The Biology of Veganism: Plasma Metabolomics Analysis Reveals Distinct Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Metabolomics Profiling
2.3. Statistical Analysis
2.3.1. Data Transformation and Linear Regression
2.3.2. Adjustment for False Discovery
2.3.3. Principal Components Analysis
2.3.4. Random Forest Analysis
2.3.5. Bootstrap Regression
2.3.6. Cross-Validation
3. Results
3.1. Baseline Characteristics
3.2. Linear Regression—Abundance of Metabolites or Biochemical Subclasses in Vegans and Non-Vegetarians
3.2.1. Linear Regression to Analyze Abundance of Individual Metabolites
3.2.2. Linear Regression to Analyze Metabolite Subclasses
3.2.3. Cross-Validation and Bootstrapped Regression for Error Analysis
3.3. Random Forest and Principal Components Analyses
3.3.1. Random Forest Analysis for Classification by Diet Group
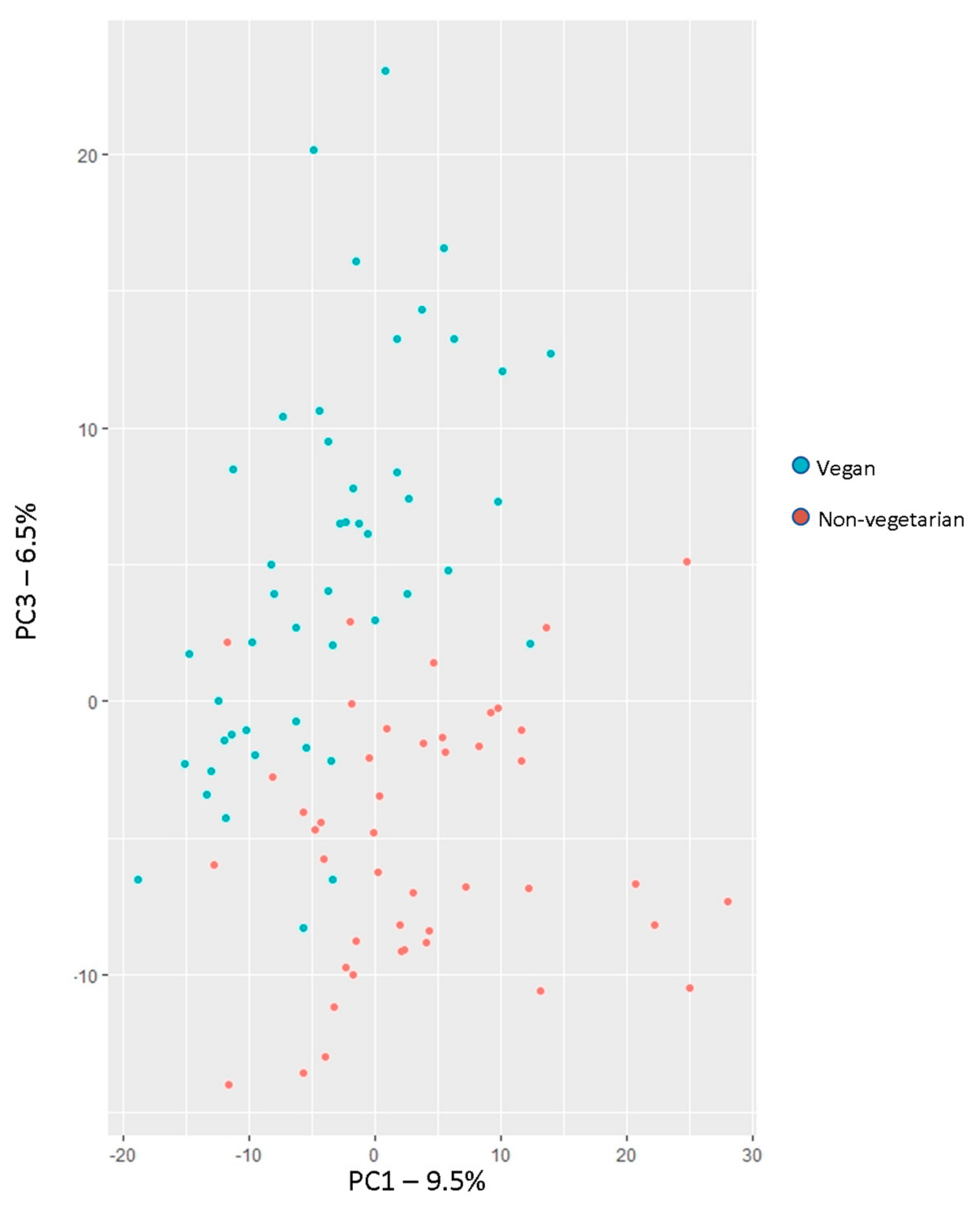
3.3.2. Principal Component Analysis for Dimension Reduction
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rizzo, N.S.; Sabate, J.; Jaceldo-Siegl, K.; Fraser, G.E. Vegetarian dietary patterns are associated with a lower risk of metabolic syndrome: The adventist health study 2. Diabetes Care 2011, 34, 1225–1227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tonstad, S.; Stewart, K.; Oda, K.; Batech, M.; Herring, R.P.; Fraser, G.E. Vegetarian diets and incidence of diabetes in the Adventist Health Study-2. Nutr. Metab. Cardiovasc. Dis. 2013, 23, 292–299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Orlich, M.J.; Singh, P.N.; Sabate, J.; Jaceldo-Siegl, K.; Fan, J.; Knutsen, S.; Beeson, W.L.; Fraser, G.E. Vegetarian dietary patterns and mortality in Adventist Health Study 2. JAMA Intern. Med. 2013, 173, 1230–1238. [Google Scholar] [CrossRef] [PubMed]
- Orlich, M.J.; Singh, P.N.; Sabate, J.; Fan, J.; Sveen, L.; Bennett, H.; Knutsen, S.F.; Beeson, W.L.; Jaceldo-Siegl, K.; Butler, T.L.; et al. Vegetarian dietary patterns and the risk of colorectal cancers. JAMA Intern. Med. 2015, 175, 767–776. [Google Scholar] [CrossRef] [PubMed]
- Tantamango-Bartley, Y.; Jaceldo-Siegl, K.; Fan, J.; Fraser, G. Vegetarian diets and the incidence of cancer in a low-risk population. Cancer Epidemiol. Biomarkers Prev. 2013, 22, 286–294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tantamango-Bartley, Y.; Knutsen, S.F.; Knutsen, R.; Jacobsen, B.K.; Fan, J.; Beeson, W.L.; Sabate, J.; Hadley, D.; Jaceldo-Siegl, K.; Penniecook, J.; et al. Are strict vegetarians protected against prostate cancer? Am. J. Clin. Nutr. 2016, 103, 153–160. [Google Scholar] [CrossRef] [Green Version]
- Bouchard-Mercier, A.; Rudkowska, I.; Lemieux, S.; Couture, P.; Vohl, M.C. The metabolic signature associated with the Western dietary pattern: A cross-sectional study. Nutr. J. 2013, 12, 158. [Google Scholar] [CrossRef] [Green Version]
- Chandler, P.D.; Balasubramanian, R.; Paynter, N.; Giulianini, F.; Fung, T.; Tinker, L.F.; Snetselaar, L.; Liu, S.M.; Eaton, C.; Tobias, D.K.; et al. Metabolic signatures associated with Western and Prudent dietary patterns in women. Am. J. Clin. Nutr. 2020, 112, 268–283. [Google Scholar] [CrossRef]
- Playdon, M.C.; Moore, S.C.; Derkach, A.; Reedy, J.; Subar, A.F.; Sampson, J.N.; Albanes, D.; Gu, F.; Kontto, J.; Lassale, C.; et al. Identifying biomarkers of dietary patterns by using metabolomics. Am. J. Clin. Nutr. 2017, 105, 450–465. [Google Scholar] [CrossRef] [Green Version]
- Tong, T.Y.N.; Koulman, A.; Griffin, J.L.; Wareham, N.J.; Forouhi, N.G.; Imamura, F. A Combination of Metabolites Predicts Adherence to the Mediterranean Diet Pattern and Its Associations with Insulin Sensitivity and Lipid Homeostasis in the General Population: The Fenland Study, United Kingdom. J. Nutr. 2020, 150, 568–578. [Google Scholar] [CrossRef] [Green Version]
- Vazquez-Fresno, R.; Llorach, R.; Urpi-Sarda, M.; Lupianez-Barbero, A.; Estruch, R.; Corella, D.; Fito, M.; Aros, F.; Ruiz-Canela, M.; Salas-Salvado, J.; et al. Metabolomic pattern analysis after mediterranean diet intervention in a nondiabetic population: A 1- and 3-year follow-up in the PREDIMED study. J. Proteome Res. 2015, 14, 531–540. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garcia-Perez, I.; Posma, J.M.; Gibson, R.; Chambers, E.S.; Hansen, T.H.; Vestergaard, H.; Hansen, T.; Beckmann, M.; Pedersen, O.; Elliott, P.; et al. Objective assessment of dietary patterns by use of metabolic phenotyping: A randomised, controlled, crossover trial. Lancet Diabetes Endocrinol. 2017, 5, 184–195. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, J.A.; Rinaldi, S.; Ferrari, P.; Carayol, M.; Achaintre, D.; Scalbert, A.; Cross, A.J.; Gunter, M.J.; Fensom, G.K.; Appleby, P.N.; et al. Metabolic profiles of male meat eaters, fish eaters, vegetarians, and vegans from the EPIC-Oxford cohort. Am. J. Clin. Nutr. 2015, 102, 1518–1526. [Google Scholar] [CrossRef] [Green Version]
- Wu, G.D.; Compher, C.; Chen, E.Z.; Smith, S.A.; Shah, R.D.; Bittinger, K.; Chehoud, C.; Albenberg, L.G.; Nessel, L.; Gilroy, E.; et al. Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut 2016, 65, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Miles, F.L.; Lloren, J.I.C.; Haddad, E.; Jaceldo-Siegl, K.; Knutsen, S.; Sabate, J.; Fraser, G.E. Plasma, Urine, and Adipose Tissue Biomarkers of Dietary Intake Differ Between Vegetarian and Non-Vegetarian Diet Groups in the Adventist Health Study-2. J. Nutr. 2019, 149, 667–675. [Google Scholar] [CrossRef]
- Miles, F.L.; Mashchak, A.; Filippov, V.; Orlich, M.J.; Duerksen-Hughes, P.; Chen, X.; Wang, C.; Siegmund, K.; Fraser, G.E. DNA Methylation Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort. Nutrients 2020, 12, 3697. [Google Scholar] [CrossRef]
- Jaceldo-Siegl, K.; Knutsen, S.F.; Sabate, J.; Beeson, W.L.; Chan, J.; Herring, R.P.; Butler, T.L.; Haddad, E.; Bennett, H.; Montgomery, S.; et al. Validation of nutrient intake using an FFQ and repeated 24 h recalls in black and white subjects of the Adventist Health Study-2 (AHS-2). Public Health Nutr. 2010, 13, 812–819. [Google Scholar] [CrossRef] [PubMed]
- Schakel, S.F. Maintaining a nutrient database in a changing marketplace: Keeping pace with changing food products—A research perspective. J. Food Compos. Anal. 2001, 14, 315–322. [Google Scholar] [CrossRef]
- Fraser, G.E.; Stram, D.O. Regression calibration when foods (measured with error) are the variables of interest: Markedly non-Gaussian data with many zeroes. Am. J. Epidemiol. 2012, 175, 325–331. [Google Scholar] [CrossRef] [Green Version]
- Fraser, G.E.; Yan, R. A multivariate method for measurement error correction using pairs of concentration biomarkers. Ann. Epidemiol. 2007, 17, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Martins, M.C.T.; Jaceldo-Siegl, K.; Orlich, M.; Fan, J.; Mashchak, A.; Fraser, G.E. A New Approach to Assess Lifetime Dietary Patterns Finds Lower Consumption of Animal Foods with Aging in a Longitudinal Analysis of a Health-Oriented Adventist Population. Nutrients 2017, 9, 1118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.W.; Morton, K.R.; Walters, J.; Bellinger, D.L.; Butler, T.L.; Wilson, C.; Walsh, E.; Ellison, C.G.; McKenzie, M.M.; Fraser, G.E. Cohort profile: The biopsychosocial religion and health study (BRHS). Int. J. Epidemiol. 2009, 38, 1470–1478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, J.; Knutsen, S.F.; Sabate, J.; Haddad, E.; Yan, R.; Fraser, G.E. Feasibility of running clinics to collect biological specimens in a nationwide cohort study--Adventist Health Study-2. Ann. Epidemiol. 2007, 17, 454–457. [Google Scholar] [CrossRef]
- Orlich, M.J.; Jaceldo-Siegl, K.; Sabate, J.; Fan, J.; Singh, P.N.; Fraser, G.E. Patterns of food consumption among vegetarians and non-vegetarians. Br. J. Nutr. 2014, 112, 1644–1653. [Google Scholar] [CrossRef] [Green Version]
- Mesnage, R.; Teixeira, M.; Mandrioli, D.; Falcioni, L.; Ducarmon, Q.R.; Zwittink, R.D.; Mazzacuva, F.; Caldwell, A.; Halket, J.; Amiel, C.; et al. Use of Shotgun Metagenomics and Metabolomics to Evaluate the Impact of Glyphosate or Roundup MON 52276 on the Gut Microbiota and Serum Metabolome of Sprague-Dawley Rats. Environ. Health Perspect. 2021, 129, 17005. [Google Scholar] [CrossRef]
- Storey, J.D.; Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 2003, 100, 9440–9445. [Google Scholar] [CrossRef] [Green Version]
- Freedman, D.; Lane, D. A Nonstochastic Interpretation of Reported Significance Levels. J. Bus. Econ. Stat. 1983, 1, 292–298. [Google Scholar] [CrossRef]
- Hastie, T.; Friedman, J. Random Forests. In The Elements of Statistical Learning, 2nd ed.; Springer Publishers: New York, NY, USA, 2009. [Google Scholar]
- Setchell, K.D.; Clerici, C. Equol: History, chemistry, and formation. J. Nutr. 2010, 140, 1355S–1362S. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Z.; Kulkarni, K.; Zhu, W.; Hu, M. Bioavailability and pharmacokinetics of genistein: Mechanistic studies on its ADME. Anticancer Agents Med. Chem. 2012, 12, 1264–1280. [Google Scholar] [CrossRef]
- Velenosi, T.J.; Hennop, A.; Feere, D.A.; Tieu, A.; Kucey, A.S.; Kyriacou, P.; McCuaig, L.E.; Nevison, S.E.; Kerr, M.A.; Urquhart, B.L. Untargeted plasma and tissue metabolomics in rats with chronic kidney disease given AST-120. Sci. Rep. 2016, 6, 22526. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.; Laaksonen, O.; Heinonen, J.; Sainio, T.; Kallio, H.; Yang, B. Sensory profile of ethyl beta-d-glucopyranoside and its contribution to quality of sea buckthorn (Hippophae rhamnoides L.). Food Chem. 2017, 233, 263–272. [Google Scholar] [CrossRef]
- Xie, X.; Yang, C.; Cui, Q.; Ma, W.; Liu, J.; Yao, Q.; Zhang, Z.; Xiao, L.; Wang, N. Stachydrine Mediates Rapid Vascular Relaxation: Activation of Endothelial Nitric Oxide Synthase Involving AMP-Activated Protein Kinase and Akt Phosphorylation in Vascular Endothelial Cells. J. Agric. Food Chem. 2019, 67, 9805–9811. [Google Scholar] [CrossRef]
- Burstein, S.; McQuain, C.; Salmonsen, R.; Seicol, B. N-Amino acid linoleoyl conjugates: Anti-inflammatory activities. Bioorg. Med. Chem. Lett. 2012, 22, 872–875. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, P.J.; Lapworth, A.L.; An, J.; Wang, L.; McGarrah, R.W.; Stevens, R.D.; Ilkayeva, O.; George, T.; Muehlbauer, M.J.; Bain, J.R.; et al. Branched-chain amino acid restriction in Zucker-fatty rats improves muscle insulin sensitivity by enhancing efficiency of fatty acid oxidation and acyl-glycine export. Mol. Metab. 2016, 5, 538–551. [Google Scholar] [CrossRef] [PubMed]
- Kaulmann, A.; Bohn, T. Carotenoids, inflammation, and oxidative stress-implications of cellular signaling pathways and relation to chronic disease prevention. Nutr. Res. 2014, 34, 907–929. [Google Scholar] [CrossRef] [PubMed]
- Saleh, H.A.; Yousef, M.H.; Abdelnaser, A. The Anti-Inflammatory Properties of Phytochemicals and Their Effects on Epigenetic Mechanisms Involved in TLR4/NF-kappaB-Mediated Inflammation. Front. Immunol. 2021, 12, 606069. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.C.; Lai, C.S.; Tsai, M.L.; Ho, C.T.; Wang, Y.J.; Pan, M.H. Chemopreventive effect of natural dietary compounds on xenobiotic-induced toxicity. J. Food Drug Anal. 2017, 25, 176–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moller, N. Ketone Body, 3-Hydroxybutyrate: Minor Metabolite—Major Medical Manifestations. J. Clin. Endocrinol. Metab. 2020, 105, 2884–2892. [Google Scholar] [CrossRef]
- Puchalska, P.; Crawford, P.A. Multi-dimensional Roles of Ketone Bodies in Fuel Metabolism, Signaling, and Therapeutics. Cell Metab. 2017, 25, 262–284. [Google Scholar] [CrossRef] [Green Version]
- Ding, Y.; Svingen, G.F.; Pedersen, E.R.; Gregory, J.F.; Ueland, P.M.; Tell, G.S.; Nygard, O.K. Plasma Glycine and Risk of Acute Myocardial Infarction in Patients With Suspected Stable Angina Pectoris. J. Am. Heart Assoc. 2015, 5, e002621. [Google Scholar] [CrossRef] [Green Version]
- Floegel, A.; Stefan, N.; Yu, Z.; Muhlenbruch, K.; Drogan, D.; Joost, H.G.; Fritsche, A.; Haring, H.U.; Hrabe de Angelis, M.; Peters, A.; et al. Identification of serum metabolites associated with risk of type 2 diabetes using a targeted metabolomic approach. Diabetes 2013, 62, 639–648. [Google Scholar] [CrossRef] [Green Version]
- Gall, W.E.; Beebe, K.; Lawton, K.A.; Adam, K.P.; Mitchell, M.W.; Nakhle, P.J.; Ryals, J.A.; Milburn, M.V.; Nannipieri, M.; Camastra, S.; et al. alpha-hydroxybutyrate is an early biomarker of insulin resistance and glucose intolerance in a nondiabetic population. PLoS ONE 2010, 5, e10883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huffman, K.M.; Shah, S.H.; Stevens, R.D.; Bain, J.R.; Muehlbauer, M.; Slentz, C.A.; Tanner, C.J.; Kuchibhatla, M.; Houmard, J.A.; Newgard, C.B.; et al. Relationships Between Circulating Metabolic Intermediates and Insulin Action in Overweight to Obese, Inactive Men and Women. Diabetes Care 2009, 32, 1678–1683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Aguiar Vallim, T.Q.; Tarling, E.J.; Edwards, P.A. Pleiotropic roles of bile acids in metabolism. Cell Metab. 2013, 17, 657–669. [Google Scholar] [CrossRef] [Green Version]
- Ridlon, J.M.; Kang, D.J.; Hylemon, P.B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 2006, 47, 241–259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trefflich, I.; Marschall, H.U.; Giuseppe, R.D.; Stahlman, M.; Michalsen, A.; Lampen, A.; Abraham, K.; Weikert, C. Associations between Dietary Patterns and Bile Acids-Results from a Cross-Sectional Study in Vegans and Omnivores. Nutrients 2019, 12, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, J.; Yang, S.; Cai, S.; Dong, J.; Li, X.; Chen, Z. Identification of biochemical changes in lactovegetarian urine using 1H NMR spectroscopy and pattern recognition. Anal. Bioanal. Chem. 2010, 396, 1451–1463. [Google Scholar] [CrossRef]
- Vessally, E.; Soleimani-Amiri, S.; Hosseinian, A.; Edjlali, L.; Bekhradnia, A. New protocols to access imidazoles and their ring fused analogues: Synthesis from N-propargylamines. Rsc Adv. 2017, 7, 7079–7091. [Google Scholar] [CrossRef] [Green Version]
- McKeran, R.O.; Halliday, D.; Purkiss, P.; Royston, P. 3-Methylhistidine excretion as an index of myofibrillar protein catabolism in neuromuscular disease. J. Neurol. Neurosurg. Psychiatry 1979, 42, 536–541. [Google Scholar] [CrossRef] [Green Version]
- Myint, T.; Fraser, G.E.; Lindsted, K.D.; Knutsen, S.F.; Hubbard, R.W.; Bennett, H.W. Urinary 1-methylhistidine is a marker of meat consumption in Black and in White California Seventh-day Adventists. Am. J. Epidemiol. 2000, 152, 752–755. [Google Scholar] [CrossRef]
- Sullards, M.C.; Lynch, D.V.; Merrill, A.H., Jr.; Adams, J. Structure determination of soybean and wheat glucosylceramides by tandem mass spectrometry. J. Mass. Spectrom. 2000, 35, 347–353. [Google Scholar] [CrossRef]
- Raj, D.S.; Ouwendyk, M.; Francoeur, R.; Pierratos, A. Plasma amino acid profile on nocturnal hemodialysis. Blood Purif. 2000, 18, 97–102. [Google Scholar] [CrossRef] [PubMed]
- Tuma, P.; Samcova, E.; Balinova, P. Determination of 3-methylhistidine and 1-methylhistidine in untreated urine samples by capillary electrophoresis. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2005, 821, 53–59. [Google Scholar] [CrossRef]
- Batch, B.C.; Shah, S.H.; Newgard, C.B.; Turer, C.B.; Haynes, C.; Bain, J.R.; Muehlbauer, M.; Patel, M.J.; Stevens, R.D.; Appel, L.J.; et al. Branched chain amino acids are novel biomarkers for discrimination of metabolic wellness. Metabolism 2013, 62, 961–969. [Google Scholar] [CrossRef] [Green Version]
- Weng, L.; Quinlivan, E.; Gong, Y.; Beitelshees, A.L.; Shahin, M.H.; Turner, S.T.; Chapman, A.B.; Gums, J.G.; Johnson, J.A.; Frye, R.F.; et al. Association of branched and aromatic amino acids levels with metabolic syndrome and impaired fasting glucose in hypertensive patients. Metab. Syndr. Relat. Disord. 2015, 13, 195–202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamaguchi, N.; Mahbub, M.H.; Takahashi, H.; Hase, R.; Ishimaru, Y.; Sunagawa, H.; Amano, H.; Kobayashi-Miura, M.; Kanda, H.; Fujita, Y.; et al. Plasma free amino acid profiles evaluate risk of metabolic syndrome, diabetes, dyslipidemia, and hypertension in a large Asian population. Environ. Health Prev. Med. 2017, 22, 35. [Google Scholar] [CrossRef] [Green Version]
- Guasch-Ferre, M.; Hruby, A.; Toledo, E.; Clish, C.B.; Martinez-Gonzalez, M.A.; Salas-Salvado, J.; Hu, F.B. Metabolomics in Prediabetes and Diabetes: A Systematic Review and Meta-analysis. Diabetes Care 2016, 39, 833–846. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, T.J.; Larson, M.G.; Vasan, R.S.; Cheng, S.; Rhee, E.P.; McCabe, E.; Lewis, G.D.; Fox, C.S.; Jacques, P.F.; Fernandez, C.; et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 2011, 17, 448–453. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Jiang, H.; Zhang, Z.; Duan, W.; Han, T.; Sun, C. Interaction between dietary branched-chain amino acids and genetic risk score on the risk of type 2 diabetes in Chinese. Genes Nutr. 2021, 16, 4. [Google Scholar] [CrossRef] [PubMed]
- Lachkar, F.; Ferre, P.; Foufelle, F.; Papaioannou, A. Dihydroceramides: Their emerging physiological roles and functions in cancer and metabolic diseases. Am. J. Physiol. Endocrinol. Metab. 2021, 320, E122–E130. [Google Scholar] [CrossRef]
- Meikle, P.J.; Summers, S.A. Sphingolipids and phospholipids in insulin resistance and related metabolic disorders. Nat. Rev. Endocrinol. 2017, 13, 79–91. [Google Scholar] [CrossRef]
- Sokolowska, E.; Blachnio-Zabielska, A. The Role of Ceramides in Insulin Resistance. Front. Endocrinol 2019, 10, 577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haus, J.M.; Kashyap, S.R.; Kasumov, T.; Zhang, R.; Kelly, K.R.; Defronzo, R.A.; Kirwan, J.P. Plasma ceramides are elevated in obese subjects with type 2 diabetes and correlate with the severity of insulin resistance. Diabetes 2009, 58, 337–343. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Filippov, V.; Song, M.A.; Zhang, K.; Vinters, H.V.; Tung, S.; Kirsch, W.M.; Yang, J.; Duerksen-Hughes, P.J. Increased ceramide in brains with Alzheimer’s and other neurodegenerative diseases. J. Alzheimers Dis. 2012, 29, 537–547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimm, M.O.W.; Michaelson, D.M.; Hartmann, T. Omega-3 fatty acids, lipids, and apoE lipidation in Alzheimer’s disease: A rationale for multi-nutrient dementia prevention. J. Lipid Res. 2017, 58, 2083–2101. [Google Scholar] [CrossRef] [Green Version]
- Kurz, J.; Parnham, M.J.; Geisslinger, G.; Schiffmann, S. Ceramides as Novel Disease Biomarkers. Trends Mol. Med. 2019, 25, 20–32. [Google Scholar] [CrossRef]
- Nakayama, H.; Nagafuku, M.; Suzuki, A.; Iwabuchi, K.; Inokuchi, J.I. The regulatory roles of glycosphingolipid-enriched lipid rafts in immune systems. FEBS Lett. 2018, 592, 3921–3942. [Google Scholar] [CrossRef] [Green Version]
- Aronov, P.A.; Luo, F.J.; Plummer, N.S.; Quan, Z.; Holmes, S.; Hostetter, T.H.; Meyer, T.W. Colonic contribution to uremic solutes. J. Am. Soc. Nephrol. 2011, 22, 1769–1776. [Google Scholar] [CrossRef] [Green Version]
- Ottosson, F.; Brunkwall, L.; Smith, E.; Orho-Melander, M.; Nilsson, P.M.; Fernandez, C.; Melander, O. The gut microbiota-related metabolite phenylacetylglutamine associates with increased risk of incident coronary artery disease. J. Hypertens. 2020, 38, 2427–2434. [Google Scholar] [CrossRef]
- Ussher, J.R.; Lopaschuk, G.D.; Arduini, A. Gut microbiota metabolism of L-carnitine and cardiovascular risk. Atherosclerosis 2013, 231, 456–461. [Google Scholar] [CrossRef] [PubMed]
- Makrecka-Kuka, M.; Sevostjanovs, E.; Vilks, K.; Volska, K.; Antone, U.; Kuka, J.; Makarova, E.; Pugovics, O.; Dambrova, M.; Liepinsh, E. Plasma acylcarnitine concentrations reflect the acylcarnitine profile in cardiac tissues. Sci. Rep. 2017, 7, 17528. [Google Scholar] [CrossRef] [Green Version]
- DiNicolantonio, J.J.; O’Keefe, J.H. Good Fats versus Bad Fats: A Comparison of Fatty Acids in the Promotion of Insulin Resistance, Inflammation, and Obesity. Mo. Med. 2017, 114, 303–307. [Google Scholar] [PubMed]
- Miller, M.; Stone, N.J.; Ballantyne, C.; Bittner, V.; Criqui, M.H.; Ginsberg, H.N.; Goldberg, A.C.; Howard, W.J.; Jacobson, M.S.; Kris-Etherton, P.M.; et al. Triglycerides and cardiovascular disease: A scientific statement from the American Heart Association. Circulation 2011, 123, 2292–2333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneda, T. Iso- and anteiso-fatty acids in bacteria: Biosynthesis, function, and taxonomic significance. Microbiol. Rev. 1991, 55, 288–302. [Google Scholar] [CrossRef] [PubMed]
- Mika, A.; Stepnowski, P.; Kaska, L.; Proczko, M.; Wisniewski, P.; Sledzinski, M.; Sledzinski, T. A comprehensive study of serum odd- and branched-chain fatty acids in patients with excess weight. Obesity 2016, 24, 1669–1676. [Google Scholar] [CrossRef] [Green Version]
- Ran-Ressler, R.R.; Khailova, L.; Arganbright, K.M.; Adkins-Rieck, C.K.; Jouni, Z.E.; Koren, O.; Ley, R.E.; Brenna, J.T.; Dvorak, B. Branched chain fatty acids reduce the incidence of necrotizing enterocolitis and alter gastrointestinal microbial ecology in a neonatal rat model. PLoS ONE 2011, 6, e29032. [Google Scholar] [CrossRef] [Green Version]
- Su, X.; Magkos, F.; Zhou, D.; Eagon, J.C.; Fabbrini, E.; Okunade, A.L.; Klein, S. Adipose tissue monomethyl branched-chain fatty acids and insulin sensitivity: Effects of obesity and weight loss. Obesity 2015, 23, 329–334. [Google Scholar] [CrossRef] [Green Version]
- Jaceldo-Siegl, K.; Haddad, E.; Knutsen, S.; Fan, J.; Lloren, J.; Bellinger, D.; Fraser, G.E. Lower C-reactive protein and IL-6 associated with vegetarian diets are mediated by BMI. Nutr. Metab. Cardiovas. 2018, 28, 787–794. [Google Scholar] [CrossRef] [PubMed]
- Orlich, M.J.; Fraser, G.E. Vegetarian diets in the Adventist Health Study 2: A review of initial published findings. Am. J. Clin. Nutr. 2014, 100 (Suppl. S1), 353S–358S. [Google Scholar] [CrossRef] [Green Version]

| Non-Vegetarian | Vegan | p-Value | |
|---|---|---|---|
| Participants, n | 46 | 47 | |
| Demographic | |||
| Sex | 1 | ||
| Male | 23 (50) | 24 (51) | |
| Female | 23 (50) | 23 (49) | |
| Age (years) | 60.8 (11.1) | 66.5 (9.9) | 0.011 |
| BMI (kg/m2) 3 | 31.3 (7.0) | 24.7 (3.4) | <0.001 |
| Ethnicity | 1 | ||
| Caucasian | 23 (50) | 23 (49) | |
| Black | 23 (50) | 24 (51) | |
| Dietary | |||
| Total kca3 | 2520.8 (902.8) | 1776.8 (744.7) | <0.001 |
| Fiber (g/d) | 19.6 (6.5) | 40.5 (8.8) | <0.001 |
| Saturated fat (g/d) | 21.8 (7.9) | 9.0 (2.7) | <0.001 |
| Total meat (g/d) | 104.9 (41.0) | 0 | < 0.001 |
| Red meat (g/d) | 54.9 (22.8) | 0 | < 0.001 |
| Processed meat (g/d) | 6.4 (6.6) | 0 | <0.001 |
| Poultry (g/d) | 31.2 (21.6) | 0 | < 0.001 |
| Fish (g/d) | 20.7 (17.2) | 0 | < 0.001 |
| Dairy (kcal/d) | 252.8 (151.7) | 17.0 (14.7) 3 | <0.001 |
| Fruit (kcal/d) | 108.3 (64.9) | 320.9 (159.3) | <0.001 |
| Vegetables (kcal/d) | 67.6 (36.3) | 134.5 (115.4) | <0.001 |
| Soy (kcal/d) | 60.06 (42.3) | 156.3 (99.2) | <0.001 |
| Legumes (kcal/d) | 46.1 (56.4) | 73.2 (56.0) | <0.001 |
| Whole grains (kcal/d) | 172.4 (130.9) | 326.6 (189.8) | <0.022 |
| Lifestyle | |||
| Exercise (min/week) 4 | 63.6 (100.8) | 132.3 (110.6) | 0.003 |
| Smoking (ever vs. never) | 12 (26.1) | 2 (4.3) | 0.008 |
| Alcohol drinking (ever vs. never) | 24 (52.2) | 12 (25.5) | 0.015 |
| Coffee (any vs. never/rarely) | 18 (39.1) | 0 | <0.001 |
| Aspirin/NSAIDS (<1/wk vs. ≥1/wk) | 11 (23.9) | 3 (6.4) | 0.038 |
| Metabolite | Fold Change | FDR | Subclass | Major Class |
|---|---|---|---|---|
| 4-ethylphenyl sulfate | 6.58 | 5.2 × 10−4 | Benzoate Metabolism | Xenobiotics |
| s-methylmethionine | 4.61 | 2.0 × 10−4 | Methionine, Cysteine, SAM, and Taurine Metabolism | Amino Acid |
| branched chain 14:0 dicarboxylic acid | 3.54 | 2.2 × 10−4 | Fatty Acid, Dicarboxylate | Lipid |
| 4-acetylphenyl sulfate | 3.20 | 2.6 × 10−4 | Benzoate Metabolism | Xenobiotics |
| glycohyocholate | 2.86 | 6.3 × 10−4 | Secondary Bile Acid Metabolism | Lipid |
| ethyl beta-glucopyranoside | 2.79 | 1.5 × 10−4 | Food Component/Plant | Xenobiotics |
| methyl glucopyranoside (alpha + beta) | 2.64 | 2.9 × 10−4 | Food Component/Plant | Xenobiotics |
| beta-cryptoxanthin | 2.52 | 1.0 × 10−3 | Vitamin A Metabolism | Cofactors and Vitamins |
| n-methylproline | 2.28 | 9.0 × 10−5 | Urea cycle; Arginine and Proline Metabolism | Amino Acid |
| stachydrine | 2.27 | 1.6 × 10−4 | Food Component/Plant | Xenobiotics |
| 4-allylcatechol sulfate | 2.21 | 6.7 × 10−5 | Benzoate Metabolism | Xenobiotics |
| indolepropionate | 2.03 | 9.8 × 10−5 | Tryptophan Metabolism | Amino Acid |
| 2-methylserine | 1.95 | 1.3 × 10−4 | Glycine, Serine, and Threonine Metabolism | Amino Acid |
| 1-linoleoyl-2-linolenoyl-gpc (18:2/18:3) | 1.93 | 4.5 × 10−4 | Phosphatidylcholine (PC) | Lipid |
| s-methylcysteine sulfoxide | 1.93 | 1.1 × 10−4 | Methionine, Cysteine, SAM, and Taurine Metabolism | Amino Acid |
| daidzein sulfate (2) | 1.91 | 9.8 × 10−3 | Food Component/Plant | Xenobiotics |
| gentisate | 1.90 | 7.8 × 10−5 | Tyrosine Metabolism | Amino Acid |
| 4-allylphenol sulfate | 1.89 | 2.4 × 10−3 | Food Component/Plant | Xenobiotics |
| dodecanedioate (c12:1-dc) | 1.88 | 1.8 × 10−4 | Fatty Acid, Dicarboxylate | Lipid |
| glycochenodeoxycholate 3-sulfate | 1.87 | 2.3 × 10−3 | Primary Bile Acid Metabolism | Lipid |
| 3-hydroxybutyrate (bhba) | 1.86 | 1.7 × 10−4 | Ketone Bodies | Lipid |
| octadecanedioate (c18:1-dc) | 1.83 | 3.5 × 10−4 | Fatty Acid, Dicarboxylate | Lipid |
| cinnamoylglycine | 1.82 | 1.7 × 10−3 | Food Component/Plant | Xenobiotics |
| n-delta-acetylornithine | 1.79 | 1.6 × 10−4 | Urea cycle; Arginine and Proline Metabolism | Amino Acid |
| 3-hydroxydodecanedioate | 1.76 | 3.9 × 10−4 | Fatty Acid, Dicarboxylate | Lipid |
| catechol sulfate | 1.75 | 1.2 × 10−4 | Benzoate Metabolism | Xenobiotics |
| octadecadienedioate (c18:2-dc) | 1.75 | 1.2 × 10−4 | Fatty Acid, Dicarboxylate | Lipid |
| tryptophan betaine | 1.74 | 1.8 × 10−3 | Tryptophan Metabolism | Amino Acid |
| n-linoleoylglycine | 1.70 | 6.2 × 10−5 | Fatty Acid Metabolism (Acyl Glycine) | Lipid |
| 4-methoxyphenol sulfate | 1.69 | 1.4 × 10−3 | Tyrosine Metabolism | Amino Acid |
| octadecenedioylcarnitine (c18:1-dc) | 1.69 | 1.4 × 10−4 | Fatty Acid Metabolism (Acyl Carnitine, Dicarboxylate) | Lipid |
| 4-ethylcatechol sulfate | 1.68 | 3.5 × 10−3 | Benzoate Metabolism | Xenobiotics |
| 12,13-dihome | 1.66 | 9.5 × 10−5 | Fatty Acid, Dihydroxy | Lipid |
| carotene diol (2) | 1.65 | 8.0 × 10−5 | Vitamin A Metabolism | Cofactors and Vitamins |
| chiro-inositol | 1.65 | 1.2 × 10−3 | Inositol Metabolism | Lipid |
| 2-acetamidophenol sulfate | 1.65 | 2.1 × 10−2 | Food Component/Plant | Xenobiotics |
| 3-hydroxysebacate | 1.63 | 2.1 × 10−4 | Fatty Acid, Monohydroxy | Lipid |
| acetoacetate | 1.63 | 1.6 × 10−4 | Ketone Bodies | Lipid |
| Cis-4-decenoate (10:1n6) | 1.63 | 1.2 × 10−4 | Medium Chain Fatty Acid | Lipid |
| pentose acid | 1.62 | 7.7 × 10−5 | Partially Characterized Molecules | Partially Characterized Molecules |
| Metabolite | Fold Change | Subclass | Major Class |
|---|---|---|---|
| 5-acetylamino-6-amino-3-methyluracil | 0.04 | Xanthine Metabolism | Xenobiotics |
| theobromine | 0.04 | Xanthine Metabolism | Xenobiotics |
| 3-carboxy-4-methyl-5-propyl-2-furanpropanoate (cmpf) | 0.05 | Fatty Acid, Dicarboxylate | Lipid |
| paraxanthine | 0.07 | Xanthine Metabolism | Xenobiotics |
| theophylline | 0.07 | Xanthine Metabolism | Xenobiotics |
| 3-methylhistidine | 0.07 | Histidine Metabolism | Amino Acid |
| 1-methyl-5-imidazoleacetate | 0.09 | Histidine Metabolism | Amino Acid |
| hydroxy-cmpf | 0.09 | Fatty Acid, Dicarboxylate | Lipid |
| 1,7-dimethylurate | 0.09 | Xanthine Metabolism | Xenobiotics |
| 4-acetaminophen sulfate | 0.10 | Drug—Analgesics, Anesthetics | Xenobiotics |
| caffeine | 0.11 | Xanthine Metabolism | Xenobiotics |
| piperine | 0.12 | Food Component/Plant | Xenobiotics |
| 7-methylxanthine | 0.17 | Xanthine Metabolism | Xenobiotics |
| 2-hydroxyacetaminophen sulfate | 0.18 | Drug—Analgesics, Anesthetics | Xenobiotics |
| 3-methylxanthine | 0.19 | Xanthine Metabolism | Xenobiotics |
| 4-acetamidophenol | 0.19 | Drug—Analgesics, Anesthetics | Xenobiotics |
| sulfate of piperine metabolite c16h19no3 (2) | 0.19 | Food Component/Plant | Xenobiotics |
| 1-methylurate | 0.20 | Xanthine Metabolism | Xenobiotics |
| 5-acetylamino-6-formylamino-3-methyluracil | 0.20 | Xanthine Metabolism | Xenobiotics |
| 3-bromo-5-chloro-2,6-dihydroxybenzoic acid | 0.22 | Chemical | Xenobiotics |
| sulfate of piperine metabolite c16h19no3 (3) | 0.23 | Food Component/Plant | Xenobiotics |
| n,n,n-trimethyl-5-aminovalerate | 0.24 | Lysine Metabolism | Amino Acid |
| glucuronide of piperine metabolite c17h21no3 (4) | 0.25 | Food Component/Plant | Xenobiotics |
| sulfate of piperine metabolite c18h21no3 (1) | 0.26 | Food Component/Plant | Xenobiotics |
| heptenedioate (c7:1-dc) | 0.27 | Fatty Acid, Dicarboxylate | Lipid |
| glucuronide of piperine metabolite c17h21no3 (3) | 0.27 | Food Component/Plant | Xenobiotics |
| glucuronide of piperine metabolite c17h21no3 (5) | 0.28 | Food Component/Plant | Xenobiotics |
| 1-methylxanthine | 0.29 | Xanthine Metabolism | Xenobiotics |
| (14 or 15)-methyl palmitate (a17:0 or i17:0) | 0.29 | Fatty Acid, Branched | Lipid |
| sulfate of piperine metabolite c18h21no3 (3) | 0.29 | Food Component/Plant | Xenobiotics |
| ibuprofen | 0.29 | Drug—Analgesics, Anesthetics | Xenobiotics |
| (12 or 13)-methyl myristate (a15:0 or i15:0) | 0.30 | Fatty Acid, Branched | Lipid |
| 1,3-dimethylurate | 0.32 | Xanthine Metabolism | Xenobiotics |
| 3-methyl catechol sulfate (1) | 0.32 | Benzoate Metabolism | Xenobiotics |
| 1-margaroylglycerol (17:0) | 0.32 | Monoacylglycerol | Lipid |
| perfluorooctanesulfonate (pfos) | 0.33 | Chemical | Xenobiotics |
| saccharin | 0.35 | Food Component/Plant | Xenobiotics |
| tridecenedioate (c13:1-dc) | 0.36 | Fatty Acid, Dicarboxylate | Lipid |
| 3,5-dichloro-2,6-dihydroxybenzoic acid | 0.38 | Chemical | Xenobiotics |
| sphingomyelin (d18:1/25:0, d19:0/24:1, d20:1/23:0, d19:1/24:0) | 0.39 | Sphingomyelins | Lipid |
| Subclass | Fold Change (95% CI) | FDR 2 | Significant Metabolites (n) | #↓ 3 | #↑ 4 | n Total Metabolites 5 |
|---|---|---|---|---|---|---|
| Ketone Bodies | 1.75 (2.16, 1.43) | 1.0 × 10−3 | 2 | 0 | 2 | 2 |
| Vitamin A Metabolism | 1.40 (1.57, 1.25) | 2.0 × 10−3 | 6 | 2 | 4 | 6 |
| Inositol Metabolism | 1.35 (1.57, 1.16) | 6.0 × 10−3 | 2 | 0 | 2 | 2 |
| Fatty Acid Metabolism (Acyl Glycine) | 1.21 (1.35, 1.08) | 2.6 × 10−2 | 5 | 1 | 4 | 7 |
| Lactosylceramides (LCER) | 1.17 (1.28, 1.07) | 2.8 × 10−2 | 2 | 0 | 2 | 3 |
| Benzoate Metabolism | 1.16 (1.27, 1.06) | 4.8 × 10−2 | 16 | 7 | 9 | 24 |
| Amino Sugar Metabolism | 0.94 (1.00, 0.88) | 4.7 × 10−2 | 2 | 2 | 0 | 5 |
| Urea cycle; Arginine and Proline Metabolism | 0.94 (0.99, 0.90) | 2.3 × 10−2 | 14 | 10 | 4 | 21 |
| Glutamate Metabolism | 0.94 (0.99, 0.89) | 3.4 × 10−2 | 8 | 5 | 3 | 12 |
| Tyrosine Metabolism | 0.92 (0.98, 0.86) | 1.1 × 10−2 | 11 | 9 | 2 | 22 |
| Pyrimidine Metabolism, Uracil containing | 0.92 (0.97, 0.88) | 1.1 × 10−3 | 5 | 5 | 0 | 12 |
| Purine Metabolism, Adenine containing | 0.92 (0.97, 0.88) | 1.4 × 10−3 | 3 | 3 | 0 | 6 |
| Fatty Acid, Dicarboxylate | 0.91 (0.98, 0.85) | 1.5 × 10−2 | 22 | 10 | 12 | 34 |
| Long-Chain Polyunsaturated Fatty Acid (n3 and n6) | 0.90 (0.97, 0.83) | 1.0 × 10−2 | 12 | 9 | 3 | 17 |
| Phospholipid Metabolism | 0.90 (0.96, 0.85) | 1.2 × 10−3 | 4 | 4 | 0 | 7 |
| Sphingolipid Synthesis | 0.89 (0.98, 0.80) | 2.1 × 10−2 | 2 | 2 | 0 | 3 |
| Alanine and Aspartate Metabolism | 0.89 (0.94, 0.85) | 1.1 × 10−4 | 6 | 5 | 1 | 9 |
| Partially Characterized Molecules | 0.88 (0.98, 0.79) | 2.8 × 10−2 | 8 | 6 | 2 | 13 |
| Creatine Metabolism | 0.86 (0.92, 0.79) | 1.7 × 10−4 | 3 | 2 | 1 | 3 |
| Diacylglycerol | 0.84 (0.97, 0.72) | 2.5 × 10−2 | 6 | 6 | 0 | 14 |
| Pyrimidine Metabolism, Orotate containing | 0.84 (0.93, 0.76) | 9.9 × 10−4 | 3 | 3 | 0 | 4 |
| Purine Metabolism, (Hypo)Xanthine/Inosine containing | 0.84 (0.90, 0.77) | <3.9 × 10−5 | 5 | 5 | 0 | 7 |
| Fructose, Mannose, and Galactose Metabolism | 0.83 (0.98, 0.70) | 3.4 × 10−2 | 2 | 2 | 0 | 4 |
| Secondary Bile Acid Metabolism | 0.83 (0.95, 0.72) | 8.1 × 10−3 | 11 | 9 | 2 | 21 |
| Purine Metabolism, Guanine containing | 0.83 (0.90, 0.76) | <3.9 × 10−5 | 3 | 3 | 0 | 3 |
| Sphingomyelins | 0.83 (0.89, 0.77) | <3.9 × 10−5 | 22 | 18 | 4 | 29 |
| Food Component/Plant | 0.83 (0.88, 0.78) | <3.9 × 10−5 | 36 | 22 | 14 | 51 |
| Fatty Acid Metabolism (also BCAA Metabolism) | 0.82 (0.93, 0.72) | 4.8 × 10−3 | 4 | 4 | 0 | 5 |
| Tryptophan Metabolism | 0.82 (0.88, 0.76) | <3.9 × 10−5 | 17 | 14 | 3 | 20 |
| Pantothenate and CoA Metabolism | 0.81 (0.97, 0.68) | 2.6 × 10−2 | 1 | 1 | 0 | 2 |
| Guanidino and Acetamido Metabolism | 0.80 (0.94, 0.69) | 1.0 × 10−2 | 2 | 2 | 0 | 2 |
| Lysine Metabolism | 0.80 (0.87, 0.74) | <3.9 × 10−5 | 11 | 11 | 0 | 18 |
| Plasmalogen | 0.80 (0.85, 0.75) | <3.9 × 10−5 | 7 | 6 | 1 | 11 |
| Glycerolipid Metabolism | 0.79 (0.88, 0.71) | 4.3 × 10−5 | 3 | 3 | 0 | 3 |
| Leucine, Isoleucine, and Valine Metabolism | 0.79 (0.84, 0.75) | <3.9 × 10−5 | 24 | 23 | 1 | 32 |
| Dihydrosphingomyelins | 0.78 (0.89, 0.69) | 3.0 × 10−4 | 5 | 4 | 1 | 5 |
| Chemical | 0.78 (0.83, 0.73) | <3.9 × 10−5 | 13 | 10 | 3 | 20 |
| Monoacylglycerol | 0.77 (0.84, 0.69) | <3.9 × 10−5 | 11 | 11 | 0 | 17 |
| Lysoplasmalogen | 0.75 (0.82, 0.68) | <3.9 × 10−5 | 4 | 4 | 0 | 4 |
| Phenylalanine Metabolism | 0.75 (0.80, 0.70) | <3.9 × 10−5 | 6 | 6 | 0 | 7 |
| Long-Chain Monounsaturated Fatty Acid | 0.74 (0.80, 0.68) | <3.9 × 10−5 | 4 | 4 | 0 | 7 |
| Long-Chain Saturated Fatty Acid | 0.73 (0.78, 0.68) | <3.9 × 10−5 | 7 | 7 | 0 | 8 |
| Fatty Acid Metabolism (Acyl Carnitine, Long-Chain Saturated) | 0.72 (0.80, 0.65) | <3.9 × 10−5 | 8 | 8 | 0 | 8 |
| Ceramides | 0.64 (0.72, 0.57) | <3.9 × 10−5 | 10 | 10 | 0 | 11 |
| Dihydroceramides | 0.62 (0.74, 0.52) | <3.9 × 10−5 | 2 | 2 | 0 | 2 |
| Acetylated Peptides | 0.59 (0.74, 0.48) | <3.9 × 10−5 | 4 | 4 | 0 | 4 |
| Histidine Metabolism | 0.59 (0.65, 0.55) | <3.9 × 10−5 | 11 | 10 | 1 | 15 |
| Fatty Acid, Branched | 0.33 (0.38, 0.29) | <3.9 × 10−5 | 3 | 3 | 0 | 3 |
| Drug—Analgesics, Anesthetics | 0.22 (0.29, 0.16) | <3.9 × 10−5 | 5 | 5 | 0 | 5 |
| Xanthine Metabolism | 0.14 (0.20, 0.10) | <3.9 × 10−5 | 13 | 13 | 0 | 13 |
| β | SE | T Value | p-Value | Correlation Coefficient | p-Value (Correlation) | |
|---|---|---|---|---|---|---|
| Vegan | −0.5 | 7.9 × 10−7 | ||||
| PC1 | −7.33 | 1.96 | −3.7 | 3.3 × 10−4 | ||
| PC3 | 8.52 | 1.88 | 4.6 | 1.7 × 10−5 | ||
| PC4 | −5.23 | 1.44 | −3.6 | 5.0 × 10−4 | ||
| BMI | 0.61 | 2.1 × 10−10 | ||||
| PC1 | 0.99 | 0.15 | 6.5 | 4.7 × 10−9 | ||
| PC2 | −0.28 | 0.13 | −2.1 | 4.0 × 10−2 | ||
| PC3 | −0.35 | 0.15 | −2.4 | 2.0 × 10−2 | ||
| Red meat | 0.52 | 3.5 × 10−7 | ||||
| PC1 | 1.89 | 0.49 | 3.9 | 3.7 × 10−3 | ||
| PC3 | −2.14 | 0.47 | −4.6 | 1.7 × 10−5 | ||
| PC4 | 1.35 | 0.36 | 3.7 | 3.5 × 10−4 | ||
| Total meat | 0.51 | 4.0 × 10−7 | ||||
| PC1 | 1.61 | 0.42 | 3.8 | 2.7 × 10−4 | ||
| PC3 | −1.86 | 0.4 | −4.6 | 1.4 × 10−5 | ||
| PC4 | 1.17 | 0.31 | 3.8 | 3.2 × 10−4 | ||
| Processed meat | ||||||
| PC1 | 1.51 | 0.37 | 4.1 | 8.9 × 10−5 | 0.44 | 2.04 × 10−5 |
| PC3 | −0.79 | 0.39 | −2.1 | 4.0 × 10−2 | ||
| PC4 | 0.75 | 0.28 | 2.7 | 9.6 × 10−3 | ||
| Poultry | ||||||
| PC1 | 1.99 | 0.57 | 3.49 | 7.6 × 10−4 | 0.45 | 1.10 x 10−5 |
| PC3 | −2.25 | 0.55 | −4.1 | 9.2 × 10−5 | ||
| PC4 | 1.45 | 0.42 | 3.46 | 8.5 × 10−4 | ||
| Fish | 0.52 | 2.6 × 10−7 | ||||
| PC1 | 1.91 | 0.61 | 3.1 | 2.0 × 10−3 | ||
| PC2 | −1.15 | 0.56 | −2.1 | 4.0 × 10−2 | ||
| PC3 | −2.27 | 0.58 | −3.9 | 2.0 × 10−4 | ||
| PC4 | 1.64 | 0.44 | 3.8 | 3.0 × 10−4 | ||
| Fiber | −0.41 | 7.2 × 10−5 | ||||
| PC1 | −6.67 | 2.18 | −3.1 | 2.9 × 10−3 | ||
| PC3 | 9.13 | 2.04 | 4.5 | 2.2 × 10−5 | ||
| PC4 | −3.32 | 1.65 | −2.0 | 5.0 × 10−2 | ||
| Soy | −0.22 | 0.04 | ||||
| PC4 | −1.7 | 0.72 | −2.4 | 2.0 × 10−2 | ||
| PC7 | 1.21 | 0.53 | 2.3 | 2.5 × 10−2 | ||
| Vegetables | −0.23 | 0.03 | ||||
| PC1 | −2.7 | 1.18 | −2.3 | 2.5 × 10−2 | ||
| PC8 | −2.4 | 0.65 | −3.7 | 4.3 × 10−4 | ||
| Fruit | ||||||
| PC1 | −4.69 | 0.94 | −5.0 | 3.4 × 10−6 | −0.47 | 3.4 × 10−6 |
| Dairy | 0.63 | 9.6 × 10−11 | ||||
| PC1 | 2.27 | 0.56 | 4.1 | 9.7 × 10−5 | ||
| PC2 | −1.03 | 0.53 | −2.0 | 5.4 × 10−2 | ||
| PC3 | −2.06 | 0.56 | −3.7 | 3.8 × 10−4 | ||
| PC4 | 1.26 | 0.42 | 3.0 | 3.9 × 10−3 | ||
| Saturated fat | ||||||
| PC1 | 7.44 | 1.63 | 4.6 | 1.7 × 10−5 | 0.55 | 3.1 × 10−8 |
| PC3 | −6.7 | 1.64 | −4.1 | 9.3 × 10−5 | ||
| PC4 | 3.24 | 1.29 | 2.5 | 1.4 × 10−2 | ||
| Whole grains | ||||||
| PC3 | 3.14 | 0.89 | 3.5 | 6.6 × 10−4 | 0.36 | 6.6 × 10−4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miles, F.L.; Orlich, M.J.; Mashchak, A.; Chandler, P.D.; Lampe, J.W.; Duerksen-Hughes, P.; Fraser, G.E. The Biology of Veganism: Plasma Metabolomics Analysis Reveals Distinct Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort. Nutrients 2022, 14, 709. https://doi.org/10.3390/nu14030709
Miles FL, Orlich MJ, Mashchak A, Chandler PD, Lampe JW, Duerksen-Hughes P, Fraser GE. The Biology of Veganism: Plasma Metabolomics Analysis Reveals Distinct Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort. Nutrients. 2022; 14(3):709. https://doi.org/10.3390/nu14030709
Chicago/Turabian StyleMiles, Fayth L., Michael J. Orlich, Andrew Mashchak, Paulette D. Chandler, Johanna W. Lampe, Penelope Duerksen-Hughes, and Gary E. Fraser. 2022. "The Biology of Veganism: Plasma Metabolomics Analysis Reveals Distinct Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort" Nutrients 14, no. 3: 709. https://doi.org/10.3390/nu14030709
APA StyleMiles, F. L., Orlich, M. J., Mashchak, A., Chandler, P. D., Lampe, J. W., Duerksen-Hughes, P., & Fraser, G. E. (2022). The Biology of Veganism: Plasma Metabolomics Analysis Reveals Distinct Profiles of Vegans and Non-Vegetarians in the Adventist Health Study-2 Cohort. Nutrients, 14(3), 709. https://doi.org/10.3390/nu14030709







