline | P-cis to P-trans | Proline Isomerases: e.g., Pin1 | Transcription |
3.1. Acetylation
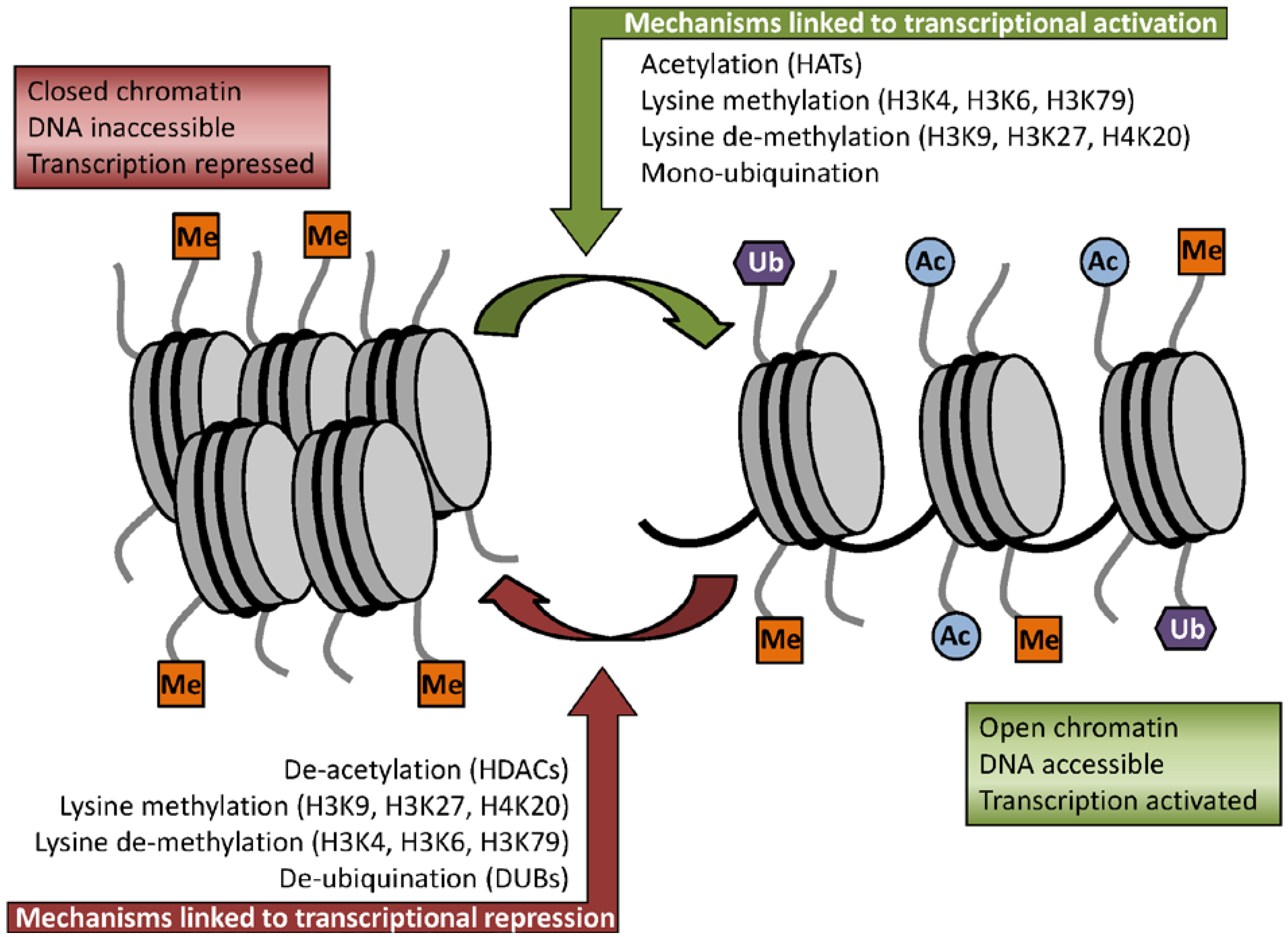
3.2. Methylation
3.3. Ubiquitination
3.4. SUMOylation
3.5. Other Modifications
4. HDACs in Health and Disease
4.1. Class I HDACs (HDACs 1–3 and HDAC8)
4.2. Class IIa HDACs (HDACs 4, 5, 7 and 9)
4.3. Class IIb HDACs (HDAC6 and 10)
4.4. Class III HDACs (SIRT1–7)
4.5. Class IV HDACs (HDAC11)
| Name | Type of dysregulation | Disease Implicated | Reference(s) | |
|---|---|---|---|---|
| Class I | HDAC1 | HDAC1 overexpression | Prostate cancer Gastric cancer Ovarian cancer Hodgkin’s lymphoma | [53,68] [54,98] [51] [52] |
| HDAC1 underexpression | Colorectal cancer | [55] | ||
| HDAC2 | Truncating HDAC2 mutation | Colonic, gastric and endometrial cancers | [55,56] | |
| HDAC2 overexpression | Ovarian cancer Hodgkin’s lymphoma | [51] [52] | ||
| Reduction in activity and expression | Chronic obstructive pulmonary disease | [57] | ||
| HDAC3 | SNP variants | Type 2 diabetes Schizophrenia | [60] [64] | |
| Liver-specific deletion | Severe hepatosteatosis and increase in insulin sensitivity | [61] | ||
| HDAC3 overexpression | Ovarian cancer | [51] | ||
| Increased HDAC3 protein expression | Hodgkin’s lymphoma Colon cancer | [52] [99] | ||
| HDAC8 | HDAC8 mutations | Cornelia de Lange disease | [65] | |
| Class IIa | HDAC4 | Splice-site/missense mutations | Breast cancer Eating disorders | [66] [73] |
| SNP variant | Lung function Schizophrenia | [69] [64] | ||
| HDAC4 overexpression | Prostate cancer Colon cancer Lung cancer Breast cancer | [68] [67] [67] [55] | ||
| Haploinsufficiency | Pyschomotor and behavioural abnormalities | [70,71] | ||
| Reduction | Huntington’s disease | [72] | ||
| HDAC5 | HDAC5 underexpression HDAC5 overexpression | Colorectal cancer Major depression | [55] [74] | |
| HDAC7 | Over expression | Colorectal cancer Pancreatic cancer | [55] [75] | |
| HDAC9 | Gene variants Gene disruption | Multiple sclerosis Peters’ anomaly | [78] [77] | |
| Class IIb | HDAC6 | HDAC6 overexpression | Neurodegenerative diseases (e.g., Alzheimer’s disease) | [3,79] |
| Increased HDAC6 protein expression | Polycystic liver disease | [80] | ||
| X-linked HDAC6 mutation | Adult-onset Alexander disease | [100] | ||
| Little or no HDAC6 expression | Hodgkin’s lymphoma | [81] | ||
| HDAC10 | HDAC10 overexpression | Chronic lymphocytic leukemia | [82] | |
| Class III | SIRT1 | Overexpression | Breast, colorectal and prostate cancer | [86] |
| SIRT1 underexpression | Colorectal cancer | [55] | ||
| SIRT2 | Polymorphism | Alzheimer’s disease | [91] | |
| SIRT3 | mRNA and protein underexpression | Gastric cancer | [94] | |
| SIRT4 | Gene variants | Multiple sclerosis | [78] | |
| SIRT5 | Gene variants SIRT5 overexpression | Multiple sclerosis Alzheimer’s disease | [78] [87] | |
| SIRT6 | Decreased SIRT6 expression | Liver cancer and cirrhotic livers | [101] | |
| SIRT7 | SIRT7 overexpression | Breast cancer | [96] | |
| Class IV | HDAC11 | Gene variants | Multiple sclerosis | [78] |
5. HDAC Inhibitors
5.1. Naturally Occurring HDAC Inhibitors
5.2. Synthetic HDAC Inhibitors
5.3. Use of HDAC Inhibitors in Treating Disease
6. Dietary HDAC Inhibitors as Therapeutics
| Dietary Component | Food Source | References |
|---|---|---|
| Allicin | Garlic | [133] |
| Bis-(4-hydroxybenzyl)sulfide | Polygonaceae (Pleuropterus ciliinervis Nakai) | [134] |
| Caffeic acid | Intestinal metabolite of nutritional polyphenols | [135] |
| Catechins (e.g., green tea polyphenols) | Tea (Camellia sinensis), particularly green tea | [136] |
| Coumaric/hydroxycinnamic acid | Cinnamon | [135] |
| Curcumin | Turmeric | [137] |
| Diallyl disulfide | Garlic | [138,139] |
| 3,3ʹ-di-indolylmethane | Cruciferous vegetables, e.g., broccoli | [129] |
| Equol | Soy | [140] |
| Flavonoids, e.g., Apigenin Chrysin | Common fruits and vegetables, e.g., grapefruit, parsley and chamomile Fruits, vegetables, olive oil and red wine | [141] [142] |
| Genistein | Soy | [140,143] |
| Isothiocyanates (e.g., sulforaphane) | Cruciferous vegetables, e.g., broccoli | [144,145] |
| MCP30 | Bitter melon | [131] |
| Organoselenium compounds e.g., Se-methyl-l-selenocysteine | Broccoli, Garlic, Onion | [125] |
| Parthenolide | Feverfew (Tanacetum parthenium) | [132] |
| Pomiferin | Osage orange/Hedge apple (Maclura pomifera) | [146] |
| Quercetin | Citrus, apple, berries | [147] |
| Resveratrol | Grapes, wine, eucalyptus | [148,149] |
| Selenium compounds | Brazil nuts | [124,127] |
| Sesquiterpenoids | Shampoo Ginger (Zingiber zerumbet) | [150] |
| Ursolic acid | Basil | [151] |
Effect of Cooking on HDACi Activity
7. Conclusions and Future Directions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Blewitt, M.; Whitelaw, E. The use of mouse models to study epigenetics. Cold Spring Harb. Perspect. Biol. 2013, 5, a017939. [Google Scholar]
- Coppede, F. The role of epigenetics in colorectal cancer. Expert Rev. Gastroenterol. Hepatol. 2014. [Google Scholar] [CrossRef]
- Simoes-Pires, C.; Zwick, V.; Nurisso, A.; Schenker, E.; Carrupt, P.-A.; Cuendet, M. HDAC6 as a target for neurodegenerative diseases: What makes it different from the other HDACs? Mol. Neurodegener. 2013, 8, 7. [Google Scholar]
- Glauben, R.; Batra, A.; Stroh, T.; Erben, U.; Fedke, I.; Lehr, H.A.; Leoni, F.; Mascagni, P.; Dinarello, C.A.; Zeitz, M.; et al. Histone deacetylases: Novel targets for prevention of colitis-associated cancer in mice. Gut 2008, 57, 613–622. [Google Scholar]
- Dokmanovic, M.; Clarke, C.; Marks, P.A. Histone deacetylase inhibitors: Overview and perspectives. Mol. Cancer Res. 2007, 5, 981–989. [Google Scholar]
- Kim, H.J.; Bae, S.C. Histone deacetylase inhibitors: Molecular mechanisms of action and clinical trials as anti-cancer drugs. Am. J. Transl. Res. 2011, 3, 166–179. [Google Scholar]
- Levene, P.A. On the chemistry of the chromatin substance of the nerve cell. J. Med. Res. 1903, 10, 204–211. [Google Scholar]
- Moore, B.; Whitley, E.; Webster, A. The basic and acidic proteins of the sperm of Echinus esculentus. Direct measurements of the osmotic pressure of a protamine or histone. Biochem. J. 1913, 7, 142–147. [Google Scholar]
- Mirsky, A.E.; Pollister, A.W. Chromosin, a desoxyribose nucleoprotein complex of the cell nucleus. J. Gen. Physiol. 1946, 30, 117–148. [Google Scholar]
- Mellor, J. Dynamic nucleosomes and gene transcription. Trends Genet. 2006, 22, 320–329. [Google Scholar]
- Gershey, E.L.; Vidali, G.; Allfrey, V.G. Chemical studies of histone acetylation. The occurrence of epsilon-N-acetyllysine in the f2a1 histone. J. Biol. Chem. 1968, 243, 5018–5022. [Google Scholar]
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar]
- Andreu-Vieyra, C.V.; Liang, G. Nucleosome occupancy and gene regulation during tumorigenesis. Adv. Exp. Med. Biol. 2013, 754, 109–134. [Google Scholar]
- Barnett, M.P.G.; Bassett, S.A.; Bermingham, E.N. Epigenetics—What role could this play in functional foods and personalized nutrition? In Nutrigenomics and Nutrigenetics in Functional Foods and Personalized Nutrition; Ferguson, L.R., Ed.; CRC Press: Florence, KY, USA, 2013; pp. 243–267. [Google Scholar]
- Haberland, M.; Montgomery, R.L.; Olson, E.N. The many roles of histone deacetylases in development and physiology: Implications for disease and therapy. Nat. Rev. Genet. 2009, 10, 32–42. [Google Scholar]
- Ho, E.; Dashwood, R.H. Dietary manipulation of histone structure and function. World Rev. Nutr. Diet. 2010, 101, 95–102. [Google Scholar]
- Xu, F.; Zhang, K.; Grunstein, M. Acetylation in histone H3 globular domain regulates gene expression in yeast. Cell 2005, 121, 375–385. [Google Scholar]
- Glozak, M.A.; Seto, E. Histone deacetylases and cancer. Oncogene 2007, 26, 5420–5432. [Google Scholar]
- Xu, W.S.; Parmigiani, R.B.; Marks, P.A. Histone deacetylase inhibitors: Molecular mechanisms of action. Oncogene 2007, 26, 5541–5552. [Google Scholar]
- Vannini, A.; Volpari, C.; Filocamo, G.; Casavola, E.C.; Brunetti, M.; Renzoni, D.; Chakravarty, P.; Paolini, C.; De Francesco, R.; Gallinari, P.; et al. Crystal structure of a eukaryotic zinc-dependent histone deacetylase, human HDAC8, complexed with a hydroxamic acid inhibitor. Proc. Natl. Acad. Sci. USA 2004, 101, 15064–15069. [Google Scholar]
- Yamamoto, H.; Schoonjans, K.; Auwerx, J. Sirtuin functions in health and disease. Mol. Endocrinol. 2007, 21, 1745–1755. [Google Scholar]
- Blander, G.; Guarente, L. The Sir2 family of protein deacetylases. Annu. Rev. Biochem. 2004, 73, 417–435. [Google Scholar]
- Howitz, K.T.; Bitterman, K.J.; Cohen, H.Y.; Lamming, D.W.; Lavu, S.; Wood, J.G.; Zipkin, R.E.; Chung, P.; Kisielewski, A.; Zhang, L.L.; et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 2003, 425, 191–196. [Google Scholar]
- Porcu, M.; Chiarugi, A. The emerging therapeutic potential of sirtuin-interacting drugs: From cell death to lifespan extension. Trends Pharmacol. Sci. 2005, 26, 94–103. [Google Scholar]
- Shoba, B.; Lwin, Z.M.; Ling, L.S.; Bay, B.H.; Yip, G.W.; Kumar, S.D. Function of sirtuins in biological tissues. Anat. Rec. 2009, 292, 536–543. [Google Scholar]
- Huang, J.Y.; Hirschey, M.D.; Shimazu, T.; Ho, L.; Verdin, E. Mitochondrial sirtuins. Biochim. Biophys. Acta 2010, 8, 7. [Google Scholar]
- Verdin, E.; Hirschey, M.D.; Finley, L.W.; Haigis, M.C. Sirtuin regulation of mitochondria: Energy production, apoptosis, and signaling. Trends Biochem. Sci. 2010, 35, 669–675. [Google Scholar]
- Adimoolam, S.; Sirisawad, M.; Chen, J.; Thiemann, P.; Ford, J.M.; Buggy, J.J. HDAC inhibitor PCI-24781 decreases RAD51 expression and inhibits homologous recombination. Proc. Natl. Acad. Sci. USA 2007, 104, 19482–19487. [Google Scholar]
- Claus, R.; Lubbert, M. Epigenetic targets in hematopoietic malignancies. Oncogene 2003, 22, 6489–6496. [Google Scholar]
- Khan, O.; La Thangue, N.B. Drug Insight: Histone deacetylase inhibitor-based therapies for cutaneous T-cell lymphomas. Nat. Clin. Prac. Oncol. 2008, 5, 714–726. [Google Scholar]
- Bannister, A.J.; Kouzarides, T. Reversing histone methylation. Nature 2005, 436, 1103–1106. [Google Scholar]
- Nottke, A.; Colaiácovo, M.P.; Shi, Y. Developmental roles of the histone lysine demethylases. Dev. Change 2009, 136, 879–889. [Google Scholar]
- Chang, B.; Chen, Y.; Zhao, Y.; Bruick, R.K. JMJD6 is a histone arginine demethylase. Science 2007, 318, 444–447. [Google Scholar]
- Cao, R.; Tsukada, Y.I.; Zhang, Y. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol. Cell 2005, 20, 845–854. [Google Scholar]
- Wang, H.; Zhai, L.; Xu, J.; Joo, H.Y.; Jackson, S.; Erdjument-Bromage, H.; Tempst, P.; Xiong, Y.; Zhang, Y. Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage. Mol. Cell 2006, 22, 383–394. [Google Scholar]
- Cao, J.; Yan, Q. Histone ubiquitination and deubiquitination in transcription, DNA damage response, and cancer. Front. Oncol. 2012, 2, 26. [Google Scholar]
- Liu, H.W.; Zhang, J.; Heine, G.F.; Arora, M.; Gulcin Ozer, H.; Onti-Srinivasan, R.; Huang, K.; Parvin, J.D. Chromatin modification by SUMO-1 stimulates the promoters of translation machinery genes. Nucleic Acids Res. 2012, 40, 10172–10186. [Google Scholar]
- Drag, M.; Salvesen, G.S. DeSUMOylating enzymes—SENPs. IUBMB Life 2008, 60, 734–742. [Google Scholar]
- Fillingham, J.; Keogh, M.C.; Krogan, N.J. GammaH2AX and its role in DNA double-strand break repair. Biochem. Cell Biol. 2006, 84, 568–577. [Google Scholar]
- Rossetto, D.; Avvakumov, N.; Côté, J. Histone phosphorylation: A chromatin modification involved in diverse nuclear events. Epigenetics 2012, 7, 1098–1108. [Google Scholar]
- Nakada, S.; Chen, G.I.; Gingras, A.C.; Durocher, D. PP4 is a γH2AX phosphatase required for recovery from the DNA damage checkpoint. EMBO Rep. 2008, 9, 1019–1026. [Google Scholar]
- Denis, H.; Deplus, R.; Putmans, P.; Yamada, M.; Métivier, R.; Fuks, F. Functional connection between deimination and deacetylation of histones. Mol. Cell. Biol. 2009, 29, 4982–4993. [Google Scholar]
- Cuthbert, G.L.; Daujat, S.; Snowden, A.W.; Erdjument-Bromage, H.; Hagiwara, T.; Yamada, M.; Schneider, R.; Gregory, P.D.; Tempst, P.; Bannister, A.J.; Kouzarides, T. Histone deimination antagonizes arginine methylation. Cell 2004, 118, 545–553. [Google Scholar]
- Wang, Y.; Wysocka, J.; Sayegh, J.; Lee, Y.H.; Perlin, J.R.; Leonelli, L.; Sonbuchner, L.S.; McDonald, C.H.; Cook, R.G.; Dou, Y.; Roeder, R.G.; Clarke, S.; Stallcup, M.R.; Allis, C.D.; Coonrod, S.A. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 2004, 306, 279–283. [Google Scholar]
- Hassa, P.O.; Haenni, S.S.; Elser, M.; Hottiger, M.O. Nuclear ADP-ribosylation reactions in mammalian cells: Where are we today and where are we going? Microbiol. Mol. Biol. Rev. 2006, 70, 789–829. [Google Scholar]
- Kassner, I.; Barandun, M.; Fey, M.; Rosenthal, F.; Hottiger, M.O. Crosstalk between SET7/9-dependent methylation and ARTD1-mediated ADP-ribosylation of histone H1.4. Epigenetics Chromatin 2013, 6, 1. [Google Scholar]
- Nelson, C.J.; Santos-Rosa, H.; Kouzarides, T. Proline isomerization of histone H3 regulates lysine methylation and gene expression. Cell 2006, 126, 905–916. [Google Scholar]
- Raghuram, N.; Strickfaden, H.; McDonald, D.; Williams, K.; Fang, H.; Mizzen, C.; Hayes, J.J.; Th’ng, J.; Hendzel, M.J. Pin1 promotes histone H1 dephosphorylation and stabilizes its binding to chromatin. J. Cell Biol. 2013, 203, 57–71. [Google Scholar]
- Newkirk, T.L.; Bowers, A.A.; Williams, R.M. Discovery, biological activity, synthesis and potential therapeutic utility of naturally occurring histone deacetylase inhibitors. Nat. Prod. Rep. 2009, 26, 1293–1320. [Google Scholar]
- Gui, C.-Y.; Ngo, L.; Xu, W.S.; Richon, V.M.; Marks, P.A. Histone deacetylase (HDAC) inhibitor activation of p21WAF1 involves changes in promoter-associated proteins, including HDAC1. Proc. Natl. Acad. Sci. USA 2004, 101, 1241–1246. [Google Scholar]
- Jin, K.L.; Pak, J.H.; Park, J.-Y.; Choi, W.H.; Lee, J.-Y.; Kim, J.-H.; Nam, J.-H. Expression profile of histone deacetylases 1, 2 and 3 in ovarian cancer tissues. J. Gynecol. Oncol. 2008, 19, 185–190. [Google Scholar]
- Adams, H.; Fritzsche, F.R.; Dirnhofer, S.; Kristiansen, G.; Tzankov, A. Class I histone deacetylases 1, 2 and 3 are highly expressed in classical Hodgkin′s lymphoma. Expert Opin. Ther. Targets 2010, 14, 577–584. [Google Scholar]
- Song, Y.; Shiota, M.; Tamiya, S.; Kuroiwa, K.; Naito, S.; Tsuneyoshi, M. The significance of strong histone deacetylase 1 expression in the progression of prostate cancer. Histopathology 2011, 58, 773–780. [Google Scholar]
- Sudo, T.; Mimori, K.; Nishida, N.; Kogo, R.; Iwaya, T.; Tanaka, F.; Shibata, K.; Fujita, H.; Shirouzu, K.; Mori, M. Histone deacetylase 1 expression in gastric cancer. Oncol. Rep. 2011, 26, 777–782. [Google Scholar]
- Ozdag, H.; Teschendorff, A.; Ahmed, A.; Hyland, S.; Blenkiron, C.; Bobrow, L.; Veerakumarasivam, A.; Burtt, G.; Subkhankulova, T.; Arends, M.; et al. Differential expression of selected histone modifier genes in human solid cancers. BMC Genomics 2006, 7, 90. [Google Scholar]
- Ropero, S.; Fraga, M.F.; Ballestar, E.; Hamelin, R.; Yamamoto, H.; Boix-Chornet, M.; Caballero, R.; Alaminos, M.; Setien, F.; Paz, M.F.; et al. A truncating mutation of HDAC2 in human cancers confers resistance to histone deacetylase inhibition. Nat. Genet. 2006, 38, 566–569. [Google Scholar]
- Barnes, P.J.; Adcock, I.M.; Ito, K. Histone acetylation and deacetylation: Importance in inflammatory lung diseases. Eur. Respir. J. 2005, 25, 552–563. [Google Scholar]
- Khan, N.; Jeffers, M.; Kumar, S.; Hackett, C.; Boldog, F.; Khramtsov, N.; Qian, X.; Mills, E.; Berghs, S.C.; Carey, N.; et al. Determination of the class and isoform selectivity of small-molecule histone deacetylase inhibitors. Biochem. J. 2008, 409, 581–589. [Google Scholar]
- Noh, H.; Oh, E.Y.; Seo, J.Y.; Yu, M.R.; Kim, Y.O.; Ha, H.; Lee, H.B. Histone deacetylase-2 is a key regulator of diabetes- and transforming growth factor-β1-induced renal injury. Am. J. Physiol. Renal Physiol. 2009, 207, F729–F739. [Google Scholar]
- Zeng, Z.; Liao, R.; Yao, Z.; Zhou, W.; Ye, P.; Zheng, X.; Li, X.; Huang, Y.; Chen, S.; Chen, Q. Three single nucleotide variants of the HDAC gene are associated with type 2 diabetes mellitus in a Chinese population: A community-based case-control study. Gene 2014, 533, 427–433. [Google Scholar]
- Sun, Z.; Miller, R.A.; Patel, R.T.; Chen, J.; Dhir, R.; Wang, H.; Zhang, D.; Graham, M.J.; Unterman, T.G.; Shulman, G.I.; et al. Hepatic Hdac3 promotes gluconeogenesis by repressing lipid synthesis and sequestration. Nat. Med. 2012, 18, 934–942. [Google Scholar]
- Yang, X.-J.; Grégoire, S. Class II histone deacetylases: From sequence to function, regulation, and clinical implication. Mol. Cell. Biol. 2005, 25, 2873–2884. [Google Scholar]
- Wang, W.-H.; Cheng, L.-C.; Pan, F.-Y.; Xue, B.; Wang, D.-Y.; Chen, Z.; Li, C.-J. Intracellular trafficking of histone deacetylase 4 regulates long-term memory formation. Anat. Rec. Hoboken 2011, 294, 1025–1034. [Google Scholar]
- Kim, T.; Park, J.K.; Kim, H.-J.; Chung, J.-H.; Kim, J.W. Association of histone deacetylase genes with schizophrenia in Korean population. Psychiatry Res. 2010, 178, 266–269. [Google Scholar]
- Deardorff, M.A.; Bando, M.; Nakato, R.; Watrin, E.; Itoh, T.; Minamino, M.; Saitoh, K.; Komata, M.; Katou, Y.; Clark, D.; et al. HDAC8 mutations in Cornelia de Lange syndrome affect the cohesin acetylation cycle. Nature 2012, 489, 313–317. [Google Scholar]
- Sjöblom, T.; Jones, S.; Wood, L.D.; Parsons, D.W.; Lin, J.; Barber, T.D.; Mandelker, D.; Leary, R.J.; Ptak, J.; Silliman, N.; et al. The consensus coding sequences of human breast and colorectal cancers. Science 2006, 314, 268–274. [Google Scholar]
- LLeonart, M.; Vidal, F.; Gallardo, D.; Diaz-Fuertes, M.; Rojo, F.; Cuatrecasas, M.; López-Vicente, L.; Kondoh, H.; Blanco, C.; Carnero, A.; et al. New p53 related genes in human tumors: Significant downregulation in colon and lung carcinomas. Oncol. Rep. 2006, 16, 603–608. [Google Scholar]
- Halkidou, K.; Gaughan, L.; Cook, S.; Leung, H.Y.; Neal, D.E.; Robson, C.N. Upregulation and nuclear recruitment of HDAC1 in hormone refractory prostate cancer. Prostate 2004, 59, 177–189. [Google Scholar]
- Collins, S.A.; Lucas, J.S.A.; Inskip, H.M.; Godfrey, K.M.; Roberts, G.; Holloway, J.W.; The Southampton Women’s Survey Study Group. HHIP, HDAC4, NCR3 and RARB polymorphisms affect fetal, childhood and adult lung function. Eur. Respir. J. 2013, 41, 756–757. [Google Scholar]
- Villavicencio-Lorini, P.; Klopocki, E.; Trimborn, M.; Koll, R.; Mundlos, S.; Horn, D. Phenotypic variant of Brachydactyly-mental retardation syndrome in a family with an inherited interstitial 2q37.3 microdeletion including HDAC4. Eur. J. Hum. Genet. 2013, 21, 743–748. [Google Scholar]
- Williams, S.R.; Aldred, M.A.; Der Kaloustian, V.M.; Halal, F.; Gowans, G.; McLeod, D.R.; Zondag, S.; Toriello, H.V.; Magenis, R.E.; Elsea, S.H. Haploinsufficiency of HDAC4 causes Brachydactyly mental retardation syndrome, with Brachydactyly type E, developmental delays, and behavioral problems. Am. J. Hum. Genet. 2010, 87, 219–228. [Google Scholar]
- Mielcarek, M.; Landles, C.; Weiss, A.; Bradaia, A.; Seredenina, T.; Inuabasi, L.; Osborne, G.F.; Wadel, K.; Touller, C.; Butler, R.; et al. HDAC4 reduction: A novel therapeutic strategy to target cytoplasmic huntingtin and ameliorate neurodegeneration. PLoS Biol. 2013, 11, e1001717. [Google Scholar]
- Cui, H.; Moore, J.; Ashimi, S.S.; Mason, B.L.; Drawbridge, J.N.; Han, S.; Hing, B.; Matthews, A.; McAdams, C.J.; Darbro, B.W.; Pieper, A.A.; Waller, D.A.; Xing, C.; Lutter, M. Eating disorder predisposition is associated with ESRRA and HDAC4 mutations. J. Clin. Investig. 2013, 123, 4706–4713. [Google Scholar]
- Iga, J.-I.; Ueno, S.-I.; Yamauchi, K.; Numata, S.; Kinouchi, S.; Tayoshi-Shibuya, S.; Song, H.; Ohmori, T. Altered HDAC5 and CREB mRNA expressions in the peripheral leukocytes of major depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 2007, 31, 628–632. [Google Scholar]
- Ouaissi, M.; Sielezneff, I.; Silvestre, R.; Sastre, B.; Bernard, J.P.; Lafontaine, J.S.; Payan, M.J.; Dahan, L.; Pirro, N.; Seitz, J.F.; Mas, E.; Lombardo, D.; Ouaissi, A. High histone deacetylase 7 (HDAC7) expression is significantly associated with adenocarcinomas of the pancreas. Ann. Surg. Oncol. 2008, 15, 2318–2328. [Google Scholar]
- Yan, K.; Cao, Q.; Reilly, C.M.; Young, N.L.; Garcia, B.A.; Mishra, N. Histone deacetylase 9 deficiency protects against effector T cell-mediated systemic autoimmunity. J. Biol. Chem. 2011, 286, 28833–28843. [Google Scholar]
- David, D.; Cardoso, J.; Marques, B.á.; Marques, R.; Silva, E.D.; Santos, H.; Boavida, M.G. Molecular characterization of a familial translocation implicates disruption of HDAC9 and possible position effect on TGFβ2 in the pathogenesis of Peters’ anomaly. Genomics 2003, 81, 489–503. [Google Scholar]
- Inkster, B.; Strijbis, E.M.M.; Vounou, M.; Kappos, L.; Radue, E.-W.; Matthews, P.M.; Uitdehaag, B.M.J.; Barkhof, F.; Polman, C.H.; Montana, G.; et al. Histone deacetylase gene variants predict brain volume changes in multiple sclerosis. Neurobiol. Aging 2013, 34, 238–247. [Google Scholar]
- Zhang, L.; Sheng, S.; Qin, C. The role of HDAC6 in Alzheimer's disease. J. Alzheimers Dis. 2013, 33, 283–295. [Google Scholar]
- Gradilone, S.A.; Habringer, S.; Masyuk, T.V.; Howard, B.N.; Masyuk, A.I.; Larusso, N.F. HDAC6 is overexpressed in cystic cholangiocytes and its inhibition reduces cystogenesis. Am. J. Pathol. 2014, 184, 600–608. [Google Scholar]
- Gloghini, A.; Buglio, D.; Khaskhely, N.M.; Georgakis, G.; Orlowski, R.Z.; Neelapu, S.S.; Carbone, A.; Younes, A. Expression of histone deacetylases in lymphoma: Implication for the development of selective inhibitors. Br. J. Haematol. 2009, 147, 515–525. [Google Scholar]
- Wang, J.C.; Kafeel, M.I.; Avezbakiyev, B.; Chen, C.; Sun, Y.; Rathnasabapathy, C.; Kalavar, M.; He, Z.; Burton, J.; Lichter, S. Histone deacetylase in chronic lymphocytic leukemia. Oncology 2011, 81, 325–329. [Google Scholar]
- Bosch-Presegue, L.; Vaquero, A. The dual role of sirtuins in cancer. Genes Cancer 2011, 2, 648–662. [Google Scholar]
- Carafa, V.; Nebbioso, A.; Altucci, L. Sirtuins & disease: The road ahead. Front. Pharmacol. 2012, 3, 4. [Google Scholar]
- Deng, C.X. SIRT1, is it a tumor promoter or tumor suppressor? Int. J. Biol. Sci. 2009, 5, 147–152. [Google Scholar]
- Kuzmichev, A.; Margueron, R.; Vaquero, A.; Preissner, T.S.; Scher, M.; Kirmizis, A.; Ouyang, X.; Brockdorff, N.; Abate-Shen, C.; Farnham, P.; et al. Composition and histone substrates of polycomb repressive group complexes change during cellular differentiation. Proc. Natl. Acad. Sci. USA 2005, 102, 1859–1864. [Google Scholar]
- Lutz, M.I.; Milenkovic, I.; Regelsberger, G.; Kovacs, G.G. Distinct patterns of sirtuin expression during progression of Alzheimer’s disease. Neuromolecular Med. 2014, 16, 405–414. [Google Scholar]
- Du, J.; Zhou, Y.; Su, X.; Yu, J.J.; Khan, S.; Jiang, H.; Kim, J.; Woo, J.; Kim, J.H.; Choi, B.H.; et al. Sirt5 is a NAD-dependent protein lysine demalonylase and desuccinylase. Science 2011, 334, 806–809. [Google Scholar]
- Peng, C.; Lu, Z.; Xie, Z.; Cheng, Z.; Chen, Y.; Tan, M.; Luo, H.; Zhang, Y.; He, W.; Yang, K.; et al. The first identification of lysine malonylation substrates and its regulatory enzyme. Mol. Cell. Proteomics 2011, 10. [Google Scholar] [CrossRef]
- Osborne, B.; Cooney, G.J.; Turner, N. Are sirtuin deacylase enzymes important modulators of mitochondrial energy metabolism? Biochim. Biophys. Acta. 2014, 1840, 1295–1302. [Google Scholar]
- Wei, W.; Xu, X.; Li, H.; Zhang, Y.; Han, D.; Wang, Y.; Yan, W.; Wang, X.; Zhang, J.; Liu, N.; You, Y. The SIRT2 Polymorphism rs10410544 and Risk of Alzheimer’s Disease: A Meta-analysis. Neuromolecular Med. 2014, 16, 448–456. [Google Scholar]
- Riccio, P. The molecular basis of nutritional intervention in multiple sclerosis: A narrative review. Complement. Ther. Med. 2011, 19, 228–237. [Google Scholar]
- Riccio, P.; Rossano, R.; Liuzzi, G.M. May diet and dietary supplements improve the wellness of multiple sclerosis patients? A molecular approach. Autoimmune Dis. 2011, 24, 249842. [Google Scholar]
- Yang, B.; Fu, X.; Shao, L.; Ding, Y.; Zeng, D. Aberrant expression of SIRT3 is conversely correlated with the progression and prognosis of human gastric cancer. Biochem. Biophys. Res. Commun. 2014, 443, 156–160. [Google Scholar]
- Vakhrusheva, O.; Braeuer, D.; Liu, Z.; Braun, T.; Bober, E. Sirt7-dependent inhibition of cell growth and proliferation might be instrumental to mediate tissue integrity during aging. J. Physiol. Pharmacol. 2008, 9, 201–212. [Google Scholar]
- Ashraf, N.; Zino, S.; Macintyre, A.; Kingsmore, D.; Payne, A.P.; George, W.D.; Shiels, P.G. Altered sirtuin expression is associated with node-positive breast cancer. Br. J. Cancer 2006, 95, 1056–1061. [Google Scholar]
- Miremadi, A.; Oestergaard, M.Z.; Pharoah, P.D.; Caldas, C. Cancer genetics of epigenetic genes. Hum. Mol. Genet. 2007, 16 (R1), R28–R49. [Google Scholar]
- Choi, J.H.; Kwon, H.J.; Yoon, B.I.; Kim, J.H.; Han, S.U.; Joo, H.J.; Kim, D.Y. Expression profile of histone deacetylase 1 in gastric cancer tissues. Jpn. J. Cancer Res. 2001, 92, 1300–1304. [Google Scholar]
- Wilson, A.J.; Byun, D.-S.; Popova, N.; Murray, L.B.; L'Italien, K.; Sowa, Y.; Arango, D.; Velcich, A.; Augenlicht, L.H.; Mariadason, J.M. Histone deacetylase 3 (HDAC3) and other Class I HDACs regulate colon cell maturation and p21 expression and are deregulated in human colon cancer. J. Biol. Chem. 2006, 281, 13548–13558. [Google Scholar]
- Melchionda, L.; Fang, M.; Wang, H.; Fugnanesi, V.; Morbin, M.; Liu, X.; Li, W.; Ceccherini, I.; Farina, L.; Savoiardo, M.; et al. Adult-onset Alexander disease, associated with a mutation in an alternative GFAP transcript, may be phenotypically modulated by a non-neutral HDAC6 variant. Orphanet J. Rare Dis. 2013, 8, 66. [Google Scholar]
- Marquardt, J.U.; Fischer, K.; Baus, K.; Kashyap, A.; Ma, S.; Krupp, M.; Linke, M.; Teufel, A.; Zechner, U.; Strand, D.; Thorgeirsson, S.S.; Galle, P.R.; Strand, S. Sirtuin-6-dependent genetic and epigenetic alterations are associated with poor clinical outcome in hepatocellular carcinoma patients. Hepatology 2013, 58, 1054–1064. [Google Scholar]
- Riggs, M.G.; Whittaker, R.G.; Neumann, J.R.; Ingram, V.M. n-Butyrate causes histone modification in HeLa and Friend erythroleukaemia cells. Nature 1977, 268, 462–464. [Google Scholar]
- Kijima, M.; Yoshida, M.; Sugita, K.; Horinouchi, S.; Beppu, T. Trapoxin, an antitumor cyclic tetrapeptide, is an irreversible inhibitor of mammalian histone deacetylase. J. Biol. Chem. 1993, 268, 22429–22435. [Google Scholar]
- Yoshida, M.; Kijima, M.; Akita, M.; Beppu, T. Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J. Biol. Chem. 1990, 265, 17174–17179. [Google Scholar]
- Yoshida, M.; Horinouchi, S.; Beppu, T. Trichostatin A and trapoxin: Novel chemical probes for the role of histone acetylation in chromatin structure and function. Bioessays 1995, 17, 423–430. [Google Scholar]
- Taunton, J.; Hassig, C.A.; Schreiber, S.L. A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 1996, 272, 408–411. [Google Scholar]
- Ververis, K.; Hiong, A.; Karagiannis, T.C.; Licciardi, P.V. Histone deacetylase inhibitors (HDACIs): Multitargeted anticancer agents. Biologics 2013, 7, 47–60. [Google Scholar]
- Rajendran, P.; Ho, E.; Williams, D.E.; Dashwood, R.H. Dietary phytochemicals, HDAC inhibition, and DNA damage/repair defects in cancer cells. Clin. Epigenetics 2011, 3, 4. [Google Scholar]
- Licciardi, P.V.; Kwa, F.A.; Ververis, K.; Di Costanzo, N.; Balcerczyk, A.; Tang, M.L.; El-Osta, A.; Karagiannis, T.C. Influence of natural and synthetic histone deacetylase inhibitors on chromatin. Antioxid. Redox Signal. 2012, 17, 340–354. [Google Scholar]
- Glauben, R.; Batra, A.; Fedke, I.; Zeitz, M.; Lehr, H.A.; Leoni, F.; Mascagni, P.; Fantuzzi, G.; Dinarello, C.A.; Siegmund, B. Histone hyperacetylation is associated with amelioration of experimental colitis in mice. J. Immunol. 2006, 176, 5015–5022. [Google Scholar]
- Banerjee, A.; Trivedi, C.M.; Damera, G.; Jiang, M.; Jester, W.; Hoshi, T.; Epstein, J.A.; Panettieri, R.A. Trichostatin A abrogates airway constriction, but not inflammation, in murine and human asthma models. Am. J. Respir. Cell Mol. Biol. 2012, 46, 132–138. [Google Scholar]
- Chuang, D.M.; Leng, Y.; Marinova, Z.; Kim, H.J.; Chiu, C.T. Multiple roles of HDAC inhibition in neurodegenerative conditions. Trends Neurosci. 2009, 32, 591–601. [Google Scholar]
- Kong, Y.; Tannous, P.; Lu, G.; Berenji, K.; Rothermel, B.A.; Olson, E.N.; Hill, J.A. Suppression of class I and II histone deacetylases blunts pressure-overload cardiac hypertrophy. Circulation 2006, 113, 2579–2588. [Google Scholar]
- Wetzel, M.; Premkumar, D.R.; Arnold, B.; Pollack, I.F. Effect of trichostatin A, a histone deacetylase inhibitor, on glioma proliferation in vitro by inducing cell cycle arrest and apoptosis. J. Neurosurg. 2005, 103, 549–556. [Google Scholar]
- Krasteva, S.; Heiss, E.; Krenn, L. Optimization and application of a fluorimetric assay for the identification of histone deacetylase inhibitors from plant origin. Pharm. Biol. 2011, 49, 658–668. [Google Scholar]
- Liu, Y.; Salvador, L.A.; Byeon, S.; Ying, Y.; Kwan, J.C.; Law, B.K.; Hong, J.; Luesch, H. Anticolon cancer activity of largazole, a marine-derived tunable histone deacetylase inhibitor. J. Pharmacol. Exp. Ther. 2010, 335, 351–361. [Google Scholar]
- Marks, P.A.; Breslow, R. Dimethyl sulfoxide to vorinostat: Development of this histone deacetylase inhibitor as an anticancer drug. Nat. Biotechnol. 2007, 25, 84–90. [Google Scholar]
- Whittle, N.; Singewald, N. HDAC inhibitors as cognitive enhancers in fear, anxiety and trauma therapy: Where do we stand? Biochem. Soc. Trans. 2014, 42, 569–581. [Google Scholar]
- Anand, P.; Kunnumakara, A.; Sundaram, C.; Harikumar, K.; Tharakan, S.; Lai, O.; Sung, B.; Aggarwal, B. Cancer is a preventable disease that requires major lifestyle changes. Pharm. Res. 2008, 25, 2097–2116. [Google Scholar]
- Gonzalez, C.A.; Riboli, E. Diet and cancer prevention: Contributions from the european prospective investigation into cancer and nutrition (EPIC) study. Eur. J. Cancer 2010, 46, 2555–2562. [Google Scholar]
- Greenwald, P.; Clifford, C.K.; Milner, J.A. Diet and cancer prevention. Eur. J. Cancer 2001, 37, 948–965. [Google Scholar]
- Yao, H.; Xu, W.; Shi, X.; Zhang, Z. Dietary flavonoids as cancer prevention agents. J. Environ. Sci. Health C Environ. Carcinog. Ecotoxicol. Rev. 2011, 29, 1–31. [Google Scholar]
- Nilsson, A.C.; Ostman, E.M.; Knudsen, K.E.; Holst, J.J.; Bjorck, I.M. A cereal-based evening meal rich in indigestible carbohydrates increases plasma butyrate the next morning. J. Nutr. 2010, 140, 1932–1936. [Google Scholar]
- Nian, H.; Bisson, W.H.; Dashwood, W.M.; Pinto, J.T.; Dashwood, R.H. Alpha-keto acid metabolites of organoselenium compounds inhibit histone deacetylase activity in human colon cancer cells. Carcinogenesis 2009, 30, 1416–1423. [Google Scholar]
- Lee, J.I.; Nian, H.; Cooper, A.J.; Sinha, R.; Dai, J.; Bisson, W.H.; Dashwood, R.H.; Pinto, J.T. Alpha-keto acid metabolites of naturally occurring organoselenium compounds as inhibitors of histone deacetylase in human prostate cancer cells. Cancer Prev. Res. Phila. 2009, 2, 683–693. [Google Scholar]
- Pinto, J.T.; Lee, J.I.; Sinha, R.; MacEwan, M.E.; Cooper, A.J. Chemopreventive mechanisms of alpha-keto acid metabolites of naturally occurring organoselenium compounds. Amino Acids 2011, 41, 29–41. [Google Scholar]
- Cao, S.; Durrani, F.A.; Toth, K.; Rustum, Y.M. Se-methylselenocysteine offers selective protection against toxicity and potentiates the antitumour activity of anticancer drugs in preclinical animal models. Br. J. Cancer 2014, 110, 1733–1743. [Google Scholar]
- Myzak, M.C.; Dashwood, R.H. Histone deacetylases as targets for dietary cancer preventive agents: Lessons learned with butyrate, diallyl disulfide, and sulforaphane. Curr. Drug Targets 2006, 7, 443–452. [Google Scholar]
- Bhatnagar, N.; Li, X.; Chen, Y.; Zhou, X.; Garrett, S.H.; Guo, B. 3,3′-Diindolylmethane enhances the efficacy of butyrate in colon cancer prevention through down-regulation of survivin. Cancer Prev. Res. Phila. 2009, 2, 581–589. [Google Scholar]
- Aggarwal, B.B.; Ichikawa, H. Molecular targets and anticancer potential of indole-3-carbinol and its derivatives. Cell Cycle 2005, 4, 1201–1215. [Google Scholar]
- Xiong, S.D.; Yu, K.; Liu, X.H.; Yin, L.H.; Kirschenbaum, A.; Yao, S.; Narla, G.; DiFeo, A.; Wu, J.B.; Yuan, Y.; Ho, S.-M.; Lam, Y.W.; Levine, A.C. Ribosome-inactivating proteins isolated from dietary bitter melon induce apoptosis and inhibit histone deacetylase-1 selectively in premalignant and malignant prostate cancer cells. Int. J. Cancer 2009, 125, 774–782. [Google Scholar]
- Gopal, Y.N.V.; Arora, T.S.; Van Dyke, M.W. Parthenolide specifically depletes histone deacetylase 1 protein and induces cell death through ataxia telangiectasia mutated. Chem. Biol. 2007, 14, 813–823. [Google Scholar]
- Lea, M.A.; Rasheed, M.; Randolph, V.M.; Khan, F.; Shareef, A.; desBordes, C. Induction of histone acetylation and inhibition of growth of mouse erythroleukemia cells by S-allylmercaptocysteine. Nutr. Cancer 2002, 43, 90–102. [Google Scholar]
- Son, I.H.; Lee, S.I.; Yang, H.D.; Moon, H.-I. Bis(4-hydroxybenzyl)sulfide: A sulfur compound inhibitor of histone deacetylase isolated from root extract of Pleuropterus ciliinervis. Molecules 2007, 12, 815–820. [Google Scholar]
- Waldecker, M.; Kautenburger, T.; Daumann, H.; Busch, C.; Schrenk, D. Inhibition of histone-deacetylase activity by short-chain fatty acids and some polyphenol metabolites formed in the colon. J. Nutr. Biochem 2008, 19, 587–593. [Google Scholar]
- Thakur, V.S.; Gupta, K.; Gupta, S. Green tea polyphenols causes cell cycle arrest and apoptosis in prostate cancer cells by suppressing class I histone deacetylases. Carcinogenesis 2012, 33, 377–384. [Google Scholar]
- Sbardella, G.; Castellano, S.; Vicidomini, C.; Rotili, D.; Nebbioso, A.; Miceli, M.; Altucci, L.; Mai, A. Identification of long chain alkylidenemalonates as novel small molecule modulators of histone acetyltransferases. Bioorg. Med. Chem. Lett. 2008, 18, 2788–2792. [Google Scholar]
- Lea, M.A.; Randolph, V.M.; Patel, M. Increased acetylation of histones induced by diallyl disulfide and structurally related molecules. Int. J. Oncol. 1999, 15, 347–352. [Google Scholar]
- Druesne, N.; Pagniez, A.; Mayeur, C.; Thomas, M.; Cherbuy, C.; Duee, P.H.; Martel, P.; Chaumontet, C. Repetitive treatments of colon HT-29 cells with diallyl disulfide induce a prolonged hyperacetylation of histone H3 K14. Ann. N. Y. Acad. Sci. 2004, 1030, 612–621. [Google Scholar]
- Hong, T.; Nakagawa, T.; Pan, W.; Kim, M.Y.; Kraus, W.L.; Ikehara, T.; Yasui, K.; Aihara, H.; Takebe, M.; Muramatsu, M.; et al. Isoflavones stimulate estrogen receptor-mediated core histone acetylation. Biochem. Biophys. Res. Commun. 2004, 317, 259–264. [Google Scholar]
- Pandey, M.; Kaur, P.; Shukla, S.; Abbas, A.; Fu, P.; Gupta, S. Plant flavone apigenin inhibits HDAC and remodels chromatin to induce growth arrest and apoptosis in human prostate cancer cells: In vitro and in vivo study. Mol. Carcinog. 2012, 51, 952–962. [Google Scholar]
- Pal-Bhadra, M.; Ramaiah, M.J.; Reddy, T.L.; Krishnan, A.; Pushpavalli, S.N.C.V.L.; Babu, K.S.; Tiwari, A.K.; Rao, J.M.; Yadav, J.S.; Bhadra, U. Plant HDAC inhibitor chrysin arrest cell growth and induce p21WAF1 by altering chromatin of STAT response element in A375 cells. BMC Cancer 2012, 12, 180. [Google Scholar]
- Majid, S.; Dar, A.A.; Ahmad, A.E.; Hirata, H.; Kawakami, K.; Shahryari, V.; Saini, S.; Tanaka, Y.; Dahiya, A.V.; Khatri, G.; et al. BTG3 tumor suppressor gene promoter demethylation, histone modification and cell cycle arrest by genistein in renal cancer. Carcinogenesis 2009, 30, 662–670. [Google Scholar]
- Lea, M.A.; Randolph, V.M.; Lee, J.E.; desBordes, C. Induction of histone acetylation in mouse erythroleukemia cells by some organosulfur compounds including allyl isothiocyanate. Int. J. Cancer 2001, 92, 784–789. [Google Scholar]
- Myzak, M.C.; Dashwood, W.M.; Orner, G.A.; Ho, E.; Dashwood, R.H. Sulforaphane inhibits histone deacetylase in vivo and suppresses tumorigenesis in Apc-minus mice. FASEB J. 2006, 20, 506–508. [Google Scholar]
- Son, I.H.; Chung, I.-M.; Lee, S.I.; Yang, H.D.; Moon, H.-I. Pomiferin, histone deacetylase inhibitor isolated from the fruits of Maclura pomifera. Bioorg. Med. Chem. Lett. 2007, 17, 4753–4755. [Google Scholar]
- Priyadarsini, R.V.; Vinothini, G.; Murugan, R.S.; Manikandan, P.; Nagini, S. The flavonoid quercetin modulates the hallmark capabilities of hamster buccal pouch tumors. Nutr. Cancer 2011, 63, 218–226. [Google Scholar]
- Wood, J.G.; Rogina, B.; Lavu, S.; Howitz, K.; Helfand, S.L.; Tatar, M.; Sinclair, D. Sirtuin activators mimic caloric restriction and delay ageing in metazoans. Nature 2004, 430, 686–689. [Google Scholar]
- Yang, S.R.; Wright, J.; Bauter, M.; Seweryniak, K.; Kode, A.; Rahman, I. Sirtuin regulates cigarette smoke-induced proinflammatory mediator release via RelA/p65 NF-kappaB in macrophages in vitro and in rat lungs in vivo: Implications for chronic inflammation and aging. Am. J. Physiol. Lung Cell. Mol. Physiol. 2007, 292, L567–L576. [Google Scholar]
- Chung, I.M.; Kim, M.Y.; Park, W.H.; Moon, H.I. Histone deacetylase inhibitors from the rhizomes of Zingiber zerumbet. Pharmazie 2008, 63, 774–776. [Google Scholar]
- Chen, I.H.; Lu, M.C.; Du, Y.C.; Yen, M.H.; Wu, C.C.; Chen, Y.H.; Hung, C.S.; Chen, S.L.; Chang, F.R.; Wu, Y.C. Cytotoxic triterpenoids from the stems of Microtropis japonica. J. Nat. Prod. 2009, 72, 1231–1236. [Google Scholar]
- Shukla, S.; Gupta, S. Apigenin: A promising molecule for cancer prevention. Pharm. Res. 2010, 27, 962–978. [Google Scholar]
- Venturelli, S.; Berger, A.; Bocker, A.; Busch, C.; Weiland, T.; Noor, S.; Leischner, C.; Schleicher, S.; Mayer, M.; Weiss, T.S.; Bischoff, S.C.; Lauer, U.M.; Bitzer, M. Resveratrol as a pan-HDAC inhibitor alters the acetylation status of histone proteins in human-derived hepatoblastoma cells. PLoS One 2013, 8, e73097. [Google Scholar]
- Lee, S.J.; Krauthauser, C.; Maduskuie, V.; Fawcett, P.T.; Olson, J.M.; Rajasekaran, S.A. Curcumin-induced HDAC inhibition and attenuation of medulloblastoma growth in vitro and in vivo. BMC Cancer 2011, 11, 144. [Google Scholar]
- Morimoto, T.; Sunagawa, Y.; Kawamura, T.; Takaya, T.; Wada, H.; Nagasawa, A.; Komeda, M.; Fujita, M.; Shimatsu, A.; Kita, T.; Hasegawa, K. The dietary compound curcumin inhibits p300 histone acetyltransferase activity and prevents heart failure in rats. J. Clin. Invest. 2008, 118, 868–878. [Google Scholar]
- Xiao, X.; Shi, D.; Liu, L.; Wang, J.; Xie, X.; Kang, T.; Deng, W. Quercetin suppresses cyclooxygenase-2 expression and angiogenesis through inactivation of P300 signaling. PLoS One 2011, 6, 8. [Google Scholar]
- Rajendran, P.; Williams, D.E.; Ho, E.; Dashwood, R.H. Metabolism as a key to histone deacetylase inhibition. Crit. Rev. Biochem. Mol. Biol. 2011, 46, 181–199. [Google Scholar]
- Myzak, M.C.; Tong, P.; Dashwood, W.M.; Dashwood, R.H.; Ho, E. Sulforaphane retards the growth of human PC-3 xenografts and inhibits HDAC activity in human subjects. Exp. Biol. Med. 2007, 232, 227–234. [Google Scholar]
- Link, A.; Balaguer, F.; Goel, A. Cancer chemoprevention by dietary polyphenols: Promising role for epigenetics. Biochem. Pharmacol. 2010, 80, 1771–1792. [Google Scholar]
- Cieślik, E.; Leszczyńska, T.; Filipiak-Florkiewicz, A.; Sikora, E.; Pisulewski, P.M. Effects of some technological processes on glucosinolate contents in cruciferous vegetables. Food Chem. 2007, 105, 976–981. [Google Scholar]
- Lin, C.H.; Chang, C.Y. Textural change and antioxidant properties of broccoli under different cooking treatments. Food Chem. 2005, 90, 9–15. [Google Scholar]
- Yuan, G.F.; Sun, B.; Yuan, J.; Wang, Q.M. Effects of different cooking methods on health-promoting compounds of broccoli. J. Zhejiang Univ. Sci. B 2009, 10, 580–588. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bassett, S.A.; Barnett, M.P.G. The Role of Dietary Histone Deacetylases (HDACs) Inhibitors in Health and Disease. Nutrients 2014, 6, 4273-4301. https://doi.org/10.3390/nu6104273
Bassett SA, Barnett MPG. The Role of Dietary Histone Deacetylases (HDACs) Inhibitors in Health and Disease. Nutrients. 2014; 6(10):4273-4301. https://doi.org/10.3390/nu6104273
Chicago/Turabian StyleBassett, Shalome A., and Matthew P. G. Barnett. 2014. "The Role of Dietary Histone Deacetylases (HDACs) Inhibitors in Health and Disease" Nutrients 6, no. 10: 4273-4301. https://doi.org/10.3390/nu6104273
APA StyleBassett, S. A., & Barnett, M. P. G. (2014). The Role of Dietary Histone Deacetylases (HDACs) Inhibitors in Health and Disease. Nutrients, 6(10), 4273-4301. https://doi.org/10.3390/nu6104273







