1. Introduction
Knowledge of tree species plays an important role in forest management and planning. The optimum output, requested by forest companies from the forest mapping process, is the species-specific size distribution of the trees. The traditional method, based on field inventory work for tree species identification, is labor intensive, time consuming, and limited by spatial extent. Therefore, remote sensing techniques were introduced, such as the interpretation of large-scale aerial color or infra-red images [
1,
2]. Although remotely-sensed data have been widely used for forest applications, traditional optical remote sensing techniques suffer from a lack of the ability to capture three-dimensional forest structures, particularly in unevenly-aged, mixed species forests with multiple canopy layers [
3]. Recent developments in active remote sensing, particularly laser scanning techniques, have shown potential in forest mapping and other applications because of the capability to capture three-dimensional (3D) information of forests [
4,
5,
6,
7,
8,
9,
10,
11].
Airborne laser scanning (ALS) is a useful tool for retrieving biophysical variables and for updating forest inventory maps. The successful use of ALS data has been demonstrated for a variety of applications. For example, ALS has been used to estimate tree height [
6,
7], identify tree species [
8,
9,
10], and estimate tree volume, biomass [
11,
12,
13], and growth [
14,
15]. Tree species information at an individual tree level is particularly useful in growth and yield estimates, and has been primarily studied for forest applications, such as updating forest inventories. Tree species classification using ALS has not been intensively studied, when compared with studies on the successful use of ALS for other forest attribute mapping, because of the lack of spectral information. Brandtberg [
9] classified three leaf-off individual deciduous tree species (oaks, red maple, and yellow poplar) in West Virginia, USA, using high density laser data, and reported 64% total accuracy. Holmgren and Persson [
8] classified Norway spruce and Scots pine in Remningstorp, Sweden, using ALS-derived point and intensity features, and achieved an accuracy of 95%. Ørka et al. [
16] classified three species (spruce, birch and aspen) at the Ostmarka natural forest in southern Norway. Suratno et al. [
17] classified ponderosa pine, Douglas-fir, western larch, and lodgepole pine, in a western North American montane forest using low density ALS data, and achieved a classification accuracy of 95% at the dominant species level, and 68% for individual trees.
Intensity was also demonstrated to be useful information for tree species identification. Ørka et al. [
18] reported an accuracy of 73% when classifying conifers and deciduous trees, solely based on intensity information. Korpela et al. [
19] classified Scots pine, Norway spruce, and birch, by using intensity variables at Hyytiälä in southern Finland, and showed that intensity features can contribute to a classification accuracy of 88% among the three species. With full-waveform (FWF) lasers, the total received power corresponding to the backscattering cross-section can be calculated, which provides information on the objects, from the intensity waveform.
Previous studies have demonstrated that FWF data and the derived metrics can be used to improve the performance of tree species classification. For example, Yao et al. [
20] demonstrated the usefulness of waveform features for the classification of deciduous and coniferous trees. Heinzel and Koch [
21] analyzed a set of waveform features and identified the most predictive features for classifying up to six tree species. Cao et al. [
22] demonstrated that full-waveform data and derived metrics have significant potential for tree species classification in the subtropical forests, and results demonstrated that all tree species were classified with relatively high accuracy (68.6% for six classes, 75.8% for four main species, and 86.2% for conifers and broadleaved trees).
Previous studies have also revealed that combining multispectral information with 3D ALS data can lead to improvement in the accuracy of tree extraction and tree species classification, as we can take advantage of both datasets. For example, Naidoo et al. [
23] concluded that the use of ALS and hyperspectral data yielded the highest classification accuracy and prediction success for the eight savanna tree species, with an overall classification accuracy of 87.68%. Zhou et al. [
24] demonstrated that the ALS intensity data can contribute to the classification of shaded areas in an urban environment where high resolution digital aerial imagery alone did not produce good results. The fusion of high resolution (satellite or aerial) remote sensing and ALS data can achieve mutual benefits for compensating the lack of 3D structure from imagery and multi-spectral information from ALS data. With respect to the success of these case studies, multi-sensor data fusion seems to be a feasible solution, especially for the mapping of land cover over large areas.
However, there are challenging factors that limit the effective operational use of the fused datasets [
25,
26]. For example, geometric and radiometric registration between two datasets is demanding, because of the fact that data are normally acquired at different times, using different sensors. It is also costly to make measurements with two sensors, particularly in the boreal forest zone where the measurements can seldom be carried out during a single flight, because ALS measurements can be taken two to four times longer than aerial/hyperspectral measurements during a day, since ALS does not depend on sun light illumination. Furthermore, in contrast to passive imagery, laser scanning always views the targets at the zero degree phase angle in a narrow off-nadir viewing geometry and the transmitted energy is also controllable, thus the interpretation of the laser intensity is less complex than in the case of passive airborne images [
27]. The recently developed multispectral laser scanning technique is therefore becoming an attractive option for forest mapping, because it can provide not only a dense point cloud, but also spectral information which can simplify data processing and facilitate the interpretation of data. There are a couple of studies that have demonstrated the potential of multispectral ALS for classifying tree species [
28,
29]. In Lindberg et al [
28], multispectral data were acquired with separate instruments and from different flights—an analogue to Titan multispectral data. The study described the characterization of tree species from ALS data, using three wavelengths: 1064 nm, 1550 nm, and 532 nm, and a point density over 20 point/m
2. However, classification accuracy was not reported. In St-Onge and Budei [
29], values for the mean and standard deviation of the intensity in three channels of Titan multispectral ALS, were used in the classification of broadleave vs. needleleaf trees (level 1), and eight genera (level 2) in a suburb of the city of Toronto, Canada. Random forest classification produced a classification error of 4.59% in the case of the level 1 classification (broadleave vs. needleleaf trees), and of 24.29% in the case of the level 2 classification. The point density of the data used was 10.6 first returns/m
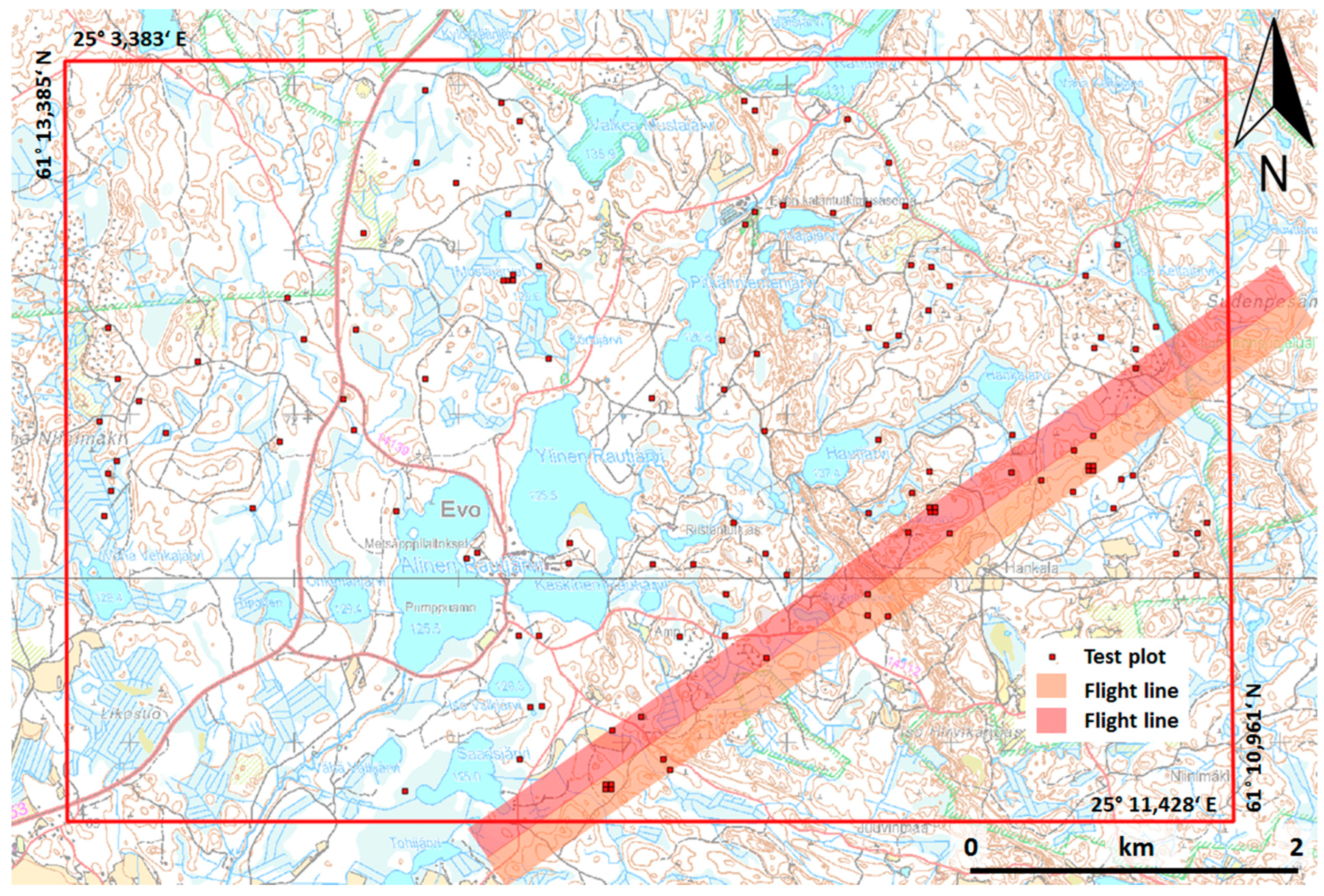
2 per channel. Currently, the cost of data acquisition of multispectral ALS is relatively higher than that of aerial images and ALS data, if they are acquired from the same flight. However, it is expected that this cost will decrease in the future, as the technology advances. Therefore, it is worth investigating the potential of multispectral laser scanning for forest inventories, particularly for tree species classification. The objectives of this study are to evaluate the feasibility of multispectral ALS data for tree species classification with intensive field measurements, and to investigate the information content of features derived from both point cloud and intensity. The study was conducted in a boreal forest using 1903 trees in 22 plots.
5. Discussion
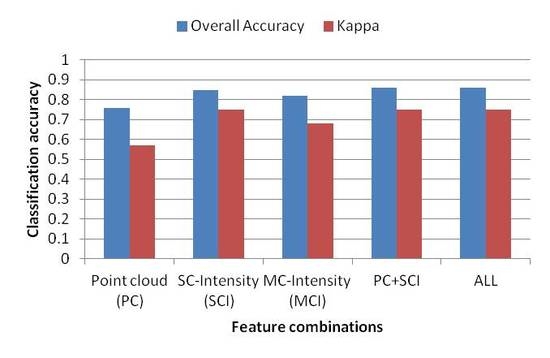
In this study, we explored the potential of multispectral ALS data in tree species classification of a boreal forest. Results showed that multispectral ALS data can be used to separate three main tree species, i.e., pine, spruce, and birch, with a high overall accuracy of 85.9% in the best case scenario, which was based on the combined use of point cloud and SCI features. Overall, the results indicated that the intensity of the three channels contains more information for tree species classification than point cloud data. When using the intensity of the three channels, both the producer’s and user’s accuracies for single tree species were improved, as well as the overall accuracy compared with the results obtained from point cloud data. However, different types of features are more influential on certain tree species. For example, intensity features are more powerful in separating birch from pine and spruce (produce’s accuracy improved from 45.8% to 71%, and user’s accuracy from 64.5% to 80.2%, when compared with those using point cloud features). With the inclusion of point cloud features, the classification accuracy of intensity features was improved by only 0.5 percentage point, while the corresponding value was 10 percentage points when adding intensity features to point cloud features.
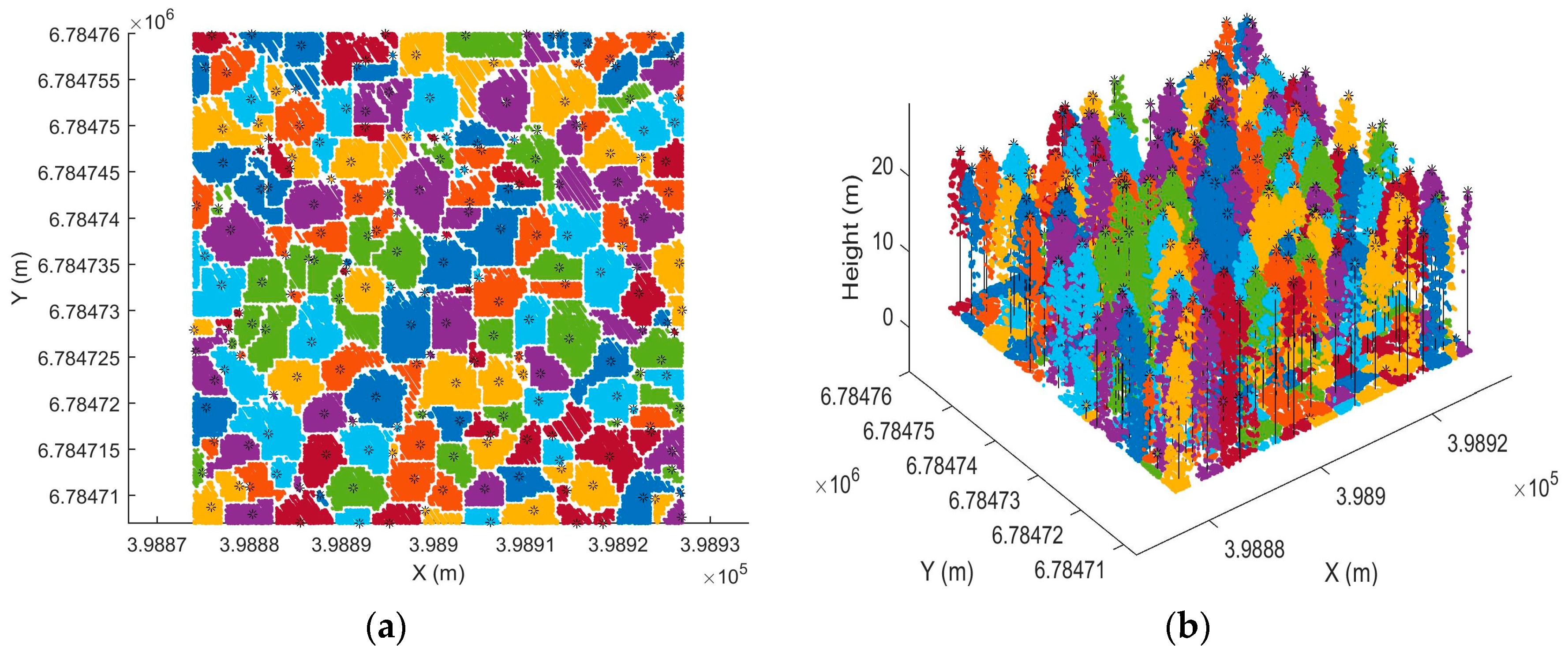
The individual tree-detection rate was not very high in this study. Two factors influenced this. Firstly, individual tree detection was based on CHM, so most of the understory trees were not detectable and 3D information of the dense point cloud was not fully utilized. Secondly, distribution of the point cloud was not optimal, as it was denser in scanning direction than flight direction. The uneven distribution of points affected the results of individual tree detection, as the detection rate tended to decrease when the plot was located near the boundary of data coverage where uneven distribution was more severe. In order to improve individual tree detection, we recommend developing methods which can fully utilize the 3D information provided by point cloud. Multispectral information could also be useful for improving the accuracy of individual tree detection. When point cloud and spectral information are used in tree detection, a simultaneous classification is possible, such that the knowledge relevant to each can aid in the analysis of the other. Ultimately, this could lead to the improvement of accuracy of individual tree detection and classification, as well as computational advantages.
A large variance in feature values can be found, due to the irregular geometry of the canopy surface and varying degrees of penetration. There were more points penetrated, thus reaching the ground, in channels 1 and 3, than in channel 2. One potential factor that contributed to this was the forward viewing geometry for channels 1 and 3. There were also more returns in channels 2 and 3, than in channel 1. For the same point cloud features, the values in channel 3 were higher than those in channels 1 and 2, while channels 1 and 2 produced similar values. This trend was observed for all three species and could be one reason why the point cloud features did not significantly improve the classification, when used with SCI features. SCI features also overlapped between species. However, the degree of overlap varied among the features and channels. In general, higher percentiles of the intensity distribution and minimum intensity value were more separated than the lower percentiles. For example, the maximum intensity was smaller for pine than spruce in both channel 1 and channel 2, while similar values were observed for pine and spruce in channel 3. There were more overlapping values and variations at lower percentiles of intensity distribution, among tree species in all channels.
MCI features have been used to reduce the radiometric effects on multispectral images and improvements in classification have been reported. In this study, the use of similar ratios and indices did not improve the classification. The reason for this could be that the laser scanner is an active instrument, and recorded intensity mainly depends on the instrument design, measurement range, and reflectance of the targets. If the same instrument has been used for data acquisition and the range effect has been corrected, the major factor affecting recorded intensity is the targets illuminated by the laser. Therefore, the intensity itself is good enough to characterize the objects.
The results in this study are in agreement with previous results, in which tree species were classified using ALS combined with multi/hyper-spectral data, although the studies cannot be compared directly because of the differences in the data used, and the number and type of species identified. For example, Dalponte et al [
40] reported a kappa accuracy of 0.89 when classifying three boreal tree species (pine, spruce and broad-leaves), using hyperspectral and ALS data with the manual detection of trees. The higher kappa coefficient obtained in their study could be a result of better delineation of individual trees by manual detection, and a higher spectral resolution. Jones et al [
41] achieved an overall accuracy of 73% for classifying 11 species in coastal south-western Canada, using hyperspectral and ALS data. The lower accuracy could be explained by the higher number of species recognised in the study. This indicates that multispectral ALS data contains similar information to the fusion of multispectral images and ALS data. Compared with the previous study, which used a multispectral ALS of similar density for tree species classification, St-onge and Budei [
29] reported a classification error of 4.59% in the case of the level 1 classification (broadleave vs. needleleaf trees), and of 24.29% in the case of level 2 classification (eight genera), using intensity features (mean and standard deviation of intensity in three channels). The different number of species could be the reason for the difference in accuracy.
The use of a single source of data apparently has advantages over the use of fused data, with respect to data processing. For example, geometric and radiometric calibrations between different data sources produce big challenges, and require much effort to compensate the changes in illumination conditions and vegetation [
26]. Furthermore, previous studies have shown that background signal reduced classification accuracies when using multispectral/hyperspectral images [
42,
43,
44]. In contrast, multispectral ALS data can easily separate the reflections of vegetation from the reflections of the ground, thus background influences on the results, like soil, could be minimised. Therefore, the accuracy of the classification could be improved with the use of multispectral ALS data. However, this issue needs to be explored further in order to investigate the extent to which the accuracy can be improved with multispectral ALS data.
The intensity values of different returns are affected by the vertical structures of trees. In theory, the intensity of only returns can be radiometrically calibrated with high reliability. The first of many returns is distorted by the signal penetrating to the second and other layers. However, there is still valuable information of all return intensities confirmed by this study. In the future, it should be studied whether it is possible to calibrate the intensities of multiple returns in a better way, by taking into account the attenuated part of the signal and the part that causes other returns.
The major drawback with applied Titan data was the inhomogeneous distribution of the point cloud. In the across track, the point spacing was significantly smaller than that in the along track. Either lower aircraft speed or higher scan frequency should be achieved to provide more homogenous point spacing. Another drawback is that the points from the three channels are not registered from the same location, which means that it is not multispectral data in the conventional sense. As a result, pixel/point wise classification cannot be performed; instead, object-based analysis has to be carried out, like in this study. The accuracy of the classification may also deteriorate, because the backscatter from different channels could come from different parts of the objects. The impact of such system design on classifications needs further investigation. Regardless of these drawbacks, multispectral ALS data are still a valuable data source for tree species classification, as shown in this study.
Currently, it is more expensive to acquire multispectral ALS data than aerial images and single-channel ALS. However, it is anticipated that the price will drop as technology develops, and the market is growing. Furthermore, ALS data can be acquired during both the day and night, which partly compensates for the cost of the data acquisition. Therefore, multispectral ALS data could be a cost-effective solution for species-specific mapping of forests in the future, and it has the potential to increase the automation of the whole processing chain.
6. Conclusions
In this study, we assessed the potential utility of single-sensor multispectral ALS data for tree species classification in mixed coniferous forests in a boreal zone. The results suggest that additional information, provided by multispectral laser scanning, may be a valuable source of information for tree species classification of pine, spruce, and birch, which are the main tree species found in boreal forest zones. The best overall classification accuracy achieved was 85.9% using point cloud and SCI features, which was not significantly different from the ones in which all features, or solely SCI features were used. Point cloud features alone achieved an accuracy of 76.0%. Channel 2 performed the best when separating pine, spruce, and birch, followed by channel 1 and channel 3, with overall accuracies of 81.9%, 78.3%, and 69.1%, respectively.
This preliminary study has demonstrated the potential of multispectral airborne laser scanning for possible future solutions for automatic single-sensor forest mapping. It is expected that multispectral airborne laser scanning can provide highly valuable data for forest mapping. However, there are many aspects of multispectral ALS that need to be investigated further, for example: how will multispectral ALS data perform in other forest zones where the number of species composition is higher? Is it possible to derive more useful features to improve the classification? From a practical point of view, future studies could explore the possibility to improve the accuracy of forest inventory mapping using species information obtained from this study.