Gut Microbiota, in the Halfway between Nutrition and Lung Function
Abstract
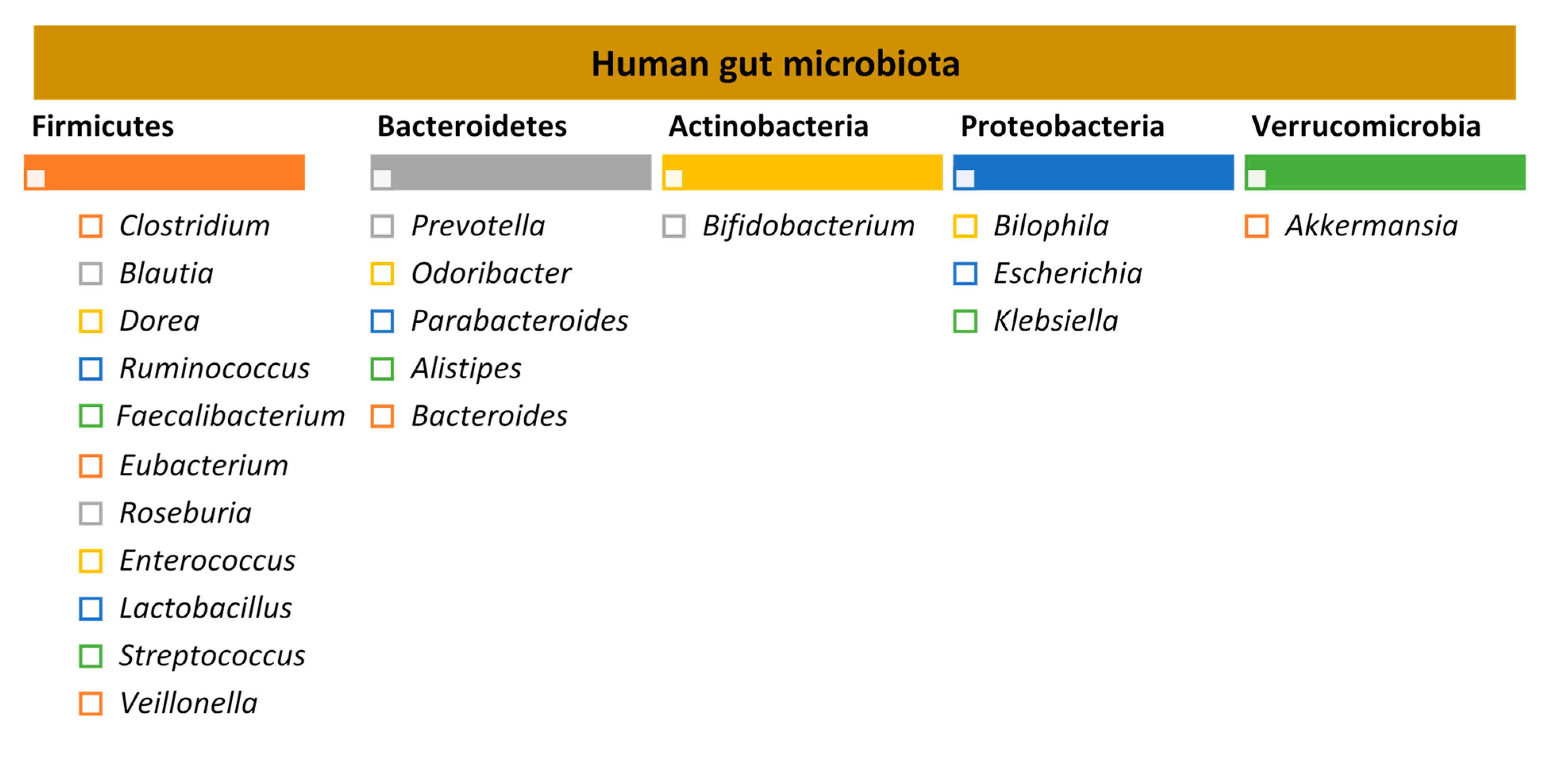
1. The Human Gut Microbiota
2. Through the Intestinal Barrier: Microbiota–Immune System Interactions
3. Diet and Gut Microbiota
4. Diet, Gut Microbiota, Immune System and Lung Diseases
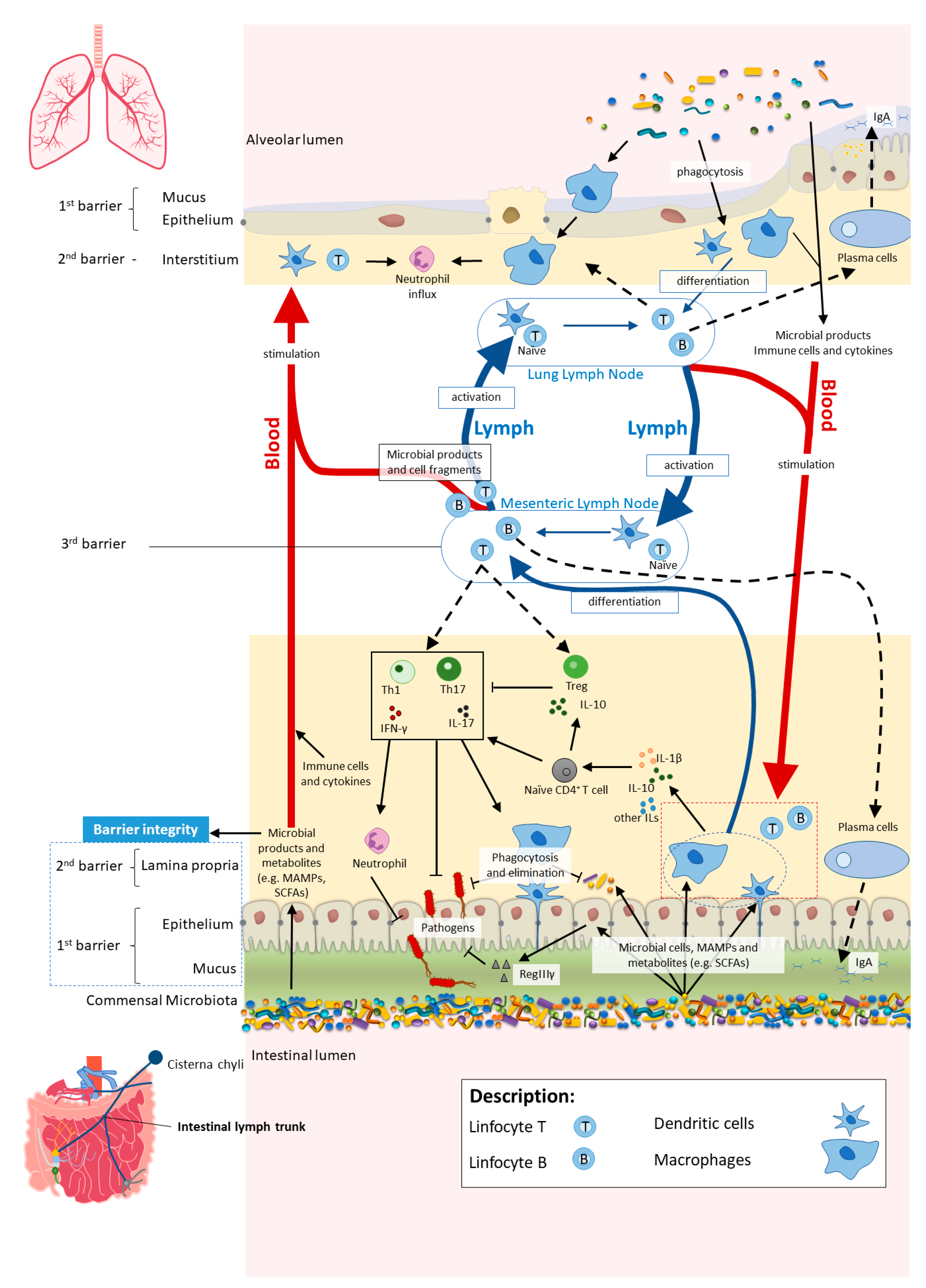
4.1. The Cross-Talk between the Gut and the Lungs: Long-Reaching Immune Modulation
4.2. The Diet–Gut–Lung Axis in Lung Diseases: Asthma, Chronic Obstructive Pulmonary Disease and Cystic Fibrosis
4.2.1. Asthma
4.2.2. Chronic Obstructive Pulmonary Disease
4.2.3. Cystic Fibrosis
5. Perspectives
Supplementary Materials
Funding
Acknowledgments
Conflicts of Interest
References
- Sekirov, I.; Russell, S.L.; Antunes, L.C.M.; Finlay, B.B. Gut Microbiota in Health and Disease. Physiol. Rev. 2010, 90, 859–904. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Peterson, D.A.; Gordon, J.I. Ecological and Evolutionary Forces Shaping Microbial Diversity in the Human Intestine. Cell 2006, 124, 837–848. [Google Scholar] [CrossRef]
- Ostaff, M.J.; Stange, E.F.; Wehkamp, J. Antimicrobial peptides and gut microbiota in homeostasis and pathology. EMBO Mol. Med. 2013, 5, 1465–1483. [Google Scholar] [CrossRef] [PubMed]
- Sender, R.; Fuchs, S.; Milo, R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLoS Biol. 2016, 14, e1002533. [Google Scholar] [CrossRef]
- Cani, P.D. Human gut microbiome: Hopes, threats and promises. Gut 2018, 67, 1716–1725. [Google Scholar] [CrossRef] [PubMed]
- Segal, J.P.; Mullish, B.H.; Quraishi, M.N.; Acharjee, A.; Williams, H.R.T.; Iqbal, T.; Hart, A.L.; Marchesi, J.R. The application of omics techniques to understand the role of the gut microbiota in inflammatory bowel disease. Ther. Adv. Gastroenterol. 2019, 12, 12. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Power, S.E.; O’Toole, P.W.; Stanton, C.; Ross, R.P.; Fitzgerald, G.F. Intestinal microbiota, diet and health. Br. J. Nutr. 2013, 111, 387–402. [Google Scholar] [CrossRef] [PubMed]
- Valdes, A.M.; Walter, J.; Segal, E.; Spector, T.D. Role of the gut microbiota in nutrition and health. BMJ 2018, 361, k2179. [Google Scholar] [CrossRef]
- Rooks, M.G.; Garrett, W.S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 2016, 16, 341–352. [Google Scholar] [CrossRef]
- Lavelle, A.; Sokol, H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 223–237. [Google Scholar] [CrossRef] [PubMed]
- Krautkramer, K.A.; Fan, J.; Bäckhed, F. Gut microbial metabolites as multi-kingdom intermediates. Nat. Rev. Microbiol. 2021, 19, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Koenig, J.E.; Spor, A.; Scalfone, N.; Fricker, A.D.; Stombaugh, J.; Knight, R.; Angenent, L.T.; Ley, R.E. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 2011, 108, 4578–4585. [Google Scholar] [CrossRef]
- Bäckhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host Microbe 2015, 17, 690–703. [Google Scholar] [CrossRef] [PubMed]
- Thursby, E.; Juge, N. Introduction to the human gut microbiota. Biochem. J. 2017, 474, 1823–1836. [Google Scholar] [CrossRef]
- Rodríguez, J.M.; Murphy, K.; Stanton, C.; Ross, R.P.; Kober, O.I.; Juge, N.; Avershina, E.; Rudi, K.; Narbad, A.; Jenmalm, M.C.; et al. The composition of the gut microbiota throughout life, with an emphasis on early life. Microb. Ecol. Health Dis. 2015, 26, 26050. [Google Scholar] [CrossRef]
- Fouhy, F.; Guinane, C.M.; Hussey, S.; Wall, R.; Ryan, C.A.; Dempsey, E.M.; Murphy, B.; Ross, R.P.; Fitzgerald, G.F.; Stanton, C.; et al. High-Throughput Sequencing Reveals the Incomplete, Short-Term Recovery of Infant Gut Microbiota following Parenteral Antibiotic Treatment with Ampicillin and Gentamicin. Antimicrob. Agents Chemother. 2012, 56, 5811–5820. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef]
- Bocci, V. The neglected organ: Bacterial flora has a crucial immunostimulatory role. Perspect. Biol. Med. 1992, 35, 251–260. [Google Scholar] [CrossRef]
- O’Hara, A.M.; Shanahan, F. The gut flora as a forgotten organ. EMBO Rep. 2006, 7, 688–693. [Google Scholar] [CrossRef]
- Stephens, R.W.; Arhire, L.; Covasa, M. Gut Microbiota: From Microorganisms to Metabolic Organ Influencing Obesity. Obesity 2018, 26, 801–809. [Google Scholar] [CrossRef] [PubMed]
- Qin, N.; Zheng, B.; Yao, J.; Guo, L.; Zuo, J.; Wu, L.; Zhou, J.; Liu, L.; Guo, J.; Ni, S.; et al. Influence of H7N9 virus infection and associated treatment on human gut microbiota. Sci. Rep. 2015, 5, 14771. [Google Scholar] [CrossRef] [PubMed]
- Goodman, B.; Gardner, H. The microbiome and cancer. J. Pathol. 2018, 244, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Mowat, A.M.; Agace, W.W. Regional specialization within the intestinal immune system. Nat. Rev. Immunol. 2014, 14, 667–685. [Google Scholar] [CrossRef]
- Bain, C.C.; Mowat, A.M. Macrophages in intestinal homeostasis and inflammation. Immunol. Rev. 2014, 260, 102–117. [Google Scholar] [CrossRef]
- Groschwitz, K.R.; Hogan, S.P. Intestinal barrier function: Molecular regulation and disease pathogenesis. J. Allergy Clin. Immunol. 2009, 124, 3–20, quiz 21-2. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; MacPherson, A.J. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat. Rev. Immunol. 2010, 10, 159–169. [Google Scholar] [CrossRef]
- Abreu, M.T.; Fukata, M.; Arditi, M. TLR Signaling in the Gut in Health and Disease. J. Immunol. 2005, 174, 4453–4460. [Google Scholar] [CrossRef]
- Philpott, D.J.; Girardin, S.E. The role of Toll-like receptors and Nod proteins in bacterial infection. Mol. Immunol. 2004, 41, 1099–1108. [Google Scholar] [CrossRef]
- Nishiya, T.; DeFranco, A.L. Ligand-regulated Chimeric Receptor Approach Reveals Distinctive Subcellular Localization and Signaling Properties of the Toll-like Receptors. J. Biol. Chem. 2004, 279, 19008–19017. [Google Scholar] [CrossRef]
- Abreu, M.T. Toll-like receptor signalling in the intestinal epithelium: How bacterial recognition shapes intestinal function. Nat. Rev. Immunol. 2010, 10, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Hand, T.W. Role of the Microbiota in Immunity and Inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef] [PubMed]
- Tsolis, R.M.; Bäumler, A.J. Gastrointestinal host-pathogen interaction in the age of microbiome research. Curr. Opin. Microbiol. 2020, 53, 78–89. [Google Scholar] [CrossRef]
- Zheng, L.; Kelly, C.J.; Battista, K.D.; Schaefer, R.; Lanis, J.M.; Alexeev, E.E.; Wang, R.X.; Onyiah, J.C.; Kominsky, D.J.; Colgan, S.P. Microbial-Derived Butyrate Promotes Epithelial Barrier Function through IL-10 Receptor–Dependent Repression of Claudin-2. J. Immunol. 2017, 199, 2976–2984. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett, W.S. The Microbial Metabolites, Short-Chain Fatty Acids, Regulate Colonic Treg Cell Homeostasis. Science 2013, 341, 569–573. [Google Scholar] [CrossRef]
- Littman, D.R.; Pamer, E.G. Role of the Commensal Microbiota in Normal and Pathogenic Host Immune Responses. Cell Host Microbe 2011, 10, 311–323. [Google Scholar] [CrossRef]
- Pandiyan, P.; Bhaskaran, N.; Zou, M.; Schneider, E.; Jayaraman, S.; Huehn, J. Microbiome Dependent Regulation of Tregs and Th17 Cells in Mucosa. Front. Immunol. 2019, 10, 426. [Google Scholar] [CrossRef]
- Lee, N.; Kim, W.-U. Microbiota in T-cell homeostasis and inflammatory diseases. Exp. Mol. Med. 2017, 49, e340. [Google Scholar] [CrossRef]
- Gaboriau-Routhiau, V.; Rakotobe, S.; Lécuyer, E.; Mulder, I.; Lan, A.; Bridonneau, C.; Rochet, V.; Pisi, A.; De Paepe, M.; Brandi, G.; et al. The Key Role of Segmented Filamentous Bacteria in the Coordinated Maturation of Gut Helper T Cell Responses. Immunity 2009, 31, 677–689. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Liu, C.H.; Tzianabos, A.O.; Kasper, D.L. An Immunomodulatory Molecule of Symbiotic Bacteria Directs Maturation of the Host Immune System. Cell 2005, 122, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of colonic regulatory T cells by indigenous clostridium species. Science 2011, 331, 337–341. [Google Scholar] [CrossRef]
- Lazar, V.; Ditu, L.-M.; Pircalabioru, G.G.; Gheorghe, I.; Curutiu, C.; Holban, A.M.; Picu, A.; Petcu, L.; Chifiriuc, M.C. Aspects of Gut Microbiota and Immune System Interactions in Infectious Diseases, Immunopathology, and Cancer. Front. Immunol. 2018, 9, 1830. [Google Scholar] [CrossRef]
- MacPherson, A.J.; Slack, E.; Geuking, M.B.; McCoy, K.D. The mucosal firewalls against commensal intestinal microbes. Semin. Immunopathol. 2009, 31, 145–149. [Google Scholar] [CrossRef]
- König, J.; Wells, J.; Cani, P.; Garcia-Rodenas, C.L.; Macdonald, T.; Mercenier, A.; Whyte, J.; Troost, F.; Brummer, R.-J. Human Intestinal Barrier Function in Health and Disease. Clin. Transl. Gastroenterol. 2016, 7, e196. [Google Scholar] [CrossRef] [PubMed]
- Cone, R.A. Barrier properties of mucus. Adv. Drug Deliv. Rev. 2009, 61, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Paone, P.; Cani, P.D. Mucus barrier, mucins and gut microbiota: The expected slimy partners? Gut 2020, 69, 2232–2243. [Google Scholar] [CrossRef]
- Chelakkot, C.; Ghim, J.; Ryu, S.H. Mechanisms regulating intestinal barrier integrity and its pathological implications. Exp. Mol. Med. 2018, 50, 1–9. [Google Scholar] [CrossRef]
- Moschen, A.R.; Wieser, V.; Tilg, H. Dietary Factors: Major Regulators of the Gut’s Microbiota. Gut Liver 2012, 6, 411–416. [Google Scholar] [CrossRef]
- Rinninella, E.; Cintoni, M.; Raoul, P.; Lopetuso, L.R.; Scaldaferri, F.; Pulcini, G.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. Food Components and Dietary Habits: Keys for a Healthy Gut Microbiota Composition. Nutrients 2019, 11, 2393. [Google Scholar] [CrossRef]
- Marsland, B.J.; Trompette, A.; Gollwitzer, E.S. The Gut-Lung Axis in Respiratory Disease. Ann. Am. Thorac. Soc. 2015, 12, S150–S156. [Google Scholar] [CrossRef]
- Bingula, R.; Filaire, M.; Radosevic-Robin, N.; Bey, M.; Berthon, J.-Y.; Bernalier-Donadille, A.; Vasson, M.-P.; Filaire, E. Desired Turbulence? Gut-Lung Axis, Immunity, and Lung Cancer. J. Oncol. 2017, 2017, 1–15. [Google Scholar] [CrossRef]
- Budden, K.F.; Gellatly, S.L.; Wood, D.L.A.; Cooper, M.A.; Morrison, M.; Hugenholtz, P.; Hansbro, P.M. Emerging pathogenic links between microbiota and the gut–lung axis. Nat. Rev. Microbiol. 2017, 15, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Trompette, A.; Gollwitzer, E.S.; Yadava, K.; Sichelstiel, A.K.; Sprenger, N.; Ngom-Bru, C.; Blanchard, C.; Junt, T.; Nicod, L.P.; Harris, N.L.; et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 2014, 20, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Halnes, I.; Baines, K.J.; Berthon, B.S.; MacDonald-Wicks, L.K.; Gibson, P.G.; Wood, L.G. Soluble Fibre Meal Challenge Reduces Airway Inflammation and Expression of GPR43 and GPR41 in Asthma. Nutrients 2017, 9, 57. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, A.; Frazer, Z.A.; Hansbro, P.; Yang, I.A. COPD and the gut-lung axis: The therapeutic potential of fibre. J. Thorac. Dis. 2019, 11, S2173–S2180. [Google Scholar] [CrossRef]
- Raftery, A.L.; Tsantikos, E.; Harris, N.L.; Hibbs, M.L. Links Between Inflammatory Bowel Disease and Chronic Obstructive Pulmonary Disease. Front. Immunol. 2020, 11, 2144. [Google Scholar] [CrossRef]
- Dumas, A.; Bernard, L.; Poquet, Y.; Lugo-Villarino, G.; Neyrolles, O. The role of the lung microbiota and the gut-lung axis in respiratory infectious diseases. Cell. Microbiol. 2018, 20, e12966. [Google Scholar] [CrossRef]
- He, Y.; Wen, Q.; Yao, F.; Xu, D.; Huang, Y.; Wang, J. Gut–lung axis: The microbial contributions and clinical implications. Crit. Rev. Microbiol. 2017, 43, 81–95. [Google Scholar] [CrossRef]
- Anand, S.; Mande, S.S. Diet, Microbiota and Gut-Lung Connection. Front. Microbiol. 2018, 9, 2147. [Google Scholar] [CrossRef]
- McGhee, J.R.; Fujihashi, K. Inside the Mucosal Immune System. PLoS Biol. 2012, 10. [Google Scholar] [CrossRef]
- Date, Y.; Ebisawa, M.; Fukuda, S.; Shima, H.; Obata, Y.; Takahashi, D.; Kato, T.; Hanazato, M.; Nakato, G.; Williams, I.R.; et al. NALT M cells are important for immune induction for the common mucosal immune system. Int. Immunol. 2017, 29, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Dang, A.T.; Marsland, B.J. Microbes, metabolites, and the gut-lung axis. Mucosal. Immunol. 2019, 12, 843–850. [Google Scholar] [CrossRef]
- Enaud, R.; Prevel, R.; Ciarlo, E.; Beaufils, F.; Wieërs, G.; Guery, B.; Delhaes, L. The Gut-Lung Axis in Health and Respiratory Diseases: A Place for Inter-Organ and Inter-Kingdom Crosstalks. Front. Cell. Infect. Microbiol. 2020, 10, 9. [Google Scholar] [CrossRef]
- Zhang, D.; Li, S.; Wang, N.; Tan, H.-Y.; Zhang, Z.; Feng, Y. The Cross-Talk Between Gut Microbiota and Lungs in Common Lung Diseases. Front. Microbiol. 2020, 11, 301. [Google Scholar] [CrossRef] [PubMed]
- Bilotta, A.J.; Cong, Y. Gut microbiota metabolite regulation of host defenses at mucosal surfaces: Implication in precision medicine. Precis. Clin. Med. 2019, 2, 110–119. [Google Scholar] [CrossRef] [PubMed]
- Macia, L.; Tan, J.; Vieira, A.T.; Leach, K.; Stanley, D.; Luong, S.; Maruya, M.; McKenzie, C.I.; Hijikata, A.; Wong, C.; et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef]
- Trompette, A.; Gollwitzer, E.S.; Pattaroni, C.; Lopez-Mejia, I.C.; Riva, E.; Pernot, J.; Ubags, N.; Fajas, L.; Nicod, L.P.; Marsland, B.J. Dietary Fiber Confers Protection against Flu by Shaping Ly6c− Patrolling Monocyte Hematopoiesis and CD8+ T Cell Metabolism. Immunity 2018, 48, 992–1005.e8. [Google Scholar] [CrossRef]
- Cait, A.; Hughes, M.R.; Antignano, F.; Cait, J.; Dimitriu, P.A.; Maas, K.R.; Reynolds, L.A.; Hacker, L.; Mohr, J.; Finlay, B.B.; et al. Microbiome-driven allergic lung inflammation is ameliorated by short-chain fatty acids. Mucosal Immunol. 2018, 11, 785–795. [Google Scholar] [CrossRef]
- Zheng, D.; Liwinski, T.; Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020, 30, 492–506. [Google Scholar] [CrossRef]
- Steed, A.L.; Christophi, G.P.; Kaiko, G.E.; Sun, L.; Goodwin, V.M.; Jain, U.; Esaulova, E.; Artyomov, M.N.; Morales, D.J.; Holtzman, M.J.; et al. The microbial metabolite desaminotyrosine protects from influenza through type I interferon. Science 2017, 357, 498–502. [Google Scholar] [CrossRef] [PubMed]
- McAleer, J.P.; Nguyen, N.L.H.; Chen, K.; Kumar, P.; Ricks, D.M.; Binnie, M.; Armentrout, R.A.; Pociask, D.A.; Hein, A.; Yu, A.; et al. Pulmonary Th17 Antifungal Immunity Is Regulated by the Gut Microbiome. J. Immunol. 2016, 197, 97–107. [Google Scholar] [CrossRef]
- Noverr, M.C.; Noggle, R.M.; Toews, G.B.; Huffnagle, G.B. Role of Antibiotics and Fungal Microbiota in Driving Pulmonary Allergic Responses. Infect. Immun. 2004, 72, 4996–5003. [Google Scholar] [CrossRef] [PubMed]
- Bradley, C.P.; Teng, F.; Felix, K.M.; Sano, T.; Naskar, D.; Block, K.E.; Huang, H.; Knox, K.S.; Littman, D.R.; Wu, H.-J.J. Segmented Filamentous Bacteria Provoke Lung Autoimmunity by Inducing Gut-Lung Axis Th17 Cells Expressing Dual TCRs. Cell Host Microbe 2017, 22, 697–704.e4. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Mao, K.; Chen, X.; Sun, M.-A.; Kawabe, T.; Li, W.; Usher, N.; Zhu, J.; Urban, J.F.; Paul, W.E.; et al. S1P-dependent interorgan trafficking of group 2 innate lymphoid cells supports host defense. Science 2018, 359, 114–119. [Google Scholar] [CrossRef]
- Ichinohe, T.; Pang, I.K.; Kumamoto, Y.; Peaper, D.R.; Ho, J.H.; Murray, T.S.; Iwasaki, A. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl. Acad. Sci. USA 2011, 108, 5354–5359. [Google Scholar] [CrossRef]
- Oh, J.Z.; Ravindran, R.; Chassaing, B.; Carvalho, F.A.; Maddur, M.S.; Bower, M.; Hakimpour, P.; Gill, K.P.; Nakaya, H.I.; Yarovinsky, F.; et al. TLR5-Mediated Sensing of Gut Microbiota Is Necessary for Antibody Responses to Seasonal Influenza Vaccination. Immunity 2014, 41, 478–492. [Google Scholar] [CrossRef]
- Tsay, T.-B.; Yang, M.-C.; Chen, P.-H.; Hsu, C.-M.; Chen, L.-W. Gut flora enhance bacterial clearance in lung through toll-like receptors 4. J. Biomed. Sci. 2011, 18. [Google Scholar] [CrossRef]
- Clarke, T.B.; Davis, K.M.; Lysenko, E.S.; Zhou, A.Y.; Yu, Y.; Weiser, J.N. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 2010, 16, 228–231. [Google Scholar] [CrossRef]
- Asthma. Available online: https://www.who.int/news-room/fact-sheets/detail/asthma (accessed on 28 April 2021).
- Elsen, L.W.J.V.D.; Garssen, J.; Burcelin, R.; Verhasselt, V. Shaping the Gut Microbiota by Breastfeeding: The Gateway to Allergy Prevention? Front. Pediatr. 2019, 7, 47. [Google Scholar] [CrossRef] [PubMed]
- Rautava, S. Early microbial contact, the breast milk microbiome and child health. J. Dev. Orig. Health Dis. 2016, 7, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Klopp, A.; Vehling, L.; Becker, A.B.; Subbarao, P.; Mandhane, P.J.; Turvey, S.E.; Lefebvre, D.L.; Sears, M.R.; Daley, D.; Silverman, F.; et al. Modes of Infant Feeding and the Risk of Childhood Asthma: A Prospective Birth Cohort Study. J. Pediatr. 2017, 190, 192–199.e2. [Google Scholar] [CrossRef]
- Johnson, A.M.F.; De Paolo, R.W. Window-of-opportunity: Neonatal gut microbiota and atopy. HepatoBiliary Surg. Nutr. 2017, 6, 190–192. [Google Scholar] [CrossRef] [PubMed]
- Arrieta, M.-C.; Stiemsma, L.T.; Dimitriu, P.A.; Thorson, L.; Russell, S.; Yurist-Doutsch, S.; Kuzeljevic, B.; Gold, M.J.; Britton, H.M.; Lefebvre, D.L.; et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 2015, 7. [Google Scholar] [CrossRef]
- Fujimura, K.E.; Sitarik, A.R.; Havstad, S.; Lin, D.L.; LeVan, S.; Fadrosh, D.; Panzer, A.R.; LaMere, B.; Rackaityte, E.; Lukacs, N.W.; et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat. Med. 2016, 22, 1187–1191. [Google Scholar] [CrossRef]
- Galazzo, G.; van Best, N.; Bervoets, L.; Dapaah, I.O.; Savelkoul, P.H.; Hornef, M.W.; Hutton, E.K.; Morrison, K.; Holloway, A.C.; McDonald, H.; et al. Development of the Microbiota and Associations with Birth Mode, Diet, and Atopic Disorders in a Longitudinal Analysis of Stool Samples, Collected from Infancy Through Early Childhood. Gastroenterology 2020, 158, 1584–1596. [Google Scholar] [CrossRef] [PubMed]
- Demirci, M.; Tokman, H.; Uysal, H.; Demiryas, S.; Karakullukcu, A.; Saribas, S.; Cokugras, H.; Kocazeybek, B. Reduced Akkermansia muciniphila and Faecalibacterium prausnitzii levels in the gut microbiota of children with allergic asthma. Allergol. Immunopathol. 2019, 47, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Anand, S.; Kaur, H.; Mande, S.S. Comparative In silico Analysis of Butyrate Production Pathways in Gut Commensals and Pathogens. Front. Microbiol. 2016, 7, 1945. [Google Scholar] [CrossRef]
- Quévrain, E.; Maubert, M.A.; Michon, C.; Chain, F.; Marquant, R.; Tailhades, J.; Miquel, S.; Carlier, L.; Bermúdez-Humarán, L.G.; Pigneur, B.; et al. Identification of an anti-inflammatory protein from Faecalibacterium prausnitzii, a commensal bacterium deficient in Crohn’s disease. Gut 2016, 65, 415–425. [Google Scholar] [CrossRef]
- Thorburn, A.N.; McKenzie, C.I.; Shen, S.; Stanley, D.; Macia, L.; Mason, L.J.; Roberts, L.K.; Wong, C.H.Y.; Shim, R.; Robert, R.; et al. Evidence that asthma is a developmental origin disease influenced by maternal diet and bacterial metabolites. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Roduit, C.; Frei, R.; Ferstl, R.; Loeliger, S.; Westermann, P.; Rhyner, C.; Schiavi, E.; Barcik, W.; Rodriguez-Perez, N.; Wawrzyniak, M.; et al. High levels of butyrate and propionate in early life are associated with protection against atopy. Allergy 2018, 74, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Ivashkin, V.; Zolnikova, O.; Potskherashvili, N.; Trukhmanov, A.; Kokina, N.; Dzhakhaya, N.; Sedova, A.; Bueverova, E. Metabolic Activity of Intestinal Microflora in Patients with Bronchial Asthma. Clin. Pract. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Brigham, E.P.; Kolahdooz, F.; Hansel, N.; Breysse, P.N.; Davis, M.; Sharma, S.; Matsui, E.C.; Diette, G.; McCormack, M.C. Association between Western diet pattern and adult asthma: A focused review. Ann. Allergy Asthma Immunol. 2015, 114, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Wypych, T.P.; Marsland, B.J.; Ubags, N.D.J. The Impact of Diet on Immunity and Respiratory Diseases. Ann. Am. Thorac. Soc. 2017, 14, S339–S347. [Google Scholar] [CrossRef]
- Wang, Q.; Li, F.; Liang, B.; Liang, Y.; Chen, S.; Mo, X.; Ju, Y.; Zhao, H.; Jia, H.; Spector, T.D.; et al. A metagenome-wide association study of gut microbiota in asthma in UK adults. BMC Microbiol. 2018, 18, 114. [Google Scholar] [CrossRef] [PubMed]
- Wood, L.G.; Garg, M.L.; Gibson, P.G. A high-fat challenge increases airway inflammation and impairs bronchodilator recovery in asthma. J. Allergy Clin. Immunol. 2011, 127, 1133–1140. [Google Scholar] [CrossRef] [PubMed]
- Pendyala, S.; Walker, J.M.; Holt, P.R. A High-Fat Diet Is Associated with Endotoxemia That Originates from the Gut. Gastroenterology 2012, 142, 1100–1101.e2. [Google Scholar] [CrossRef]
- Erridge, C.; Attina, T.; Spickett, C.M.; Webb, D.J. A high-fat meal induces low-grade endotoxemia: Evidence of a novel mechanism of postprandial inflammation. Am. J. Clin. Nutr. 2007, 86, 1286–1292. [Google Scholar] [CrossRef] [PubMed]
- Araújo, J.R.; Tomas, J.; Brenner, C.; Sansonetti, P.J. Impact of high-fat diet on the intestinal microbiota and small intestinal physiology before and after the onset of obesity. Biochimie 2017, 141, 97–106. [Google Scholar] [CrossRef]
- Lee-Sarwar, K.A.; Kelly, R.S.; Lasky-Su, J.; Zeiger, R.S.; O’Connor, G.T.; Sandel, M.T.; Bacharier, L.B.; Beigelman, A.; Laranjo, N.; Gold, D.R.; et al. Integrative analysis of the intestinal metabolome of childhood asthma. J. Allergy Clin. Immunol. 2019, 144, 442–454. [Google Scholar] [CrossRef]
- Mayhew, D.; Devos, N.; Lambert, C.; Brown, J.R.; Clarke, S.C.; Kim, V.L.; Magid-Slav, M.; Miller, B.E.; Ostridge, K.K.; Patel, R.; et al. Longitudinal profiling of the lung microbiome in the AERIS study demonstrates repeatability of bacterial and eosinophilic COPD exacerbations. Thorax 2018, 73, 422–430. [Google Scholar] [CrossRef]
- Sprooten, R.T.; Lenaerts, K.; Braeken, D.C.; Grimbergen, I.; Rutten, E.P.; Wouters, E.F.; Rohde, G.G. Increased Small Intestinal Permeability during Severe Acute Exacerbations of COPD. Respiration 2018, 95, 334–342. [Google Scholar] [CrossRef]
- Bowerman, K.L.; Rehman, S.F.; Vaughan, A.; Lachner, N.; Budden, K.F.; Kim, R.Y.; Wood, D.L.A.; Gellatly, S.L.; Shukla, S.D.; Wood, L.G.; et al. Disease-associated gut microbiome and metabolome changes in patients with chronic obstructive pulmonary disease. Nat. Commun. 2020, 11. [Google Scholar] [CrossRef]
- Shaheen, S.O.; Jameson, K.A.; Syddall, H.E.; Sayer, A.A.; Dennison, E.M.; Cooper, C.; Robinson, S.M. The relationship of dietary patterns with adult lung function and COPD. Eur. Respir. J. 2010, 36, 277–284. [Google Scholar] [CrossRef]
- Varraso, R.; Chiuve, S.E.; Fung, T.T.; Barr, R.G.; Hu, F.B.; Willett, W.C.; Camargo, C.A. Alternate Healthy Eating Index 2010 and risk of chronic obstructive pulmonary disease among US women and men: Prospective study. BMJ 2015, 350. [Google Scholar] [CrossRef] [PubMed]
- Szmidt, M.K.; Kaluza, J.; Harris, H.R.; Linden, A.; Wolk, A. Long-term dietary fiber intake and risk of chronic obstructive pulmonary disease: A prospective cohort study of women. Eur. J. Nutr. 2020, 59, 1869–1879. [Google Scholar] [CrossRef] [PubMed]
- Rafeeq, M.M.; Murad, H.A.S. Cystic fibrosis: Current therapeutic targets and future approaches. J. Transl. Med. 2017, 15, 84. [Google Scholar] [CrossRef] [PubMed]
- Bruzzese, E.; Callegari, M.L.; Raia, V.; Viscovo, S.; Scotto, R.; Ferrari, S.; Morelli, L.; Buccigrossi, V.; Vecchio, A.L.; Ruberto, E.; et al. Disrupted Intestinal Microbiota and Intestinal Inflammation in Children with Cystic Fibrosis and Its Restoration with Lactobacillus GG: A Randomised Clinical Trial. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Manor, O.; Levy, R.; Pope, C.E.; Hayden, H.S.; Brittnacher, M.J.; Carr, R.; Radey, M.C.; Hager, K.R.; Heltshe, S.L.; Ramsey, B.W.; et al. Metagenomic evidence for taxonomic dysbiosis and functional imbalance in the gastrointestinal tracts of children with cystic fibrosis. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Antosca, K.M.; Chernikova, D.A.; Price, C.E.; Ruoff, K.L.; Li, K.; Guill, M.F.; Sontag, N.R.; Morrison, H.G.; Hao, S.; Drumm, M.L.; et al. Altered Stool Microbiota of Infants with Cystic Fibrosis Shows a Reduction in Genera Associated with Immune Programming from Birth. J. Bacteriol. 2019, 201. [Google Scholar] [CrossRef] [PubMed]
- Madan, J.C.; Koestler, D.C.; Stanton, B.A.; Davidson, L.; Moulton, L.A.; Housman, M.L.; Moore, J.H.; Guill, M.F.; Morrison, H.G.; Sogin, M.L.; et al. Serial Analysis of the Gut and Respiratory Microbiome in Cystic Fibrosis in Infancy: Interaction between Intestinal and Respiratory Tracts and Impact of Nutritional Exposures. mBio 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Poulimeneas, D.; Grammatikopoulou, M.G.; Devetzi, P.; Petrocheilou, A.; Kaditis, A.G.; Papamitsou, T.; Doudounakis, S.E.; Vassilakou, T. Adherence to Dietary Recommendations, Nutrient Intake Adequacy and Diet Quality among Pediatric Cystic Fibrosis Patients: Results from the GreeCF Study. Nutrients 2020, 12, 3126. [Google Scholar] [CrossRef]
- Gutiérrez-Carrasquilla, L.; Sánchez, E.; Hernández, M.; Polanco, D.; Salas-Salvadó, J.; Betriu, À.; Gaeta, A.M.; Carmona, P.; Purroy, F.; Pamplona, R.; et al. Effects of Mediterranean Diet and Physical Activity on Pulmonary Function: A Cross-Sectional Analysis in the ILERVAS Project. Nutrients 2019, 11, 329. [Google Scholar] [CrossRef] [PubMed]
- Yeoh, Y.K.; Zuo, T.; Lui, G.C.-Y.; Zhang, F.; Liu, Q.; Li, A.Y.; Chung, A.C.; Cheung, C.P.; Tso, E.Y.; Fung, K.S.; et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 2021. [Google Scholar] [CrossRef]
- Conti, P.; Caraffa, A.; Gallenga, C.E.; Ross, R.; Kritas, S.K.; Frydas, I.; Younes, A.; Ronconi, G. Coronavirus-19 (SARS-CoV-2) induces acute severe lung inflammation via IL-1 causing cytokine storm in COVID-19: A promising inhibitory strategy. J. Biol. Regul. Homeost. Agents 2020, 34, 1971–1975. [Google Scholar] [CrossRef] [PubMed]
- Conti, P.; Ronconi, G.; Caraffa, A.; Gallenga, C.; Ross, R.; Frydas, I.; Kritas, S. Induction of pro-inflammatory cytokines (IL-1 and IL-6) and lung inflammation by Coronavirus-19 (COVI-19 or SARS-CoV-2): Anti-inflammatory strategies. J. Biol. Regul. Homeost. Agents 2020, 34. [Google Scholar] [CrossRef]
- Bousquet, J.; Anto, J.M.; Iaccarino, G.; Czarlewski, W.; Haahtela, T.; Anto, A.; Akdis, C.A.; Blain, H.; Canonica, G.W.; Cardona, V.; et al. Is diet partly responsible for differences in COVID-19 death rates between and within countries? Clin. Transl. Allergy 2020, 10. [Google Scholar] [CrossRef]
- Day, R.L.; Harper, A.J.; Woods, R.M.; Davies, O.G.; Heaney, L.M. Probiotics: Current landscape and future horizons. Futur. Sci. OA 2019, 5. [Google Scholar] [CrossRef]
- Anderson, J.L.; Miles, C.; Tierney, A.C. Effect of probiotics on respiratory, gastrointestinal and nutritional outcomes in patients with cystic fibrosis: A systematic review. J. Cyst. Fibros. 2017, 16, 186–197. [Google Scholar] [CrossRef]
- Durack, J.; Kimes, N.E.; Lin, D.L.; Rauch, M.; Mckean, M.; McCauley, K.; Panzer, A.R.; Mar, J.S.; Cabana, M.D.; Lynch, S.V. Delayed gut microbiota development in high-risk for asthma infants is temporarily modifiable by Lactobacillus supplementation. Nat. Commun. 2018, 9, 707. [Google Scholar] [CrossRef]
- Du, X.; Wang, L.; Wu, S.; Yuan, L.; Tang, S.; Xiang, Y.; Qu, X.; Liu, H.; Qin, X.; Liu, C. Efficacy of probiotic supplementary therapy for asthma, allergic rhinitis, and wheeze: A meta-analysis of randomized controlled trials. Allergy Asthma Proc. 2019, 40, 250–260. [Google Scholar] [CrossRef] [PubMed]
- Wypych, T.P.; Wickramasinghe, L.C.; Marsland, B.J. The influence of the microbiome on respiratory health. Nat. Immunol. 2019, 20, 1279–1290. [Google Scholar] [CrossRef] [PubMed]
- Saint-Criq, V.; Lugo-Villarino, G.; Thomas, M. Dysbiosis, malnutrition and enhanced gut-lung axis contribute to age-related respiratory diseases. Ageing Res. Rev. 2021, 66, 101235. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Espírito Santo, C.; Caseiro, C.; Martins, M.J.; Monteiro, R.; Brandão, I. Gut Microbiota, in the Halfway between Nutrition and Lung Function. Nutrients 2021, 13, 1716. https://doi.org/10.3390/nu13051716
Espírito Santo C, Caseiro C, Martins MJ, Monteiro R, Brandão I. Gut Microbiota, in the Halfway between Nutrition and Lung Function. Nutrients. 2021; 13(5):1716. https://doi.org/10.3390/nu13051716
Chicago/Turabian StyleEspírito Santo, Christophe, Catarina Caseiro, Maria João Martins, Rosário Monteiro, and Inês Brandão. 2021. "Gut Microbiota, in the Halfway between Nutrition and Lung Function" Nutrients 13, no. 5: 1716. https://doi.org/10.3390/nu13051716
APA StyleEspírito Santo, C., Caseiro, C., Martins, M. J., Monteiro, R., & Brandão, I. (2021). Gut Microbiota, in the Halfway between Nutrition and Lung Function. Nutrients, 13(5), 1716. https://doi.org/10.3390/nu13051716






