The Microbiome, Epigenome, and Diet in Adults with Obesity during Behavioral Weight Loss
Abstract
:1. Introduction
2. Materials and Methods
2.1. Participants and Study Design
2.2. Data Collection
2.2.1. Clinical Assessments and Anthropometrics
2.2.2. DNA Methylation and Data Pre-Processing
2.2.3. Microbiome Sequencing and Pre-Processing
2.2.4. Dietary Intake Assessment and Data Pre-Processing
2.2.5. Plasma Targeted Metabolomics
2.3. Statistical Analysis
3. Results
3.1. Participant Demographic and Clinical Characteristics
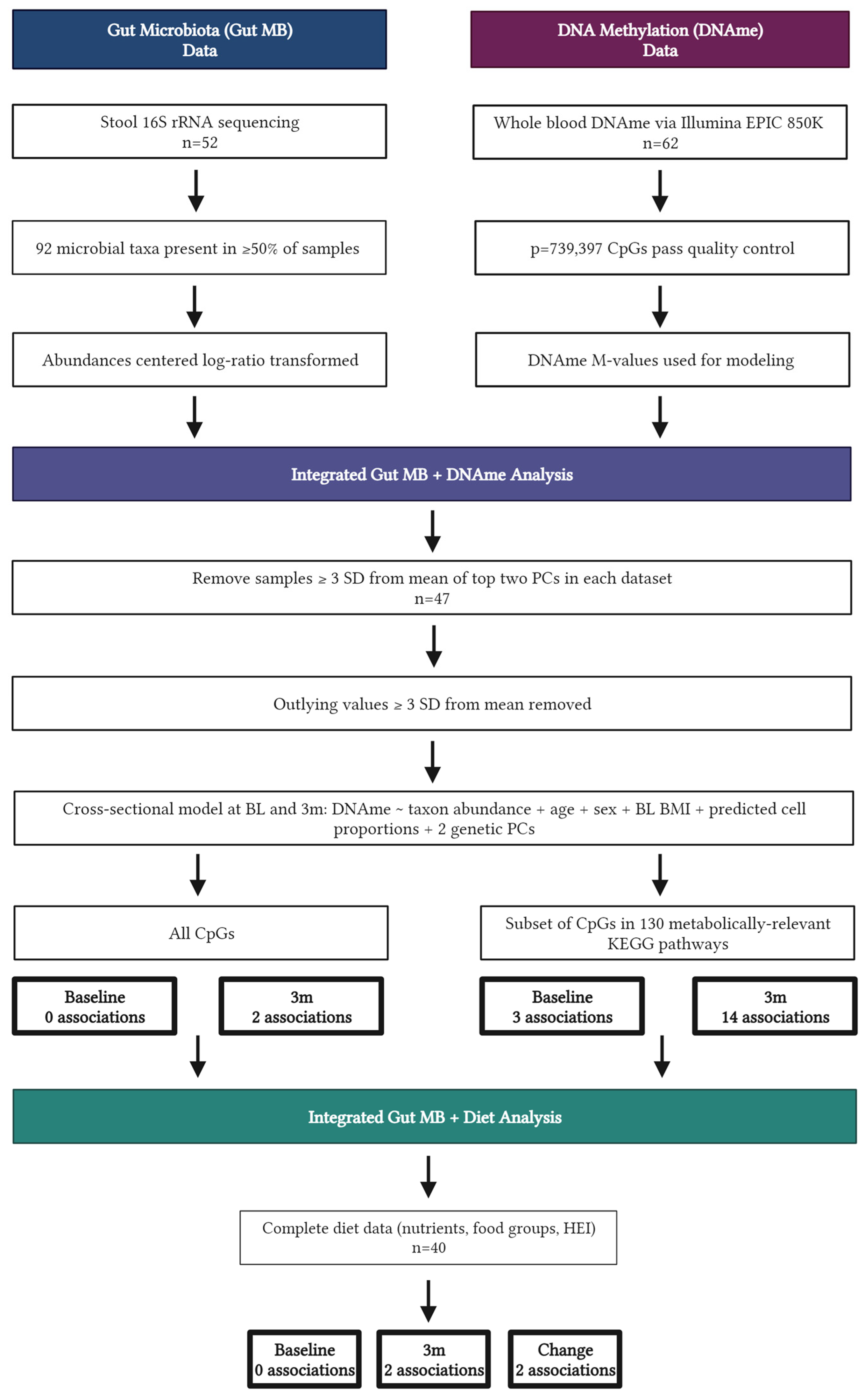
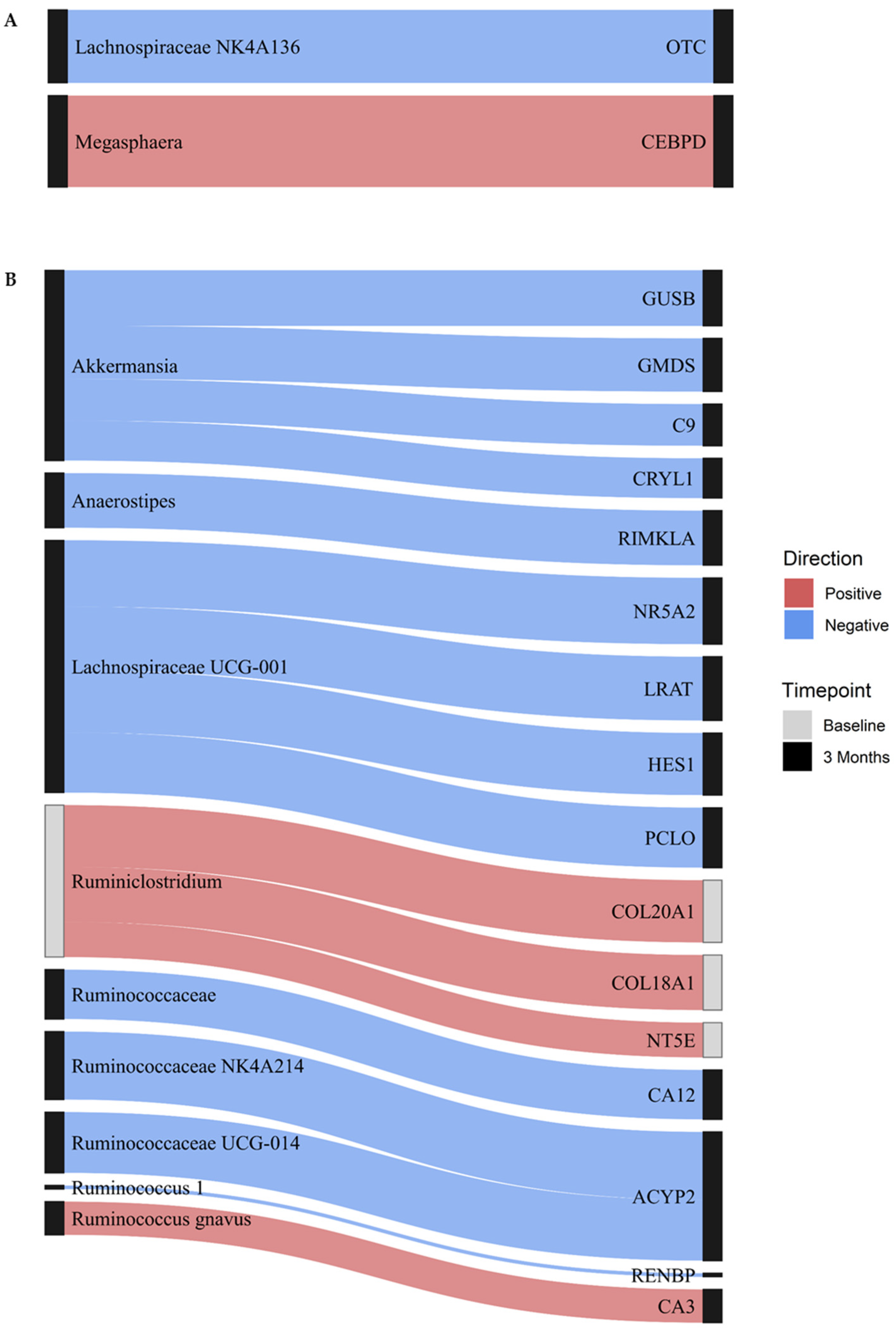
3.2. Integrated Microbiome and DNAme Analysis
3.3. Integrated Dietary Analysis
3.4. Targeted Metabolomics Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- van Dijk, S.J.; Molloy, P.L.; Varinli, H.; Morrison, J.L.; Muhlhausler, B.S.; Buckley, M.; Clark, S.J.; McMillen, I.C.; Noakes, M.; Samaras, K.; et al. Epigenetics and human obesity. Int. J. Obes. 2015, 39, 85–97. [Google Scholar] [CrossRef]
- Ling, C.; Rönn, T. Epigenetics in Human Obesity and Type 2 Diabetes. Cell Metab. 2019, 29, 1028–1044. [Google Scholar] [CrossRef] [PubMed]
- Rohde, K.; Keller, M.; la Cour Poulsen, L.; Blüher, M.; Kovacs, P.; Böttcher, Y. Genetics and epigenetics in obesity. Metabolism 2019, 92, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Meijnikman, A.S.; Gerdes, V.E.; Nieuwdorp, M.; Herrema, H. Evaluating Causality of Gut Microbiota in Obesity and Diabetes in Humans. Endocr. Rev. 2018, 39, 133–153. [Google Scholar] [CrossRef]
- Sonnenburg, J.L.; Backhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 2016, 535, 56–64. [Google Scholar] [CrossRef]
- Marzullo, P.; Di Renzo, L.; Pugliese, G.; De Siena, M.; Barrea, L.; Muscogiuri, G.; Colao, A.; Savastano, S. From obesity through gut microbiota to cardiovascular diseases: A dangerous journey. Int. J. Obes. Suppl. 2020, 10, 35–49. [Google Scholar] [CrossRef]
- Muscogiuri, G.; Cantone, E.; Cassarano, S.; Tuccinardi, D.; Barrea, L.; Savastano, S.; Colao, A.; on behalf of the Obesity Programs of nutrition, E.R.; Assessment, g. Gut microbiota: A new path to treat obesity. Int. J. Obes. Suppl. 2019, 9, 10–19. [Google Scholar] [CrossRef]
- Aron-Wisnewsky, J.; Warmbrunn, M.V.; Nieuwdorp, M.; Clément, K. Metabolism and Metabolic Disorders and the Microbiome: The Intestinal Microbiota Associated With Obesity, Lipid Metabolism, and Metabolic Health—Pathophysiology and Therapeutic Strategies. Gastroenterology 2021, 160, 573–599. [Google Scholar] [CrossRef]
- Maruvada, P.; Leone, V.; Kaplan, L.M.; Chang, E.B. The Human Microbiome and Obesity: Moving beyond Associations. Cell Host. Microbe 2017, 22, 589–599. [Google Scholar] [CrossRef] [PubMed]
- Seganfredo, F.B.; Blume, C.A.; Moehlecke, M.; Giongo, A.; Casagrande, D.S.; Spolidoro, J.V.N.; Padoin, A.V.; Schaan, B.D.; Mottin, C.C. Weight-loss interventions and gut microbiota changes in overweight and obese patients: A systematic review. Obes. Rev. 2017, 18, 832–851. [Google Scholar] [CrossRef]
- Stanislawski, M.A.; Frank, D.N.; Borengasser, S.J.; Ostendorf, D.M.; Ir, D.; Jambal, P.; Bing, K.; Wayland, L.; Siebert, J.C.; Bessesen, D.H.; et al. The Gut Microbiota during a Behavioral Weight Loss Intervention. Nutrients 2021, 13, 3248. [Google Scholar] [CrossRef] [PubMed]
- Carbonero, F. Human epigenetics and microbiome: The potential for a revolution in both research areas by integrative studies. Future Sci. OA 2017, 3, Fso207. [Google Scholar] [CrossRef] [PubMed]
- Peterson, L.W.; Artis, D. Intestinal epithelial cells: Regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 2014, 14, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Vallianou, N.; Stratigou, T.; Christodoulatos, G.S.; Dalamaga, M. Understanding the Role of the Gut Microbiome and Microbial Metabolites in Obesity and Obesity-Associated Metabolic Disorders: Current Evidence and Perspectives. Curr. Obes. Rep. 2019, 8, 317–332. [Google Scholar] [CrossRef]
- Bhat, M.I.; Kapila, R. Dietary metabolites derived from gut microbiota: Critical modulators of epigenetic changes in mammals. Nutr. Rev. 2017, 75, 374–389. [Google Scholar] [CrossRef] [PubMed]
- Woo, V.; Alenghat, T. Epigenetic regulation by gut microbiota. Gut Microbes 2022, 14, 2022407. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Li, Y.; Stoll, M.L.; Tollefsbol, T.O. The Epigenetic Connection Between the Gut Microbiome in Obesity and Diabetes. Front. Genet. 2019, 10, 1329. [Google Scholar] [CrossRef]
- Ostendorf, D.M.; Caldwell, A.E.; Zaman, A.; Pan, Z.; Bing, K.; Wayland, L.T.; Creasy, S.A.; Bessesen, D.H.; MacLean, P.; Melanson, E.L.; et al. Comparison of weight loss induced by daily caloric restriction versus intermittent fasting (DRIFT) in individuals with obesity: Study protocol for a 52-week randomized clinical trial. Trials 2022, 23, 718. [Google Scholar] [CrossRef] [PubMed]
- Siebert, J.C.; Stanislawski, M.A.; Zaman, A.; Ostendorf, D.M.; Konigsberg, I.R.; Jambal, P.; Ir, D.; Bing, K.; Wayland, L.; Scorsone, J.J.; et al. Multiomic Predictors of Short-Term Weight Loss and Clinical Outcomes During a Behavioral-Based Weight Loss Intervention. Obesity 2021, 29, 859–869. [Google Scholar] [CrossRef]
- Zhou, W.; Triche, T.J.; Laird, P.W.; Shen, H. SeSAMe: Reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nucleic Acids Res. 2018, 46, e123. [Google Scholar] [CrossRef]
- Zhou, W.; Laird, P.W.; Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 2017, 45, e22. [Google Scholar] [CrossRef]
- Houseman, E.A.; Accomando, W.P.; Koestler, D.C.; Christensen, B.C.; Marsit, C.J.; Nelson, H.H.; Wiencke, J.K.; Kelsey, K.T. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform. 2012, 13, 86. [Google Scholar] [CrossRef] [PubMed]
- Du, P.; Zhang, X.; Huang, C.-C.; Jafari, N.; Kibbe, W.A.; Hou, L.; Lin, S.M. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinform. 2010, 11, 587. [Google Scholar] [CrossRef]
- Hara, N.; Alkanani, A.K.; Ir, D.; Robertson, C.E.; Wagner, B.D.; Frank, D.N.; Zipris, D. Prevention of virus-induced type 1 diabetes with antibiotic therapy. J. Immunol. 2012, 189, 3805–3814. [Google Scholar] [CrossRef]
- Markle, J.G.; Frank, D.N.; Mortin-Toth, S.; Robertson, C.E.; Feazel, L.M.; Rolle-Kampczyk, U.; von Bergen, M.; McCoy, K.D.; Macpherson, A.J.; Danska, J.S. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 2013, 339, 1084–1088. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Janssen, S.; McDonald, D.; Gonzalez, A.; Navas-Molina, J.A.; Jiang, L.; Xu, Z.Z.; Winker, K.; Kado, D.M.; Orwoll, E.; Manary, M.; et al. Phylogenetic Placement of Exact Amplicon Sequences Improves Associations with Clinical Information. mSystems 2018, 3, 00021-18. [Google Scholar] [CrossRef] [PubMed]
- Krebs-Smith, S.M.; Pannucci, T.E.; Subar, A.F.; Kirkpatrick, S.I.; Lerman, J.L.; Tooze, J.A.; Wilson, M.M.; Reedy, J. Update of the Healthy Eating Index: HEI-2015. J. Acad. Nutr. Diet. 2018, 118, 1591–1602. [Google Scholar] [CrossRef] [PubMed]
- Koeth, R.A.; Wang, Z.; Levison, B.S.; Buffa, J.A.; Org, E.; Sheehy, B.T.; Britt, E.B.; Fu, X.; Wu, Y.; Li, L.; et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 2013, 19, 576–585. [Google Scholar] [CrossRef]
- Kirsch, S.H.; Herrmann, W.; Rabagny, Y.; Obeid, R. Quantification of acetylcholine, choline, betaine, and dimethylglycine in human plasma and urine using stable-isotope dilution ultra performance liquid chromatography-tandem mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2010, 878, 3338–3344. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Cheng, J.; Duncan, A.E.; Kau, A.L.; Griffin, N.W.; Lombard, V.; Henrissat, B.; Bain, J.R.; et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 2013, 341, 1241214. [Google Scholar] [CrossRef]
- Tang, Z.Z.; Chen, G.; Hong, Q.; Huang, S.; Smith, H.M.; Shah, R.D.; Scholz, M.; Ferguson, J.F. Multi-Omic Analysis of the Microbiome and Metabolome in Healthy Subjects Reveals Microbiome-Dependent Relationships Between Diet and Metabolites. Front. Genet. 2019, 10, 454. [Google Scholar] [CrossRef]
- Tan, J.; McKenzie, C.; Potamitis, M.; Thorburn, A.N.; Mackay, C.R.; Macia, L. The role of short-chain fatty acids in health and disease. Adv. Immunol. 2014, 121, 91–119. [Google Scholar] [CrossRef]
- Errera, F.I.; Canani, L.H.; Yeh, E.; Kague, E.; Armelin-Corrêa, L.M.; Suzuki, O.T.; Tschiedel, B.; Silva, M.E.; Sertié, A.L.; Passos-Bueno, M.R. COL18A1 is highly expressed during human adipocyte differentiation and the SNP c.1136C > T in its “frizzled” motif is associated with obesity in diabetes type 2 patients. Acad. Bras. Cienc. 2008, 80, 167–177. [Google Scholar] [CrossRef]
- Aikio, M.; Elamaa, H.; Vicente, D.; Izzi, V.; Kaur, I.; Seppinen, L.; Speedy, H.E.; Kaminska, D.; Kuusisto, S.; Sormunen, R.; et al. Specific collagen XVIII isoforms promote adipose tissue accrual via mechanisms determining adipocyte number and affect fat deposition. Proc. Natl. Acad. Sci. USA 2014, 111, E3043–E3052. [Google Scholar] [CrossRef]
- Sandholm, N.; Cole, J.B.; Nair, V.; Sheng, X.; Liu, H.; Ahlqvist, E.; van Zuydam, N.; Dahlström, E.H.; Fermin, D.; Smyth, L.J.; et al. Genome-wide meta-analysis and omics integration identifies novel genes associated with diabetic kidney disease. Diabetologia 2022, 65, 1495–1509. [Google Scholar] [CrossRef]
- Kieffer, D.A.; Piccolo, B.D.; Marco, M.L.; Kim, E.B.; Goodson, M.L.; Keenan, M.J.; Dunn, T.N.; Knudsen, K.E.B.; Adams, S.H.; Martin, R.J. Obese Mice Fed a Diet Supplemented with Enzyme-Treated Wheat Bran Display Marked Shifts in the Liver Metabolome Concurrent with Altered Gut Bacteria. J. Nutr. 2016, 146, 2445–2460. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.D.; Walkey, C.J.; Puigserver, P.; Spiegelman, B.M. Transcriptional regulation of adipogenesis. Genes Dev. 2000, 14, 1293–1307. [Google Scholar] [CrossRef] [PubMed]
- Merrett, J.E.; Bo, T.; Psaltis, P.J.; Proud, C.G. Identification of DNA response elements regulating expression of CCAAT/enhancer-binding protein (C/EBP) β and δ and MAP kinase-interacting kinases during early adipogenesis. Adipocyte 2020, 9, 427–442. [Google Scholar] [CrossRef] [PubMed]
- Rossmeislová, L.; Malisová, L.; Kracmerová, J.; Tencerová, M.; Kovácová, Z.; Koc, M.; Siklová-Vítková, M.; Viquerie, N.; Langin, D.; Stich, V. Weight loss improves the adipogenic capacity of human preadipocytes and modulates their secretory profile. Diabetes 2013, 62, 1990–1995. [Google Scholar] [CrossRef] [PubMed]
- Isakson, P.; Hammarstedt, A.; Gustafson, B.; Smith, U. Impaired preadipocyte differentiation in human abdominal obesity: Role of Wnt, tumor necrosis factor-alpha, and inflammation. Diabetes 2009, 58, 1550–1557. [Google Scholar] [CrossRef]
- Palmas, V.; Pisanu, S.; Madau, V.; Casula, E.; Deledda, A.; Cusano, R.; Uva, P.; Vascellari, S.; Loviselli, A.; Manzin, A.; et al. Gut microbiota markers associated with obesity and overweight in Italian adults. Sci. Rep. 2021, 11, 5532. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Jayachandran, M.; Chung, S.S.M.; Xu, B. A critical review of the relationship between dietary components, the gut microbe Akkermansia muciniphila, and human health. Crit. Rev. Food Sci. Nutr. 2020, 60, 2265–2276. [Google Scholar] [CrossRef]
- Yoon, H.S.; Cho, C.H.; Yun, M.S.; Jang, S.J.; You, H.J.; Kim, J.H.; Han, D.; Cha, K.H.; Moon, S.H.; Lee, K.; et al. Akkermansia muciniphila secretes a glucagon-like peptide-1-inducing protein that improves glucose homeostasis and ameliorates metabolic disease in mice. Nat. Microbiol. 2021, 6, 563–573. [Google Scholar] [CrossRef]
- Collado, M.C.; Derrien, M.; Isolauri, E.; de Vos, W.M.; Salminen, S. Intestinal integrity and Akkermansia muciniphila, a mucin-degrading member of the intestinal microbiota present in infants, adults, and the elderly. Appl. Environ. Microbiol. 2007, 73, 7767–7770. [Google Scholar] [CrossRef]
- Zhao, Q.; Yu, J.; Hao, Y.; Zhou, H.; Hu, Y.; Zhang, C.; Zheng, H.; Wang, X.; Zeng, F.; Hu, J.; et al. Akkermansia muciniphila plays critical roles in host health. Crit. Rev. Microbiol. 2023, 49, 82–100. [Google Scholar] [CrossRef]
- Cani, P.D.; de Vos, W.M. Next-Generation Beneficial Microbes: The Case of Akkermansia muciniphila. Front. Microbiol. 2017, 8, 1765. [Google Scholar] [CrossRef] [PubMed]
- Everard, A.; Belzer, C.; Geurts, L.; Ouwerkerk, J.P.; Druart, C.; Bindels, L.B.; Guiot, Y.; Derrien, M.; Muccioli, G.G.; Delzenne, N.M.; et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. USA 2013, 110, 9066–9071. [Google Scholar] [CrossRef] [PubMed]
- Dao, M.C.; Everard, A.; Aron-Wisnewsky, J.; Sokolovska, N.; Prifti, E.; Verger, E.O.; Kayser, B.D.; Levenez, F.; Chilloux, J.; Hoyles, L.; et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: Relationship with gut microbiome richness and ecology. Gut 2016, 65, 426–436. [Google Scholar] [CrossRef] [PubMed]
- Davenport, E.R.; Cusanovich, D.A.; Michelini, K.; Barreiro, L.B.; Ober, C.; Gilad, Y. Genome-Wide Association Studies of the Human Gut Microbiota. PLoS ONE 2015, 10, e0140301. [Google Scholar] [CrossRef]
- Vacca, M.; Celano, G.; Calabrese, F.M.; Portincasa, P.; Gobbetti, M.; De Angelis, M. The Controversial Role of Human Gut Lachnospiraceae. Microorganisms 2020, 8, 573. [Google Scholar] [CrossRef]
- Vital, M.; Karch, A.; Pieper, D.H. Colonic Butyrate-Producing Communities in Humans: An Overview Using Omics Data. mSystems 2017, 2, 00130-17. [Google Scholar] [CrossRef]
- Muralidharan, J.; Moreno-Indias, I.; Bulló, M.; Lopez, J.V.; Corella, D.; Castañer, O.; Vidal, J.; Atzeni, A.; Fernandez-García, J.C.; Torres-Collado, L.; et al. Effect on gut microbiota of a 1-y lifestyle intervention with Mediterranean diet compared with energy-reduced Mediterranean diet and physical activity promotion: PREDIMED-Plus Study. Am. J. Clin. Nutr. 2021, 114, 1148–1158. [Google Scholar] [CrossRef]
- Machado Arruda, S.P.; da Silva, A.A.M.; Kac, G.; Vilela, A.A.F.; Goldani, M.; Bettiol, H.; Barbieri, M.A. Dietary patterns are associated with excess weight and abdominal obesity in a cohort of young Brazilian adults. Eur. J. Nutr. 2016, 55, 2081–2091. [Google Scholar] [CrossRef] [PubMed]
- Couchet, M.; Breuillard, C.; Corne, C.; Rendu, J.; Morio, B.; Schlattner, U.; Moinard, C. Ornithine Transcarbamylase From Structure to Metabolism: An Update. Front. Physiol. 2021, 12, 748249. [Google Scholar] [CrossRef]
- Song, X.; Zhong, L.; Lyu, N.; Liu, F.; Li, B.; Hao, Y.; Xue, Y.; Li, J.; Feng, Y.; Ma, Y.; et al. Inulin Can Alleviate Metabolism Disorders in ob/ob Mice by Partially Restoring Leptin-related Pathways Mediated by Gut Microbiota. Genom. Proteom. Bioinform. 2019, 17, 64–75. [Google Scholar] [CrossRef] [PubMed]
- Atzeni, A.; Galié, S.; Muralidharan, J.; Babio, N.; Tinahones, F.J.; Vioque, J.; Corella, D.; Castañer, O.; Vidal, J.; Moreno-Indias, I.; et al. Gut Microbiota Profile and Changes in Body Weight in Elderly Subjects with Overweight/Obesity and Metabolic Syndrome. Microorganisms 2021, 9, 346. [Google Scholar] [CrossRef] [PubMed]
- Baldeon, A.D.; McDonald, D.; Gonzalez, A.; Knight, R.; Holscher, H.D. Diet Quality and the Fecal Microbiota in Adults in the American Gut Project. J. Nutr. 2023, 153, 2004–2015. [Google Scholar] [CrossRef]
- Cotillard, A.; Cartier-Meheust, A.; Litwin, N.S.; Chaumont, S.; Saccareau, M.; Lejzerowicz, F.; Tap, J.; Koutnikova, H.; Lopez, D.G.; McDonald, D.; et al. A posteriori dietary patterns better explain variations of the gut microbiome than individual markers in the American Gut Project. Am. J. Clin. Nutr. 2022, 115, 432–443. [Google Scholar] [CrossRef] [PubMed]
- Johnson, A.J.; Vangay, P.; Al-Ghalith, G.A.; Hillmann, B.M.; Ward, T.L.; Shields-Cutler, R.R.; Kim, A.D.; Shmagel, A.K.; Syed, A.N.; Walter, J.; et al. Daily Sampling Reveals Personalized Diet-Microbiome Associations in Humans. Cell Host. Microbe. 2019, 25, 789–802.e785. [Google Scholar] [CrossRef]
- Healey, G.R.; Murphy, R.; Brough, L.; Butts, C.A.; Coad, J. Interindividual variability in gut microbiota and host response to dietary interventions. Nutr. Rev. 2017, 75, 1059–1080. [Google Scholar] [CrossRef]
- Scalbert, A.; Brennan, L.; Manach, C.; Andres-Lacueva, C.; Dragsted, L.O.; Draper, J.; Rappaport, S.M.; Van Der Hooft, J.J.; Wishart, D.S. The food metabolome: A window over dietary exposure. Am. J. Clin. Nutr. 2014, 99, 1286–1308. [Google Scholar] [CrossRef]
- Hullar, M.A.J.; Jenkins, I.C.; Randolph, T.W.; Curtis, K.R.; Monroe, K.R.; Ernst, T.; Shepherd, J.A.; Stram, D.O.; Cheng, I.; Kristal, B.S.; et al. Associations of the gut microbiome with hepatic adiposity in the Multiethnic Cohort Adiposity Phenotype Study. Gut Microbes 2021, 13, 1965463. [Google Scholar] [CrossRef]
- Henke, M.T.; Kenny, D.J.; Cassilly, C.D.; Vlamakis, H.; Xavier, R.J.; Clardy, J. Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc. Natl. Acad. Sci. USA 2019, 116, 12672–12677. [Google Scholar] [CrossRef]
- Ma, E.; Maskarinec, G.; Lim, U.; Boushey, C.J.; Wilkens, L.R.; Setiawan, V.W.; Le Marchand, L.; Randolph, T.W.; Jenkins, I.C.; Curtis, K.R.; et al. Long-term association between diet quality and characteristics of the gut microbiome in the multiethnic cohort study. Br. J. Nutr. 2022, 128, 93–102. [Google Scholar] [CrossRef]
- Lozano, C.P.; Wilkens, L.R.; Shvetsov, Y.B.; Maskarinec, G.; Park, S.-Y.; Shepherd, J.A.; Boushey, C.J.; Hebert, J.R.; Wirth, M.D.; Ernst, T.; et al. Associations of the Dietary Inflammatory Index with total adiposity and ectopic fat through the gut microbiota, LPS, and C-reactive protein in the Multiethnic Cohort–Adiposity Phenotype Study. Am. J. Clin. Nutr. 2021, 115, 1344–1356. [Google Scholar] [CrossRef] [PubMed]
- Ravelli, M.N.; Schoeller, D.A. Traditional Self-Reported Dietary Instruments Are Prone to Inaccuracies and New Approaches Are Needed. Front. Nutr. 2020, 7, 90. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Baseline Mean ± SD or % (n) | 3 Months Mean ± SD or % (n) | Change (p Value) | |
|---|---|---|---|---|
| Age, years | 40.9 ± 9.7 | - | - | |
| Sex | Male | 23 (11) | - | - |
| Female | 77 (36) | - | - | |
| Race | White | 89 (42) | - | - |
| Black | 6 (3) | - | - | |
| Multiracial | 4 (2) | - | - | |
| Ethnicity | Hispanic | 9 (19) | - | - |
| Non-Hispanic | 81 (38) | - | - | |
| Income | <$25,000 USD | 11 (5) | - | - |
| $25,000–$45,000 USD | 4 (2) | - | - | |
| $45,001–$70,000 USD | 23 (11) | - | - | |
| $70,001–$110,000 USD | 25 (12) | - | - | |
| >$110,000 USD | 36 (17) | - | - | |
| Education | Some college | 11 (5) | - | - |
| Four-year degree | 45 (21) | - | - | |
| Master’s degree | 34 (16) | - | - | |
| Doctorate degree | 11 (5) | - | - | |
| Weight, kg | 96.1 ± 16.1 | 90.2 ± 15.3 | −6.0 ± 3.9 | |
| (<0.001) | ||||
| Body mass index, kg/m2 | 33.5 ± 4.5 | 31.5 ± 4.3 | −2.1 ± 1.4 | |
| (<0.001) | ||||
| Waist circumference, cm | 109.4 ± 10.3 | 100.9 ± 10.5 | −8.5 ± 6.0 | |
| (<0.001) | ||||
| Systolic blood pressure, mmHg | 117 ± 14 | 114 ± 12 | −3 ± 12 | |
| (0.104) | ||||
| Diastolic blood pressure, mmHg | 74 ± 8 | 76 ± 9 | 2 ± 12 | |
| (0.173) | ||||
| Total cholesterol, mg/dL a | 180 ± 34 | 165 ± 30 | −15 ± 26 | |
| (<0.001) | ||||
| High-density lipoprotein (HDL) cholesterol, mg/dL a | 48 ± 12 | 47 ± 12 | −1 ± 6 | |
| (0.288) | ||||
| Triglycerides, mg/dL a | 136 ± 79 | 107 ± 56 | −29 ± 61 | |
| (0.002) | ||||
| Glucose, mg/dL a | 93 ± 11 | 88 ± 8 | −5 ± 11 | |
| (0.002) | ||||
| Insulin, uIU/mL a | 12 ± 8 | 7 ± 5 | −4 ± 6 | |
| (<0.001) |
| Characteristic | Baseline Mean ± SD | 3 Months Mean ± SD | Change (p Value) |
|---|---|---|---|
| Energy (kcal/day) | 1764 ± 338 | 1284 ± 380 | −479 ± 445 (<0.001) |
| Carbohydrate (% kcal) | 42 ± 8 | 42 ± 7 | 0 ± 6 (0.797) |
| Fat (% kcal) | 39 ± 7 | 35 ± 5 | −4 ± 6 (<0.001) |
| Protein (% kcal) | 17 ± 3 | 21 ± 4 | 4 ± 5 (<0.001) |
| Fiber (g/day) | 16 ± 5 | 14 ± 6 | −2 ± 6 (0.015) |
| Diet quality (total HEI score) | 57 ± 12 | 62 ± 12 | 4 ± 12 (0.022) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hill, E.B.; Konigsberg, I.R.; Ir, D.; Frank, D.N.; Jambal, P.; Litkowski, E.M.; Lange, E.M.; Lange, L.A.; Ostendorf, D.M.; Scorsone, J.J.; et al. The Microbiome, Epigenome, and Diet in Adults with Obesity during Behavioral Weight Loss. Nutrients 2023, 15, 3588. https://doi.org/10.3390/nu15163588
Hill EB, Konigsberg IR, Ir D, Frank DN, Jambal P, Litkowski EM, Lange EM, Lange LA, Ostendorf DM, Scorsone JJ, et al. The Microbiome, Epigenome, and Diet in Adults with Obesity during Behavioral Weight Loss. Nutrients. 2023; 15(16):3588. https://doi.org/10.3390/nu15163588
Chicago/Turabian StyleHill, Emily B., Iain R. Konigsberg, Diana Ir, Daniel N. Frank, Purevsuren Jambal, Elizabeth M. Litkowski, Ethan M. Lange, Leslie A. Lange, Danielle M. Ostendorf, Jared J. Scorsone, and et al. 2023. "The Microbiome, Epigenome, and Diet in Adults with Obesity during Behavioral Weight Loss" Nutrients 15, no. 16: 3588. https://doi.org/10.3390/nu15163588
APA StyleHill, E. B., Konigsberg, I. R., Ir, D., Frank, D. N., Jambal, P., Litkowski, E. M., Lange, E. M., Lange, L. A., Ostendorf, D. M., Scorsone, J. J., Wayland, L., Bing, K., MacLean, P. S., Melanson, E. L., Bessesen, D. H., Catenacci, V. A., Stanislawski, M. A., & Borengasser, S. J. (2023). The Microbiome, Epigenome, and Diet in Adults with Obesity during Behavioral Weight Loss. Nutrients, 15(16), 3588. https://doi.org/10.3390/nu15163588






