Abstract
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), the causative agent of Coronavirus Disease 2019 (COVID-19), poses a serious global public health threat for which there is currently no satisfactory treatment. This study examines the efficacy of Biobran/MGN-3 against SARS-CoV-2. Biobran is an arabinoxylan rice bran that has been shown to significantly inhibit the related influenza virus in geriatric subjects. Here, Biobran’s anti-SARS-CoV-2 activity was assessed using MTT and plaque reduction assays, RT-PCR, ELISA techniques, and measurements of SARS-CoV-2-related gene expression and protein levels. For Vero E6 cells infected with SARS-CoV-2, Biobran reduced the viral load by 91.9% at a dose of 100 μg/mL, it reduced viral counts (PFU/mL) by 90.6% at 50 μg/mL, and it exhibited a significant selectivity index (EC50/IC50) of 22.5. In addition, Biobran at 10 μg/mL inhibited papain-like proteinase (PLpro) by 87% and ACE2 SARS-CoV-2 S-protein RBD by 90.5%, and it significantly suppressed SARS-CoV-2 gene expression, down-regulating E-gene and RdRp gene expression by 93% each at a dose of 50 μg/mL and inhibiting the E-protein by 91.3%. An in silico docking study was also performed to examine the protein–protein interaction (PPI) between SARS-CoV-2 RBD and DC-SIGN as well as between serine carboxypeptidase and papain-like protease PLpro. Serine carboxypeptidase, an active ingredient in Biobran, was found to interfere with the binding of SARS-CoV-2 to its receptor DC-SIGN on Vero cells, thus preventing the cell entry of SARS-CoV-2. In addition, it impairs the viral replication cycle by binding to PLpro. We conclude that Biobran possesses potent antiviral activity against SARS-CoV-2 in vitro and suggest that Biobran may be able to prevent SARS-CoV-2 infection. This warrants further investigation in clinical trials.
1. Introduction
Coronaviruses have emerged as significant global health threats. They had caused two earlier pandemics—the severe acute respiratory syndrome that appeared in China in 2003 [1] and the Middle East respiratory syndrome [2]—before causing the current Coronavirus Disease 2019 (COVID-19) pandemic that arose from Wuhan, Hubei Province, China. Bats are known to be the primary animal reservoir for coronaviruses, and the viruses have been able to mutate and adapt to infect humans, resulting in an animal-to-human species barrier jump [3]. The emergence of novel coronaviruses continually poses a serious global public health threat and carries the potential for causing major pandemic outbreaks in the naïve human population. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) is the coronavirus causing the disease COVID-19, a disease whose outbreak has resulted in over 239 million confirmed cases of infection and killed over 4.8 million people across the globe [4]. This novel virus has been globally transmitted to 228 countries and territories.
SARS-CoV-2 is thought to have been transmitted from a zoonotic source and spreads via contact and direct transmission. The disease manifests symptomatically with cough, myalgia, fever, and severe respiratory failure. Reverse transcriptase PCR is used to confirm diagnosis, and managing COVID-19 is mainly carried out via supportive therapy, along with mechanical ventilation in severe cases. Attempts to reduce the public spread of SARS-CoV-2 consist mainly of preventive strategies, community containment, and disease isolation. Vaccine developments for eliminating the virus from the host remain an ongoing challenge [5,6]. Inhibiting the entry of viruses into host cells and/or preventing viral replication may represent therapeutic approaches for SARS-CoV-2. Angiotensin-converting enzyme 2 (ACE2) enzyme is a key receptor for SARS-CoV-2 viral entry, and CD209L (also known as L-SIGN) and CD209 (also known as DC-SIGN) are members of the C-type lectin superfamily that are significantly expressed in human endothelial cells, i.e., in macrophages and dendritic cells of the lymph nodes, liver, and lung, and are recognized by viral glycoproteins of SARS-CoV-2 via their C-terminal carbohydrate domain [7]. After entry into host cells, SARS-CoV-2 replicates and spreads, taking advantage of compounds such as papain-like protease (PLpro) to process viral proteins and generate a functional replicase complex [8]. New compounds which disrupt the activity of ACE2 and/or PLpro may be effective at mitigating SARS-CoV-2′s detrimental actions.
26 COVID-19 vaccines have been reported by the World Health Organization (WHO) to have been developed and evaluated in phase III clinical trials [9]. Several of these vaccines have been shown to be highly effective against the original strain of COVID-19 and its variants, including BNT162b2, AZD1222, mRNA-1273, and Sputnik V. In phase III trials, these vaccines were well tolerated and the most effective vaccines (>90%) in the prevention of symptomatic cases. However, protection against infection was observed to decline at 6 months for BNT162b2 and AZD1222, and serious adverse effects and myocarditis have been reported in a few cases [10]. There is no satisfactory treatment for the disease. Thus, it is essential to investigate therapeutic products, including the application of therapeutic natural products that are nontoxic, affordable, and capable of exerting a strong anti-COVID-19 effect.
Different types of natural products and herbal components have been examined for their potential use against SARS-CoV-2 [11,12,13,14,15,16,17,18,19,20,21,22,23,24,25]. These include scutellarein, caffeic acid, myricetin, quercetin, saikosaponin B2, silvestrol, tryptanthrin, psoralidin, lectins such as griffithsin, isobavachalcone, and steroids [16,17]. In addition, other natural products have been reported to exert anti-COVID-19 activity, including flavonoids [18], polyphenols [19], Agathis robusta bark essential oil [20], and propolis, bee honey, and their components [21]. S. japonica stem extract was shown to significantly inhibit coronavirus CoNL63 attachment in mammalian cells, indicating that it could be effective against infection in its early stage [22]; S. nigra berry extract was shown to significantly reduce virus titers in Vero cells infected with infectious bronchitis virus (a pathogenic poultry coronavirus) [23]; S. nigra fruit and flower extracts were shown to inhibit SARS-CoV-2 RBD and ACE2 receptor binding [24]; and Resveratrol, a phenolic compound produced by a variety of spermatophytes such as peanuts, mulberry, and grapes, has been shown to inhibit SARS-CoV-2 replication in cultured Vero cells [25].
Biobran/MGN-3, an arabinoxylan rice bran, is another natural product that potentially holds great promise. Biobran supplementation has been shown to reduce the incidence of influenza-like illnesses (ILI) in elderly subjects [26,27]. It is an arabinoxylan extracted from rice bran treated with hydrolyzing enzymes from Shiitake mushrooms [28], and in addition to reducing ILI in elderly subjects, it has been characterized as an antiviral agent with the ability to exert anti-HIV activity in vitro [28] and significantly reduce the viral load in patients with chronic hepatitis C virus infection [29]. In addition, Biobran has shown potential as an anticancer agent. It exhibited antitumor activity in mice bearing a solid Ehrlich carcinoma tumor [30] and synergized with conventional therapies for the treatment of hepatocellular carcinoma in human patients [31]. Biobran’s antiviral and anticancer activity has been attributed to its ability to act as a potent biological response modifier (BRM). It is known to activate different arms of the immune system such as natural killer (NK) cells [30,32,33,34,35,36], increase human T and B cell mitogen response [32], activate dendritic cells [37,38,39], and enhance the phagocytic activity by macrophages [40]. Furthermore, Biobran has been shown to exhibit a psychoneuroimmune modulatory effect by enhancing health-related quality of life in healthy older adults [41] and in cancer patients [35,42].
Based on these characteristics, we hypothesized that Biobran would exert antiviral activity against SARS-CoV-2. No previous study has explored the therapeutic potential of Biobran against coronaviruses, nor the mechanisms by which its active ingredients might counteract a coronavirus such as SARS-CoV-2 by inhibiting viral entry into host cells and/or preventing viral replication. The results of the present in vitro study demonstrate that our hypothesis is correct, namely that Biobran has a very potent antiviral effect against SARS-CoV-2. Furthermore, the potential mechanisms behind Biobran’s anti-SARS-CoV-2 effect are demonstrated via an in silico study of the docking of Ferulic acid and serine carboxypeptidase. These compounds are present in crude rice bran and wheat bran, and many studies have shown that these bran product compounds play a key role in the products’ anti-oxidant, anti-inflammatory, and anti-cancer properties [43,44,45]. Specifically with respect to the potential anti-SARS-CoV-2 activity of Biobran, we studied the binding of Ferulic acid and serine carboxypeptidase to PLpro and DC-SIGN, bindings which would have the positive effect of interfering with SARS-CoV-2′s cell entry and replication capabilities.
2. Materials and Methods
2.1. Biobran
Biobran is a denatured hemicellulose that is extracted from rice bran by reacting rice bran hemicellulose with carbohydrate-hydrolyzing enzymes obtained from Shiitake mushrooms. Biobran’s main chemical structure consists of arabinoxylan with an arabinose polymer in its side chain and a xylose in its main chain [28]. Daiwa Pharmaceutical Co. Ltd. (Tokyo, Japan) kindly provided the Biobran for this study. The Biobran was prepared in saline (0.9% w/v), and Biobran solutions were freshly prepared each day.
2.2. Anti-SARS-CoV-2 Assays
2.2.1. Protein Interactions
The Biobran was examined for its ability to inhibit papain-like proteinase, the spike protein receptor-binding domain, and ACE2 binding [46]. To achieve this aim, the protein–protein interaction (PPI) between SARS-CoV-2 RBD (PDB:7e5o) and DC-SIGN (PDB:6ghv) was examined to evaluate whether SARS-CoV-2 RBD can enter host cells by interacting with DC-SIGN. In addition, the PPI between serine carboxypeptidase (PDB:3sc2) and papain-like protease PLpro (PDB:7nfv) was examined to explore the efficacy of Biobran’s active ingredients in counteracting SARS-CoV-2 infectivity.
Papain-Like Proteinase (PLpro)
40 μL of 142 nM PLpro in buffer A (50 mM HEPES, pH 7.5; 0.1 mg/mL bovine serum albumin (BSA), and 5 mM Dithiothreitol (DTT)) was dispensed in a 96-well plate and then incubated with 100 μL of different concentrations of Biobran for 5 min. Reactions were triggered by adding the fluorogenic substrate Arg-Leu-Arg-Gly-Gly-AMC (Enzo Biochem, USA) (RLRGG-AMC, 10 μL of 250 μM) to buffer A, forcefully shaking for 30 s, and incubating for 6 min. Next, 10 μL acetic acid (0.5 M) was used to halt the reactions, the solution was shaken for 30 s, and the fluorescence emission intensity was measured (wavelength of excitation: 360 nm; wavelength of emission: 460 nm). This allowed the measurement of the inhibition percentage (%).
Spike Protein Receptor-Binding Domain (RBD)
SARS-CoV-2 RBD (Abcam, UK) (1 μg/mL) was incubated with Biobran at 37 °C for 2 h, and the mixture was added to a 96-well plate, incubated overnight at 4 °C, and then blocked with 2% fat-free milk in phosphate-buffered saline with Tween® detergent (PBST) at 37 °C for 2 h. Diluted ACE2 protein (Abcam, UK) was added to the plates and incubated again at 37 °C for 2 h. After four washes, bound protein was detected using hACE2-specific goat antibody (0.5 μg/mL, R&D system, McKinley, MN, USA) that was incubated for 2 h at 37 °C, followed by the incubation of horseradish peroxidase (HRP) conjugated anti-goat IgG antibody (1:5000, Thermo Fisher Scientific, Dreieich, Germany) for 1 h at 37 °C. The reaction was visualized by adding the substrate 3,3′,5,5′-Tetramethylbenzidine (TMB) (Sigma, St. Louis, MO, USA) and stopped using H2SO4 (1 N). An ELISA plate reader (Tecan, San Jose, CA, USA) was used to measure absorbance at 450 nm and thereby estimate the change in RBD–ACE2 complex formation.
2.2.2. In Vitro Antiviral Activity
Vero E6 Toxicity
Biobran was diluted with Dulbecco’s Modified Eagle’s Medium (DMEM). Stock solutions of Biobran were prepared in 10% DMSO in ddH2O. Vero E6 cells (ATCC, CCL-81) were used to test cytotoxic activity by using the 3-(4, 5-dimethylthiazol -2-yl)-2, 5-diphenyltetrazolium bromide (MTT) method [47], modified slightly. Briefly, cells were seeded into 96-well plates (100 μL/well; 3 × 105 cells/mL density) and incubated at 37 °C for 24 h in 5% CO2. After 24 h, various concentrations of Biobran were used to treat the cells in triplicates, and these were incubated for 24 h. After discarding the supernatant, sterile phosphate buffer saline (PBS) was used to wash the cell monolayers three times. MTT solution (20 μL of 5 mg/mL stock solution) was added to each well and incubated for 4 h at 37 °C, and then the medium was aspirated. Formazan crystals from each well were dissolved with 200 μL of acidified isopropanol (0.04 M HCl in absolute isopropanol, 0.073 mL HCl in 50 mL isopropanol). Using a multi-well plate reader, the absorbance of formazan solutions was measured at a maximum wavelength of 540 nm with a reference wavelength of 620 nm. The cytotoxicity percentage compared with untreated cells was determined using the following equation:
The concentration exhibiting 50% cytotoxicity (IC50) was calculated using a plot of percentage cytotoxicity versus sample concentration.
Anti-COVID-19 Activity (Plaque Reduction Assay)
Anti-COVID-19 activity was assessed via plaque reduction assay following the method of [48]. Vero E6 cells (1 × 106 cells/mL) were cultivated in a six-well plate at 37 °C for 24 h. SARS-CoV-2 was diluted to 1 × 104 plaque-forming unit (PFU)/well, mixed with 100 μL of Biobran’s safe concentration, and incubated at 37 °C for 1 h before being added to the cells. Growth medium was removed from the cell culture plates and the cells were inoculated with Biobran (100 μL/well). One hour of contact was allowed for virus adsorption, following which 3 mL of DMEM, 2% agarose, and the Biobran and virus were added onto the cell monolayer. The plates were left to solidify and incubated at 37 °C until viral plaques formed (3 days). Formalin (10%) was added for 2 h. Next, 0.1% crystal violet in distilled water was used to stain plates. The control wells were wells in which untreated virus was incubated with Vero E6 cells. Finally, the plaques were counted and the percentage reduction in plaque formation (% reduction) in comparison with the control wells was recorded according to the following equation:
2.3. Docking
2.3.1. Drug Docking
ChemSketch was used to draw the drug Ferulic, Avogadro energy was used for drug optimization, Spdbv was used for energy optimization of the proteins, Discovery Studio 2021 was used to visualize the ligand-receptor, and iGemodock was used for drug docking [53].
2.3.2. Molecular Docking
ClusPro 2.0 was used to examine the protein–protein interaction (PPI) between SARS-CoV-2 RBD (PDB:7e5o) and DC-SIGN (PDB:6ghv) as well as the PPI between serine carboxypeptidase (PDB:3sc2) and papain-like protease PLpro (PDB:7nfv). The data were visualized and analyzed with the PyMol and Prodigy programs [54].
2.4. Statistical Analysis
Data were analyzed using one- and two-way analysis of variance (ANOVA) with the Sidak–Holm test, two-tailed unpaired Student’s t-test, and multiple t-test. p < 0.05 was used to determine statistical significance. GraphPad Prism 7.0 (San Diego, CA, USA) was used as the statistical software.
3. Results
3.1. The Binding/Inhibitory Effect of Biobran on Papain-Like Proteinase (PLpro), the Spike Protein Receptor-Binding Domain, and ACE2
The results depicted in Table 1 show that Biobran exerts a potent inhibitory effect against PLpro. Treatment with Biobran at a concentration of 10 μg/mL inhibited PLpro by 87%. Table 1 also shows that Biobran exerts a potent inhibitory effect against ACE2 SARS-CoV-2 S-protein RBD in a dose-dependent manner, with inhibition of 90.5% and 2.32% at 10 μg/mL and 0.625 μg/mL respectively.

Table 1.
The inhibitory/binding effect of Biobran on papain-like proteinase, the spike protein receptor-binding domain, and ACE2.
3.2. Biobran Reduces Viral Load in Vero Cells Infected with SARS-CoV-2
Treatment with Biobran caused significant reduction in the viral load in Vero cells infected with SARS-CoV-2 (Table 2). A wide range of Biobran concentrations (3.1–100 μg/mL) were used to establish the dose response–relationship of its efficacy against SARS-CoV-2. Biobran exerts its effect in a dose-dependent manner. There was a 91.9% viral reduction in Vero cells post exposure to Biobran at a dose of 100 μg/mL, and a 16.1% viral reduction at a concentration of 3.125 μg/mL.

Table 2.
Cytotoxic effect of Biobran on Vero E6.
3.3. Vero E6 Toxicity and Plaque Reduction Assay
The antiviral activity of Biobran determined using Vero E6 toxicity and plaque reduction assays against SARS-CoV-2 is shown in Table 3. Compared with the initial viral counts, a significant reduction in viral counts (PFU/mL) following exposure to Biobran was observed. The toxic effect was dose dependent, with an inhibition of 90.6% at 50 μg/mL and 27% at 6.25 μg/mL.

Table 3.
Antiviral activity of Biobran using in vitro Vero E6 toxicity and plaque reduction assays against SARS-CoV-2.
3.4. IC50 and EC50 of Biobran
In order to evaluate the antiviral activity of Biobran, we examined the 50% inhibitory concentration (IC50) and the half maximal response (EC50). The results in Table 4 show that the Biobran Vero E6 IC50 = 15.36 μg/mL and the anti-COVID-19 EC50 = 3.46 μg/mL. These results indicate that Biobran exerts significant antiviral activity against SARS-CoV-2 with a high selectivity index of 22.5 for antiviral activity relative to cellular toxicity.

Table 4.
IC50 and EC50 of Biobran.
3.5. Biobran Suppresses SARS-CoV-2 Gene Expression and Protein Levels
The results in Table 5 show that treatment with Biobran causes significant suppression of SARS-CoV-2 gene expression for the E-gene and RdRp gene. Treatment with Biobran at 50 μg/mL, 25 μg/mL, 12.5 μg/mL, and 6.25 μg/mL caused down-regulation of the E-gene by 92.7%, 73.4%, 56.0%, and 21%, respectively. Similarly, treatment with Biobran resulted in the significant down regulation of RdRp by 93%, 81%, 64%, and 32%, respectively. In addition, the E-protein was significantly inhibited by 91.3%, 75.3%, 49.6%, and 19.4% post exposure to Biobran.

Table 5.
Effect of Biobran on SARS-CoV-2 gene expression and protein levels.
3.6. SARS-CoV-2 Docking with DC-SIGN
To confirm the specificity of the inhibitory effect of Biobran on SARS-CoV-2 in cultured Vero cells, we performed an in silico simulation and modeling of the structure of Biobran and the structures of the key nodes in this pathway.
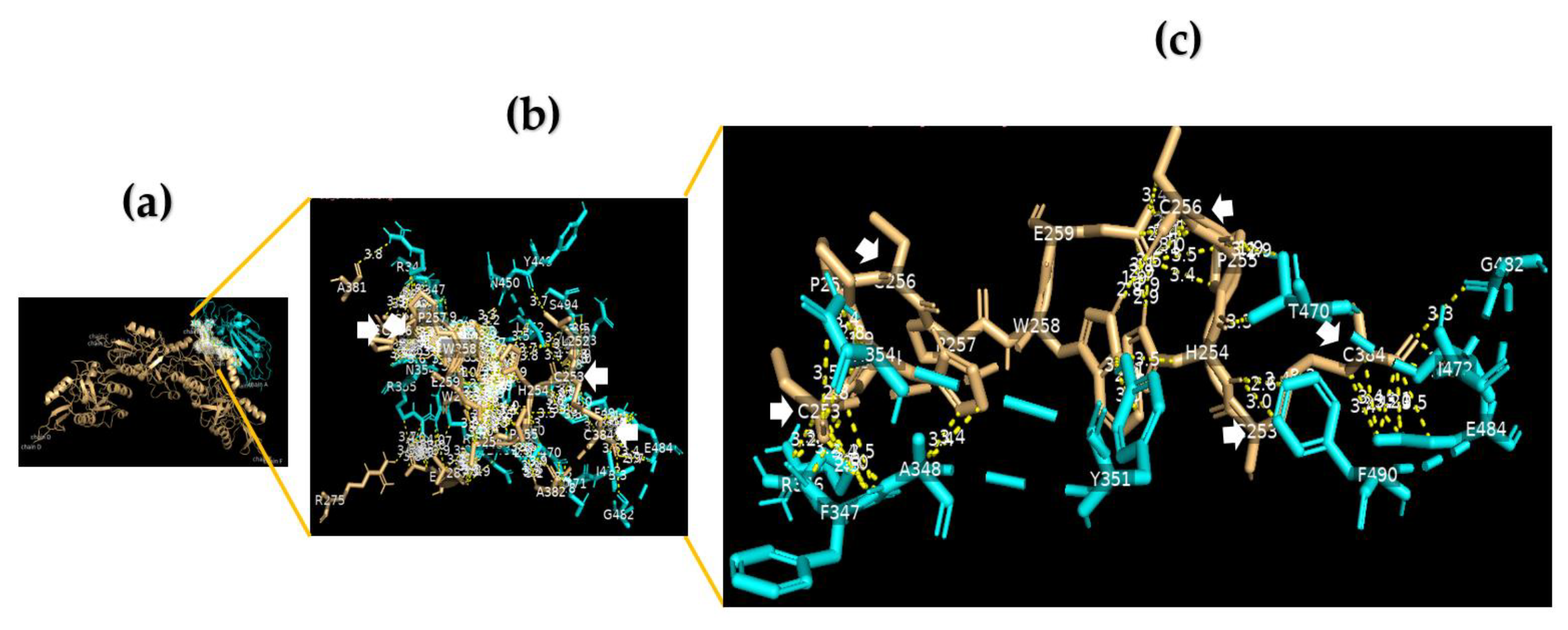
Using a cut-off of 4.0 Å, we found that the RBD domain of the SARS-CoV-2 spike protein to DC-SIGN showed a protein–protein interaction (PPI) via the binding of V382 S383 T385 L387 N388 F392 F515 E516 L517 of chain A to L252 H254 P255 C256 P257 W258 of chain D. In addition, we found a PPI between E340 R346 A347 F348 N354 (chain A) of the SARS-CoV-2 RBD and E259 (chain E) and H254 and C384 (chain C) of DC-SIGN. Furthermore, G381 V382 S383 P384 L387 N388 F392 T430 L517 A522 of the SARS-CoV-2 RBD can bind DC-SIGN at many critical residues, including L252 C253 H254 P255 P257 W258 E259 V287 A382 S383 C384 of chain D. Moreover, we identified multiple bonds between R346 F347 A348 S349 V350 Y351 A352 N354 R355 Y449 N450 L452 E465 R466 D467 I468 T470 E471 I472 G482 E484 F490 L492 Q493 S494 of the SARS-CoV-2 RBD and C253 H254 P255 C256 P257 W258 E259 A381 of chain B and L252 C253 H254 P255 C256 W258 R275 E286 V287 A382 C384 of chain E of DC-SIGN (Figure 1, Figures S1–S4).
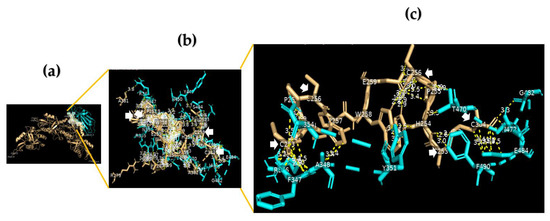
Figure 1.
The RBD domain of the SARS-CoV-2 spike protein binds with DC-SIGN. (a) A 3D image for position (model 3) showing the molecular docking of the RBD domain of the SARS-CoV-2 spike protein (chain A) (cyan) to DC-SIGN CRD (light orange); (b) a 3D image showing protein–protein interaction (PPI) via binding of R346 F347 A348 S349 V350 Y351 A352 N354 R355 Y449 N450 L452 E465 R466 D467 I468 T470 E471 I472 G482 E484 F490 L492 Q493 S494 of chain A of the RBD domain of the SARS-CoV-2 spike protein (cyan) to C253 H254 P255 C256 P257 W258 E259 A381 of chain B and L252 C253 H254 P255 C256 W258 R275 E286 V287 A382 C384 of chain E of DC-SIGN CRD (light orange), using a cut-off of 4.0 Å; (c) a 3D image showing the binding sites between R346 F347 A348 N354 Y351 T470 G482 E484 F490 of the RBD domain of the SARS-CoV-2 spike protein (cyan) and C253 C256 of chain B and C253 C256 C384 of chain E of DC-SIGN CRD (light orange) using a cut-off of 3.5 Å.
3.7. Drug Docking of DC-SIGN
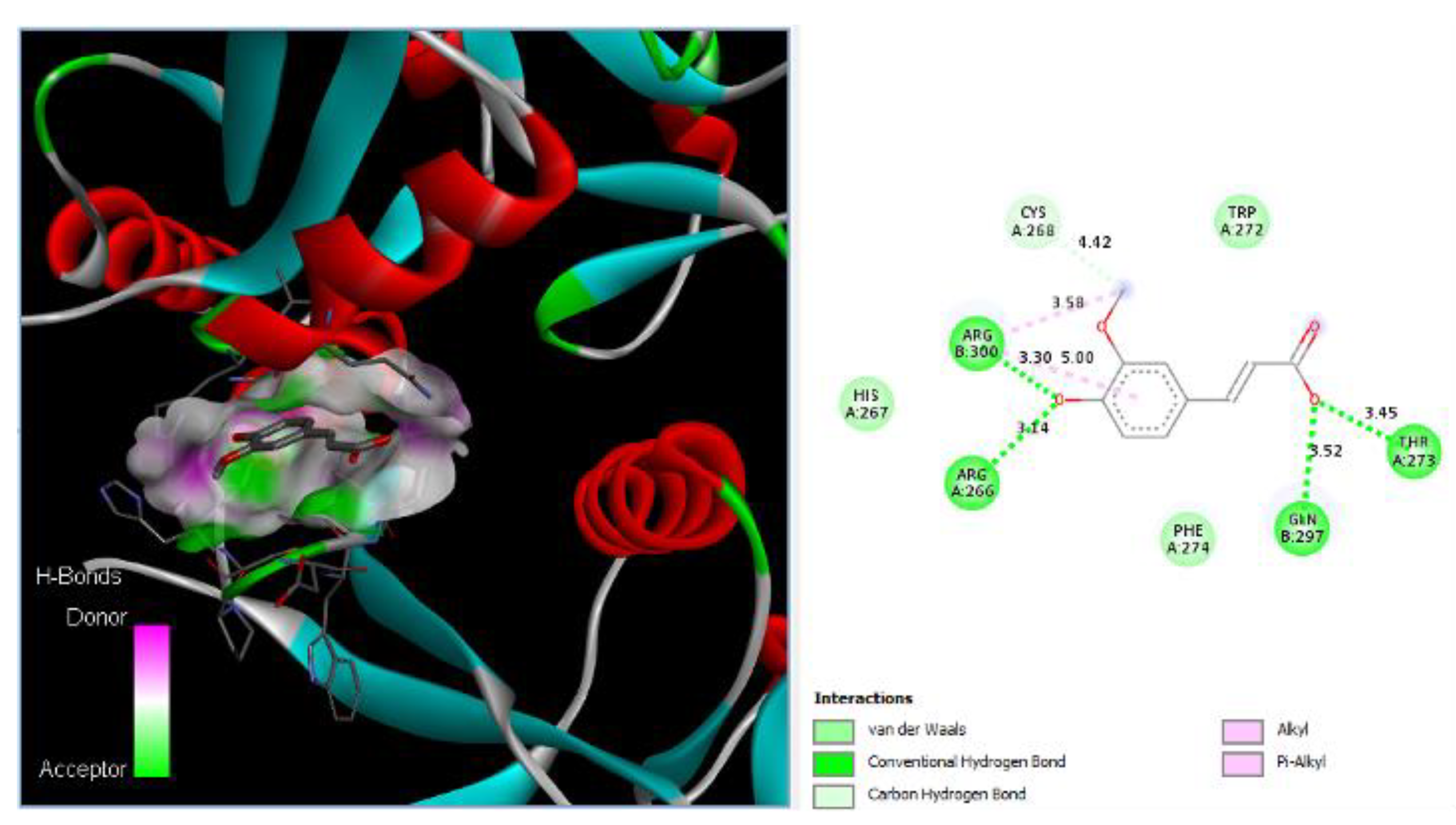
Ferulic acid has been found to bind to proteins in DC-SIGN. Figure 2 shows that Ferulic can bind to Arg266, Thr273, Gln297, and Arg300 with hydrogen bonds with distances of 3.14 Å, 3.45 Å, 3.52 Å, and 3.30 Å, respectively. Additionally, Ferulic can bind to Cys268 with carbon HBs with a distance of 4.42 Å, and it can bind to Arg300 via 2 alkyl bonds with distances of 3.58 Å and 5.00 Å.
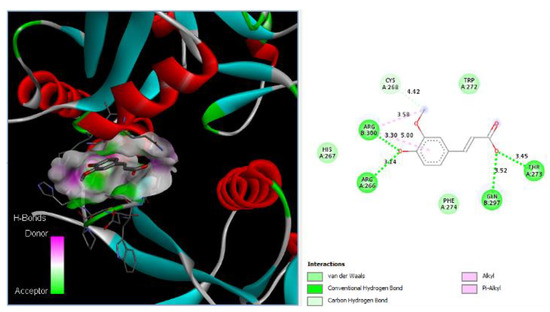
Figure 2.
Ferulic acid binds to DC-SIGN: 3D image of Ferulic showing the hydrogen donor and acceptor (left) and 2D image showing the types of bonds and distances (right) between Ferulic and DC-SIGN where Ferulic can bind to Arg266, Thr273, Gln297, and Arg300 of DC-SIGN via hydrogen bonds with distances of 3.14 Å, 3.45 Å, 3.52 Å, and 3.30 Å, respectively. In addition, Ferulic can bind to Cys268 with carbon HBs with a distance of 4.42 Å, and Ferulic can bind to Arg300 via 2 alkyl bonds with distances of 3.58 Å and 5.00 Å.
3.8. Molecular Docking of DC-SIGN
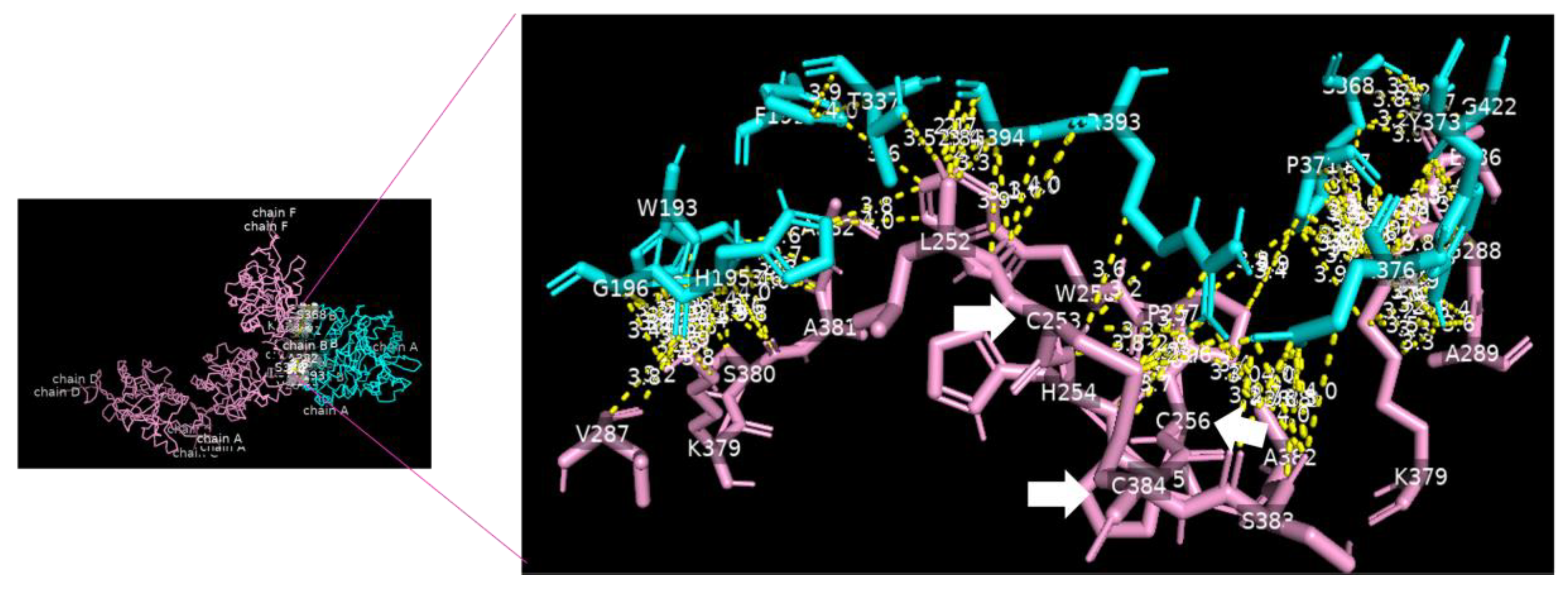
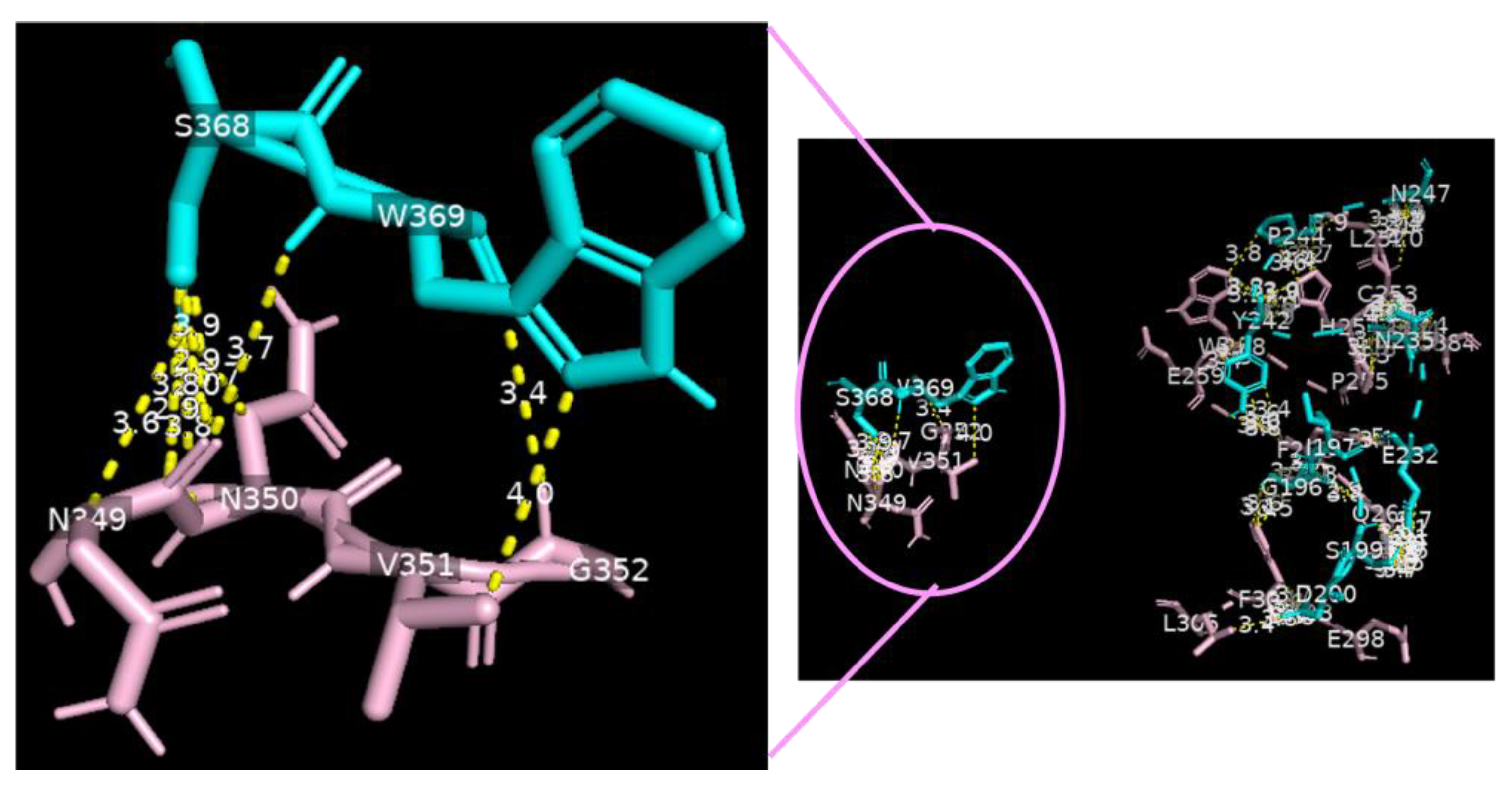
Analysis of the data collected from the molecular docking showed that serine carboxypeptidase can dock DC-SIGN with a binding affinity (ΔG) equal to −11.8 kcal mol−1 and a dissociation constant (Kd) equal to 2.3 × 10−9 M, RMSD = 7.27 Å, and TM-score = 0.26528 if normalized by a length of 3sc2, or TM-score = 0.16107 if normalized by a length of 6ghv. Using a cut-off of 4.0 Å, we detected multiple binding sites between serine carboxypeptidase and DC-SIGN. As shown in Figure 3 and Figure 4, N294 N250 V251 G252 of serine carboxypeptidase can bind to C253 C265 and C384 of DC-SIGN, and I197 S199 D200 E232 N235 Y242 P244 N247 E259 S368 W369 of serine carboxypeptidase can bind to L252 C253 W258 E259 E298 F302 L305 N349 N350 V351G352 of DC-SIGN.
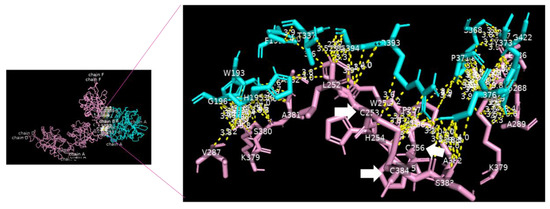
Figure 3.
Serine carboxypeptidase binding to DC-SIGN: 3D image showing the interface between the whole structure of serine carboxypeptidase (cyan) and DC-SIGN (pink) (left side), with multiple binding sites between F191 W193 H195 G196 T337 S368 P371 Y373 I376 R393 G422 of serine carboxypeptidase (cyan) binding to L252 C253 H254 C256 W259 V287 G288 A289 K379 S380 A381 A382 S383 and C384 of DC-SIGN (pink) (right side) (cut-off: 4.0 Å).
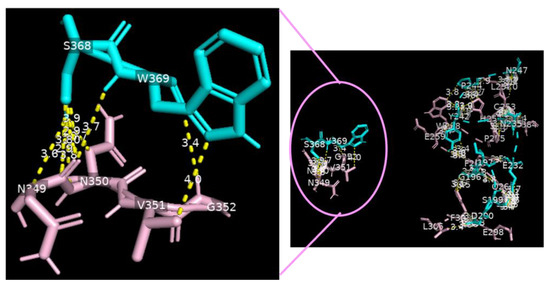
Figure 4.
Serine carboxypeptidase binding to DC-SIGN: 3D image showing multiple binding sites between serine carboxypeptidase (cyan) and DC-SIGN (pink), including I197 S199 D200 E232 N235 Y242 P244 N247 E259 S368 W369 of serine carboxypeptidase (cyan) binding to L252 C253 W258 E259 E298 F302 L305 N349 N350 V351 G352 of DC-SIGN (pink) (right side) and a focus on binding between S368 W369 of serine carboxypeptidase (cyan) and N349 N350 V351 G352 of DC-SIGN (pink) (left side) (cut-off: 4.0 Å).
3.9. Drug Docking of PLpro
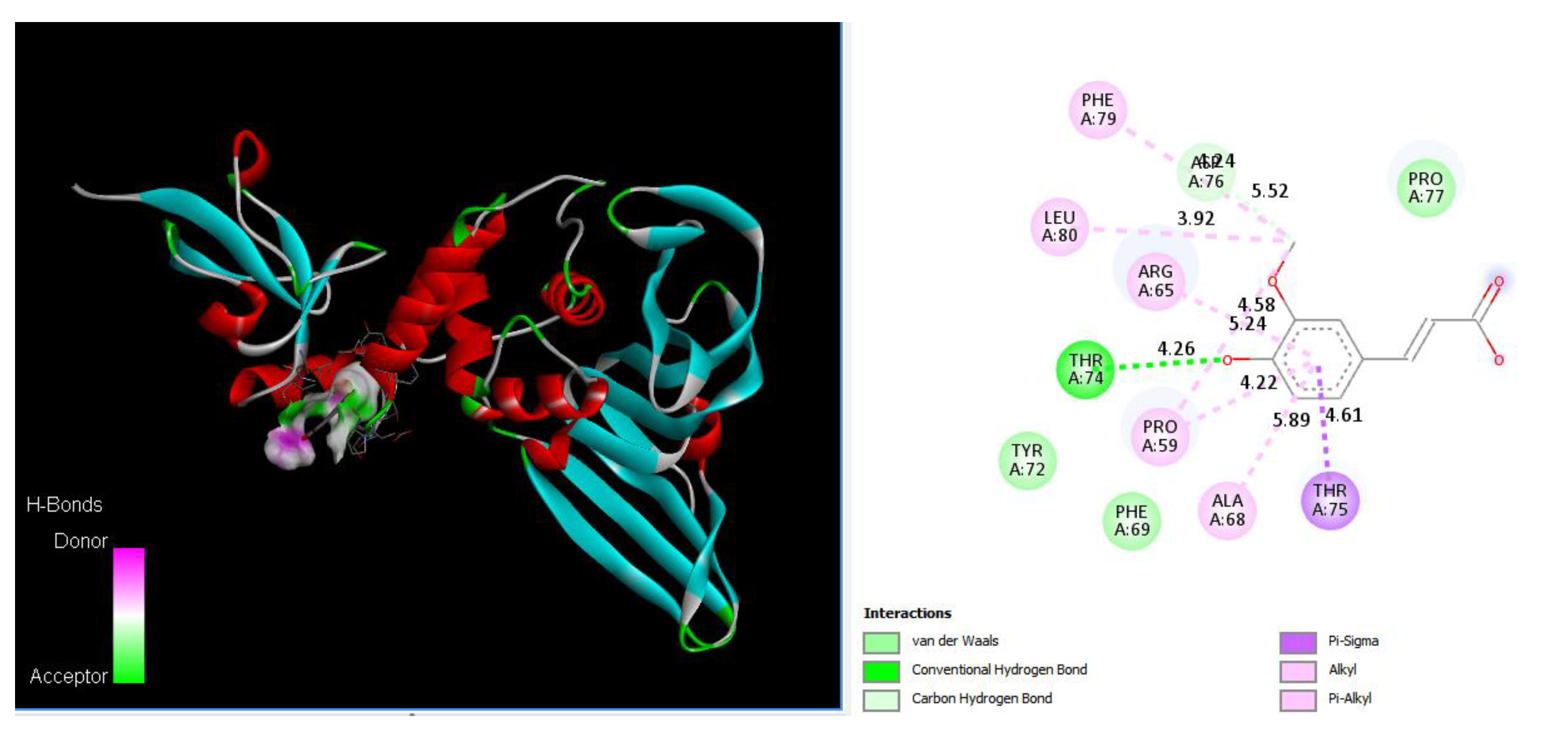
As shown in Figure 5, Ferulic acid binds to Thr74 via a hydrogen bond with a distance 4.26 Å, binds to Asp76 via a carbon HB with a distance 5.52 Å, and binds to Pro59 (4.22 Å, 5.24 Å), Arg65 (4.58 Å), Ala68 (5.89 Å), Phe79 (4.04 Å), and Leu80 (3.92 Å) via alkyl bonds. We also found that Chain B1 (N297 T299), Chain B2 (S303), and Chain B3 (D303 T304 N306 H308) of serine carboxypeptidase (PDB: 3sc2) can bind with V188 K190 K218 G219 Q221 Q229 E252 K315 of PLpro in SARS-CoV-2.
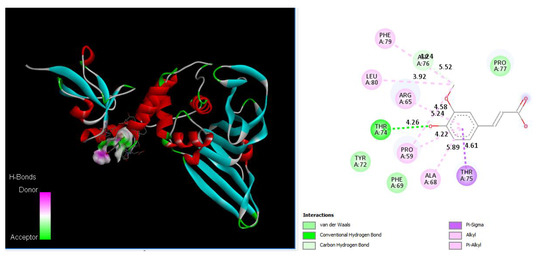
Figure 5.
Ferulic acid docking PLpro by binding to Thr74 via a hydrogen bond (4.26 Å), binding to Asp76 via a carbon HB (3.42 Å), binding to Pro59 (4.22 Å, 4.58 Å), Arg65 (5.24 Å), Ala68 (5.89 Å), and Phe79 (5.52 Å), and Leu80 (3.92 Å) via alkyl bonds, and binding to Thr75 via a pi-sigma bond (4.61 Å). A 3D image showing hydrogen donor and acceptor (left) and a 2D image showing types of bonds and distances (right).
3.10. Molecular Docking of PLpro
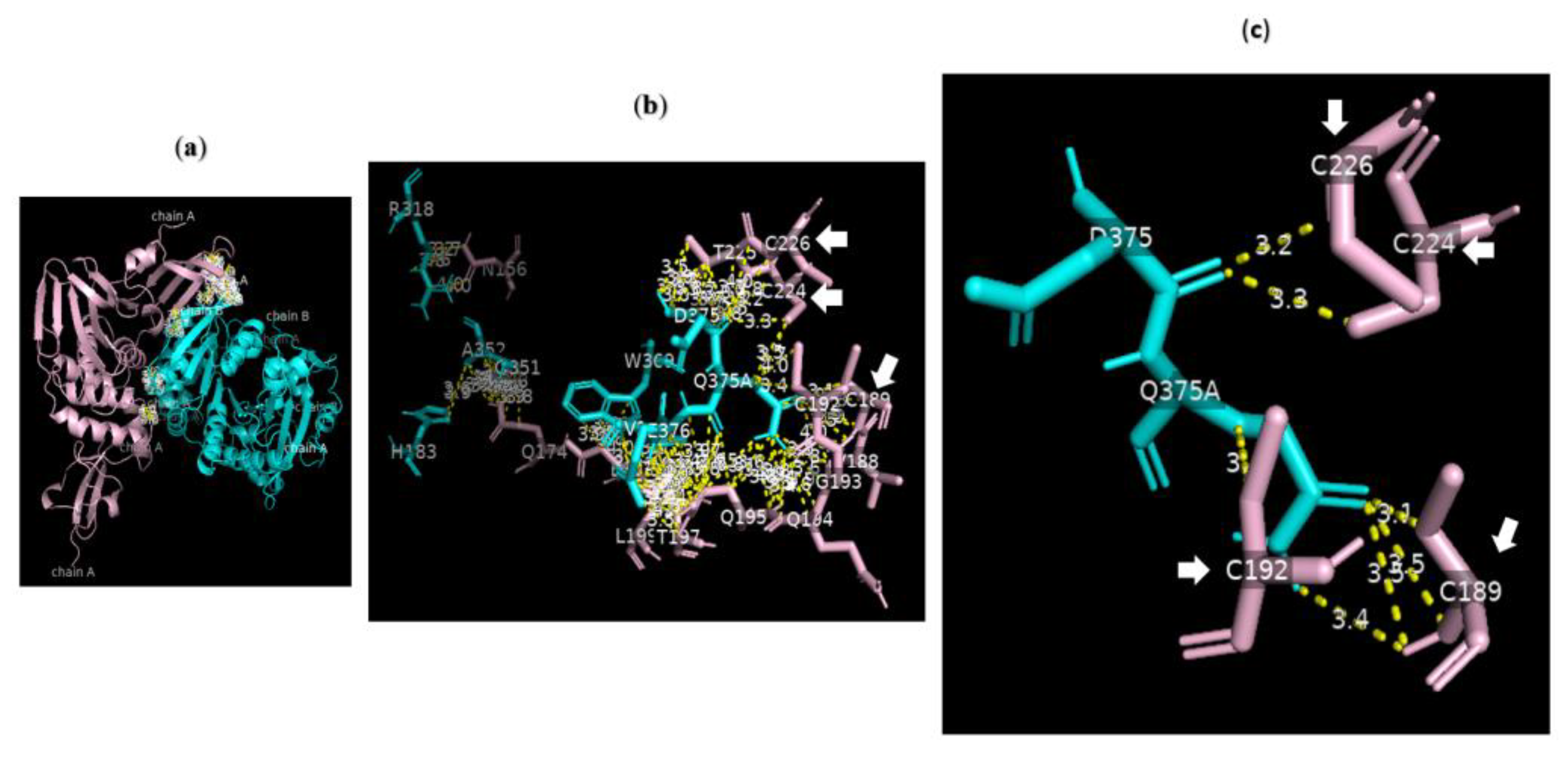
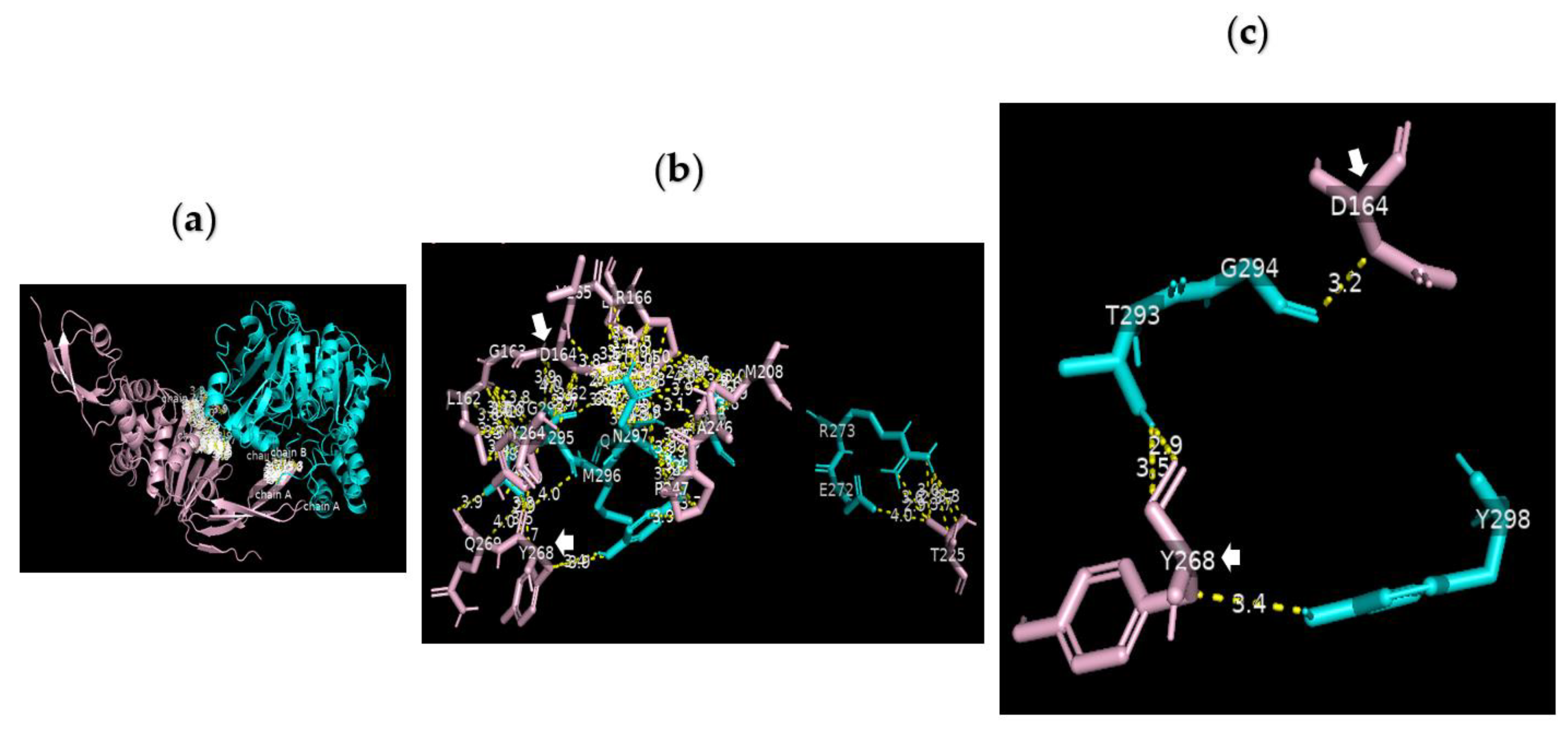
Analysis of the data collected from the molecular docking showed that serine carboxypeptidase can dock PLpro with a binding affinity (ΔG) equal to −7.8 kcal mol−1 and a dissociation constant (Kd) equal to 2.0 × 10−6 M, RMSD = 6.73 Å, and TM-score = 0.092 if normalized by a length of 7nfv, or TM-score = 0.21743 if normalized by a length of 3sc2 (Figure 6 and Figure 7). The data showed that H183, R318, G351 A352 W369 D375 Q375A E376 V377 of serine carboxypeptidase can bind to multiple residues including N156 Q174 V188 C189 C192 G193 Q194 Q195 T197 L199 E203 C224 T225 C226 of PLpro in SARS-CoV-2. Additionally, E272 R273 Q285 N291 T293 G294 A295 M296 N297 Y298 T299 of serine carboxypeptidase can bind to L162 G163 D164 V165 R166 E167 M208 T225 A246 P247 Y264 Y268 Q269 Y273 of PLpro in SARS-CoV-2.
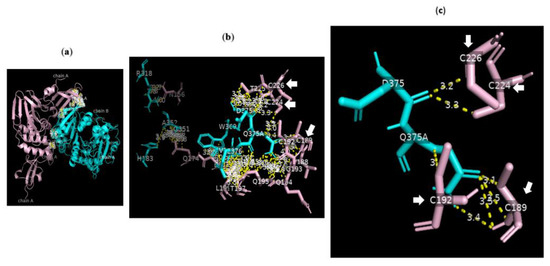
Figure 6.
Serine carboxypeptidase can bind to PLpro: 3D images showing (a) the PPI between the whole structure of serine carboxypeptidase (cyan) and that of PLpro (pink); (b) the key residues in the interface between H183, R318, G351 A352 W369 D375 Q375A E376 V377 of serine carboxypeptidase (cyan) and N156 Q174 V188 C189 C192 G193 Q194 Q195 T197 L199 E203 C224 T225 C226 of PLpro (pink) in SARS-CoV-2 using a cut-off of 4.0 Å; and (c) the binding sites between D375 of chain B8 of serine carboxypeptidase and C224 and C226 of PLpro as well as those between Q375 of chain B9 of serine carboxypeptidase and C189 and C192 of PLpro using a cut-off of 3.5 Å.
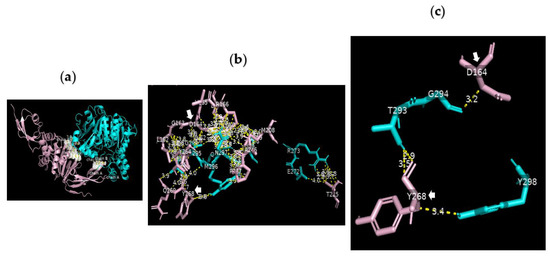
Figure 7.
Serine carboxypeptidase can bind to PLpro: 3D images showing (a) the PPI between the whole structure of serine carboxypeptidase (cyan) and that of PLpro (pink); (b) the interface between E272 R273 Q285 N291 T293 G294 A295 M296 N297 Y298 T299 of serine carboxypeptidase (cyan) and L162 G163 D164 V165 R166 E167 M208 T225 A246 P247 Y264 Y268 Q269 Y273 of PLpro (pink) in SARS-CoV-2 using a cut-off of 4.0 Å; and (c) the binding sites between T293 and Y298 of serine carboxypeptidase and the key residues of PLpro (D164 and Y268) using a cut-off of 3.5 Å.
4. Discussion
The COVID-19 pandemic has been ongoing for almost three years, yet there remains an urgent need for new therapies that can curtail the spread of the disease and reduce mortality. The present study examined the anti-COVID-19 potential of Biobran/MGN-3 and the mechanisms underlying its effects. The results showed that Biobran is a potent anti-COVID-19 agent: it significantly inhibits ACE2–SARS-CoV-2 S-protein RBD binding, it significantly suppresses SARS-CoV-2 gene expression and protein levels, and it has a significant cytotoxic effect on Vero E6 cells with a high selectivity index.
For Vero cells infected with SARS-CoV-2, Biobran reduced the viral load by 91.9% at a dose of 100 μg/mL. This highly significant viral reduction is in accordance with other recent studies. Whole aqueous extract of Adhatoda Vasica reduced the viral load in Vero cells infected with SARS-CoV-2 [55], the combination of hydroxychloroquine and azithromycin shortened the duration of SARS-CoV-2 viral load in COVID-19 patients [56], and the oral disinfectant povidone-iodine (PVP-I) inhibited SARS-CoV-2 in vitro [57]. A wide range of Biobran concentrations (3.1–100 μg/mL) were studied here to establish the dose–response relationship of its efficacy against SARS-CoV-2. Biobran caused a 91.9% viral reduction in Vero cells at a dose of 100 μg/mL and a 16.1% viral reduction at a concentration of 3.125 μg/mL.
These data are also consistent with Biobran’s effect on SARS-CoV-2 replication, as analyzed with RT-PCR assays. Biobran treatment inhibited SARS-CoV-2 replication with an EC50 (half-maximal effective concentration) of 3.46 μg/mL and an IC50 (half-maximal inhibitory concentration) against Vero E6 of 15.36 μg/mL, resulting in a high selectivity index (EC50/IC50) of 22.5. This ratio quantifies antiviral activity relative to cellular toxicity. Recent studies have reported that drugs with high SI values (>10) could potentially be used as antiviral drugs, and, theoretically, the more elevated the SI ratio, the more efficient and safer a drug is [20,58].
Biobran strongly inhibited PLpro activity and downregulated the expression of RdRp, resulting in the downregulating of the viral structural protein and the preventing of viral replication. Treatment with Biobran at a concentration of 10 μg/mL caused an 87% inhibition of PLpro. Recent work has studied several SARS-CoV-2 PLpro inhibitors. For example, a topical antiseptic Acriflavine solution has been shown to inhibit PLpro and have antiviral effects [59], cysteine reactive Ebselen and Disulfiram compounds have been identified as SARS-CoV and MERS-CoV PLpro inhibitors [60], and several drugs for Hepatitis C virus (HCV) have shown inhibitory effects against SARS-CoV-2 PLpro. The latter is based on the observation that the substrate binding cleft and active site of the SARS-CoV-2 Mpro are structurally similar to the active site of the HCV NS3/4A protease [61,62]. HCV protease inhibitors show differential potency based on the detection of SARS-CoV-2 spike protein in Vero E6 cells [63].
Furthermore, Biobran suppressed SARS-CoV-2 gene expression and protein levels. Treatment with Biobran at different doses (50 μg/mL, 25 μg/mL, 12.5 μg/mL, and 6.25 μg/mL) caused significant E-gene down regulation (92.7%, 73.4%, 56.0%, and 21%, respectively) and E-protein inhibition (91.3%, 75.3%, 49.6%, and 19.4%, respectively). These results are in accordance with other studies on the anti-SARS-CoV-2 activity of plant extracts. For example, S. nigra berries and flowers have been shown to exert inhibitory activity against ACE2 SARS-CoV-2 RBD binding in vitro [24]; S. nigra berry extract can reduce virus titers in Vero cells infected with infectious bronchitis virus (a pathogenic poultry coronavirus) [23]; Cannabis sativa extract has been found to decrease ACE2 protein levels [64]; and Agathis robusta bark essential oil has shown effectiveness against COVID-19 [20].
ACE2–SARS-CoV-2 RBD binding is key to viral entry into host cells, and its inhibition may be a potent therapeutic approach for SARS-CoV-2. The ACE2 enzyme has been reported to be the main receptor for SARS-CoV-2 viral entry [65]. Complex formation between the host cell receptor and viral particles is mediated by the receptor binding domain (RBD) of the viral spike proteins and the extracellular peptidase domain of the ACE2 receptor [66]. In this study, Biobran demonstrated a potent inhibitory capacity against ACE2–SARS-CoV-2 S-protein RBD binding in a dose-dependent manner, suggesting that Biobran can interfere with the entry of SARS-CoV-2 into host cells.
To investigate the binding further, the present in vitro study was complemented with in silico research in which it was found that Ferulic acid and serine carboxypeptidase, both of which are active ingredients of Biobran, can bind to DC-SIGN and PLpro at critical residues, thus impairing their actions. Our docking study collectively showed that cysteine residues in DC-SIGN are the main target of the SARS-CoV-2 RBD protein, and that Ferulic can bind to Arg266, Thr273, Gln297 and Arg300 Cys268 of DC-SIGN. DC-SIGN is a C-type lectin-like molecule, and a previous study has shown that cysteine residues are crucial in interactions with viral glycoproteins at the binding sites of C-type lectin-like molecules [67]. We found that among the multiple binding sites, S368 W369 of serine carboxypeptidase can bind multiple residues in DC-SIGN, especially critical peptides that extend from N349 N350 V351 G352, which are crucial residues in the binding site. Thus, serine carboxypeptidase can bind to and block the binding site of DC-SIGN, subsequently interfering with the ability of SARS-CoV-2 to enter cells.
Furthermore, Ferulic acid can dock PLpro in SARS-CoV-2 by binding to Thr74, Asp76, Pro59, Arg65, Ala68, Phe79 and by binding to Leu80 via alkyl bonds. For serine carboxypeptidase, we found that N297 T299 of Chain B1, S303 of Chain B1, and D303 T304 N306 H308 of Chain B3 can bind with V188 K190 K218 G219 Q221 Q229 E252 K315 of PLpro in SARS-CoV-2. We also detected multiple binding sites between serine carboxypeptidase and the three catalytic zinc finger domains (finger, palm, and thumb) of PLpro in SARS-CoV-2. Of these, we found a bond between C192 and C226 which is among the four essential cysteine residues (C189, C192, C224, and C226) that form a zinc binding site of the finger domain of PLpro in SARS-CoV-2. Thus, Ferulic acid and serine carboxypeptidase can hinder the replication cycle of SARS-CoV-2 by binding to PLpro, subsequently impairing the assembly of SARS-CoV-2 proteins and subgenomic RNA. We found that Asp375 and Gln375 (chain B8 and chain B9, respectively) of serine carboxypeptidase chelated the four essential cysteine residues (Cys224, 226, 189, and 192) that are critical sites for zinc binding in PLpro. A previous study indicated that zinc binding is essential for the structural integrity and protease activity of PLpro [68]. We also found that T293 G294 Y298 of serine carboxypeptidase can bind to D164 and Y268 of PLpro in SARS-CoV-2 (D164 and Y268 are essential for the maintenance of PLpro) [8]. It is of interest to note that a recent study indicated that the binding site sequence of SARS-CoV-2′s PLpro is 100% identitical to that in SARS-CoV [69]. This indicates that Biobran could be an efficient inhibitor across the entire range of the SARS-CoV family.
Biobran’s effect against a range of coronaviruses was also indicated by our previous study in which we reported that Biobran supplementation significantly reduces the incidence of influenza-like illnesses (ILI) in elderly subjects [26]. That study showed that Biobran significantly upregulated the expression levels of retinoic-acid inducible gene I (RIG-I) and melanoma differentiation-associated protein 5 (MDA-5), as well as downstream ISG15 and MX1 in the human pulmonary epithelial BEAS-2B cell lines. The induction of RIG-1 and MDA-5 post exposure to Biobran has significant implications, since recent studies have reported the crucial roles of RIG-1 and MDA-5 in sensing and mounting effective antiviral responses against SARS-CoV-2 [70,71]. In addition, ISG-ylation of the caspase activation and recruitment domains of MDA-5 promotes its oligomerization and thereby triggers the activation of innate immunity against a range of viruses, including coronaviruses [72].
The current study is limited in its exploration of Biobran’s active ingredients. Further study is required to examine the mechanisms of additional active ingredients that may be involved in Biobran’s protection against SARS-CoV-2. The study is also limited in scope to in vitro measurements. The applicability of the current observations to live subjects must be studied to draw conclusions about the generalizability of the current results. Nevertheless, Biobran holds great promise, and it is a truly unique biological response modifier. Biobran does not exhibit hyporesponsiveness [35], making it a unique and attractive agent for long-term preventive and/or therapeutic purposes. Biobran has also been shown to be a completely safe and nontoxic agent, its biosafety has been demonstrated in many studies [35,73,74], it has been shown to reduce chemotoxic effects in animals [75,76] and to improve quality of life parameters in cancer patients [42,77], and it has not shown any adverse side effects in humans or animals after long periods of treatment [35,78,79].
5. Conclusions
Vero cells cultured with SARS-CoV-2 were used to study Biobran’s anti-COVID-19 effect in vitro. Biobran significantly reduced the viral load and viral count in infected Vero cells with a high selectivity index, it strongly inhibited viral entry by interacting with the ACE2 enzyme on the cell surface and inhibiting spike protein interaction, and it prevented viral replication by inhibiting PLpro activity and downregulating the expression of RdRp. Computational modeling further showed that serine carboxypeptidase, an active ingredient in Biobran, interferes with the binding of SARS-CoV-2 to its receptor DC-SIGN on Vero cells, thus preventing the cell entry of SARS-CoV-2. In addition, it binds to PLpro, thus impairing the viral replication cycle. Biobran may be a very useful and safe product for preventing and treating COVID-19 as an adjunct therapy, and further validations of its anti-SARS-CoV-2 applications are warranted.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/nu15020453/s1, Figure S1–S4. Figure S1. 3D image for position (model 0) of docking of RBD domain of SARS-CoV-2 spike protein (cyan color) to DC-SIGN CRD (light orange color) showing PPI via binding of V382 S383 T385 L387 N388 F392 F515 E516 L517 of chain A to L252 H254 P255 C256 P257 W258 of chain D, respectively, using cut-off 4.0 Å.; Figure S2. 3D image for position (model 1) of docking of RBD domain of SARS-CoV-2 spike protein (cyan color) to DC-SIGN CRD (light orange color) showing PPI via binding of E340 R346 A347 F348 N354 chain A to E259 of chain E and H254 and C384 of chain C, respectively, using cut-off 4.0 Å.; Figure S3. 3D image for position (model 2) of docking of RBD domain of SARS-CoV-2 spike protein (cyan color) to DC-SIGN CRD (light orange color) showing PPI via binding of N360 R466 D467 I468 S469 T470 I472 G482 V483 E484 T523 of chain A to N276 H278 D279 E286 of chain B and E286 V287 K379 A382 S383 C384 of chain C, respectively, using cut-off 4.0 Å.; Figure S4. 3D image for position (model 3) of docking of RBD domain of SARS-CoV-2 spike protein (cyan color) to DC-SIGN CRD (light orange color) showing PPI via binding of G381 V382 S383 P384 L387 N388 F392 T430 L517 A522 of chain A to L252 C253 H254 P255 P257 W258 E259 V287 A382 S383 C384 of chain D, respectively, using cut-off 4.0 Å.
Author Contributions
Conceptualization, M.G. and S.A.; methodology, S.A. and H.H.F.; investigation, S.A. and H.H.F.; in vitro study, S.A.; in silico study, H.H.F.; writing—original draft preparation, M.G., S.A. and H.H.F.; writing—review and editing, M.G.; visualization, H.H.F. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from S.A. (for in vitro data) and from H.H.F. (for in silico data).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Hui, D.S.C.; Zumla, A. Severe acute respiratory synrome: Historical, epidemiologic, and clinical features. Infect. Dis. Clin. North Am. 2019, 33, 869–889. [Google Scholar] [CrossRef] [PubMed]
- De Groot, R.J.; Baker, S.C.; Baric, R.S.; Brown, C.S.; Drosten, C.; Enjuanes, L.; Fouchier, R.A.; Galiano, M.; Gorbalenya, A.E.; Memish, Z.A.; et al. Commentary: Middle East respiratory syndrome coronavirus (MERS-CoV): Announcement of the Coronavirus Study Group. J. Virol. 2013, 87, 7790–7792. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Tiwari, S.; Deb, M.K.; Marty, J.L. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): A global pandemic and treatment strategies. Int. J. Antimicrob. Agents 2020, 56, 106054. [Google Scholar] [CrossRef]
- WHO. World Health Organization. WHO Coronavirus Disease (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 15 October 2021).
- Umakanthan, S.; Sahu, P.; Ranade, A.V.; Bukelo, M.M.; Rao, J.S.; Abrahao-Machado, L.F.; Dahal, S.; Kumar, H.; Kv, D. Origin, transmission, diagnosis and management of coronavirus disease 2019 (COVID-19). Postgrad. Med. J. 2020, 96, 753–758. [Google Scholar]
- Sharma, A.; Ahmad Farouk, I.; Lal, S.K. COVID-19: A Review on the Novel Coronavirus Disease Evolution, Transmission, Detection, Control and Prevention. Viruses 2021, 13, 202. [Google Scholar] [CrossRef] [PubMed]
- Amraei, R.; Yin, W.; Napoleon, M.A.; Suder, E.L.; Berrigan, J.; Zhao, Q.; Olejnik, J.; Chandler, K.B.; Xia, C.; Feldman, J.; et al. CD209L/L-SIGN and CD209/DC-SIGN Act as Receptors for SARS-CoV-2. ACS Cent. Sci. 2021, 7, 1156–1165. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.; Mukherjee, R.; Grewe, D.; Bojkova, D.; Baek, K.; Bhattacharya, A.; Schulz, L.; Widera, M.; Mehdipour, A.R.; Tascher, G.; et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 2020, 587, 657–662. [Google Scholar] [CrossRef]
- World Health Organization Draft Landscape and Tracker of COVID-19 Candidate Vaccines. Available online: https://www.who.int/publications/m/item/draft-landscape-of-covid-19-candidate-vaccines (accessed on 30 May 2021).
- Fiolet, T.; Kherabi, Y.; MacDonald, C.J.; Ghosn, J.; Peiffer-Smadja, N. Comparing COVID-19 vaccines for their characteristics, efficacy and effectiveness against SARS-CoV-2 and variants of concern: A narrative review. Clin. Microbiol. Infect. 2022, 28, 202–221. [Google Scholar] [CrossRef]
- Kim, Y.S.; Kim, B.; Kwon, E.B.; Chung, H.S.; Choi, J.G. Mulberrofuran G, a mulberry component, prevents SARS-CoV-2 infection by blocking the interaction between SARS-CoV-2 spike protein S1 receptor-binding domain and human angiotensin-converting enzyme 2 receptor. Nutrients 2022, 14, 4170. [Google Scholar] [CrossRef]
- Yang, L.; Wang, Z. Natural products, alone or in combination with FDA-approved drugs, to treat COVID-19 and lung cancer. Biomedicines 2021, 9, 689. [Google Scholar] [CrossRef]
- Wang, D.; Huang, J.; Yeung, A.W.K.; Tzvetkov, N.T.; Horbanczuk, J.O.; Willschke, H.; Gai, Z.; Atanasov, A.G. The significance of natural product derivatives and traditional medicine for COVID-19. Processes 2020, 8, 937. [Google Scholar] [CrossRef]
- Wang, Z.; Yang, L.; Song, X.Q. Oral GS-441524 derivatives: Next-generation inhibitors of SARS-CoV-2 RNA-dependent RNA polymerase. Front. Immunol. 2022, 13, 1015355. [Google Scholar] [CrossRef]
- Brahmi, F.; Vejux, A.; Ghzaiel, I.; Ksila, M.; Zarrouk, A.; Ghrairi, T.; Essadek, S.; Mandard, S.; Leoni, V.; Poli, G.; et al. Role of diet and nutrients in SARS-CoV-2 infection: Incidence on oxidative stress, inflammatory status and viral production. Nutrients 2022, 14, 2194. [Google Scholar] [CrossRef] [PubMed]
- Mani, J.S.; Johnson, J.B.; Steel, J.C.; Broszczak, D.A.; Neilsen, P.M.; Walsh, K.B.; Naiker, M. Natural product-derived phytochemicals as potential agents against coronaviruses: A review. Virus Res. 2020, 284, 197989. [Google Scholar] [CrossRef]
- Orhan, I.E.; Senol Deniz, F.S. Natural products as potential leads against coronaviruses: Could they be encouraging structural models against SARS-CoV-2? Nat. Prod. Bioprospect. 2020, 10, 171–186. [Google Scholar] [CrossRef]
- Russo, M.; Moccia, S.; Spagnuolo, C.; Tedesco, I.; Russo, G.L. Roles of flavonoids against coronavirus infection. Chem. Biol. Interact. 2020, 328, 109211. [Google Scholar] [CrossRef]
- Paraiso, I.L.; Revel, J.S.; Stevens, J.F. Potential use of polyphenols in the battle against COVID-19. Curr. Opin. Food Sci. 2020, 32, 149–155. [Google Scholar] [CrossRef]
- Mohamed, M.E.; Tawfeek, N.; Elbaramawi, S.S.; Fikry, E. Agathis robusta Bark Essential Oil Effectiveness against COVID-19: Chemical Composition, In Silico and In Vitro Approaches. Plants 2022, 11, 663. [Google Scholar] [CrossRef]
- Ali, A.M.; Kunugi, H. Propolis, Bee Honey, and Their Components Protect against Coronavirus Disease 2019 (COVID-19): A Review of In Silico, In Vitro, and Clinical Studies. Molecules 2021, 26, 1232. [Google Scholar] [CrossRef] [PubMed]
- Weng, J.R.; Lin, C.S.; Lai, H.C.; Lin, Y.P.; Wang, C.Y.; Tsai, Y.C.; Wu, K.C.; Huang, S.H.; Lin, C.W. Antiviral activity of Sambucus Formosan Nakai ethanol extract and related phenolic acid constituents against human coronavirus NL63. Virus Res. 2019, 273, 197767. [Google Scholar] [CrossRef]
- Chen, C.; Zuckerman, D.M.; Brantley, S.; Sharpe, M.; Childress, K.; Hoiczyk, E.; Pendleton, A.R. Sambucus nigra extracts inhibit infectious bronchitis virus at an early point during replication. BMC Vet. Res. 2014, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Boroduske, A.; Jekabsons, K.; Riekstina, U.; Muceniece, R.; Rostoks, N.; Nakurte, I. Wild Sambucus nigra L. from north-east edge of the species range: A valuable germplasm with inhibitory capacity against SARS-CoV2 S-protein RBD and hACE2 binding in vitro. Ind. Crops Prod. 2021, 165, 113438. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Wei, J.; Huang, T.; Lei, L.; Shen, C.; Lai, J.; Yang, M.; Liu, L.; Yang, Y.; Liu, G.; et al. Resveratrol inhibits the replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in cultured Vero cells. Phytother. Res. 2021, 35, 1127–1129. [Google Scholar] [CrossRef]
- Elsaid, A.F.; Agrawal, S.; Agrawal, A.; Ghoneum, M. Dietary Supplementation with Biobran/MGN-3 Increases Innate Resistance and Reduces the Incidence of Influenza-like Illnesses in Elderly Subjects: A Randomized, Double-Blind, Placebo-Controlled Pilot Clinical Trial. Nutrients 2021, 13, 4133. [Google Scholar] [CrossRef] [PubMed]
- Tazawa, K.; Ichihashi, K.; Fujii, T.; Omura, K.; Anazawa, M.; Maeda, H. The orally administration of the Hydrolysis Rice Bran prevents a common cold syndrome for the elderly people based on immunomodulatory function. J. Trad. Med. 2003, 20, 132–141. [Google Scholar]
- Ghoneum, M. Anti-HIV activity in vitro of MGN-3, an activated arabinoxylan from rice bran. Biochem. Biophys. Res. Commun. 1998, 243, 25–29. [Google Scholar] [CrossRef]
- Salama, H.; Medhat, E.; Shaheen, M.; Zekri, A.N.; Darwish, T.; Ghoneum, M. Arabinoxylan rice bran (Biobran) suppresses the viremia level in patients with chronic HCV infection: A randomized trial. Int. J. Immunopathol. Pharmacol. 2016, 29, 647–653. [Google Scholar] [CrossRef]
- Badr El-Din, N.K.; Noaman, E.; Ghoneum, M. In vivo tumor inhibitory effects of nutritional rice bran supplement MGN-3/Biobran on Ehrlich carcinoma-bearing mice. Nutr. Cancer 2008, 60, 235–244. [Google Scholar] [CrossRef]
- Bang, M.H.; Van Riep, T.; Thinh, N.T.; Song, L.H.; Dung, T.T.; Truong, L.V.; Don, L.V.; Ky, T.D.; Pan, D.; Shaheen, M.; et al. Arabinoxylan rice bran (MGN-3) enhances the effects of interventional therapies for the treatment of hepatocellular carcinoma: A three year randomized clinical trial. Anticancer Res. 2010, 30, 5145–5151. [Google Scholar]
- Ghoneum, M. Enhancement of human natural killer cell activity by modified arabinoxylan from rice bran (MGN-3). Int. J. Immunother. 1998, 14, 89–99. [Google Scholar]
- Ghoneum, M.; Jewett, A. Production of tumor necrosis factor alpha and interferon-gamma from human peripheral blood lymphocytes by MGN-3, a modified arabinoxylan from rice bran, and its synergy with interleukin-2 in vitro. Cancer Detect. Prev. 2000, 24, 314–324. [Google Scholar] [PubMed]
- Ghoneum, M.; Abedi, S. Enhancement of natural killer cell activity of aged mice by modified arabinoxylan rice bran (MGN-3/Biobran). J. Pharm. Pharmacol. 2004, 56, 1581–1588. [Google Scholar] [CrossRef] [PubMed]
- Ghoneum, M.; Brown, J. NK immunorestoration of cancer patients by, M.G.N.-3.; a modified arabinoxylan rice bran (study of 32 patients followed for up to 4 years). In Anti-Aging Medical Therapeutics; Klatz, R., Goldman, R., Eds.; Health Quest Publications Marina del Rey: Belmont, CA, USA, 1999; Volume III, pp. 217–226. [Google Scholar]
- Cholujova, D.; Jakubikova, J.; Czako, B.; Martisova, M.; Hunakova, L.; Duraj, J.; Mistrik, M.; Sedlak, J. MGN-3 arabinoxylan rice bran modulates innate immunity in multiple myeloma patients. Cancer Immunol. Immunother. 2013, 62, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Cholujova, D.; Jakubikova, J.; Sedlak, J. Biobran-augmented maturation of human monocyte-derived dendritic cells. Neoplasma 2009, 56, 89–95. [Google Scholar] [CrossRef]
- Ghoneum, M.; Agrawal, S. Activation of human monocyte derived dendritic cells in vitro by the biological response modifier arabinoxylan rice bran (MGN-3/Biobran). Int. J. Immunopathol. Pharmacol. 2011, 24, 941–948. [Google Scholar] [CrossRef]
- Ghoneum, M.; Agrawal, S. MGN-3/biobran enhances generation of cytotoxic CD8+ T cells via upregulation of dec-205 expression on dendritic cells. Int. J. Immunopathol. Pharmacol. 2014, 27, 523–530. [Google Scholar] [CrossRef]
- Ghoneum, M.; Matsuura, M. Augmentation of macrophage phagocytosis by modified arabinoxylan rice bran (MGN-3/biobran). Int. J. Immunopathol. Pharmacol. 2004, 17, 283–292. [Google Scholar] [CrossRef]
- Elsaid, A.F.; Fahmi, R.M.; Shaheen, M.; Ghoneum, M. The enhancing effects of Biobran/MGN-3, an arabinoxylan rice bran, on healthy old adults’ health-related quality of life: A randomized, double-blind, placebo-controlled clinical trial. Qual. Life Res. 2020, 29, 357–367. [Google Scholar] [CrossRef]
- Takahara, K.; Sano, K. The life prolongation and QOL improvement effect of rice bran arabinoxylan derivative (MGN-3. Bio-Bran) for progressive cancer. Clin. Pharmacol. Ther. 2004, 14, 267–271. [Google Scholar]
- Yin, Z.N.; Wu, W.J.; Sun, C.Z.; Liu, H.F.; Chen, W.B.; Zhan, Q.P.; Lei, Z.G.; Xin, X.; Ma, J.J.; Yao, K.; et al. Antioxidant and Anti-inflammatory Capacity of Ferulic Acid Released from Wheat Bran by Solid-state Fermentation of Aspergillus niger. Biomed. Environ. Sci. 2019, 32, 11–21. [Google Scholar] [CrossRef]
- Liao, D.I.; Remington, S.J. Structure of wheat II at 3.5-A resolution. A new class of serine proteinase. J. Biol. Chem. 1990, 265, 6528–6531. [Google Scholar] [CrossRef] [PubMed]
- Mendez-Encinas, M.A.; Carvajal-Millan, E.; Rascon-Chu, A.; Astiazaran-Garcia, H.F.; Valencia-Rivera, D.E. Ferulated Arabinoxylans and Their Gels: Functional Properties and Potential Application as Antioxidant and Anticancer Agent. Oxidative Med. Cell. Longev. 2018, 2018, 2314759. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef]
- Liu, Y.; Peterson, D.A.; Kimura, H.; Schubert, D. Mechanism of cellular 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) reduction. J. Neurochem. 1997, 69, 581–593. [Google Scholar] [CrossRef]
- Landry, M.L.; Stanat, S.; Biron, K.; Brambilla, D.; Britt, W.; Jokela, J.; Chou, S.; Drew, W.L.; Erice, A.; Gilliam, B.; et al. A standardized plaque reduction assay for determination of drug susceptibilities of cytomegalovirus clinical isolates. Antimicrob. Agents Chemother. 2000, 44, 688–692. [Google Scholar] [CrossRef] [PubMed]
- Caly, L.; Druce, J.D.; Catton, M.G.; Jans, D.A.; Wagstaff, K.M. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antiviral Res. 2020, 178, 104787. [Google Scholar] [CrossRef]
- Xia, S.; Liu, M.; Wang, C.; Xu, W.; Lan, Q.; Feng, S.; Qi, F.; Bao, L.; Du, L.; Liu, S.; et al. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 2020, 30, 343–355. [Google Scholar] [CrossRef]
- Chu, D.K.W.; Pan, Y.; Cheng, S.M.S.; Hui, K.P.Y.; Krishnan, P.; Liu, Y.; Ng, D.Y.M.; Wan, C.K.C.; Yang, P.; Wang, Q.; et al. Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of pneumonia. Clin. Chem. 2020, 66, 549–555. [Google Scholar] [CrossRef]
- Hong, J.; Wang, Q.; Wu, Q.; Chen, J.; Wang, X.; Wang, Y.; Chen, Y.; Xia, N. Rabbit Monoclonal Antibody Specifically Recognizing a Linear Epitope in the RBD of SARS-CoV-2 Spike Protein. Vaccines 2021, 9, 829. [Google Scholar] [CrossRef] [PubMed]
- Hsu, K.C.; Chen, Y.F.; Lin, S.R.; Yang, J.M. iGEMDOCK: A graphical environment of enhancing GEMDOCK using pharmacological interactions and post-screening analysis. BMC Bioinform. 2011, 12 (Suppl. S1), S33. [Google Scholar] [CrossRef]
- Kozakov, D.; Hall, D.R.; Xia, B.; Porter, K.A.; Padhorny, D.; Yueh, C.; Beglov, D.; Vajda, S. The ClusPro web server for protein-protein docking. Nat. Protoc. 2017, 12, 255–278. [Google Scholar] [CrossRef]
- Gheware, A.; Dholakia, D.; Kannan, S.; Panda, L.; Rani, R.; Pattnaik, B.R.; Jain, V.; Parekh, Y.; Enayathullah, M.G.; Bokara, K.K.; et al. Adhatoda Vasica attenuates inflammatory and hypoxic responses in preclinical mouse models: Potential for repurposing in COVID-19-like conditions. Respir. Res. 2021, 22, 99. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B.; Le Bideau, M.; Andreani, J.; Hoang, V.T.; Grimaldier, C.; Colson, P.; Gautret, P.; Raoult, D. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 1059–1061. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, Y.; Wang, Q.; Zhu, J.; Shi, W.; Han, Z.; Zhang, Y.; Chen, K. Virucidal effect of povidone-iodine against SARS-CoV-2 in vitro. J. Int. Med. Res. 2021, 49, 3000605211063695. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Twilley, D.; Esmear, T.; Oosthuizen, C.B.; Reid, A.M.; Nel, M.; Lall, N. Anti-SARS-CoV natural products with the potential to inhibit SARS-CoV-2 (COVID-19). Front. Pharmacol. 2020, 11, 1514. [Google Scholar] [CrossRef]
- Napolitano, V.; Dabrowska, A.; Schorpp, K.; Mourão, A.; Barreto-Duran, E.; Benedyk, M.; Botwina, P.; Brandner, S.; Bostock, M.; Chykunova, Y.; et al. Acriflavine, a Clinically Approved Drug, Inhibits SARS-CoV-2 and Other Betacoronaviruses. Cell Chem. Biol. 2022, 29, 774–784. e8. [Google Scholar] [CrossRef]
- Lin, M.H.; Moses, D.C.; Hsieh, C.H.; Cheng, S.C.; Chen, Y.H.; Sun, C.Y.; Chou, C.Y. Disulfiram Can Inhibit MERS and SARS Coronavirus Papain-like Proteases via Different Modes. Antiviral Res. 2018, 150, 155–163. [Google Scholar] [CrossRef]
- Bafna, K.; Cioffi, C.L.; Krug, R.M.; Montelione, G.T. Structural similarities between SARS-CoV2 3CLpro and other viral proteases suggest potential lead molecules for developing broad spectrum antivirals. Front. Chem. 2022, 10, 948553. [Google Scholar] [CrossRef]
- Bafna, K.; White, K.; Harish, B.; Rosales, R.; Ramelot, T.A.; Acton, T.B.; Moreno, E.; Kehrer, T.; Miorin, L.; Royer, C.A.; et al. Hepatitis C virus drugs that inhibit SARS-CoV-2 papain-like protease synergize with remdesivir to suppress viral replication in cell culture. Cell Rep. 2021, 35, 109133. [Google Scholar] [CrossRef]
- Gammeltoft, K.A.; Zhou, Y.; Duarte Hernandez, C.R.; Galli, A.; Offersgaard, A.; Costa, R.; Pham, L.V.; Fahnøe, U.; Feng, S.; Scheel, T.K.H.; et al. Hepatitis C Virus Protease Inhibitors Show Differential Efficacy and Interactions with Remdesivir for Treatment of SARS-CoV-2 In Vitro. Antimicrob. Agents Chemother. 2021, 65, e02680-20. [Google Scholar] [CrossRef]
- Wang, B.; Kovalchuk, A.; Li, D.; Rodriguez-Juarez, R.; Ilnytskyy, Y.; Kovalchuk, I.; Kovalchuk, O. In search of preventive strategies: Novel high-CBD Cannabis sativa extracts modulate ACE2 expression in COVID-19 gateway tissues. Aging 2020, 12, 22425–22444. [Google Scholar]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [PubMed]
- Chaipan, C.; Soilleux, E.J.; Simpson, P.; Hofmann, H.; Gramberg, T.; Marzi, A.; Geier, M.; Stewart, E.A.; Eisemann, J.; Steinkasserer, A.; et al. DC-SIGN and CLEC-2 mediate human immunodeficiency virus type 1 capture by platelets. J. Virol. 2006, 80, 8951–8960. [Google Scholar] [CrossRef] [PubMed]
- Barretto, N.; Jukneliene, D.; Ratia, K.; Chen, Z.; Mesecar, A.D.; Baker, S.C. The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity. J. Virol. 2005, 79, 15189–15198. [Google Scholar] [CrossRef]
- Ibrahim, T.M.; Ismail, M.I.; Bauer, M.R.; Bekhit, A.A.; Boeckler, F.M. Supporting SARS-CoV-2 Papain-Like Protease Drug Discovery: In silico Methods and Benchmarking. Front. Chem. 2020, 8, 592289. [Google Scholar] [CrossRef]
- Yamada, T.; Sato, S.; Sotoyama, Y.; Orba, Y.; Sawa, H.; Yamauchi, H.; Sasaki, M.; Takaoka, A. RIG-I triggers a signaling-abortive anti-SARS-CoV-2 defense in human lung cells. Nat. Immunol. 2021, 22, 820–828. [Google Scholar] [CrossRef] [PubMed]
- Sampaio, N.G.; Chauveau, L.; Hertzog, J.; Bridgeman, A.; Fowler, G.; Moonen, J.P.; Dupont, M.; Russell, R.A.; Noerenberg, M.; Rehwinkel, J. The RNA sensor MDA5 detects SARS-CoV-2 infection. Sci. Rep. 2021, 11, 13638. [Google Scholar] [CrossRef]
- Liu, G.; Lee, J.H.; Parker, Z.M.; Acharya, D.; Chiang, J.J.; van Gent, M.; Riedl, W.; Davis-Gardner, M.E.; Wies, E.; Chiang, C.; et al. ISG15-dependent activation of the sensor MDA5 is antagonized by the SARS-CoV-2 papain-like protease to evade host innate immunity. Nat. Microbiol. 2021, 6, 467–478. [Google Scholar] [CrossRef] [PubMed]
- Daiwa Pharmaceutical Co., Ltd. BioBran Rice Bran Arabinoxylan Compound. Available online: http://www.daiwa-pharm.com/english/product/biobran.html (accessed on 22 October 2017).
- Tazawa, K. BioBran/MGN-3 (Rice Bran Arabinoxylan Deritive): Basic and Clinical Application to Integrative Medicine; Iyakushuppan Co. Publishers: Tokyo, Japan, 2006; pp. 18–22. [Google Scholar]
- Endo, Y.; Kanbayashi, H. Modified rice bran beneficial for weight loss of mice as a major and acute adverse effect of cisplatin. Pharmacol Toxicol. 2003, 92, 300–303. [Google Scholar] [CrossRef]
- Jacoby, H.I.; Wnorowski, G.; Sakata, K.; Maeda, H. The effect of MGN-3 on cisplatin and doxorubicin induced toxicity in the rat. J. Nutraceuticals Funct. Med. Foods 2001, 3, 3–11. [Google Scholar] [CrossRef]
- Hajto, T.; Horvath, A.; Papp, S. Improvement of quality of life in tumor patients after an immunomodulatory treatment with standardized mistletoe lectin and arabinoxylan plant extracts. Int. J. Neurorehabilit. 2016, 3, 2376-0281. [Google Scholar] [CrossRef]
- Ghoneum, M. Immunostimulation and cancer prevention. In Proceedings of the 7th International Congress on Anti-aging and Biomedical Technologies, Las Vegas, NV, USA, 11–13 December 1999. [Google Scholar]
- Badr El-Din, N.K.; Abdel Fattah, S.M.; Pan, D.; Tolentino, L.; Ghoneum, M. Chemopreventive Activity of MGN-3/Biobran Against Chemical Induction of Glandular Stomach Carcinogenesis in Rats and Its Apoptotic Effect in Gastric Cancer Cells. Integr. Cancer Ther. 2016, 15, NP26–NP34. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).