Non-Biopsy Serology-Based Diagnosis of Celiac Disease in Adults Is Accurate with Different Commercial Kits and Pre-Test Probabilities
Abstract
1. Introduction
2. Materials and Methods
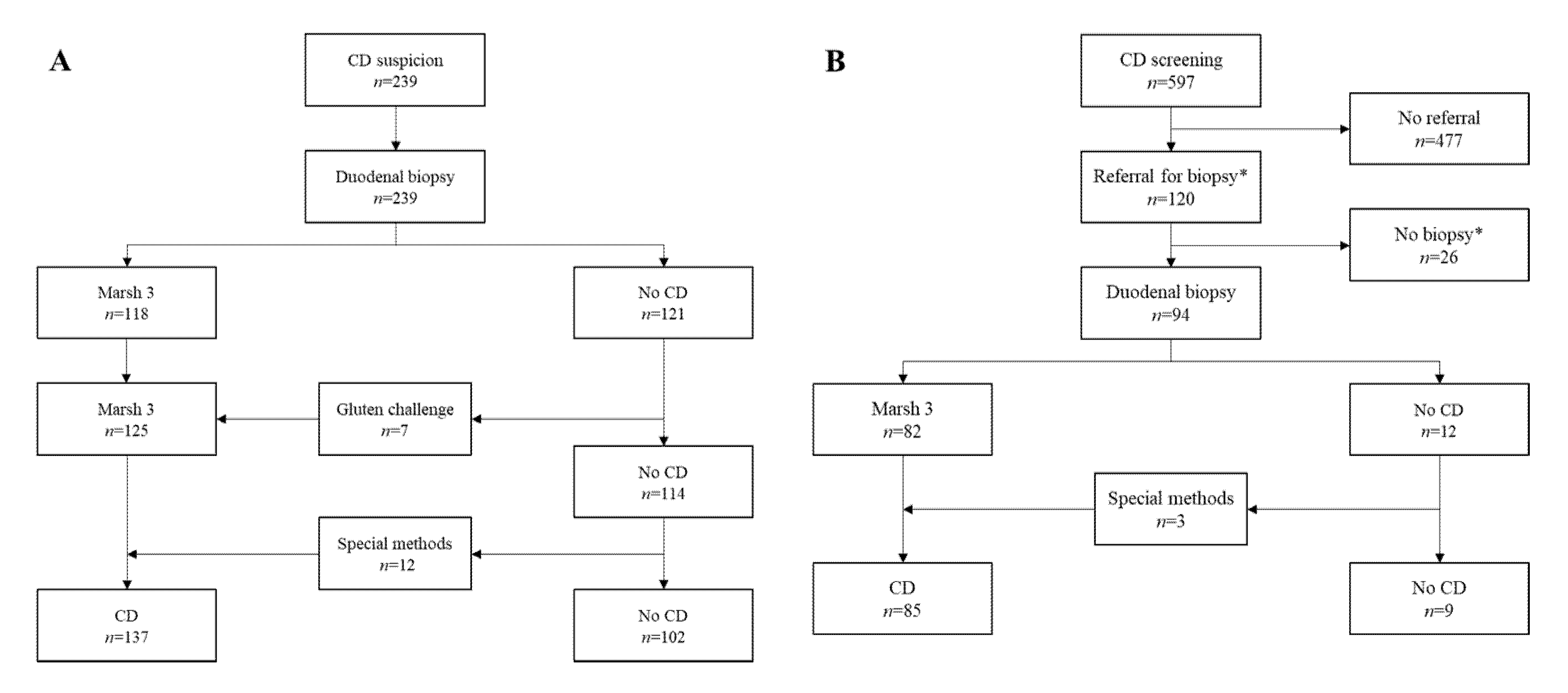
2.1. Patients and Study Design
2.2. Ethics
2.3. Serological and Genetic Testing
2.4. Histology
2.5. Statistics
3. Results
4. Discussion
4.1. Strengths and Weaknesses
4.2. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Vilppula, A.; Kaukinen, K.; Luostarinen, L.; Krekelä, I.; Patrikainen, H.; Valve, R.; Mäki, M.; Collin, P. Increasing prevalence and high incidence of celiac disease in elderly people: A population-based study. BMC Gastroenterol. 2009, 9, 1–5. [Google Scholar] [CrossRef]
- Makharia, G.K.; Catassi, C. Celiac disease in Asia. Gastroenterol. Clin. N. Am. 2019, 48, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Taavela, J.; Koskinen, O.; Huhtala, H.; Lähdeaho, M.-L.; Popp, A.; Laurila, K.; Collin, P.; Kaukinen, K.; Kurppa, K.; Mäki, M. Validation of morphometric analyses of small-intestinal biopsy readouts in celiac disease. PLoS ONE 2013, 8, e76163. [Google Scholar] [CrossRef] [PubMed]
- Freeman, H.J. Pearls and pitfalls in the diagnosis of adult celiac disease. Can. J. Gastroenterol. 2008, 22, 273–280. [Google Scholar] [CrossRef]
- Husby, S.; Koletzko, S.; Korponay-Szabó, I.R.; Kurppa, K.; Mearin, M.L.; Ribes-Koninckx, C.; Shamir, R.; Troncone, R.; Auricchio, R.; Castillejo, G.; et al. European Society Paediatric Gastroenterology, Hepatology and Nutrition guidelines for diagnosing celiac disease 2020. J. Pediatr. Gastroenterol. Nutr. 2020, 70, 141–156. [Google Scholar] [CrossRef]
- Tortora, R.; Imperatore, N.; Capone, P.; De Palma, G.D.; De Stefano, G.; Gerbino, N.; Caporaso, N.; Rispo, A. The presence of anti-endomysial antibodies and the level of anti-tissue transglutaminases can be used to diagnose adult celiac disease without duodenal biopsy. Aliment. Pharmacol. Ther. 2014, 40, 1223–1229. [Google Scholar] [CrossRef]
- Efthymakis, K.; Serio, M.; Milano, A.; Laterza, F.; Bonitatibus, A.; Di Nicola, M.; Neri, M. Application of the biopsy-sparing ESPGHAN guidelines for celiac disease diagnosis in adults: A real-life study. Dig. Dis. Sci. 2017, 62, 2433–2439. [Google Scholar] [CrossRef]
- Fuchs, V.; Kurppa, K.; Huhtala, H.; Laurila, K.; Mäki, M.; Collin, P.; Salmi, T.; Luostarinen, L.; Saavalainen, P.; Kaukinen, K. Serology-based criteria for adult celiac disease have excellent accuracy across the range of pre-test probabilities. Aliment. Pharmacol. Ther. 2019, 49, 277–284. [Google Scholar] [CrossRef]
- Werkstetter, K.J.; Korponay-Szabó, I.R.; Popp, A.; Villanacci, V.; Salemme, M.; Heilig, G.; Lillevang, S.T.; Mearin, M.L.; Ribes-Koninckx, C.; Thomas, A.; et al. Accuracy in diagnosis of celiac disease without biopsies in clinical practice. Gastroenterology 2017, 153, 924–935. [Google Scholar] [CrossRef]
- Kauma, S.; Kaukinen, K.; Huhtala, H.; Kivelä, L.; Pekki, H.; Salmi, T.; Saavalainen, P.; Lindfors, K.; Kurppa, K. The phenotype of celiac disease has low concordance between siblings, despite a similar distribution of HLA haplotypes. Nutrients 2019, 25, 11. [Google Scholar] [CrossRef]
- Ladinser, B.; Rossipal, E.; Pittschieler, K. Endomysium antibodies in celiac disease: An improved method. Gut 1994, 35, 776–778. [Google Scholar] [CrossRef] [PubMed]
- Kurppa, K.; Collin, P.; Viljamaa, M.; Haimila, K.; Saavalainen, P.; Partanen, J.; Laurila, K.; Huhtala, H.; Paasikivi, K.; Mäki, M.; et al. Diagnosing mild enteropathy celiac disease: A randomized, controlled clinical study. Gastroenterology 2009, 136, 816–823. [Google Scholar] [CrossRef] [PubMed]
- Marsh, M.N. Gluten, major histocompatibility complex, and the small intestine. A molecular and immunobiologic approach to the spectrum of gluten sensitivity (‘celiac sprue’). Gastroenterology 1992, 102, 330–354. [Google Scholar] [CrossRef]
- Järvinen, T.T.; Kaukinen, K.; Laurila, K.; Kyrönpalo, S.; Rasmussen, M.; Mäki, M.; Korhonen, H.; Reunala, T.; Collin, P. Intraepithelial lymphocytes in celiac disease. Am. J. Gastroenterol. 2003, 98, 1332–1337. [Google Scholar] [CrossRef] [PubMed]
- Kaukinen, K.; Peräaho, M.; Collin, P.; Partanen, J.; Woolley, N.; Kaartinen, T.; Nuutinen, T.; Halttunen, T.; Mäki, M.; Korponay-Szabo, I. Small-bowel mucosal transglutaminase 2-specific IgA deposits in coeliac disease without villous atrophy: A prospective and randomized clinical study. Scand. J. Gastroenterol. 2005, 40, 564–572. [Google Scholar] [CrossRef]
- Salmi, T.; Hervonen, K.; Kurppa, K.; Collin, P.; Kaukinen, K.; Reunala, T. Celiac disease evolving into dermatitis herpetiformis in patients adhering to normal or gluten-free diet. Scand. J. Gastroenterol. 2015, 50, 387–392. [Google Scholar] [CrossRef]
- Klapp, G.; Masip, E.; Bolonio, M.; Donat, E.; Polo, B.; Ramos, D.; Ribes-Koninckx, C. Celiac disease: The new proposed ESPGHAN diagnostic criteria do work well in a selected population. J. Pediatr. Gastroenterol. Nutr. 2013, 56, 251–256. [Google Scholar] [CrossRef]
- Mubarak, A.; Wolters, V.M.; Gmelig-Meyling, F.H.; Kate, F.J.T.; Houwen, R.H. Tissue transglutaminase levels above 100 U/mL and celiac disease: A prospective study. World J. Gastroenterol. 2012, 28, 4399–4403. [Google Scholar] [CrossRef]
- Wolf, J.; Petroff, D.; Richter, T.; Auth, M.K.; Uhlig, H.H.; Laass, M.W.; Lauenstein, P.; Krahl, A.; Händel, N.; De Laffolie, J.; et al. Validation of antibody-based strategies for diagnosis of pediatric celiac disease without biopsy. Gastroenterology 2017, 153, 410–419. [Google Scholar] [CrossRef]
- Popp, A.; Arvola, T.; Taavela, J.; Kivelä, L.; Ene, A.; Laurila, K.; Saavalainen, P.; Mäki, M.; Kurppa, K. Nonbiopsy approach for celiac disease is accurate when using exact duodenal histomorphometry: Prospective study in 2 countries. J. Clin. Gastroenterol. 2020. [Google Scholar] [CrossRef]
- Holmes, G.; Forsyth, J.; Knowles, S.; Seddon, H.; Hill, P.G.; Austin, A.S. Coeliac disease: Further evidence that biopsy is not always necessary for diagnosis. Eur. J. Gastroenterol. Hepatol. 2017, 29, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Zanini, B.; Magni, A.; Caselani, F.; Lanzarotto, F.; Carabellese, N.; Villanacci, V.; Ricci, C.; Lanzini, A. High tissue-transglutaminase antibody level predicts small intestinal villous atrophy in adult patients at high risk of celiac disease. Dig. Liver Dis. 2012, 44, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Beltran, L.; Koenig, M.; Egner, W.; Howard, M.; Butt, A.; Austin, M.R.; Patel, D.; Sanderson, R.R.; Goubet, S.; Saleh, F.; et al. High-titre circulating tissue transglutaminase-2 antibodies predict small bowel villous atrophy, but decision cut-off limits must be locally validated. Clin. Exp. Immunol. 2014, 176, 190–198. [Google Scholar] [CrossRef] [PubMed]
- Rozenberg, O.; Rinawi, F.; Haritan, Y.; Yerushalmi, B.; Kori, M.; Morgenstern, S.; Peleg, S.; Osyntsov, L.; Colodner, R.; Shamir, R. Automated analyzers are suited for diagnosing celiac disease without a biopsy. J. Pediatr. Gastroenterol. Nutr. 2020, 71, 64–70. [Google Scholar] [CrossRef]
- Trovato, C.M.; Montuori, M.; Anania, C.; Barbato, M.; Vestri, A.R.; Guida, S.; Oliva, S.; Mainiero, F.; Cucchiara, S.; Valitutti, F. Are ESPGHAN “Biopsy-Sparing” guidelines for celiac disease also suitable for asymptomatic patients? Am. J. Gastroenterol. 2015, 110, 1485–1489. [Google Scholar] [CrossRef] [PubMed]
- Elitsur, Y.; Sigman, T.; Watkins, R.; Porto, A.F.; Puppa, E.L.L.; Foglio, E.J.; Preston, D.L. Tissue transglutaminase levels are not sufficient to diagnose celiac disease in North American practices without intestinal biopsies. Dig. Dis. Sci. 2017, 62, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Zanini, B.; Caselani, F.; Magni, A.; Turini, D.; Ferraresi, A.; Lanzarotto, F.; Villanacci, V.; Carabellese, N.; Ricci, C.; Lanzini, A. Celiac disease with mild enteropathy is not mild disease. Clin. Gastroenterol. Hepatol. 2013, 11, 253–258. [Google Scholar] [CrossRef]
- Fernandez-Bañares, F.; Alsina, M.; Modolell, I.; Andújar, X.; Piqueras, M.; García-Puig, R.; Martín, B.; Rosinach, M.; Salas, A.; Viver, J.M.; et al. Are positive serum-IgA-tissue-transglutaminase antibodies enough to diagnose coeliac disease without a small bowel biopsy? Post-test probability of coeliac disease. J. Crohns Colitis 2012, 6, 861–866. [Google Scholar] [CrossRef][Green Version]
- Adelman, D.C.; Murray, J.A.; Wu, T.-T.; Mäki, M.; Green, P.H.; Kelly, C.P. Measuring change in small intestinal histology in patients with celiac disease. Am. J. Gastroenterol. 2018, 113, 339–347. [Google Scholar] [CrossRef]
- Agardh, D.; Lee, H.-S.; Kurppa, K.; Simell, V.; Aronsson, C.A.; Jörneus, O.; Hummel, M.; Liu, E.; Koletzko, S. Clinical features of celiac disease: A prospective birth cohort. Pediatrics 2015, 135, 627–634. [Google Scholar] [CrossRef]
- Kivelä, L.; Kaukinen, K.; Huhtala, H.; Lähdeaho, M.-L.; Mäki, M.; Kurppa, K. At-risk screened children with celiac disease are comparable in disease severity and dietary adherence to those found because of clinical suspicion: A large cohort study. J. Pediatr. 2017, 183, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Taavela, J.; Kurppa, K.; Collin, P.; Lähdeaho, M.; Salmi, T.; Saavalainen, P.; Haimila, K.; Huhtala, H.; Laurila, K.; Sievänen, H.; et al. Degree of damage to the small bowel and serum antibody titers correlate with clinical presentation of patients with celiac disease. Clin. Gastroenterol. Hepatol. 2013, 11, 166–171. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.P.; Sandhu, B.; Spray, C.H.; Basude, D.; Ramani, P. Evidence supporting serology-based pathway for diagnosing celiac disease in asymptomatic children from high-risk groups. J. Pediatr. Gastroenterol. Nutr. 2018, 66, 641–644. [Google Scholar] [CrossRef] [PubMed]
- Gustafsson, I.; Repo, M.; Popp, A.; Kaukinen, K.; Hiltunen, P.; Arvola, T.; Taavela, J.; Vornanen, M.; Kivelä, L.; Kurppa, K. Prevalence and diagnostic outcomes of children with duodenal lesions and negative celiac serology. Dig. Liver Dis. 2020, 52, 289–295. [Google Scholar] [CrossRef]
- Egner, W.; Shrimpton, A.; Sargur, R.; Patel, D.; Swallow, K. ESPGHAN guidance on coeliac disease 2012: Multiples of ULN for decision making do not harmonise assay performance across centres. J. Pediatr. Gastroenterol. Nutr. 2012, 55, 733–735. [Google Scholar] [CrossRef]
- Celiac Disease. Current Care Guidelines. Suomalainen Lääkäriseura Duodecim: Helsinki, 2018. Available online: www.kaypahoito.fi (accessed on 2 June 2020).
| Clinical Cohort | Family Cohort | |||
|---|---|---|---|---|
| All Subjects n = 239 (%) | Celiac Disease n = 137 (%) | All Subjects n = 597 (%) | Celiac Disease n = 85 (%) | |
| Baseline Data | ||||
| Age, median (range) | 45 (17–83) | 45 (18–74) | 48 (18–96) | 44 (18–80) |
| Females | 72.8 | 79.6 | 53.4 | 50.6 |
| Affected relative | 21.1 | 25.8 | 100 | 100 |
| HLA DQ2/DQ8 | 82.8 | 100 | 74.4 | 100 |
| TGA positivity | ||||
| Celikey | 48.5 | 84.7 | 15.4 | 76.5 |
| Orgentec | 51.8 | 90.8 | 18.6 | 88.8 |
| Eurospital | 55.5 | 91.6 | 23.8 | 96.3 |
| Inova | 62.3 | 96.6 | 43.0 | 98.8 |
| EmA positivity | 51.5 | 89.8 | 19.3 | 98.8 |
| ≥10× ULN a | ≥1× ULN a | |||||||
|---|---|---|---|---|---|---|---|---|
| Positive Subjects (n) | Celiac Disease (n) | PPV (%) | 95% CI (%) | Positive Subjects (n) | Celiac Disease (n) | PPV (%) | 95% CI (%) | |
| Clinical cohort | ||||||||
| Celikey | 56 | 56 | 100 | 92.0–100 | 116 | 116 | 100 | 96.0–100 |
| Orgentec | 36 | 36 | 100 | 88.0–100 | 113 | 108 | 95.6 | 89.5–98.4 |
| Eurospital | 51 | 51 | 100 | 91.3–100 | 121 | 109 | 90.1 | 83.0–94.5 |
| Inova | 54 | 54 | 100 | 91.7–100 | 134 | 112 | 83.6 | 76.0–89.2 |
| Family cohort | ||||||||
| Celikey | 18 | 18 | 100 | 78.1–100 | 66 | 65 | 98.5 | 90.7–99.9 |
| Orgentec | 26 | 26 | 100 | 84.0–100 | 78 | 72 | 92.3 | 83.4–96.8 |
| Eurospital | 33 | 33 | 100 | 87.0–100 | 84 | 78 | 92.9 | 84.5–97.1 |
| Inova | 21 | 21 | 100 | 80.8–100 | 93 | 84 | 90.3 | 82.0–95.2 |
| Clinical Cohort | Family Cohort | |||
|---|---|---|---|---|
| Value, U/mL | ×ULN a | Value, U/mL | ×ULN a | |
| Celikey | 4.8 | 1.0 | 6.6 | 1.3 |
| Orgentec | 32 | 3.2 | 24 | 2.4 |
| Eurospital | 38 | 3.8 | 38 | 3.8 |
| Inova | 102 | 5.1 | 98 | 4.9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ylönen, V.; Lindfors, K.; Repo, M.; Huhtala, H.; Fuchs, V.; Saavalainen, P.; Musikka, A.; Laurila, K.; Kaukinen, K.; Kurppa, K. Non-Biopsy Serology-Based Diagnosis of Celiac Disease in Adults Is Accurate with Different Commercial Kits and Pre-Test Probabilities. Nutrients 2020, 12, 2736. https://doi.org/10.3390/nu12092736
Ylönen V, Lindfors K, Repo M, Huhtala H, Fuchs V, Saavalainen P, Musikka A, Laurila K, Kaukinen K, Kurppa K. Non-Biopsy Serology-Based Diagnosis of Celiac Disease in Adults Is Accurate with Different Commercial Kits and Pre-Test Probabilities. Nutrients. 2020; 12(9):2736. https://doi.org/10.3390/nu12092736
Chicago/Turabian StyleYlönen, Venla, Katri Lindfors, Marleena Repo, Heini Huhtala, Valma Fuchs, Päivi Saavalainen, Alex Musikka, Kaija Laurila, Katri Kaukinen, and Kalle Kurppa. 2020. "Non-Biopsy Serology-Based Diagnosis of Celiac Disease in Adults Is Accurate with Different Commercial Kits and Pre-Test Probabilities" Nutrients 12, no. 9: 2736. https://doi.org/10.3390/nu12092736
APA StyleYlönen, V., Lindfors, K., Repo, M., Huhtala, H., Fuchs, V., Saavalainen, P., Musikka, A., Laurila, K., Kaukinen, K., & Kurppa, K. (2020). Non-Biopsy Serology-Based Diagnosis of Celiac Disease in Adults Is Accurate with Different Commercial Kits and Pre-Test Probabilities. Nutrients, 12(9), 2736. https://doi.org/10.3390/nu12092736





