Hyperspectral Data for Mangrove Species Mapping: A Comparison of Pixel-Based and Object-Based Approach
Abstract
:1. Introduction
2. Data and Methods
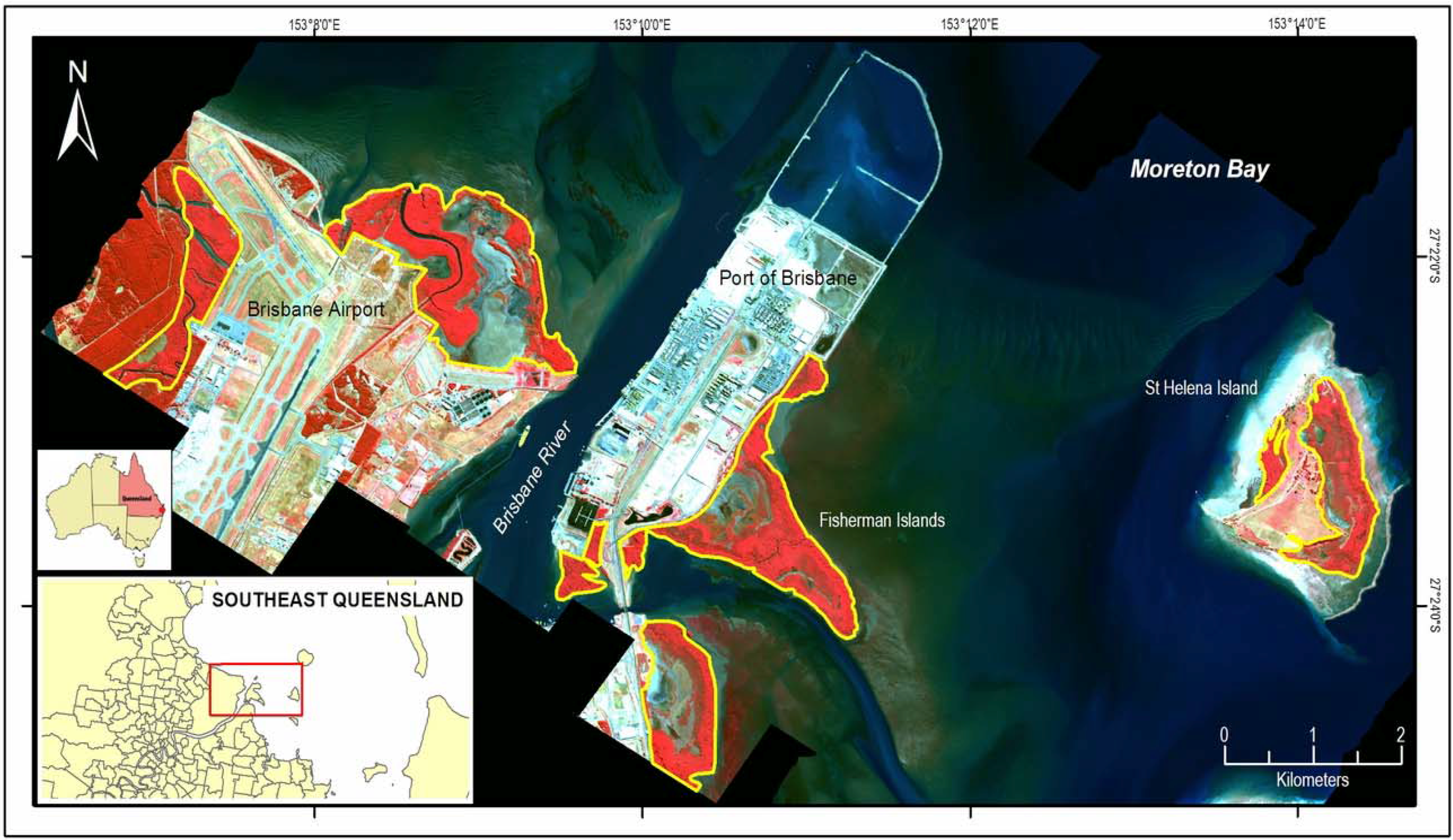
2.1. Study Area
2.2. Image Data and Reference Map
| Acquisition Parameters | |
|---|---|
| Sensor altitude: ~1,200 m Acquisition date and time: -Date: 29 July 2004 -UTC: 23:56:29 -Local: 09:56:29 Meteorological conditions at acquisition: - Temperature: 16.7 °C - Atmospheric pressure: 66 % - Humidity: 1,022.4 hPa Acquisition and Solar Geometry: Off-NADIR view = 2.6 (ideal = 0) Satellite Azimuth = 97.6 Satellite Elevation= 87.3 Sun Azimuth = 35.0 Sun Elevation = 36.5 | Image size (pixels, rows): 4,971, 4,108
Pixel size: 4.0 m × 4.0 m Geometric Attributes: WGS84 in Decimal degrees for Lat/Lon Spectral band for vegetation mapping: 30 bands Radiometric resolution (dynamic range): 14 bit (16,384 levels) |
2.3. Endmember Selection
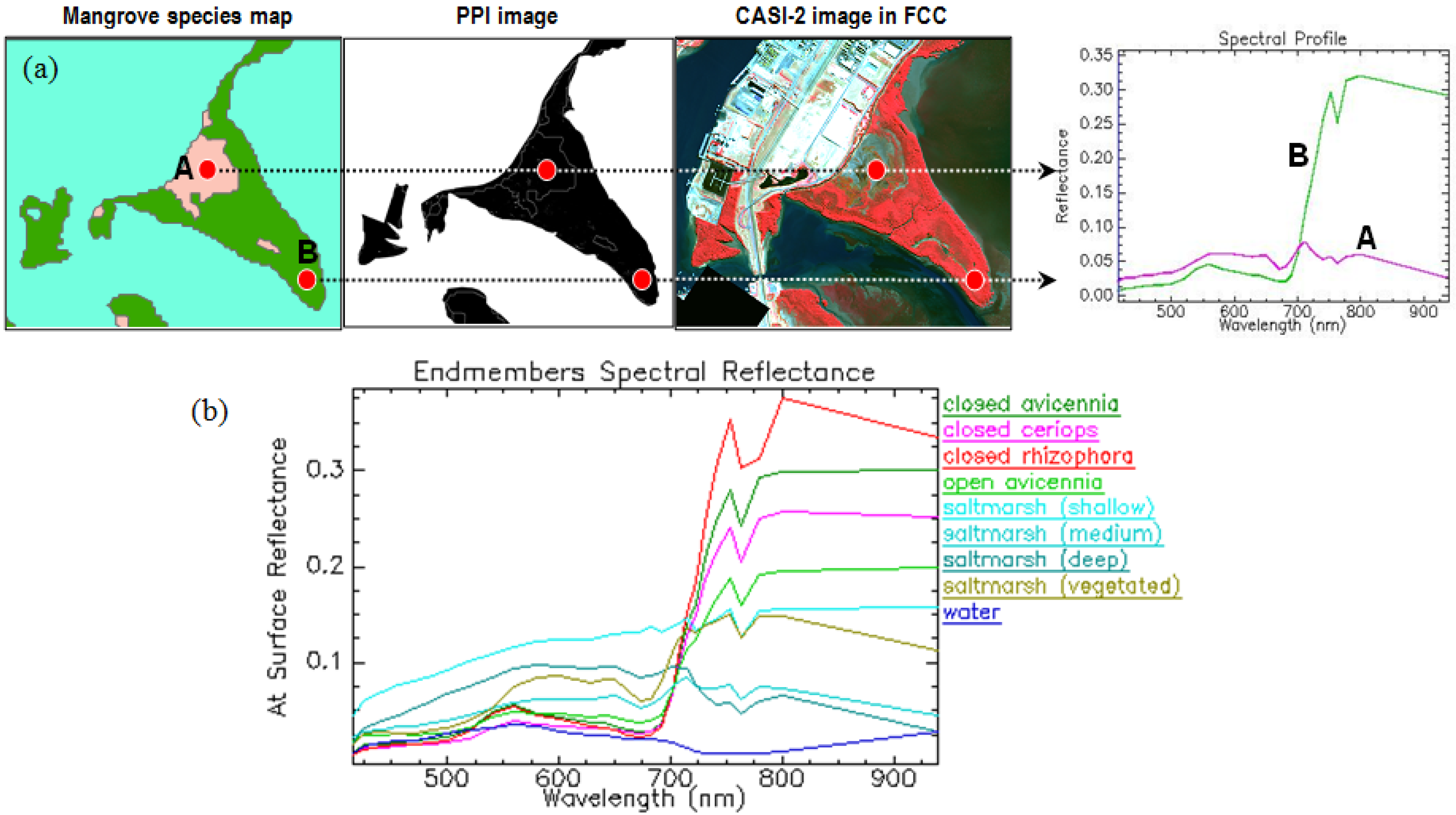
2.4. Image Classification
2.4.1. Spectral Angle Mapper (Per-Pixel Mapping)

2.4.2. Linear Spectral Unmixing (Sub-Pixel Mapping)
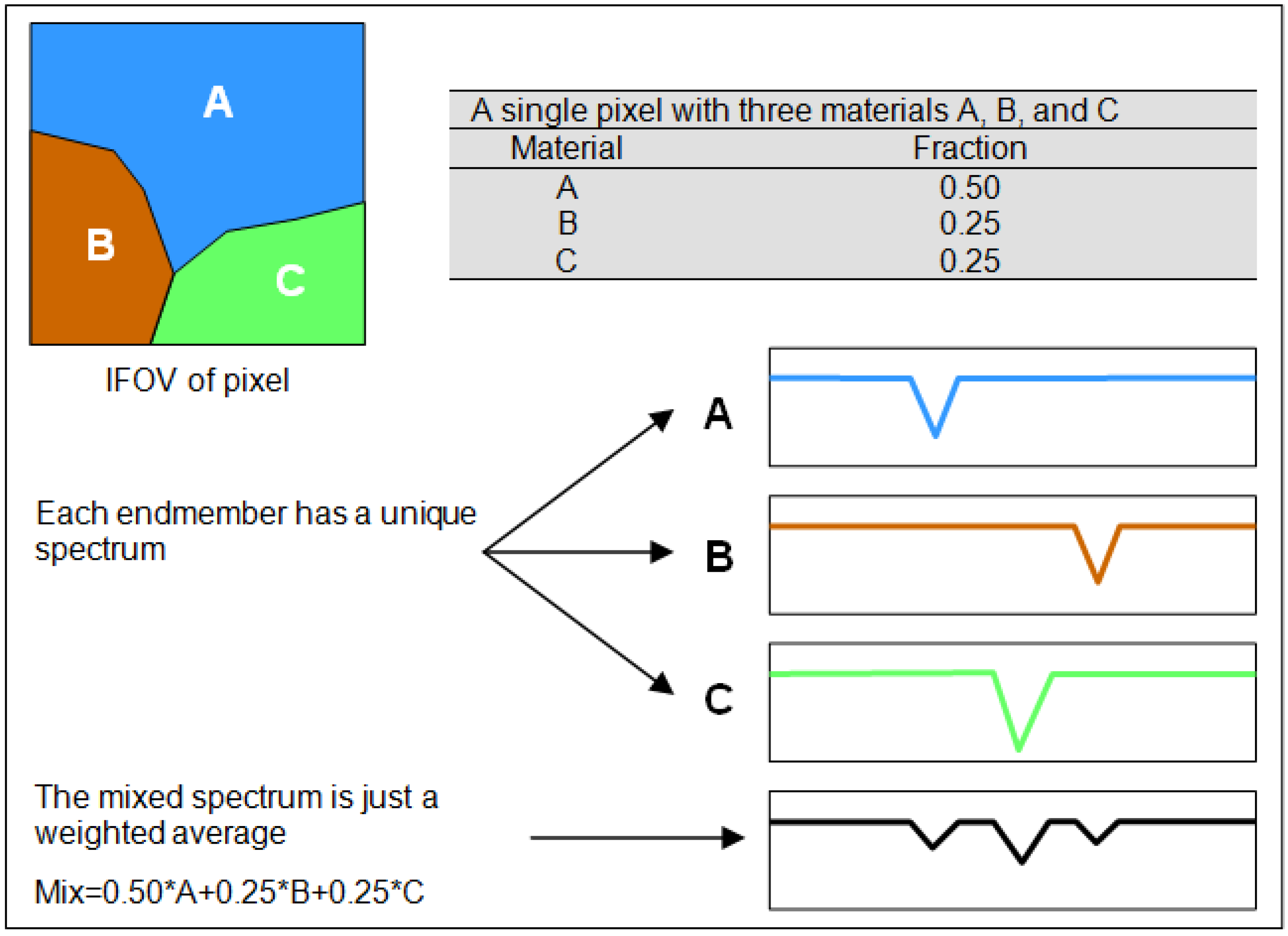
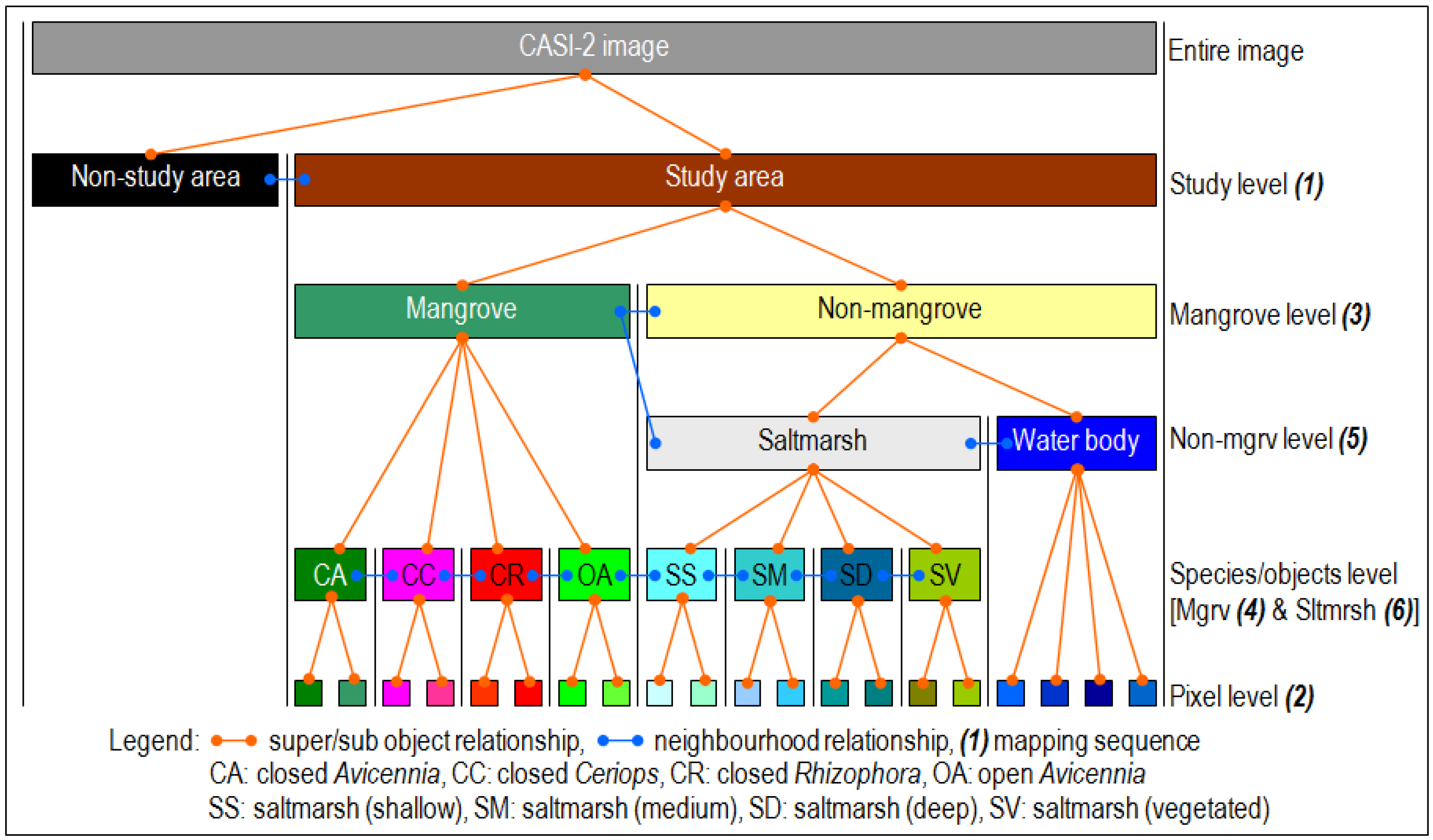
2.4.3. Object-Based Mapping

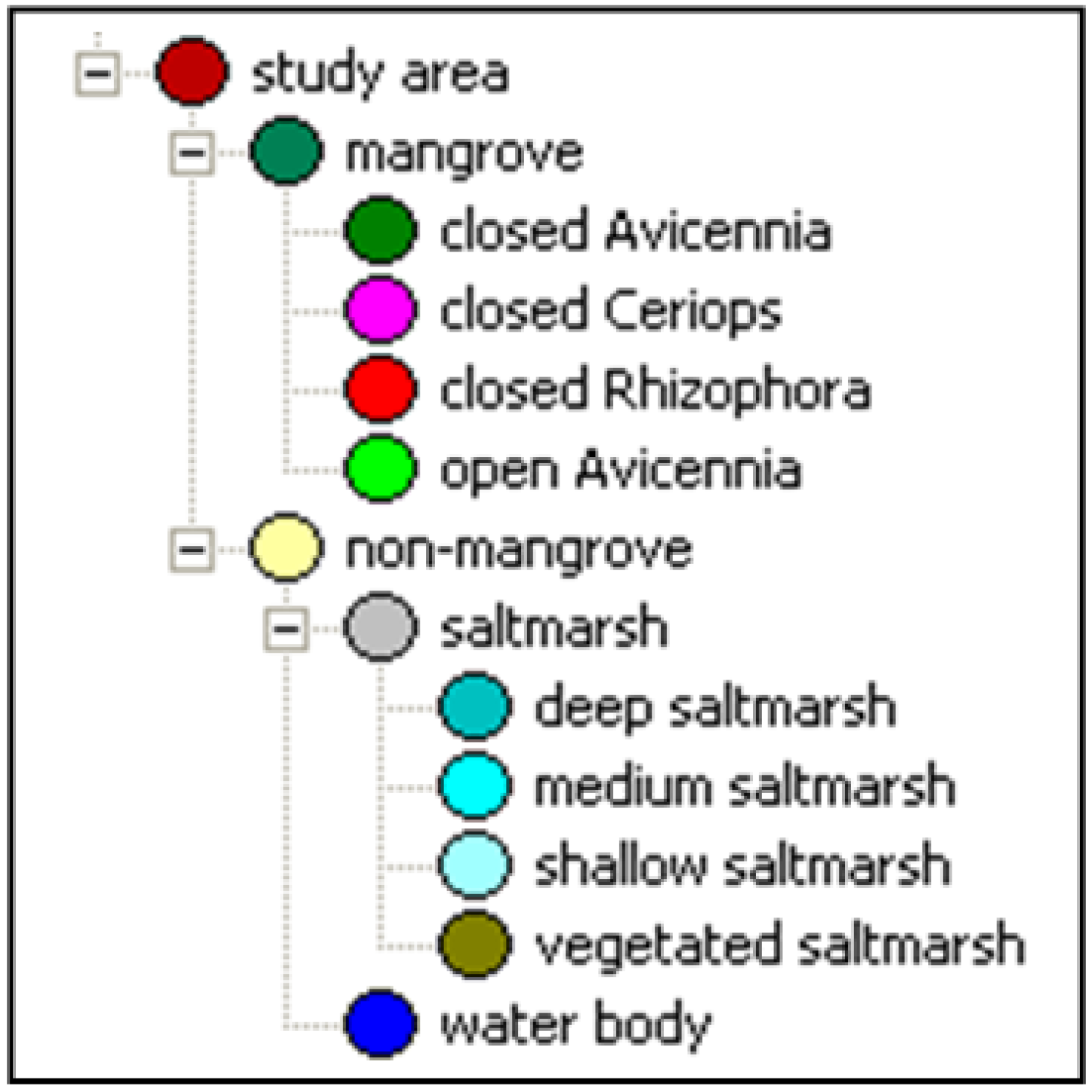
2.5. Error and Accuracy Assessment
3. Results and Discussion
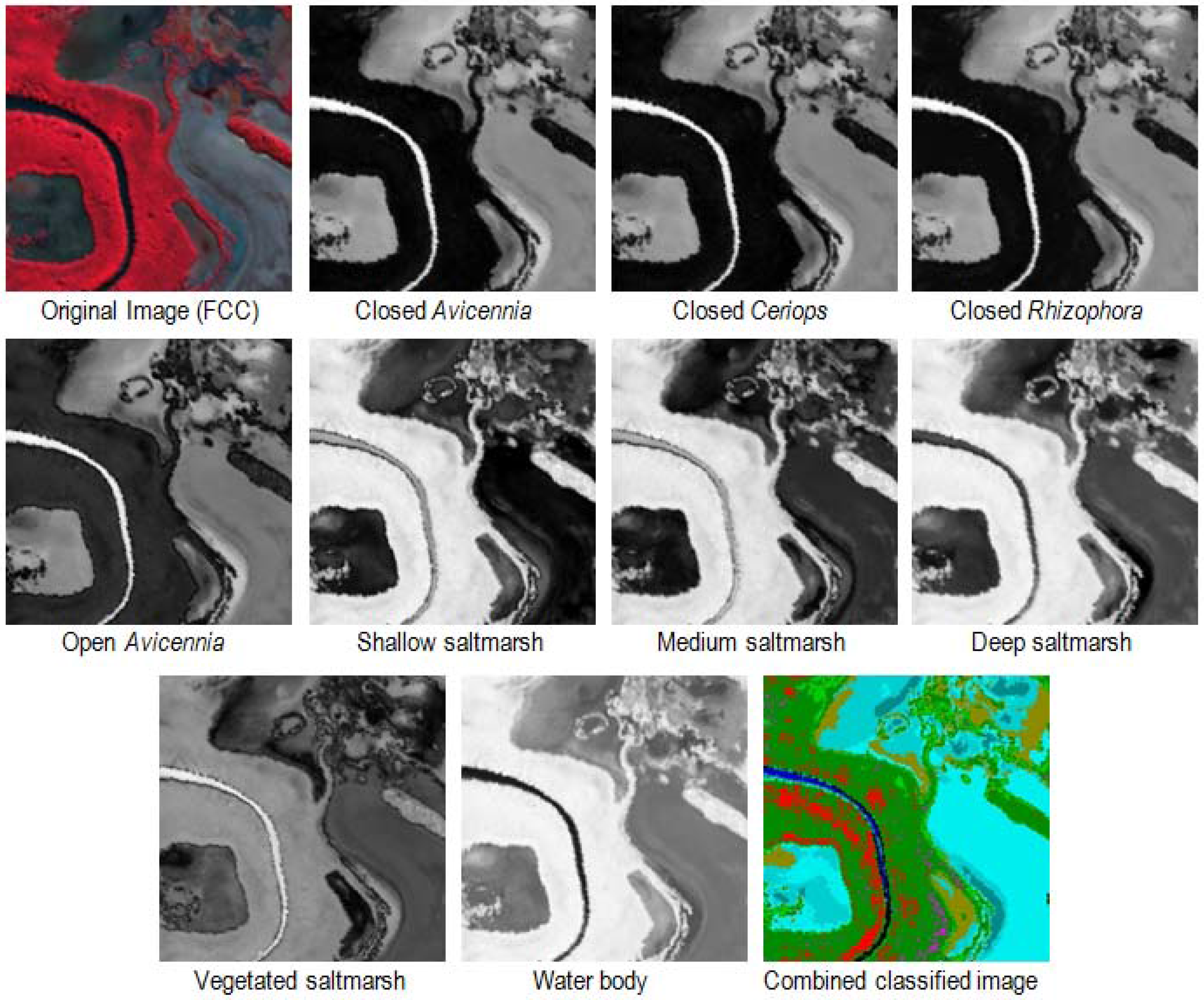
3.1. Image Classification Results
3.1.1. Spectral Angle Mapper
| Wetland Class | SAM | LSU | Object-Based | |||||
|---|---|---|---|---|---|---|---|---|
| Pixel | Area (ha) | % | Pixel | Area (ha) | % | Area (ha) | % | |
| Unclassified | 3,101 | 4.96 | 0.65 | 906 | 1.45 | 0.19 | 9.85 | 1.28 |
| Closed Avicennia | 265,894 | 425.43 | 55.44 | 155,560 | 248.90 | 32.43 | 347.24 | 45.25 |
| Closed Ceriops | 8,187 | 13.10 | 1.71 | 20,992 | 33.59 | 4.38 | 18.37 | 2.39 |
| Closed Rhizophora | 20,117 | 32.19 | 4.19 | 2,809 | 4.49 | 0.59 | 30.23 | 3.94 |
| Open Avicennia | 69,515 | 111.22 | 14.49 | 142,675 | 228.28 | 29.75 | 122.16 | 15.92 |
| Shallow saltmarsh | 59,863 | 95.78 | 12.48 | 35,180 | 56.29 | 7.33 | 40.69 | 5.30 |
| Medium saltmarsh | 21,436 | 34.30 | 4.47 | 95,059 | 152.09 | 19.82 | 72.48 | 9.44 |
| Deep saltmarsh | 5,313 | 8.50 | 1.11 | 4,231 | 6.77 | 0.88 | 29.68 | 3.87 |
| Vegetated saltmarsh | 24,647 | 39.44 | 5.14 | 6,983 | 11.17 | 1.46 | 81.67 | 10.64 |
| Water body (river) | 1,565 | 2.50 | 0.33 | 15,243 | 24.39 | 3.18 | 15.04 | 1.96 |
| Total | 479,638 | 767.42 | 100.00 | 479,638 | 767.42 | 100.00 | 767.42 | 100.00 |
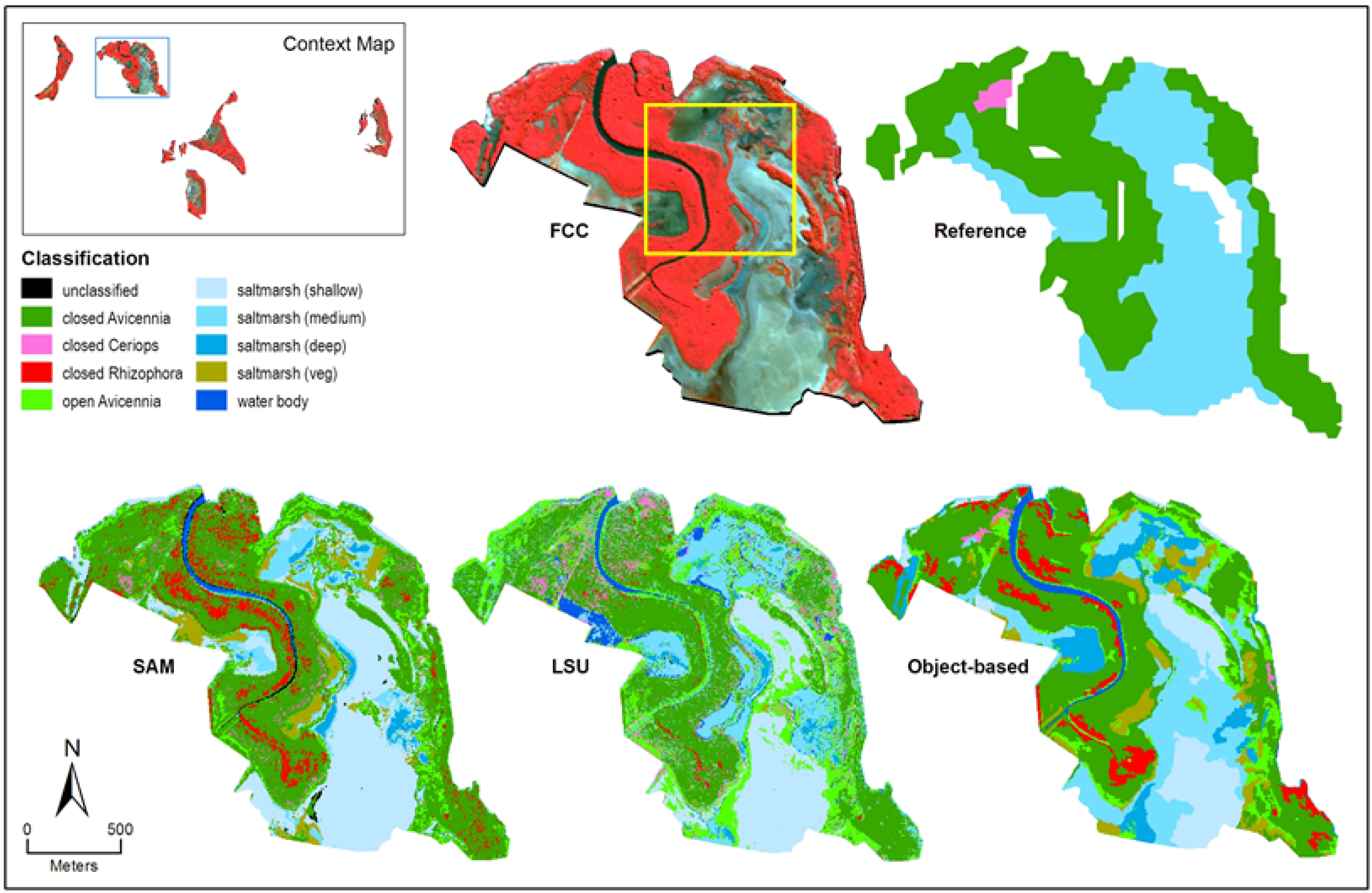

3.1.2. Linear Spectral Unmixing
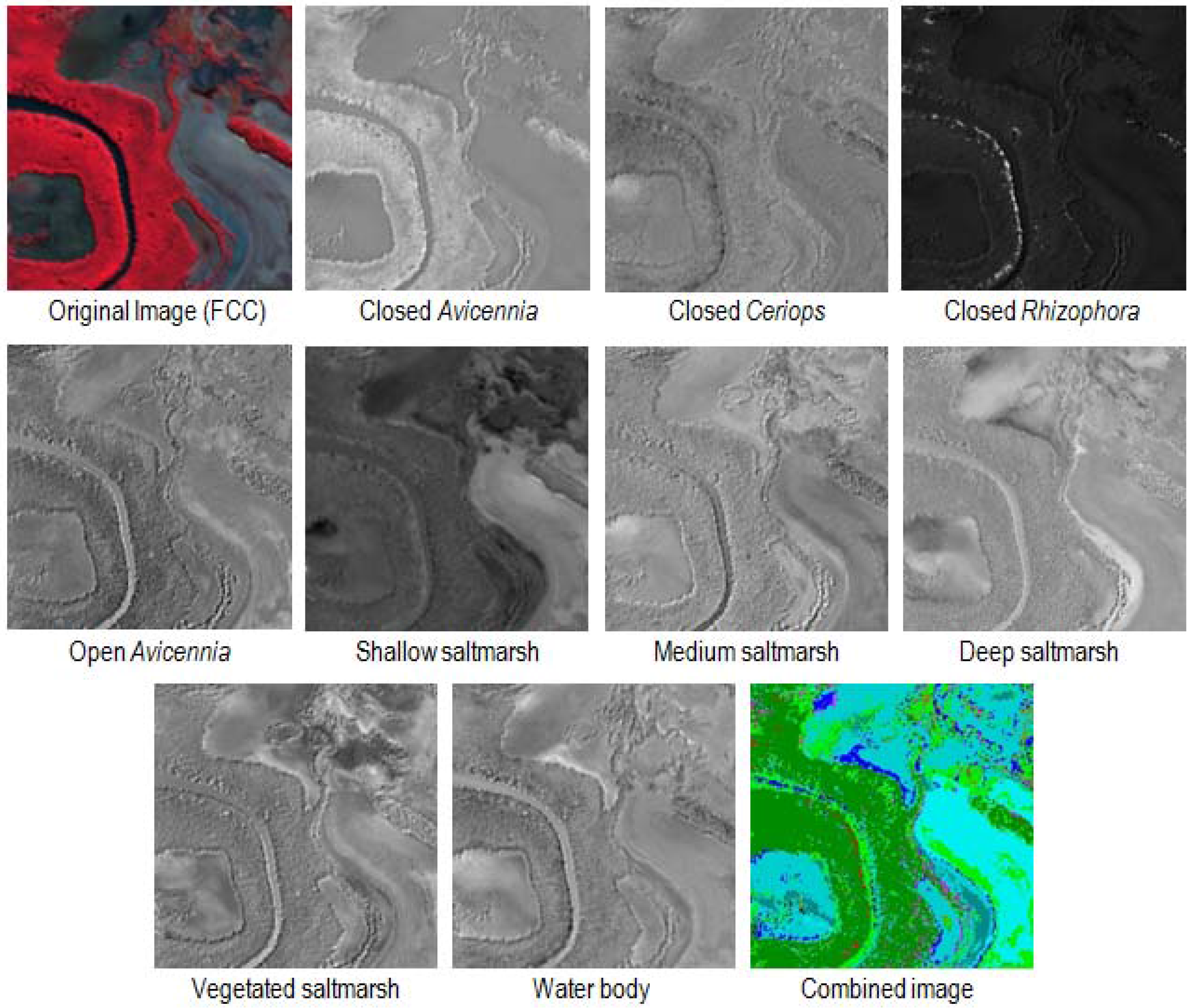
3.1.3. Object-Based Classification
3.2. Comparison between Classification Approaches
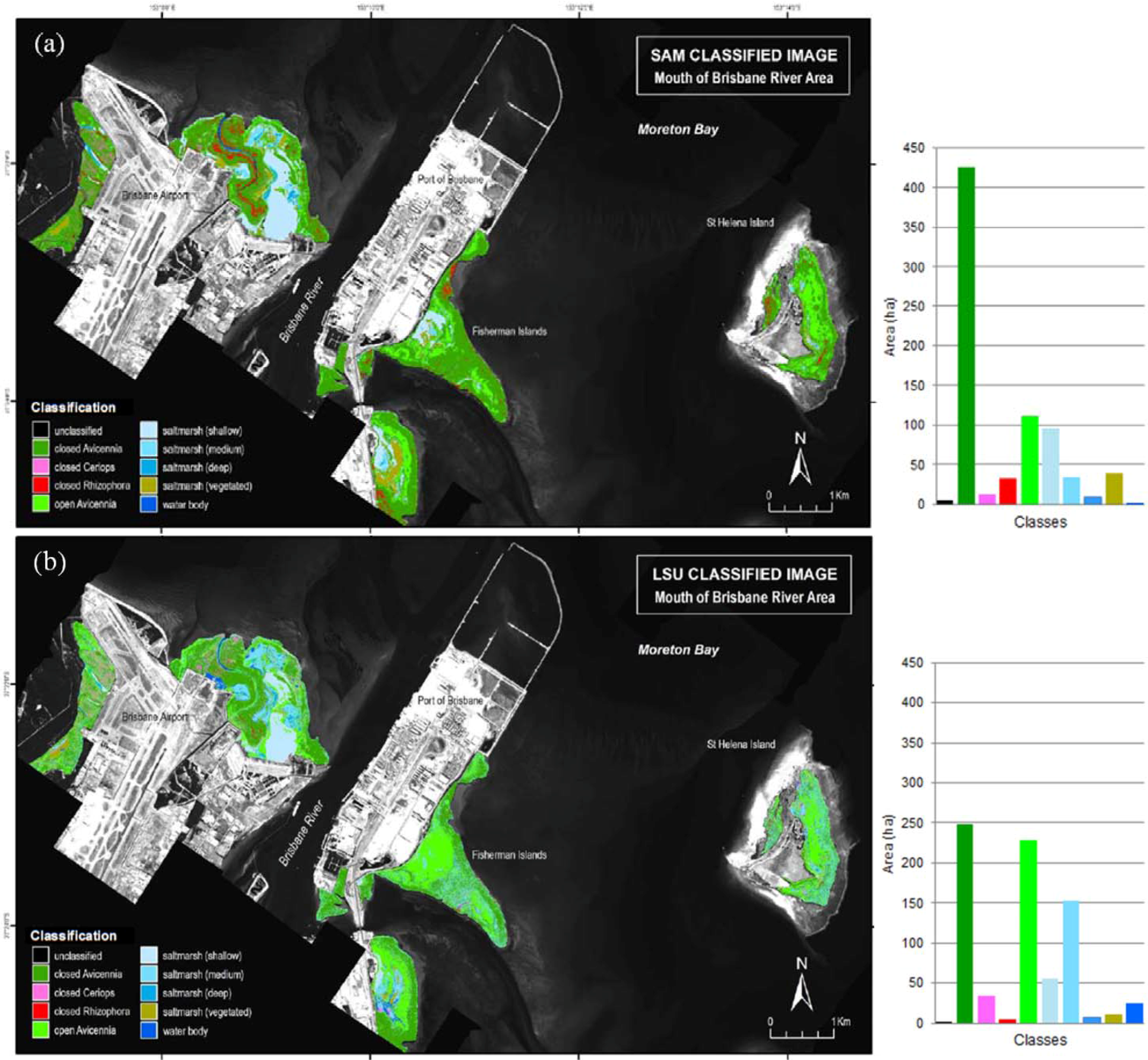
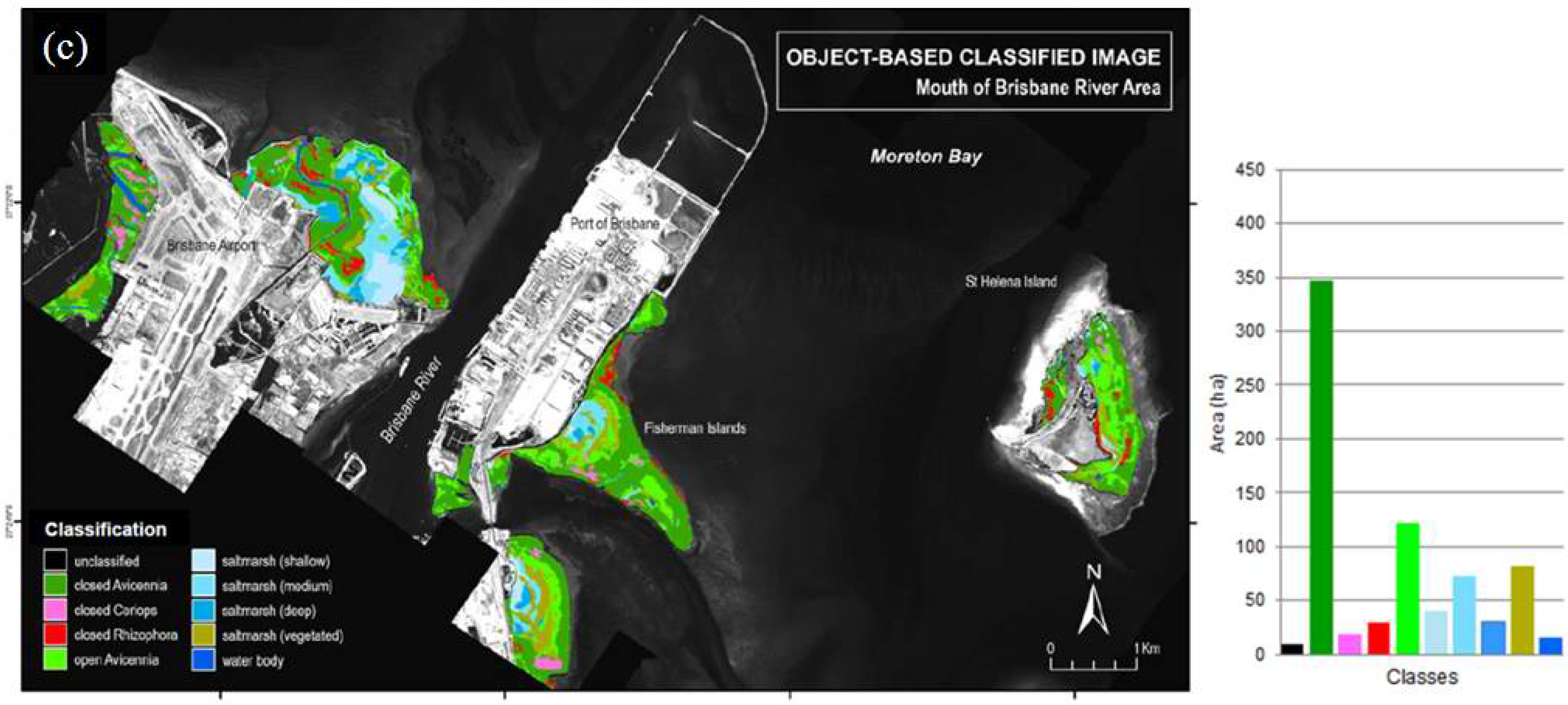
3.3. Error and Accuracy Assessment
| Reference Map | Producer’s Accuracy | User’s Accuracy | |||||||
| Class | CA | CC | CR | OA | SM | Total | |||
| Classified (SAM) | CA | 122 | 24 | 22 | 27 | 11 | 206 | 81% | 59% |
| CC | 3 | 21 | 0 | 2 | 0 | 26 | 42% | 81% | |
| CR | 10 | 0 | 25 | 0 | 0 | 35 | 50% | 71% | |
| OA | 13 | 5 | 3 | 21 | 2 | 44 | 42% | 48% | |
| SM | 2 | 0 | 0 | 0 | 87 | 89 | 87% | 98% | |
| Total | 150 | 50 | 50 | 50 | 100 | 400 | |||
| Reference Map | Producer’s Accuracy | User’s Accuracy | |||||||
| Class | CA | CC | CR | OA | SM | Total | |||
| Classified (LSU) | CA | 87 | 7 | 25 | 10 | 0 | 129 | 58% | 67% |
| CC | 2 | 33 | 1 | 3 | 0 | 39 | 66% | 85% | |
| CR | 2 | 0 | 6 | 0 | 0 | 8 | 12% | 75% | |
| OA | 24 | 10 | 1 | 29 | 33 | 97 | 58% | 30% | |
| SM | 35 | 0 | 17 | 8 | 67 | 127 | 67% | 53% | |
| Total | 150 | 50 | 50 | 50 | 100 | 400 | |||
| Reference Map | Producer’s Accuracy | User’s Accuracy | |||||||
| Class | CA | CC | CR | OA | SM | Total | |||
| Classified (OBIA) | CA | 114 | 18 | 18 | 12 | 3 | 165 | 76% | 69% |
| CC | 7 | 27 | 0 | 0 | 0 | 34 | 54% | 79% | |
| CR | 6 | 0 | 30 | 0 | 0 | 36 | 60% | 83% | |
| OA | 19 | 3 | 2 | 36 | 1 | 61 | 72% | 59% | |
| SM | 4 | 2 | 0 | 2 | 96 | 104 | 96% | 92% | |
| Total | 150 | 50 | 50 | 50 | 100 | 400 | |||
4. Conclusions
Acknowledgements
References
- Kathiresan, K.; Bingham, B.L. Biology of mangroves and mangrove ecosystems. Adv. Mar. Biol. 2001, 40, 81–251. [Google Scholar]
- Green, E.P.; Clark, C.D.; Mumby, P.J.; Edwards, A.J.; Ellis, A.C. Remote sensing techniques for mangrove mapping. Int. J. Remote Sens. 1998, 19, 935–956. [Google Scholar] [CrossRef]
- Saenger, P.; Hegerl, E.; Davis, J. Global status of mangrove ecosystems. The Environmentalist 1983, 3, 1–88. [Google Scholar]
- Robertson, A.I.; Duke, N.C. Mangroves as nursery sites, comparisons of the abundance of fish and crustaceans in mangroves and other near shore habitats in tropical Australia. Mar. Biol. 1987, 96, 193–205. [Google Scholar] [CrossRef]
- Green, E.P.; Mumby, P.J.; Edwards, A.J.; Clark, C.D.; Ellis, A.C. The assessment of mangrove areas using high resolution multispectral airborne imagery. J. Coast. Res. 1998, 14, 433–443. [Google Scholar]
- Phinn, S.R.; Hess, L.; Finlayson, C.M. An assessment of the usefulness of remote sensing for wetland inventory and monitoring in Australia. In Techniques for Enhanced Wetland Inventory and Monitoring; Supervising Scientist Report 147; Finlayson, C.M., Speirs, A.G., Eds.; Supervising Scientist, Environment Australia: Canberra, ACT, Australia, 1999; pp. 44–83. [Google Scholar]
- Davis, B.A.; Jensen, J.R. Remote sensing of mangrove biophysical characteristics. Geocarto Int. 1998, 13, 55–64. [Google Scholar] [CrossRef]
- Kuenzer, C.; Bluemel, A.; Gebhardt, S.; Quoc, T.V.; Dech, S. Remote sensing of mangrove ecosystems: A review. Remote Sens. 2011, 3, 878–928. [Google Scholar] [CrossRef]
- Green, E.P.; Clark, C.D.; Mumby, P.J.; Edwards, A.J.; Ellis, A.C. Remote sensing techniques for mangrove mapping. Int. J. Remote Sens. 1999, 19, 935–956. [Google Scholar] [CrossRef]
- Held, A.; Ticehurst, C.; Lymburner, L.; Williams, N. High resolution mapping of tropical mangrove ecosystem using hyperspectral and radar remote sensing. Int. J. Remote Sens. 2003, 24, 2739–2759. [Google Scholar] [CrossRef]
- Heumann, B.W. Satellite remote sensing of mangrove forests: Recent advances and future opportunities. Prog. Phys. Geog. 2011, 35, 87–108. [Google Scholar] [CrossRef]
- Yang, C.; Everitt, J.H.; Fletcher, R.S.; Jensen, R.R.; Mausel, P.W. Evaluating AISA+ hyperspectral imagery for mapping black mangrove along the South Texas Gulf Coast. Photogramm. Eng. Remote Sensing 2009, 75, 425–435. [Google Scholar] [CrossRef]
- Hirano, A.; Madden, M.; Welch, R. Hyperspectral image data for mapping wetland vegetation. Wetlands 2003, 23, 436–448. [Google Scholar] [CrossRef]
- Demuro, M.; Chisholm, L. Assessment of Hyperion for Characterizing Mangrove Communities. In Proceedings of the 12th Earth Science Airborne Workshop, Pasadena, CA, USA, 25–28 February 2003; Available online: ftp://popo.jpl.nasa.gov/pub/docs/workshops/03_docs/Demuro_AVIRIS _2003_web.pdf (accessed on 5 August 2007).
- Wang, L.; Sousa, W.P. Distinguishing mangrove species with laboratory measurements of hyperspectral leaf reflectance. Int. J. Remote Sens. 2009, 30, 1267–1281. [Google Scholar] [CrossRef]
- Jensen, R.; Mausel, P.; Dias, N.; Gonser, R.; Yang, C.; Everitt, J.; Fletcher, R. Spectral analysis of coastal vegetation and land cover using AISA+ hyperspectral data. Geocarto Int. 2007, 22, 17–28. [Google Scholar] [CrossRef]
- Van Der Meer, F.; De Jong, S.; Bakker, W. Imaging Spectrometry: Basic analytical techniques. In Imaging Spectrometry: Basic Principles and Prospective Applications; Van Der Meer, F., De Jong, S., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2001; pp. 17–61. [Google Scholar]
- Apan, A.; Phinn, S. Special feature hyperspectral remote sensing. J. Spat. Sci. 2006, 52, 47–48. [Google Scholar] [CrossRef]
- Held, A.; Ticehurst, C.; Lymburner, L.; Williams, N. High resolution mapping of tropical mangrove ecosystems using hyperspectral and radar remote sensing. Int. J. Remote Sens. 2003, 24, 2739–2759. [Google Scholar] [CrossRef]
- Baatz, M.; Schape, A. Multiresolution Segmentation: an optimization approach for high quality multiscale image segmentation. In Angewandte Geographische Informationsverarbeitung XII; Strobl, J., Blaschke, T., Griesebner, G., Eds.; Wichmann-Verlag: Heidelberg, Germany, 2000; pp. 12–23. [Google Scholar]
- Blaschke, T. Object based image analysis for remote sensing. ISPRS J. Photogramm. 2010, 65, 2–16. [Google Scholar] [CrossRef]
- Dowling, R.M. The mangrove vegetation of Moreton Bay. Queensland Bot. Bull. 1986, 6, 1–45. [Google Scholar]
- Abal, E.G.; Dennison, W.C.; O’Dononhue, M.H. Seagrass and mangroves in Moreton Bay. In Moreton Bay and Catchment; Tibbetts, I.R., Hall, N.J., Dennison, W.C., Eds.; School of Marine Science, University of Queensland: Brisbane, QLD, Australia, 1998; pp. 269–278. [Google Scholar]
- Duke, N.C.; Lawn, P.; Roelfsema, C.M.; Phinn, S.; Zahmel, K.N.; Pedersen, D.; Haris, C.; Steggles, N.; Tack, C. Assessing Historical Change in Coastal Environments; Port Curtis, Fitzroy River Estuary and Moreton Bay Regions; CRC for Coastal Zone Estuary & Waterway Management: Brisbane, QLD, Australia, 2003. [Google Scholar]
- Manson, F.J.; Loneragan, N.R.; Phinn, S.R. Spatial and temporal variation in distribution of mangroves in Moreton Bay, subtropcal Australia: A comparison of pattern metrics and change detection analyses based on aerial photographs. Estuar. Coast. Shelf Sci. 2003, 57, 657–670. [Google Scholar] [CrossRef]
- Brando, V.E.; Dekker, A.G. Satellite hyperspectral remote sensing for estimating estuarine and coastal water quality. IEEE Trans. Geosci. Remote Sens. 2003, 41, 1378–1387. [Google Scholar] [CrossRef]
- Phinn, S.; Roelfsema, C.; Dekker, A.; Brando, V.; Anstee, J. Mapping seagrass species, cover and biomass in shallow waters: An assessment of satellite multi-spectral and airborne hyperspectral imaging systems in Moreton Bay (Australia). Remote Sens. Environ. 2008, 112, 3413–3425. [Google Scholar] [CrossRef]
- Dowling, R.M.; Stephen, K. Coastal Wetlands of South-Eastern Queensland Maroochy Shire to New South Waler Border; Queensland Herbarium, Environmental Protection Agency: Brisbane, QLD, Australia, 1999. [Google Scholar]
- Asner, G.P.; Heidebrecht, K.B. Spectral unmixing of vegetation, soil and dry carbon cover in arid regions: Comparing multispectral and hyperspectral observations. Int. J. Remote Sens. 2002, 23, 3939–3958. [Google Scholar] [CrossRef]
- Kruse, F.A.; Boardman, J.W.; Lefkoff, A.B.; Young, J.M.; Kierein-Young, K.S. HyMap: An Australian Hyperspectral Sensor Solving Global Problems Results from USA HyMap Data Acquisition. In Proceedings of the 10th Australasian Remote Sensing and Photogrammetry Conference, Adelaide, SA, Australia, 21–25 August 2000; Available online: : www.hgimaging.com/PDF/kruse_10ARSPC_hymap.pdf (accessed on 5 August 2007).
- Jensen, J.R. Introductory Digital Image Processing: A Remote Sensing Perspective, 3rd ed.; Pearson Prentice Hall: Sydney, NSW, Australia, 2005. [Google Scholar]
- Borengasser, M.; Hungate, W.S.; Watkins, R. Hyperspectral Remote Sensing: Principles and Applications; Taylor & Francis in Remote Sensing Applications; CRC Press: New York, NY, USA, 2008. [Google Scholar]
- Okin, G.S.; Roberts, D.A.; Murray, B.; Okin, W.J. Practical limit on hyperspectral vegetation discrimination in arid and semiarid environment. Remote Sens. Environ. 2001, 77, 212–225. [Google Scholar] [CrossRef]
- Rosso, P.H.; Ustin, S.L.; Hasting, A. Mapping marshland vegetation of San Fransisco Bay, California, using hyperspectral data. Int. J. Remote Sens. 2005, 26, 5169–5191. [Google Scholar] [CrossRef]
- ITT. ENVI Online Manuals and Tutorials. ITT Visual Information Solution. 2003. Available online: http://www.ittvis.com/language/en-us/productsservices/envi/tutorials.aspx (accessed on 26 August 2008).
- Navulur, K. Multispectral Image Analysis Using the Object-Oriented Paradigm; CRC Press: Boca Raton, FL, USA, 2007. [Google Scholar]
- Definiens. Definiens Developer 7 User Guide; Definiens AG: Munich, Germany, 2007; Available online: http://www.pcigeomatics.com/products/pdfs/definiens/ReferenceBook.pdf (accessed on 26 March 2008).
- Baatz, M.; Benz, U.; Dehghani, S.; Heynen, M.; Holtje, A.; Hofmann, P.; Lingenfelder, I.; Mimler, M.; Sohlbach, M.; Weber, M.; Willhauck, G. eCognition Elements User Guide 4; Definiens AG: Munich, Germany, 2004. [Google Scholar]
- Congalton, R.G. A review of assessing the accuracy of classifications of remotely sensed data. Remote Sens. Environ. 1999, 35, 35–46. [Google Scholar] [CrossRef]
- Congalton, R.G.; Green, K. Assessing the Accuracy of Remotely Sensed Data: Principles and Practices; Lewis Pulishers: Boca Raton, FL, USA, 1999. [Google Scholar]
- Duke, N. Australia’s Mangroves: The Authoritative Guide to Australia’s Mangrove Plants; University of Queensland: Brisbane, QLD, Australia, 2006. [Google Scholar]
- Congalton, R.G. Putting the map back in map accuracy assessment. In Remote Sensing and GIS Accuracy Assessment; Lunetta, R.S., Lyon, J.G., Eds.; CRC Press: Boca Raton, FL, USA, 2004. [Google Scholar]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kamal, M.; Phinn, S. Hyperspectral Data for Mangrove Species Mapping: A Comparison of Pixel-Based and Object-Based Approach. Remote Sens. 2011, 3, 2222-2242. https://doi.org/10.3390/rs3102222
Kamal M, Phinn S. Hyperspectral Data for Mangrove Species Mapping: A Comparison of Pixel-Based and Object-Based Approach. Remote Sensing. 2011; 3(10):2222-2242. https://doi.org/10.3390/rs3102222
Chicago/Turabian StyleKamal, Muhammad, and Stuart Phinn. 2011. "Hyperspectral Data for Mangrove Species Mapping: A Comparison of Pixel-Based and Object-Based Approach" Remote Sensing 3, no. 10: 2222-2242. https://doi.org/10.3390/rs3102222
APA StyleKamal, M., & Phinn, S. (2011). Hyperspectral Data for Mangrove Species Mapping: A Comparison of Pixel-Based and Object-Based Approach. Remote Sensing, 3(10), 2222-2242. https://doi.org/10.3390/rs3102222






