Curcumin Ameliorates Nonalcoholic Fatty Liver Disease through Inhibition of O-GlcNAcylation
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Immunoblot Analysis
2.3. Immunohistochemistry (IHC) and Oil Red O Staining
2.4. Quantitative Reverse Transcription–Polymerase Chain Reaction
2.5. Cell Culture and Transfection
2.6. The ROS Production Assay
2.7. Immunoprecipitation (IP)
2.8. Statistical Analysis
3. Results
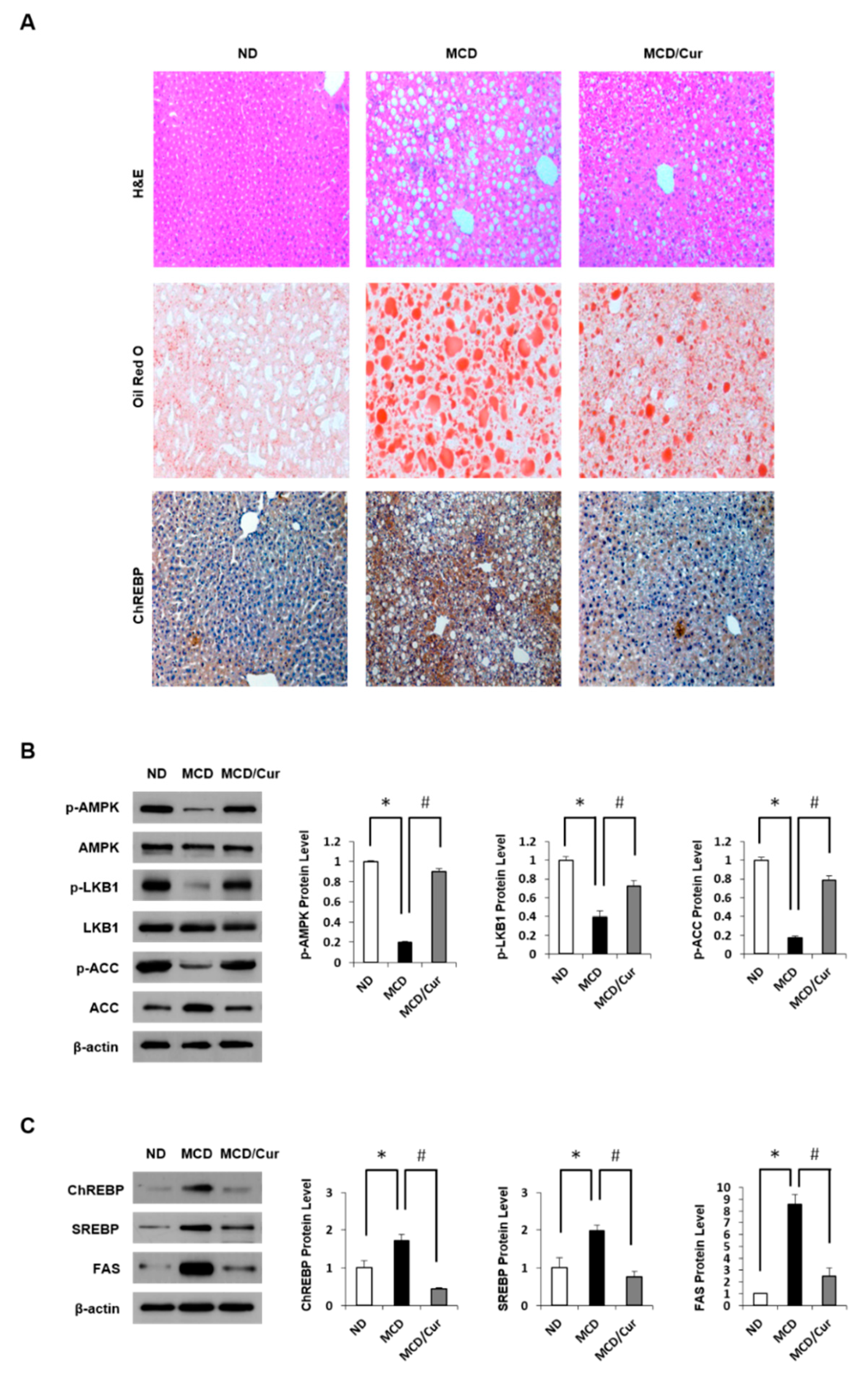
3.1. Effect of Curcumin on Hepatic Steatosis in MCD Diet Mice
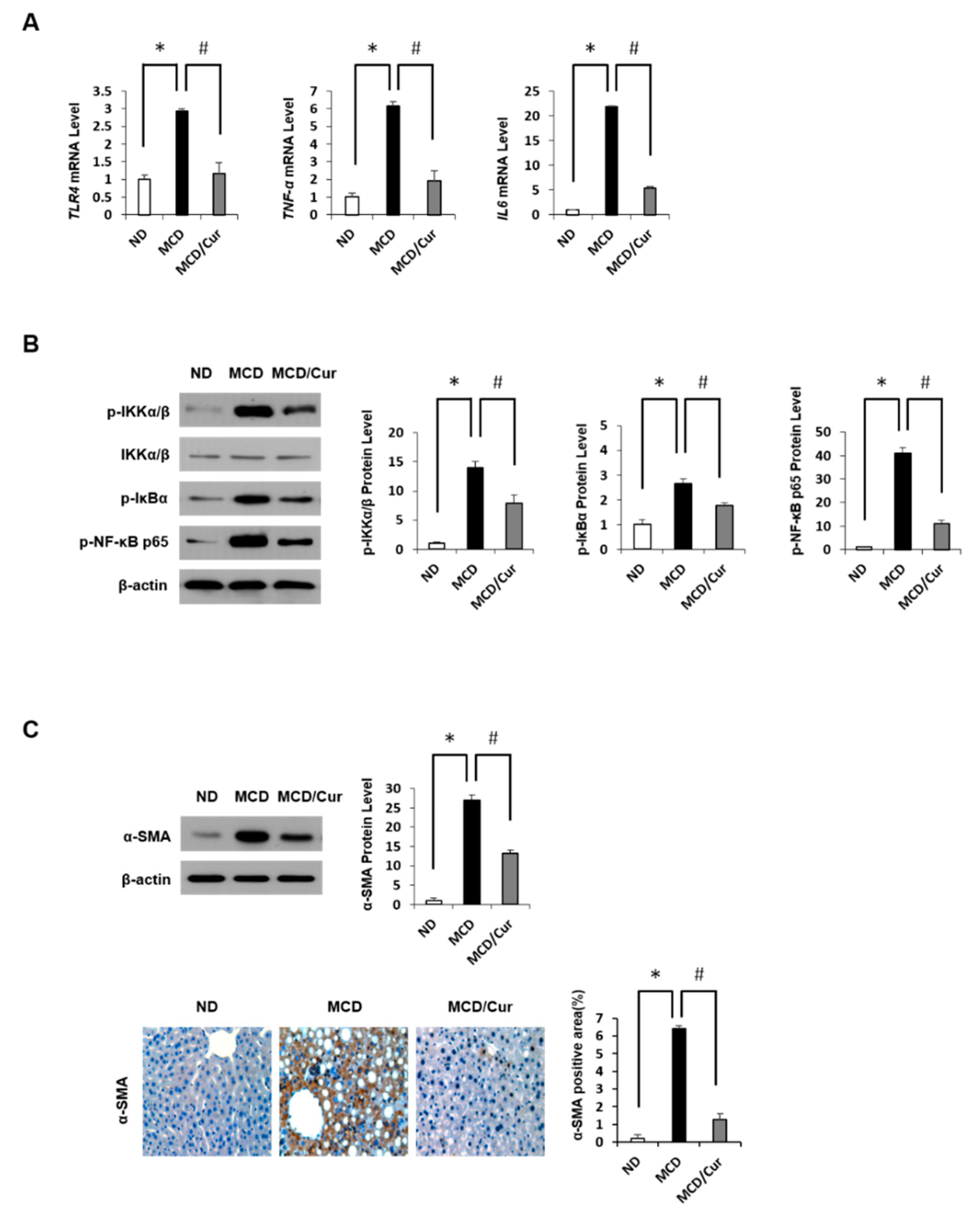
3.2. Curcumin Ameliorates Inflammation Induced in MCD Diet Mice
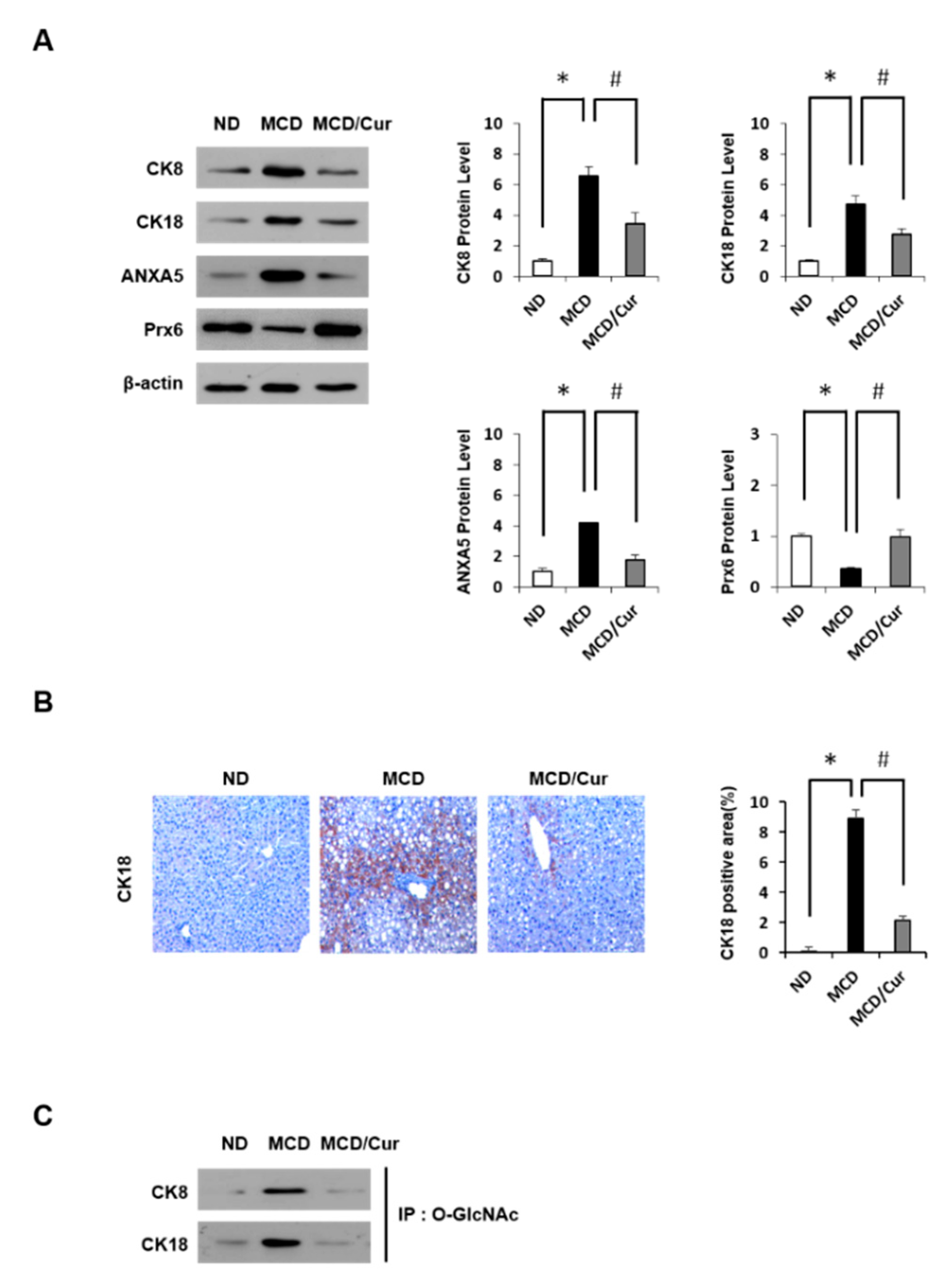
3.3. DE Analysis of Differentially Expressed Proteins
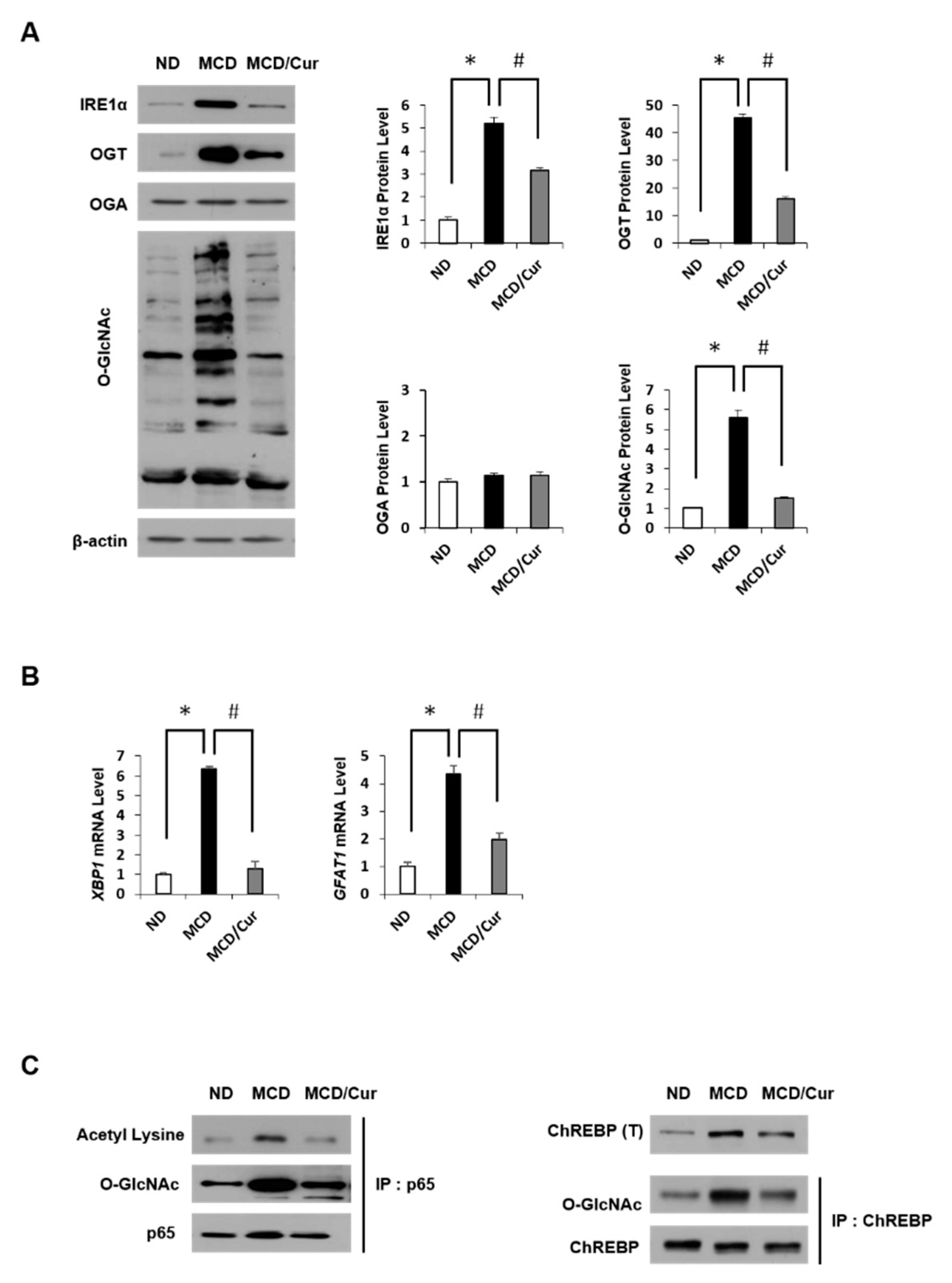
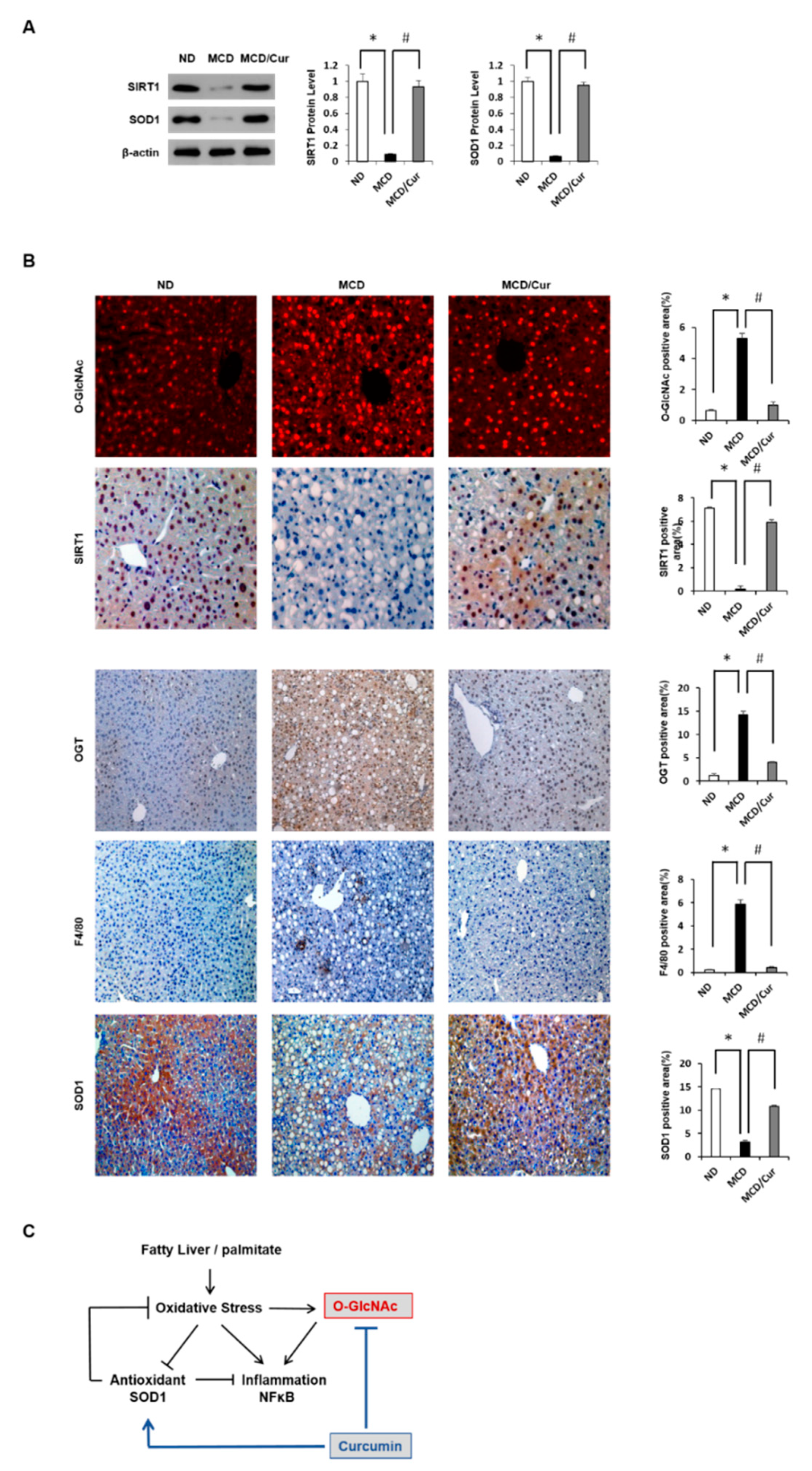
3.4. Curcumin Suppresses O-GlcNAcylation in Steatohepatitis
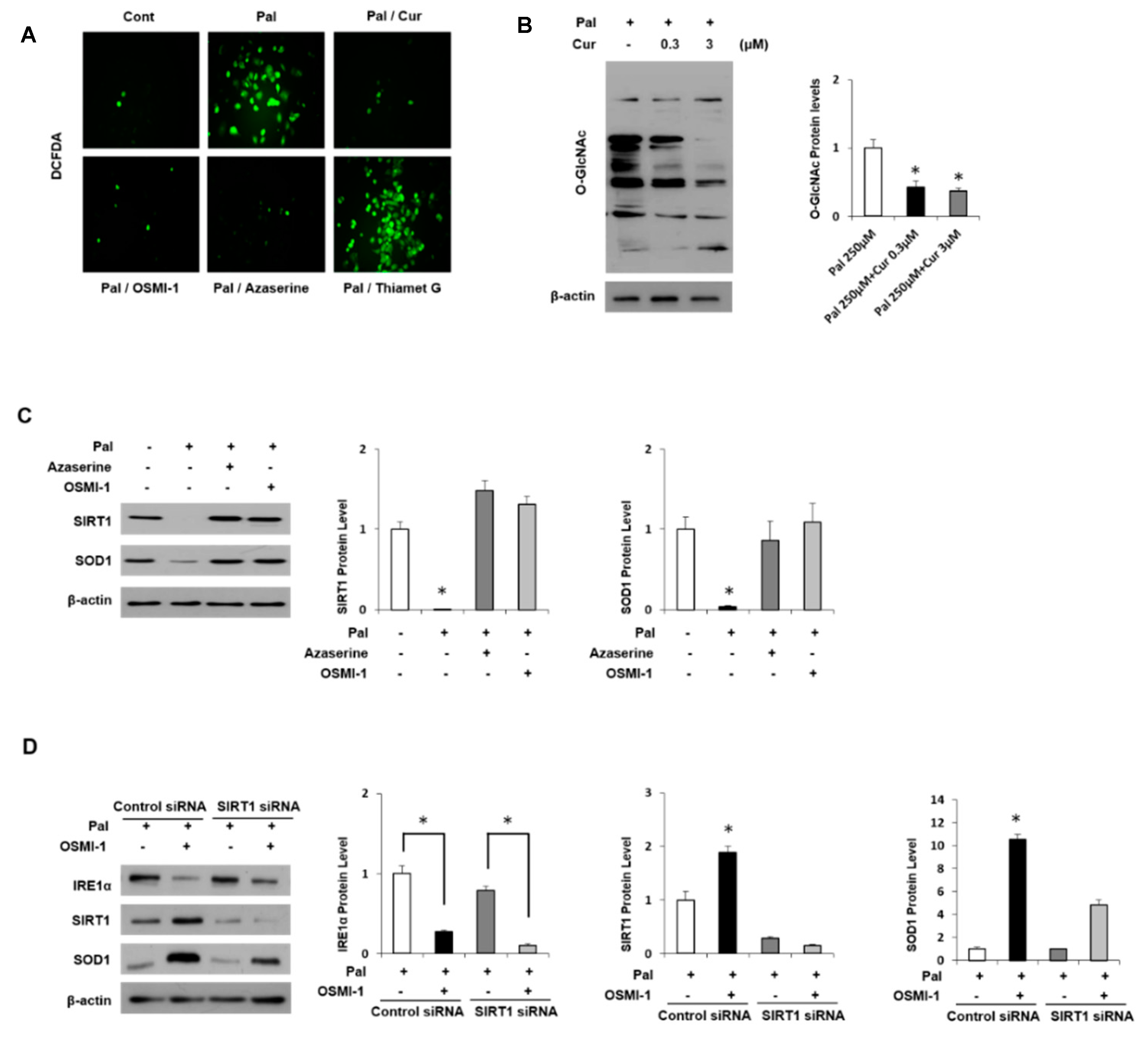
3.5. Regulation of Antioxidant Proteins in Palmitate-Induced AML12 Cells
3.6. Curcumin Ameliorates Inflammation by Inhibiting of O-GlcNAcylation and Activates Antioxidant Protein
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Perla, F.M.; Prelati, M.; Lavorato, M.; Visicchio, D.; Anania, C. The Role of Lipid and Lipoprotein Metabolism in Non-Alcoholic Fatty Liver Disease. Children 2017, 4, 46. [Google Scholar] [CrossRef]
- Mencin, A.A.; Lavine, J.E. Advances in pediatric nonalcoholic fatty liver disease. Pediatr. Clin. N. Am. 2011, 58, 1375–1392. [Google Scholar] [CrossRef] [PubMed]
- Jump, D.B.; Tripathy, S.; Depner, C.M. Fatty acid-regulated transcription factors in the liver. Annu. Rev. Nutr. 2013, 33, 249–269. [Google Scholar] [CrossRef] [PubMed]
- Cho, M.C.; Lee, K.; Paik, S.G.; Yoon, D.Y. Peroxisome Proliferators-Activated Receptor (PPAR) Modulators and Metabolic Disorders. PPAR Res. 2008, 2008, 679137. [Google Scholar] [CrossRef]
- Angulo, P. Nonalcoholic fatty liver disease. N. Engl. J. 2002, 346, 1221–1231. [Google Scholar] [CrossRef] [PubMed]
- Angelico, F.; Del Ben, M.; Conti, R.; Francioso, S.; Feole, K.; Fiorello, S.; Cavallo, M.G.; Zalunardo, B.; Lirussi, F.; Alessandri, C.; et al. Insulin resistance, the metabolic syndrome, and nonalcoholic fatty liver disease. J. Clin. Endocrinol. Metab. 2005, 90, 1578–1582. [Google Scholar] [CrossRef]
- Sanchez-Valle, V.; Chavez-Tapia, N.C.; Uribe, M.; Mendez-Sanchez, N. Role of oxidative stress and molecular changes in liver fibrosis: A review. Curr. Med. 2012, 19, 4850–4860. [Google Scholar] [CrossRef]
- Sakaguchi, S.; Takahashi, S.; Sasaki, T.; Kumagai, T.; Nagata, K. Progression of alcoholic and non-alcoholic steatohepatitis: Common metabolic aspects of innate immune system and oxidative stress. Drug Metab. Pharmacokinet. 2011, 26, 30–46. [Google Scholar] [CrossRef]
- Fridovich, I. Superoxide radical and superoxide dismutases. Annu. Rev. Biochem. 1995, 64, 97–112. [Google Scholar] [CrossRef]
- Wang, Y.; Branicky, R.; Noe, A.; Hekimi, S. Superoxide dismutases: Dual roles in controlling ROS damage and regulating ROS signaling. J. Cell Biol. 2018, 217, 1915–1928. [Google Scholar] [CrossRef]
- Esworthy, R.S.; Ho, Y.S.; Chu, F.F. The Gpx1 gene encodes mitochondrial glutathione peroxidase in the mouse liver. Arch. Biochem. Biophys. 1997, 340, 59–63. [Google Scholar] [CrossRef] [PubMed]
- Finkel, T.; Deng, C.X.; Mostoslavsky, R. Recent progress in the biology and physiology of sirtuins. Nature 2009, 460, 587–591. [Google Scholar] [CrossRef] [PubMed]
- Domitrovic, R.; Potocnjak, I. A comprehensive overview of hepatoprotective natural compounds: Mechanism of action and clinical perspectives. Arch. Toxicol. 2016, 90, 39–79. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Takikawa, Y.; Satoh, T.; Yoshioka, Y.; Kosaka, K.; Tatemichi, Y.; Suzuki, K. Carnosic acid prevents obesity and hepatic steatosis in ob/ob mice. Hepatol. Res. 2011, 41, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Ron, D.; Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 2007, 8, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Ni, M.; Zhang, Y.; Lee, A.S. Beyond the endoplasmic reticulum: Atypical GRP78 in cell viability, signalling and therapeutic targeting. Biochem. J. 2011, 434, 181–188. [Google Scholar] [CrossRef]
- Malhi, H.; Kaufman, R.J. Endoplasmic reticulum stress in liver disease. J. Hepatol. 2011, 54, 795–809. [Google Scholar] [CrossRef]
- Lake, A.D.; Novak, P.; Hardwick, R.N.; Flores-Keown, B.; Zhao, F.; Klimecki, W.T.; Cherrington, N.J. The adaptive endoplasmic reticulum stress response to lipotoxicity in progressive human nonalcoholic fatty liver disease. Toxicol. Sci. 2014, 137, 26–35. [Google Scholar] [CrossRef]
- Kim, I.; Xu, W.; Reed, J.C. Cell death and endoplasmic reticulum stress: Disease relevance and therapeutic opportunities. Nat. Rev. Drug Discov. 2008, 7, 1013–1030. [Google Scholar] [CrossRef]
- Xiong, Q.; Hase, K.; Tezuka, Y.; Namba, T.; Kadota, S. Acteoside inhibits apoptosis in D-galactosamine and lipopolysaccharide-induced liver injury. Life Sci. 1999, 65, 421–430. [Google Scholar] [CrossRef]
- Friedman, S.L. Liver fibrosis—From bench to bedside. J. Hepatol. 2003, 38, S38–S53. [Google Scholar] [CrossRef]
- Elsharkawy, A.M.; Mann, D.A. Nuclear factor-kappaB and the hepatic inflammation-fibrosis-cancer axis. Hepatology 2007, 46, 590–597. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.H.; Park, S.Y.; Nam, H.W.; Kim, D.H.; Kang, J.G.; Kang, E.S.; Kim, Y.S.; Lee, H.C.; Kim, K.S.; Cho, J.W. NFkappaB activation is associated with its O-GlcNAcylation state under hyperglycemic conditions. Proc. Natl. Acad. Sci. USA 2008, 105, 17345–17350. [Google Scholar] [CrossRef] [PubMed]
- Hart, G.W. Dynamic O-linked glycosylation of nuclear and cytoskeletal proteins. Annu. Rev. Biochem. 1997, 66, 315–335. [Google Scholar] [CrossRef]
- Hart, G.W.; Housley, M.P.; Slawson, C. Cycling of O-linked beta-N-acetylglucosamine on nucleocytoplasmic proteins. Nature 2007, 446, 1017–1022. [Google Scholar] [CrossRef]
- Liu, Y.; Cao, Y.; Pan, X.; Shi, M.; Wu, Q.; Huang, T.; Jiang, H.; Li, W.; Zhang, J. O-GlcNAc elevation through activation of the hexosamine biosynthetic pathway enhances cancer cell chemoresistance. Cell Death Dis. 2018, 9, 485. [Google Scholar] [CrossRef]
- Cairns, R.A.; Harris, I.S.; Mak, T.W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 2011, 11, 85–95. [Google Scholar] [CrossRef]
- Slawson, C.; Copeland, R.J.; Hart, G.W. O-GlcNAc signaling: A metabolic link between diabetes and cancer? Trends Biochem. Sci. 2010, 35, 547–555. [Google Scholar] [CrossRef]
- Hawkins, M.; Barzilai, N.; Liu, R.; Hu, M.; Chen, W.; Rossetti, L. Role of the glucosamine pathway in fat-induced insulin resistance. J. Clin. Investig. 1997, 99, 2173–2182. [Google Scholar] [CrossRef]
- Buse, M.G. Hexosamines, insulin resistance, and the complications of diabetes: Current status. Am. J. Physiol. Endocrinol. Metab. 2006, 290, E1–E8. [Google Scholar] [CrossRef]
- Li, H.L.; Liu, C.; de Couto, G.; Ouzounian, M.; Sun, M.; Wang, A.B.; Huang, Y.; He, C.W.; Shi, Y.; Chen, X.; et al. Curcumin prevents and reverses murine cardiac hypertrophy. J. Clin. Investig. 2008, 118, 879–893. [Google Scholar] [CrossRef]
- Yarru, L.P.; Settivari, R.S.; Gowda, N.K.; Antoniou, E.; Ledoux, D.R.; Rottinghaus, G.E. Effects of turmeric (Curcuma longa) on the expression of hepatic genes associated with biotransformation, antioxidant, and immune systems in broiler chicks fed aflatoxin. Poult. Sci. 2009, 88, 2620–2627. [Google Scholar] [CrossRef]
- Schneider, C.; Gordon, O.N.; Edwards, R.L.; Luis, P.B. Degradation of Curcumin: From Mechanism to Biological Implications. J. Agric. Food Chem. 2015, 63, 7606–7614. [Google Scholar] [CrossRef]
- Weisberg, S.P.; Leibel, R.; Tortoriello, D.V. Dietary curcumin significantly improves obesity-associated inflammation and diabetes in mouse models of diabesity. Endocrinology 2008, 149, 3549–3558. [Google Scholar] [CrossRef]
- Shakibaei, M.; Schulze-Tanzil, G.; John, T.; Mobasheri, A. Curcumin protects human chondrocytes from IL-l1beta-induced inhibition of collagen type II and beta1-integrin expression and activation of caspase-3: An immunomorphological study. Ann. Anat. 2005, 187, 487–497. [Google Scholar] [CrossRef]
- Rinella, M.E.; Green, R.M. The methionine-choline deficient dietary model of steatohepatitis does not exhibit insulin resistance. J. Hepatol. 2004, 40, 47–51. [Google Scholar] [CrossRef]
- Sahai, A.; Malladi, P.; Melin-Aldana, H.; Green, R.M.; Whitington, P.F. Upregulation of osteopontin expression is involved in the development of nonalcoholic steatohepatitis in a dietary murine model. Am. J. Physiol. Gastrointest. Liver Physiol. 2004, 287, G264–G273. [Google Scholar] [CrossRef]
- Liu, Z.; Cui, C.; Xu, P.; Dang, R.; Cai, H.; Liao, D.; Yang, M.; Feng, Q.; Yan, X.; Jiang, P. Curcumin Activates AMPK Pathway and Regulates Lipid Metabolism in Rats Following Prolonged Clozapine Exposure. Front. Neurosci. 2017, 11, 558. [Google Scholar] [CrossRef]
- Johnson, A.R.; Milner, J.J.; Makowski, L. The inflammation highway: Metabolism accelerates inflammatory traffic in obesity. Immunol. Rev. 2012, 249, 218–238. [Google Scholar] [CrossRef]
- Han, C.; Gu, Y.; Shan, H.; Mi, W.; Sun, J.; Shi, M.; Zhang, X.; Lu, X.; Han, F.; Gong, Q.; et al. O-GlcNAcylation of SIRT1 enhances its deacetylase activity and promotes cytoprotection under stress. Nat. Commun. 2017, 8, 1491. [Google Scholar] [CrossRef]
- Ferrer, C.M.; Lu, T.Y.; Bacigalupa, Z.A.; Katsetos, C.D.; Sinclair, D.A.; Reginato, M.J. O-GlcNAcylation regulates breast cancer metastasis via SIRT1 modulation of FOXM1 pathway. Oncogene 2017, 36, 559–569. [Google Scholar] [CrossRef]
- Ngoh, G.A.; Hamid, T.; Prabhu, S.D.; Jones, S.P. O-GlcNAc signaling attenuates ER stress-induced cardiomyocyte death. Am. J. Physiol. Heart Circ. Physiol. 2009, 297, H1711–H1719. [Google Scholar] [CrossRef]
- Zhang, K.; Yin, R.; Yang, X. O-GlcNAc: A Bittersweet Switch in Liver. Front. Endocrinol. 2014, 5, 221. [Google Scholar] [CrossRef]
- Hardiville, S.; Hart, G.W. Nutrient regulation of signaling, transcription, and cell physiology by O-GlcNAcylation. Cell Metab. 2014, 20, 208–213. [Google Scholar] [CrossRef]
- Farzaei, M.H.; Zobeiri, M.; Parvizi, F.; El-Senduny, F.F.; Marmouzi, I.; Coy-Barrera, E.; Naseri, R.; Nabavi, S.M.; Rahimi, R.; Abdollahi, M. Curcumin in Liver Diseases: A Systematic Review of the Cellular Mechanisms of Oxidative Stress and Clinical Perspective. Nutrients 2018, 10, 855. [Google Scholar] [CrossRef]
- Guinez, C.; Filhoulaud, G.; Rayah-Benhamed, F.; Marmier, S.; Dubuquoy, C.; Dentin, R.; Moldes, M.; Burnol, A.F.; Yang, X.; Lefebvre, T.; et al. O-GlcNAcylation increases ChREBP protein content and transcriptional activity in the liver. Diabetes 2011, 60, 1399–1413. [Google Scholar] [CrossRef]
- Baker, R.G.; Hayden, M.S.; Ghosh, S. NF-kappaB, inflammation, and metabolic disease. Cell Metab. 2011, 13, 11–22. [Google Scholar] [CrossRef]
- Baudoin, L.; Issad, T. O-GlcNAcylation and Inflammation: A Vast Territory to Explore. Front. Endocrinol. 2014, 5, 235. [Google Scholar] [CrossRef]
- Yang, Y.; Duan, W.; Lin, Y.; Yi, W.; Liang, Z.; Yan, J.; Wang, N.; Deng, C.; Zhang, S.; Li, Y.; et al. SIRT1 activation by curcumin pretreatment attenuates mitochondrial oxidative damage induced by myocardial ischemia reperfusion injury. Free Radic. Biol. Med. 2013, 65, 667–679. [Google Scholar] [CrossRef]
- de Freitas Carvalho, M.M.; Lage, N.N.; de Souza Paulino, A.H.; Pereira, R.R.; de Almeida, L.T.; da Silva, T.F.; de Brito Magalhaes, C.L.; de Lima, W.G.; Silva, M.E.; Pedrosa, M.L.; et al. Effects of acai on oxidative stress, ER stress, and inflammation-related parameters in mice with high fat diet-fed induced NAFLD. Sci. Rep. 2019, 9, 8107. [Google Scholar] [CrossRef]
- Ahmad, M.; Taweel, G.M.A.; Hidayathulla, S. Nano-composites chitosan-curcumin synergistically inhibits the oxidative stress induced by toxic metal cadmium. Int. J. Biol. Macromol. 2018, 108, 591–597. [Google Scholar] [CrossRef]
- Mota, M.; Banini, B.A.; Cazanave, S.C.; Sanyal, A.J. Molecular mechanisms of lipotoxicity and glucotoxicity in nonalcoholic fatty liver disease. Metabolism 2016, 65, 1049–1061. [Google Scholar] [CrossRef]
| No a | Identified Protein | Accession No. | MW (KDa) | PI | MOWSE Score | Coverage (%) | Fold Change | |
|---|---|---|---|---|---|---|---|---|
| MCD | MCD + Cur | |||||||
| Metabolism | ||||||||
| 1 | 14-3-3 protein zeta/delta | P63101 | 27.8 | 4.7 | 30,049 | 44.9 | 4 | 3.2 |
| 2 | Annexin A5 | P48036 | 35.8 | 4.8 | 4.29 × 108 | 47.3 | 6 | 3.5 |
| 3 | 2’-5’-oligoadenylate synthase-like protein 1 | Q8VI94 | 59.1 | 6.7 | 139,169 | 22.9 | 3 | 1 |
| Transport | ||||||||
| 4 | 40S ribosomal protein SA * | P14206 | 32.8 | 4.8 | 21,692 | 40.3 | 3 | 1 |
| 5 | Integrin beta-like protein 1 | Q8VDV0 | 54.0 | 5.3 | 14,545 | 20.0 | 2 | 0.8 |
| Oxidative Stress/Detoxification processes | ||||||||
| 6/7 | AOP2(Peroxiredoxin-6) * | O08709 | 26.3 | 6.2 | 2350 | 18.5 | 0.4 | 1 |
| 8 | Glutathione peroxidase 1 (GPx 1) * | P11352 | 22.3 | 6.7 | 141,506 | 42.3 | 0.6 | 1.3 |
| 9 | SODC(Superoxide dismutase [Cu-Zn]) * | P08228 | 15.9 | 6.0 | 1996 | 41.6 | 0.3 | 0.8 |
| 10 | 60 kDa heat shock protein, mitochondrial * | P63038 | 61.0 | 5.9 | 1.60 × 106 | 43.8 | 2 | 1 |
| Structural Proteins | ||||||||
| 11/12 | Keratin, type I cytoskeletal 18 * | P05784 | 47.5 | 5.2 | 3.72 × 107 | 36.6 | 5 | 1.5 |
| 13/14/15 | Keratin, type II cytoskeletal 8 * | P11679 | 54.6 | 5.7 | 1.25 ×107 | 42.0 | 3 | 1 |
| 16 | Actin-related protein T1 | Q9D9J3 | 42.2 | 5.2 | 701 | 18.6 | 5 | 3 |
| 17 | Rho GDP-dissociation inhibitor 1 * | Q99PT1 | 23.4 | 5.1 | 2545 | 41.7 | 4 | 2.5 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, D.E.; Lee, S.J.; Kim, S.J.; Lee, H.-S.; Kwon, O.-S. Curcumin Ameliorates Nonalcoholic Fatty Liver Disease through Inhibition of O-GlcNAcylation. Nutrients 2019, 11, 2702. https://doi.org/10.3390/nu11112702
Lee DE, Lee SJ, Kim SJ, Lee H-S, Kwon O-S. Curcumin Ameliorates Nonalcoholic Fatty Liver Disease through Inhibition of O-GlcNAcylation. Nutrients. 2019; 11(11):2702. https://doi.org/10.3390/nu11112702
Chicago/Turabian StyleLee, Da Eun, Su Jin Lee, Su Ji Kim, Hyun-Shik Lee, and Oh-Shin Kwon. 2019. "Curcumin Ameliorates Nonalcoholic Fatty Liver Disease through Inhibition of O-GlcNAcylation" Nutrients 11, no. 11: 2702. https://doi.org/10.3390/nu11112702
APA StyleLee, D. E., Lee, S. J., Kim, S. J., Lee, H.-S., & Kwon, O.-S. (2019). Curcumin Ameliorates Nonalcoholic Fatty Liver Disease through Inhibition of O-GlcNAcylation. Nutrients, 11(11), 2702. https://doi.org/10.3390/nu11112702





