The Pre-Analytical CEN/TS Standard for Microbiome Diagnostics—How Can Research and Development Benefit?
Abstract
:1. Introduction
2. Need for Microbiome Standards in Clinical Practice and Diagnostics
3. What Is a European/International Pre-Analytical Standard
3.1. Considerations Regarding a Diagnostic Pre-Analytic Standard for Isolated Human Microbiome DNA
3.2. Structure of the Diagnostic Pre-Analytic Standard for Isolated Human Microbiome DNA
3.3. Benefit of Adhering to Standards in Research and Development
3.4. Standards as Drivers of Innovation
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Baker, M. 1500 scientists lift the lid on reproducibility. Nature 2016, 533, 452–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De la Salle, B. Pre- and postanalytical errors in haematology. Int. J. Lab. Hematol. 2019, 41 (Suppl. 1), 170–176. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plebani, M. The detection and prevention of errors in laboratory medicine. Ann. Clin. Biochem. 2010, 47, 101–110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holub, P.; Kohlmayer, F.; Prasser, F.; Mayrhofer, M.T.; Schlunder, I.; Martin, G.M.; Casati, S.; Koumakis, L.; Wutte, A.; Kozera, L.; et al. Enhancing Reuse of Data and Biological Material in Medical Research: From FAIR to FAIR-Health. Biopreserv. Biobank. 2018, 16, 97–105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- European Commission. In-vitro Diagnostics Regulation. Available online: https://eur-lex.europa.eu/eli/reg/2017/746/oj (accessed on 25 March 2022).
- Prados-Bo, A.; Casino, G. Microbiome research in general and business newspapers: How many microbiome articles are published and which study designs make the news the most? PLoS ONE 2021, 16, e0249835. [Google Scholar] [CrossRef] [PubMed]
- Schlaberg, R. Microbiome Diagnostics. Clin. Chem. 2020, 66, 68–76. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turnbaugh, P.J.; Ley, R.E.; Hamady, M.; Fraser-Liggett, C.M.; Knight, R.; Gordon, J.I. The human microbiome project. Nature 2007, 449, 804–810. [Google Scholar] [CrossRef]
- Integrative HMP (iHMP) Research Network Consortium. The Integrative Human Microbiome Project: Dynamic analysis of microbiome-host omics profiles during periods of human health and disease. Cell Host. Microbe 2014, 16, 276–289. [Google Scholar] [CrossRef] [Green Version]
- Proctor, L. Priorities for the next 10 years of human microbiome research. Nature 2019, 569, 623–625. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stulberg, E.; Fravel, D.; Proctor, L.M.; Murray, D.M.; LoTempio, J.; Chrisey, L.; Garland, J.; Goodwin, K.; Graber, J.; Harris, M.C.; et al. An assessment of US microbiome research. Nat. Microbiol. 2016, 1, 15015. [Google Scholar] [CrossRef] [PubMed]
- Human Microbiome Market by Product (Prebiotics, Probiotics, Food, Diagnostic Tests, Drugs), Application (Therapeutic, Diagnostic), Disease (Infectious, Metabolic/Endocrine), Research Technology (Genomics, Proteomics, Metabolomics)—Global Forecast to 2028. Available online: https://www.marketsandmarkets.com/Market-Reports/human-microbiome-market-37621904.html (accessed on 25 March 2022).
- Schloss, P.D. Identifying and Overcoming Threats to Reproducibility, Replicability, Robustness, and Generalizability in Microbiome Research. mBio 2018, 9, e00525-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hiergeist, A.; Reischl, U.; Priority Program 1656 Intestinal Microbiota Consortium/Quality Assessment Participants; Gessner, A. Multicenter quality assessment of 16S ribosomal DNA-sequencing for microbiome analyses reveals high inter-center variability. Int. J. Med. Microbiol. 2016, 306, 334–342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eck, A.; de Groot, E.F.J.; de Meij, T.G.J.; Welling, M.; Savelkoul, P.H.M.; Budding, A.E. Robust Microbiota-Based Diagnostics for Inflammatory Bowel Disease. J. Clin. Microbiol. 2017, 55, 1720–1732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sinha, R.; Abu-Ali, G.; Vogtmann, E.; Fodor, A.A.; Ren, B.; Amir, A.; Schwager, E.; Crabtree, J.; Ma, S.; The Microbiome Quality Control Project Consortium; et al. Assessment of variation in microbial community amplicon sequencing by the Microbiome Quality Control (MBQC) project consortium. Nat. Biotechnol. 2017, 35, 1077–1086. [Google Scholar] [CrossRef] [Green Version]
- Knight, R.; Vrbanac, A.; Taylor, B.C.; Aksenov, A.; Callewaert, C.; Debelius, J.; Gonzalez, A.; Kosciolek, T.; McCall, L.I.; McDonald, D.; et al. Best practices for analysing microbiomes. Nat. Rev. Microbiol. 2018, 16, 410–422. [Google Scholar] [CrossRef] [Green Version]
- Bharti, R.; Grimm, D.G. Current challenges and best-practice protocols for microbiome analysis. Brief. Bioinform. 2021, 22, 178–193. [Google Scholar] [CrossRef] [Green Version]
- Mirzayi, C.; Renson, A.; Genomic Standards Consortium; Massive, A.; The Microbiome Quality Control Project Consortium; Zohra, F.; Elsafoury, S.; Geistlinger, L.; Kasselman, L.J.; Eckenrode, K.; et al. Reporting guidelines for human microbiome research: The STORMS checklist. Nat. Med. 2021, 27, 1885–1892. [Google Scholar] [CrossRef]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [Green Version]
- Shanahan, F.; Ghosh, T.S.; O’Toole, P.W. The Healthy Microbiome-What Is the Definition of a Healthy Gut Microbiome? Gastroenterology 2021, 160, 483–494. [Google Scholar] [CrossRef]
- Berg, G.; Rybakova, D.; Fischer, D.; Cernava, T.; Verges, M.C.; Charles, T.; Chen, X.; Cocolin, L.; Eversole, K.; Corral, G.H.; et al. Microbiome definition re-visited: Old concepts and new challenges. Microbiome 2020, 8, 103. [Google Scholar] [CrossRef] [PubMed]
- Selway, C.A.; Eisenhofer, R.; Weyrich, L.S. Microbiome applications for pathology: Challenges of low microbial biomass samples during diagnostic testing. J. Pathol. Clin. Res. 2020, 6, 97–106. [Google Scholar] [CrossRef] [PubMed]
- ISO 4307:2021; Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for Human Specimen—Isolated Microbiome DNA. ISO: Genva, Switzerland, 2021.
- ISO/IEC GUIDE 2:2004; Standardization and Related Activities—General Vocabulary. ISO: Genva, Switzerland, 2004.
- Ryan, M.J.; Schloter, M.; Berg, G.; Kostic, T.; Kinkel, L.L.; Eversole, K.; Macklin, J.A.; Schelkle, B.; Kazou, M.; Sarand, I.; et al. Development of Microbiome Biobanks—Challenges and Opportunities. Trends Microbiol. 2021, 29, 89–92. [Google Scholar] [CrossRef] [PubMed]
- Kong, H.H.; Andersson, B.; Clavel, T.; Common, J.E.; Jackson, S.A.; Olson, N.D.; Segre, J.A.; Traidl-Hoffmann, C. Performing Skin Microbiome Research: A Method to the Madness. J. Investig. Dermatol. 2017, 137, 561–568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bao, L.; Chandra, P.K.; Moroz, K.; Zhang, X.; Thung, S.N.; Wu, T.; Dash, S. Impaired autophagy response in human hepatocellular carcinoma. Exp. Mol. Pathol. 2014, 96, 149–154. [Google Scholar] [CrossRef] [Green Version]
- Wu, W.K.; Chen, C.C.; Panyod, S.; Chen, R.A.; Wu, M.S.; Sheen, L.Y.; Chang, S.C. Optimization of fecal sample processing for microbiome study—The journey from bathroom to bench. J. Formos. Med. Assoc. 2019, 118, 545–555. [Google Scholar] [CrossRef]
- Pereira-Marques, J.; Hout, A.; Ferreira, R.M.; Weber, M.; Pinto-Ribeiro, I.; van Doorn, L.J.; Knetsch, C.W.; Figueiredo, C. Impact of Host DNA and Sequencing Depth on the Taxonomic Resolution of Whole Metagenome Sequencing for Microbiome Analysis. Front. Microbiol. 2019, 10, 1277. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Hofstaedter, C.E.; Zhao, C.; Mattei, L.; Tanes, C.; Clarke, E.; Lauder, A.; Sherrill-Mix, S.; Chehoud, C.; Kelsen, J.; et al. Optimizing methods and dodging pitfalls in microbiome research. Microbiome 2017, 5, 52. [Google Scholar] [CrossRef]
- Salter, S.J.; Cox, M.J.; Turek, E.M.; Calus, S.T.; Cookson, W.O.; Moffatt, M.F.; Turner, P.; Parkhill, J.; Loman, N.J.; Walker, A.W. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014, 12, 87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stinson, L.F.; Keelan, J.A.; Payne, M.S. Identification and removal of contaminating microbial DNA from PCR reagents: Impact on low-biomass microbiome analyses. Lett. Appl. Microbiol. 2019, 68, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors—occurrence, properties and removal. J. Appl. Microbiol. 2012, 113, 1014–1026. [Google Scholar] [CrossRef]
- Costea, P.I.; Zeller, G.; Sunagawa, S.; Pelletier, E.; Alberti, A.; Levenez, F.; Tramontano, M.; Driessen, M.; Hercog, R.; Jung, F.E.; et al. Towards standards for human fecal sample processing in metagenomic studies. Nat. Biotechnol. 2017, 35, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Santiago, A.; Panda, S.; Mengels, G.; Martinez, X.; Azpiroz, F.; Dore, J.; Guarner, F.; Manichanh, C. Processing faecal samples: A step forward for standards in microbial community analysis. BMC Microbiol. 2014, 14, 112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stammler, F.; Glasner, J.; Hiergeist, A.; Holler, E.; Weber, D.; Oefner, P.J.; Gessner, A.; Spang, R. Adjusting microbiome profiles for differences in microbial load by spike-in bacteria. Microbiome 2016, 4, 28. [Google Scholar] [CrossRef] [Green Version]
- ISO 20166-1:2018; Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for Formalin-Fixed and Paraffin-Embedded (FFPE) Tissue—Part 1: Isolated RNA. ISO: Genva, Switzerland, 2018.
- ISO 20166-3:2018; Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for Formalin-Fixed and Paraffin-Embedded (FFPE) Tissue—Part 3: Isolated DNA. ISO: Genva, Switzerland, 2018.
- ISO 20166-2:2018; Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for Formalin-Fixed and Paraffin-Embedded (FFPE) Tissue—Part 2: Isolated protein. ISO: Genva, Switzerland, 2018.
- ISO 20166-4:2021; Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for Formalin-Fixed and Paraffin-Embedded (FFPE) Tissue—Part 4: In-Situ Detection Techniques. ISO: Genva, Switzerland, 2018.
- Freedman, L.P.; Cockburn, I.M.; Simcoe, T.S. The Economics of Reproducibility in Preclinical Research. PLoS Biol. 2015, 13, e1002165. [Google Scholar] [CrossRef] [PubMed]
- McGlynn, E.A.; McDonald, K.M.; Cassel, C.K. Measurement Is Essential for Improving Diagnosis and Reducing Diagnostic Error: A Report From the Institute of Medicine. JAMA 2015, 314, 2501–2502. [Google Scholar] [CrossRef] [PubMed]
| Pre-Analytical Standards (CEN/TS, ISO) | Standard Operating Procedures (SOP) |
|---|---|
| CEN/TS or ISO standards for sample pre-analytics are official documents that specify requirements (‘mandatory’; no deviation permitted; expressed as ‘shall’) and give recommendations (‘non-mandatory’; possible suitable choices, expressed as ‘should’) for the pre-analytical workflow of certain specimen types. | SOPs are written documents containing step-by-step instructions for laboratory procedures specific to a certain laboratory that the laboratory staff needs to follow. |
| They are evidence-based, written and agreed on by experts on a European (CEN) or international (ISO) level. They provide the state-of-the-art for national and international regulation (e.g., IVDR) and are considered by regulators to reduce the risk for IVD developers and users. They are applicable to a wider range of user-groups and are vendor neutral (i.e., they do not refer to specific products). | They are generated, reviewed, and approved by a particular laboratory. The information an SOP contains is more detailed and specific for the laboratory and its equipment, chemicals, reagents and procedure (e.g., defines the centrifugation speeds, temperature and duration). |
| CEN/TS and ISO pre-analytics standards provide the basis for SOPs. | SOPs are part of a laboratory’s quality management system and have to comply with requirements of standards. |
| ISO Standards and CEN Technical Specifications (CEN/TS) Molecular In Vitro Diagnostic Examinations—Specifications for Pre-Examination Processes for: |
|---|
| Published |
| EN ISO 20166-1: 2018, FFPE tissue—Part 1: Isolated RNA EN ISO 20166-2: 2018, FFPE tissue—Part 2: Isolated proteins EN ISO 20166-3: 2018, FFPE tissue—Part 3: Isolated DNA EN ISO 20166-4: 2021 FFPE tissues—Part 3: In-situ detection techniques EN ISO 20184-1: 2018, Frozen tissue—Part 1: Isolated RNA EN ISO 20184-2: 2018, Frozen tissue—Part 2: Isolated proteins EN ISO 20184-3: 2021, frozen tissue—Part 3: Isolated DNA EN ISO 20186-1: 2019, Venous whole blood—Part 1: Isolated cellular RNA EN ISO 20186-2: 2019, Venous whole blood—Part 2: Isolated genomic DNA EN ISO 20186-3: 2019, Venous whole blood—Part 3: Isolated circulating cell free DNA from plasma CEN/TS 17626: 2021, Human specimens—microbiome DNA CEN/TS 17390-1: 2020, Circulating tumor cells (CTCs)—Part 1: Isolated RNA CEN/TS 17390-2: 2020, Circulating tumor cells (CTCs)—Part 2: Isolated DNA CEN/TS 17390-3: 2020, Circulating tumor cells (CTCs)—Part 3: Preparation for analytical CTC staining EN ISO 23118: 2021, Urine, plasma, serum for metabolomics EN ISO 4307:2021, Saliva—Isolated human DNA CEN/TS 17688-1: 2021, Fine needle aspirates—Part 1: Isolated cellular RNA CEN/TS 17688-2: 2021, Fine needle aspirates—Part 2: Isolated proteins CEN/TS 17688-3: 2021, Fine needle aspirates—Part 3: Isolated genomic DNA |
| Upcoming |
| CEN/TS for exosomes and extracellular vesicles in venous whole blood—Isolated DNA /RNA/proteins CEN/TS for venous whole blood—Isolated circulating cell free RNA from plasma CEN/TS for urine and other body fluids—Isolated cell free DNA |
| Source: CEN/TC 140—In vitro diagnostic medical devices |
| Structure (Major Chapters & Topics) | Topics or Examples of Pre-Analytical Factors Addressed | |
|---|---|---|
| Introduction | Main information on microbiome (DNA) and the pre-analytical phase | |
| 1 Scope | Purpose/content: Requirements & recommendations for pre-examination phase of human specimens, such as stool, saliva, skin and urogenital specimens, intended for microbiome DNA examination Target group: Applicable to medical laboratories, in vitro diagnostics developers and manufacturers biobanks, biomedical research performing institutions/organizations, and regulatory authorities etc. | |
| 2 Normative references | Referral to other relevant standards:1 EN ISO 15189, ISO 15190, ISO/TS 20658 | |
| 3 Terms and definitions | Definition of relevant terms | |
| 4 General considerations | Overarching information on relevance | |
| 5 Outside the laboratory | Patient/specimen donor | Demographics (e.g. age, gender, geography), disease/health condition, medication and treatment (incl. e.g. antibiotics), nutrition (incl. prebiotics, habits), frequency, life style, (smoking, personal care habits, stress, physical activity) |
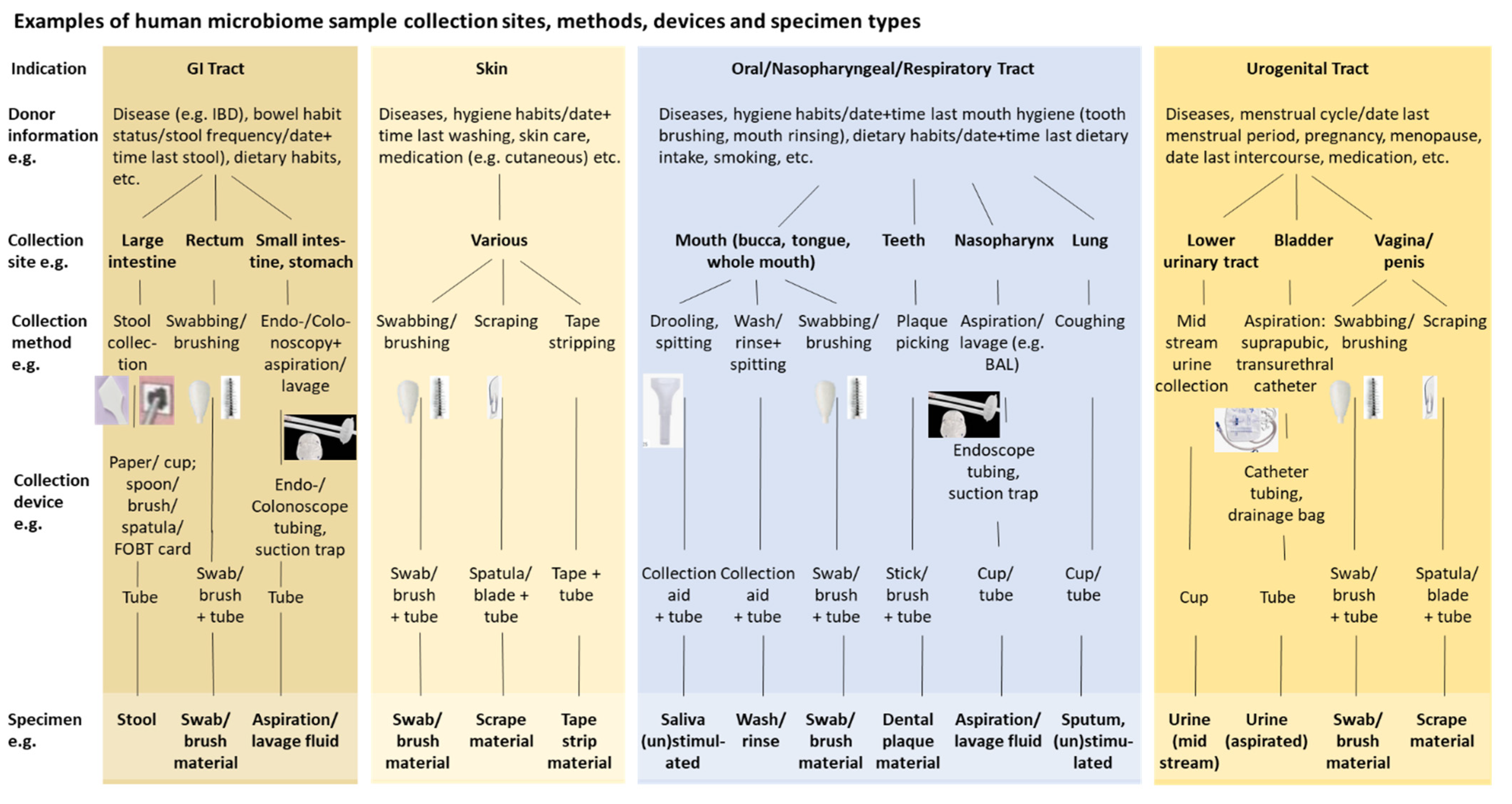
| Selection of specimen collection method & device(s) Specimen collection & stabilization | Collection method & device(s) (e.g. swab, tape, tube, collection site, spatula), process of collecting (incl. self-collection by donors), contamination, (pre-)treatment of collection site, labelling, stabilization (chemical, physical) | |
| Specimen storage & transport | Intermediate storage, with/without stabilizer, transport container, temperature, duration, oxygen, UV-light | |
| 6 Inside the laboratory | Specimen reception | Identification |
| Sample preparation | Homogenization, enrichment (e.g. centrifugation), aliquoting, labelling | |
| Sample Storage | Duration, temperature, humidity, oxygen, UV-light | |
| Microbiome DNA isolation Quantity and quality assessment of microbiome DNA | Isolation method (e.g. cell lysis), reagents/kit (e.g. type, lot.no., contaminants), process of the isolation, host DNA content, quantity/quality assessment (e.g. method, process) | |
| Storage of microbiome DNA | Duration, temperature | |
| Annex | Informative evidence-based information related to pre-analytical variables and the recommendations & requirements in the standard | |
| Bibliography | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stumptner, C.; Stadlbauer, V.; O’Neil, D.; Gessner, A.; Hiergeist, A.; Zatloukal, K.; Abuja, P.M. The Pre-Analytical CEN/TS Standard for Microbiome Diagnostics—How Can Research and Development Benefit? Nutrients 2022, 14, 1976. https://doi.org/10.3390/nu14091976
Stumptner C, Stadlbauer V, O’Neil D, Gessner A, Hiergeist A, Zatloukal K, Abuja PM. The Pre-Analytical CEN/TS Standard for Microbiome Diagnostics—How Can Research and Development Benefit? Nutrients. 2022; 14(9):1976. https://doi.org/10.3390/nu14091976
Chicago/Turabian StyleStumptner, Conny, Vanessa Stadlbauer, Dominic O’Neil, André Gessner, Andreas Hiergeist, Kurt Zatloukal, and Peter M. Abuja. 2022. "The Pre-Analytical CEN/TS Standard for Microbiome Diagnostics—How Can Research and Development Benefit?" Nutrients 14, no. 9: 1976. https://doi.org/10.3390/nu14091976
APA StyleStumptner, C., Stadlbauer, V., O’Neil, D., Gessner, A., Hiergeist, A., Zatloukal, K., & Abuja, P. M. (2022). The Pre-Analytical CEN/TS Standard for Microbiome Diagnostics—How Can Research and Development Benefit? Nutrients, 14(9), 1976. https://doi.org/10.3390/nu14091976








