1. Introduction
Forest inventory is a key process for scientists and forest managers to assess accurate information about forest resources, structure, health, and biodiversity. The single tree is the basic structural unit of the forest, and its 3D structure, growth status, spatial distribution, and other characteristics are the critical factors required for forest resource investigation, supervision, and ecological modeling research [
1]. There is a close relationship between tree segmentation and forest inventory, and it has always been confronted with significant challenges [
2,
3]. The individual tree segmentation algorithm provides pixel-level classification and object-level delineation [
4]. Researchers can characterize the forest on the scale of a single tree, which is of considerable importance for accurate estimation of the height of a single tree, above-ground biomass, and carbon storage in a precise forest inventory [
5]. A substantial body of research has shown that remote sensing technology could monitor and estimate forest resources effectively and reliably [
6].
The earliest application of forest inventory was multi-temporal analysis based on image radiation values or multi-spectral image classification maps to assess forest characteristics such as forest disturbance area, forest succession and development, or the sustainability of forest management practices [
7]. With the increasing complexity of information in remote sensing images, numerous studies have shown that forest inventory management based on instance segmentation has achieved high accuracy and considerable results in remote sensing images [
8,
9,
10,
11,
12]. However, due to the increasing demand for more precise forest resource inventories, LiDAR (Light Detection and Ranging) technology, which offers more detailed spatial information, has been increasingly applied to forest instance segmentation [
13,
14,
15,
16,
17]. Researchers could monitor wood volume or accumulation by measuring individual sample tree variables such as species, diameter at breast height (DBH), tree height, and upper diameter [
18,
19]. Typically, forest point clouds were collected by airborne laser scanner (ALS), terrestrial laser scanner (TLS), and mobile laser scanner (MLS). As one of the first applications applied to forest inventories, ALS has become an important source of auxiliary data for estimating tree height, diameter distribution, wood stock, and forest biomass [
20,
21,
22]. TLS enabled a detailed and non-destructive volume estimate of a single tree, which was then converted to biomass with a fundamental density of wood [
23,
24,
25]. MLS provides a kinematic method for collecting 3D environmental data. In particular, the single-scanner MLS system integrated with GNSS-IMU, as developed by Antero Kukko et al. [
26], has contributed significantly to practical forestry applications and enabled high-precision terrain and structural modeling under boreal forest conditions. Similarly, Malladi et al. [
27] further advanced the automated forest inventory generation process with tree instance segmentation and trait estimation using LiDAR data collected by the MLS platform in Switzerland and Finland. For the forest point cloud generated by LiDAR, current research is divided into two-dimensional methods based on the canopy height model (CHM) [
28] generated after rasterizing the point cloud, and methods based on point spatial clustering and deep learning (
Table 1). Although networks based on these methods have achieved several results, sparse 3D convolutional networks [
29] provided higher accuracy and computational advantages for most high-density forest point clouds, such as artificial limited forests, making them more suitable for the sparse characteristics of point clouds.
Following previous research, Henrich et al. recently proposed a pipeline called TreeLearn based on deep learning sparse convolutional neural networks (CNN) and clustering algorithms [
38,
39]. Similarly, Xiang et al. [
40] proposed a deep learning framework called ForAINet. Its network model mainly extracted semantic prediction and offset prediction of the forest point cloud by the deep learning method, and used the clustering algorithm to separate the point cloud into a single instance. Both networks enabled instance segmentation of forest point cloud data, driving the automation of forest inventory and management. ForAINet and TreeLearn similarly employ 3D U-Net architectures for voxel point cloud processing. However, conventional sparse CNNs have two critical limitations in forest point cloud segmentation that hinder their modeling of tree structural features: 1. Poor long-range dependency modeling (local convolutions fail to correlate non-adjacent points in overlapping canopies, a typical tree structural feature); 2. Limited local perception field, lacking coherence in modeling vertical trunk features and horizontal canopy features, leading to unclear boundaries between adjacent trees.
In response, Oktay et al. [
41] proposed a novel attention gate (AG) model for medical imaging that aims to be easily integrated into standard CNN architectures (e.g., U-Net model) with minimal computational overhead while improving model sensitivity and prediction accuracy. However, Gu et al. [
42] put forward an innovative deep learning model called Mamba, which is based on the attention architecture and its core attention module. Traditional self-attention mechanisms dealt mainly with one-dimensional or two-dimensional sequence data (such as text or images). However, when applied to high-dimensional data structures, such as 3D point clouds or spatiotemporal data, the computational complexity will increase dramatically, and they have limitations in capturing the complex relationships between different dimensions. Recent studies validate the effectiveness of Mamba in addressing these 3D segmentation challenges: Li et al. demonstrated that Mamba enables efficient long-range modeling in 3D point clouds, a capability critical to our task of single-tree segmentation, where capturing extensive contextual dependencies is essential for distinguishing overlapping crowns [
43]; Xu et al. showed that their CDA-Mamba model enhances multi-dimensional feature consistency [
44], which aligns with our goal of robustness in heterogeneous forest structures; similarly, Zhang et al. verified the feasibility of Mamba-CNN hybrids with their ConvMamba [
45], supporting the rationale behind our pre-convolution Mamba integration. Based on the possibilities of the above research, we embedded Mamba before the convolutional backbone network. By globally mixing voxel features, we enhanced the contextual awareness of the sparse convolution and helped the network align the information of the trunk and the canopy. This design is specifically tailored to the sparse forest point cloud data, improving the robustness of the method to forestry data from different regions.
As evidenced by
Table 1, current laser point cloud datasets for forest inventory exhibit notable limitations in three key aspects: (1) geographic diversity (multi-region coverage), (2) species representation (multi-type inclusion), and (3) measurement precision. To this end, the study proposes an innovative forest point cloud simulation pipeline based on FOR-instance [
46] to improve the diversity of data. The contributions of our research are as follows.
- 1.
Enhanced feature extraction through mamba integration
Our research shows that the ability of Mamba to model high-dimensional feature dependencies effectively addresses key challenges in 3D forest point cloud instance segmentation, including ambiguous boundary delineation between adjacent trees and instance misclassification due to occluded or overlapping canopies. By integrating Mamba with a sparse convolutional neural network, the study develops a robust module validated across multiple heterogeneous and composite datasets, confirming its generalizability and improved segmentation accuracy.
- 2.
Improved segmentation accuracy in complex scenes
To mitigate the high error rates of existing methods in dense and structurally complex forest scenes, we introduce a novel offset prediction module coupled with an optimized clustering algorithm within our enhanced network architecture. The proposed method significantly reduces segmentation errors due to canopy crossing and overlapping, especially in low-canopy and high-density environments.
- 3.
Synthetic data generation for real-world forest inventory
This research recognizes the limitations of existing datasets, such as limited sample diversity and high noise levels in real-world forest point clouds. It proposes a scalable procedural pipeline for generating synthetic forest scenes. This method produces biologically plausible tree distributions and structural variations, closely mimicking limited-scale forest inventory conditions and facilitating more reliable model training and evaluation.
3. Synthesis Pipeline and Method
3.1. Simulated Forest Point Cloud Synthesis Pipeline
Although forest inventory has traditionally relied on sampling within small and fixed-area plots for statistical inference [
49], recent technological advancements have shifted the focus towards analyzing larger sample sets to improve the representativeness and accuracy of forest attribute estimations [
50]. This evolution underscores the need for datasets that support the development of robust methods capable of handling the scale and variability of modern inventory tasks. For this reason, our study proposes a novel pipeline to solve the problem of insufficient samples faced by the FOR-instance dataset (
Figure 2). The purpose of this pipeline is to process each plot of the FOR-instance dataset. The CULS, NIBIO, and SCION datasets represent managed forests characterized by uniform canopy structures and greater tree height consistency, along with higher block counts and point density. In contrast, the TUWIEN and RMIT plots exhibit substantially lower values across these parameters. The processing pipeline is principally designed to execute instance trees expansion and instance re-encoding across all five blocks of the FOR-instance dataset.
Firstly, for an original point cloud plot, the pipeline randomly crops it on the original plot and divides it into two outputs: noise and denoising. Among them, the denoised output is achieved by trimming the cluster size and outliers of the edge point cloud. More specifically, we arrange the point clouds in a regular square pattern. The hyperparameter , which determines the size of each square, is set to 25–28. The hyperparameter is usually set around 500 (may increase or decrease depending on the sparsity of the forest), and point clouds with fewer than this number of clusters are typically classified as noise. The size of the point cloud after pipeline processing is usually 80 × 80, corresponding to the real scanning scene. The experimental results show that outliers in the noisy forest point cloud can improve the robustness of the model and prevent overfitting. Secondly, the pipeline rotates the cut point cloud, re-encodes each instance on the plot, and concatenates in the horizontal direction to form a limited boundary forestry point cloud.
The advantages of the synthetic point cloud over the original forest are as follows: 1. The synthetic point cloud algorithmically increases the number of tree instances per plot (targeting > 90 at least). This expansion better approximates real-world stand densities and complexity, which is critical for developing and validating robust perception models such as those employed in mobile LiDAR-based frameworks that successfully map thousands of trees across large, continuous forest stands [
51]. 2. To some extent, it solves the problem of the lack of large and fine real forest land point clouds in current research, making it more consistent with the detection of limited woodland in the real forest inventory process. The real situation is the scene where the operator’s walking trajectory distance is about 100 m. Finally, the pipeline also randomly splices the forests of the five blocks to form a more complex, limited boundary forest point cloud that is more consistent with a non-plantation forest. The purpose of this project is to investigate the segmentation performance of the model in composite forest environments while simulating inventory scenarios for more complex mixed forest conditions encountered in real-world applications. It is emphasized that during comparative experiments with baseline and state-of-the-art models, the original datasets are retained as the test benchmark. Our pipeline is only activated when the performance of the proposed method is evaluated independently, ensuring a fair comparison.
3.2. Overview of the Instance Segmentation Pipeline
To achieve an automated forest inventory, the proposed model applies 3D local forest point cloud instance segmentation as the main research content. In order to segment a large area of forest point cloud into individual instances, first, our study aims to detect tree-related points within 3D forest point clouds and subsequently cluster them into distinct individual units. Specifically, for a 3D point set
, the proposed approach simultaneously performs semantic and instance segmentation on each point. Semantic segmentation categorizes every point into either tree or non-tree categories, where non-tree points encompass ground surfaces and low-lying vegetation such as grasses and scattered bushes. The tree class includes medium-sized shrubs and points with a tree structure. The objective of instance segmentation is to assign each point in the split tree dataset to one of k mutually exclusive tree instances. Specifically, our model improves on the TreeLearn model pipeline proposed by Henrich et al. [
38].
Figure 3 illustrates the specific instance segmentation process of the model for each slice of the original input point cloud. In order to solve the problem of memory limitation, the forest point cloud is divided into small overlapping rectangular blocks based on the x- and y-coordinates. These segmented inputs must undergo voxelization before being fed into the network model. The voxelization process transforms the unstructured point cloud into a sparse volumetric representation. It is parameterized by a voxel size
and a maximum point count per voxel
. The latter acts as an upper bound for point retention or aggregation within each voxel, serving to control memory footprint and computational load while ensuring consistent tensor dimensions across the dataset. The voxel size determines the level of detail preserved in the 3D structure, while the per-voxel point limit prevents memory overflow when processing dense forest canopies.
After voxelization, an improved Mamba-based sparse convolutional neural network (M3Unet) is used for prediction. For each point inside the rectangular block, the model makes the following two predictions: 1. Predict the offset of all the tree points from the center of their respective trunks. This offset prediction is crucial because it focuses on accurately locating the position of all the points that belong to one instance in the coordinate system. By calculating the offset of each point from the predefined center of the trunk, the model provides a better estimate of the actual position of the tree, taking into account both vertical and horizontal displacements.
2. Perform semantic prediction on points to classify them as tree points or non-tree points. Non-tree points are directly counted in the final segmentation result. This semantic classification simplifies subsequent processing steps by immediately identifying and leaving out points that are not relevant for individual tree segmentation. This operation ensures that each point inherits the contextual features learned by the network, enabling point-wise semantic and offset predictions. Following feature extraction in the sparse voxel domain, the learned representations are projected back to the original point cloud via the voxel-to-point mapping established during voxelization. This mapping associates each point with its corresponding voxel , allowing the assignment of voxel-level features to all points within that voxel: . Crucially, all subsequent clustering and final evaluation are performed directly on the original point cloud.
Thereafter, the semantic and offset predictions of the entire input point cloud are obtained by aggregating the predictions of the rectangular patches of the individual cut point clouds. For the overall forest point cloud, the network projects point coordinates by adding offset values. Then, the appropriate points are selected from these projection coordinates, and the HDBSCAN (Hierarchical density-based spatial clustering of applications with noise) clustering algorithm is used to divide different point clusters into different instances. The HDBSCAN algorithm demonstrates robust clustering capabilities, effectively handling non-spherical cluster geometries while simultaneously identifying and filtering noise points within the dataset. In the context of forest point cloud processing, points belonging to the same instance can be accurately grouped based on the density distribution in the projected coordinate space. Finally, the weighted K-nearest neighbors (W-KNN) algorithm is used to assign the remaining unassigned tree points in the cluster to neighboring tree instances. The weighted nature of the k-nearest neighbor algorithm takes into account the distance of the nearest neighbor points and other related factors, which makes the assignment of the unassigned points more accurate. This step ensures that all tree points are grouped correctly and improves the completeness and accuracy of the individual tree segmentation results.
3.3. Offset Prediction Method for Instance Segmentation of Forest
For each point within an instance, the model will predict its offset. The objective of this study is to predict an offset vector pointing towards the center of the tree trunk (
Figure 4). The core of this method is to define a robust and representative reference point for each tree instance, from which all tree points’ offsets are calculated. The key innovation lies in how this reference point, or “trunk center”, is determined.
For a given tree instance, its trunk center is not defined as a single fixed height but as a vertical segment bounded by
and
. These bounds are derived from the statistical distribution of the instance’s points along the vertical (Z) axis:
and
, where
and
are the mean and standard deviation of the Z-coordinates of all points within that instance, respectively. This statistical range typically corresponds to the middle section of the tree trunk, which is more stable and less prone to occlusion or extreme curvature than the base or crown. Within this vertically defined segment, we further refine the candidate points used for calculating the final instance center. Points must satisfy two criteria: (1) their verticality
V, defined as
, where
is the Z-component of the point’s normal vector, must exceed a threshold of 0.6; and (2) their Z-coordinate must lie within
. The verticality threshold, adapted from the work of Henrich et al., ensures that the selected points belong to predominantly vertical structures like the trunk and major branches, which are reliable indicators of the tree’s central axis. Empirical results confirm that this setting achieves effective segmentation while accommodating a moderate degree of stem inclination [
38].
The final instance center for a given tree instance
l, denoted as
, is computed as the three-dimensional centroid of all points in the filtered set
, which satisfy both the verticality and the statistical height-range criteria. This center is calculated as follows:
where
is the number of points in
and
represents the 3D coordinates of the
i-th point. For each original point
belonging to instance
l, its offset vector
is then defined as follows:
This offset vector describes the specific direction and distance from any point in the tree to its calculated trunk center. The primary purpose of predicting these per-point offsets is to provide the network with a consistent, instance-specific spatial reference. This enables the network to aggregate points belonging to the same tree and to distinguish them from points of other trees or outliers (whose offsets point to different centers), forming the foundation for the subsequent instance grouping step.
Algorithm 1 details the complete procedure for generating these offset vectors for all tree points in an input forest point cloud. The algorithm takes as input a point cloud with per-point instance labels, semantic labels, and pre-computed verticality values. It processes each unique tree instance independently. For each instance, it first calculates
and
from all its points. Then it applies the verticality and height-range filters to obtain a subset of representative points. If this subset is non-empty, its centroid becomes the instance center; otherwise, a null center is assigned. Finally, the offset for every point in the instance is computed as the vector difference between this center and the point’s location. This method maximizes the preservation of the tree’s structural information (verticality and spatial distribution) while computationally focusing on the most representative parts of the trunk, thereby enhancing both the robustness and efficiency of the offset prediction.
| Algorithm 1: Offset vector calculation |
![Remotesensing 18 00664 i001 Remotesensing 18 00664 i001]() |
3.4. Mamba-Enhanced Sparse Convolutional Neural Network
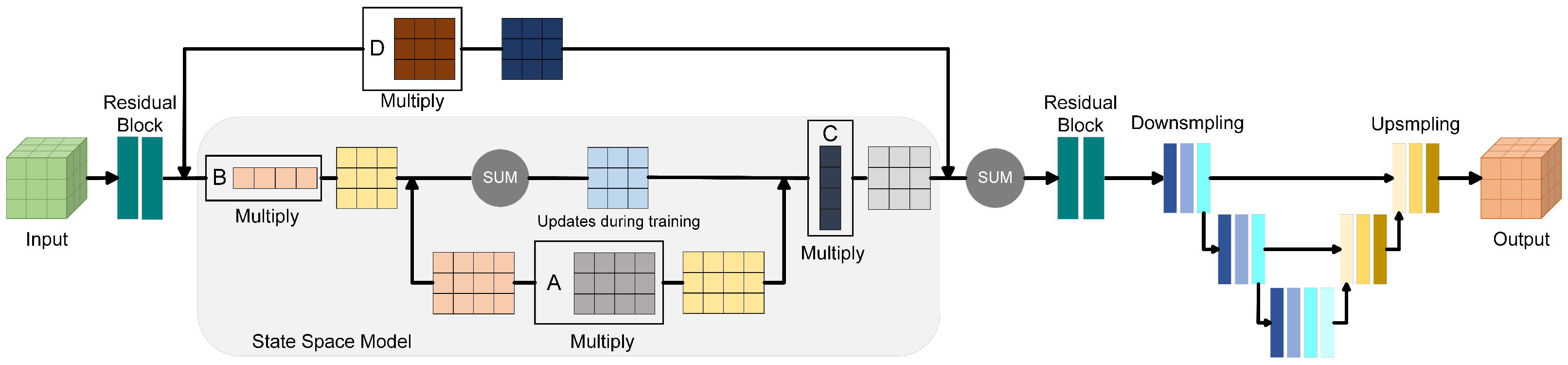
The enhancement of sparse convolutional neural networks through Mamba is primarily implemented within the 3D U-Net architecture. For an input voxel set, feature extraction begins with a residual structure. Subsequently, point cloud features are refined by Mamba through its structured State Space Model (SSM). As a sequential modeling framework, SSM efficiently captures long-range dependencies within the data by maintaining a hidden state that evolves dynamically with the input sequence. This mechanism allows the model to integrate contextual information across distant regions, which is especially beneficial for distinguishing overlapping canopies and resolving ambiguous tree boundaries in forest point clouds.
Figure 5 describes the principle of SSM to enhance the feature extraction of the 3D U-Net in detail. SSM determines the input
at the current next time and the state
at the current time by introducing a latent state
for the state
at the next time.
where
A and
B represent the state transition matrices, which similarly affect the latent state
h. And its output
depends only on
:
where
C stands for the projection matrix. Furthermore, point cloud data are usually discrete. Therefore, Mamba discretizes the model by introducing the time scale parameter
; that is, the continuous parameters
A,
B, and
are transformed into discrete parameters
,
through a fixed formula.
After discretization, the output
y is finally calculated by convolution, which is expressed as follows.
where
denotes the input feature sequence, ∗ is the convolution operator, and
is the structured convolution kernel derived from the discretized SSM parameters. The kernel length is defined by the exponent
k.
In more detail, Algorithm 2 delineates the process by which the Mamba mechanism refines features within the input point cloud. The process first aggregates the sparse forest point cloud into structured voxels through the voxelization step. Each voxel retains the corresponding point coordinates, verticality features, and mapping relationship with the origin. Subsequently, the processed voxel point cloud (
) will be input into Algorithm 3 for feature enhancement. After the instance point cloud information is extracted by Mamba, the point features are processed by a residual block connected with the upsampling-downsampling 3D U-Net coding structure to obtain the final output.
| Algorithm 2: Feature extraction with Voxel-Mamba |
Input: Point coordinates ; Point features ; Batch indices ; Batch size Output: Voxel-level features ; Voxel-to-point mapping - 1
- 2
- 3
- 4
- 5
return
|
| Algorithm 3: Tensor Mamba |
Input: Sequence X [Tensor, shape: ]; Projection weights: State matrix: A [Tensor, shape: ] State matrix: D [Tensor, shape: ] Output: Transformed Sequence Y [Tensor, shape: ] - 1
- 2
- 3
, - 4
, - 5
- 6
- 7
, , - 8
, - 9
return
Y
|
Algorithm 3 describes the basic principle and process of Mamba feature enhancement. This algorithm efficiently models sequential dependencies while maintaining linear scalability in sequence length. The following additions should be incorporated: Function applies a linear transformation to the input X. For the transformed input X, Mamba first applies a 1D convolution along the sequence dimension using , followed by a activation. For the output Y, is adopted to apply the state-space recursion to update the hidden state.
A detailed analysis shows that the algorithm leverages the sequential modeling of Mamba to process voxel feature sequences. This design enables it to effectively learn and align with the inherent structure of trees. Through its internal state-space recursion mechanism embodied in , it is plausible to capture the inherent connections between different voxels, such as those linking the trunk and canopy of a single tree or spanning non-adjacent regions in overlapping canopies. The state matrix A is generated via and , and this negative exponential transformation may assign decaying weights to historical sequence states, potentially enabling the retention of feature correlations between spatially non-adjacent but structurally related voxels. Meanwhile, the time scale parameter calibrated by might dynamically adjust the sensitivity of state updates, which could help distinguish voxel features in overlapping canopy regions by adapting to the density differences of such regions. This global feature association learning is likely to enhance the contextual information of sparse convolutions, making the network more robust to density variations (e.g., sparse understory regions and dense upper canopy) and height differences in forest environments. Additionally, the SSM mechanism in the algorithm may promote multi-dimensional feature coherence by integrating the sequential processing capability of and the non-linear activation of , which could be conducive to aligning trunk (vertical) and canopy (horizontal) structural information. This alignment might reduce ambiguities in tree boundary delineation caused by the independent feature processing of conventional sparse CNNs.
3.5. Tree Instance Clustering Method
Upon obtaining the semantic and offset predictions for each point, the model unfolds the feature map into a two-dimensional space. Points meeting criteria based on voxelization features, verticality, and predictions are selected and subsequently processed through hierarchical density clustering to handle clusters of varying densities. HDBSCAN can automatically discover clusters in the data without explicitly setting the number of clusters and is more robust to changes in cluster density.
Figure 6 illustrates the clustering mechanism of HDBSCAN via core distance computation. Formally, the core distance for any point
p is computed as its Euclidean distance to the
nearest neighbor
.
In our clustering method, HDBSCAN first processes the filtered points that meet the feature conditions to identify the main structural clusters of each tree instance, primarily focusing on the trunk and core canopy regions. Following this initial clustering, a subset of points—typically those in sparse areas or boundaries—remain unassigned. In that case, the approach taken by the network is weighted K-nearest neighbors (W-KNN). The main advantage of W-KNN over traditional KNN is that it assigns a weight to each neighbor instead of treating all neighbors equally.
Figure 6 also depicts that on the basis of the clustering results of HDBSCAN, W-KNN introduces a weight for each of its neighbors, usually the weight is based on the distance between the neighbor and the target point. The form of weighting is the inverse of the distance, where
is the distance between the target point x and its
nearest neighbor
.
In the following classification task, W-KNN performs weighted voting according to the weight of the neighbors, such that the neighbors that are closer to the target point have more influence on the prediction.
where
is the label of the neighbor of the
instance, and
is an indicator function to identify the neighbor.
3.6. Evaluation Metrics and Implementation Details for Instance Segmentation
Evaluation index of single tree example result detection: For instance-level detection evaluation of the tree instance results of the segmentation of the pipeline, we define the point-wise intersection over union (IoU) ratio between the predicted results of all points and the real instance (GT) of the woodland tree. The Hungarian matching algorithm [
52] is subsequently utilized to establish optimal assignments between detected and reference tree instances. In order to achieve the detection evaluation of segmentation results, the model set the minimum IoU of 0.3 as the matching criterion. Our study defines the number of real instances that match the prediction result as MT, the instances that are not matched by the prediction result as NMT, and the subset of unmatched real instances extracted from the prediction result as NMP. Unlike NMT, NMP retains predictions that have a certain connection with a certain GT, which may be false detections. Completeness, Commission error, Omission Error, and F1-score are used as the tree instance result detection evaluation indicators (
Table 3).
Tree instance result segmentation evaluation metrics To evaluate segmentation results at the instance level, the study first considers evaluation criteria based on true predictions (TP), false negative predictions (FN), and false positive predictions (FP), as defined by Fu et al. [
53]. The segmentation results are defined as three types. If a tree is correctly segmented from the woodland point cloud data, it is called a true positive (TP). A tree is classified as a false negative (FN) when most of its constituent points are not correctly grouped into a single instance, but are instead assigned to one or more neighboring trees. When an instance does not actually exist but is separated from a true instance, it is called a false positive (FP). After obtaining the basic prediction results, our research applies the detection rate
r (recall) and accuracy
p (precision) as the individual-tree segmentation evaluation metrics at the instance segmentation. To extend the evaluation beyond the detection of tree instances, we adopt the coverage metric proposed by Xiang et al. [
40], a measure that quantifies how accurately the predicted instance boundaries align with those of the ground truth. In more detail, given a set of ground truth trees
and a set of predicted trees as
, we compare the two sets on a per-instance basis. For each ground truth tree, the predicted instance achieving the maximum Intersection-over-Union (IoU) score is identified:
The coverage range is then defined as the average of the maximum IoU derived from all real instances. Its corresponding mathematical expression is provided in
Table 3.
4. Results and Analysis
This section presents a comprehensive experimental analysis, structured to validate the proposed method from multiple perspectives. We begin with implementation details (
Section 4.1) and baseline comparisons (
Section 4.2.1 and
Section 4.2.2), followed by evaluations against state-of-the-art methods (
Section 4.3) and ablation studies (
Section 4.4). To assess practical applicability, we analyze computational efficiency and memory usage (
Section 4.5), noise robustness (
Section 4.6), and parameter sensitivity (
Section 4.7). Finally, we conduct a comparative analysis of forest structural metrics between raw and processed data (
Section 4.8) to verify the data fidelity of our synthetic pipeline.
4.1. Implementation Specifications
The computational platform for all experimental evaluations consisted of an Intel Core i7-14700KF processor (16 cores) from Intel Corporation (Santa Clara, CA, USA), configured with 16 GB RAM per core, coupled with an Nvidia RTX 4080 graphics card featuring 16 GB dedicated memory manufactured by NVIDIA Corporation (Santa Clara, CA, USA). For all datasets, the search radius parameters for verticality were calibrated using their respective validation sets.
4.2. Comparison Experiment with Baseline
To verify the effectiveness of our optimized approach, we conduct comparisons with baseline, including quantitative performance evaluation on key metrics and visual analysis of segmentation results across various forest types.
4.2.1. Quantitative Results
In order to research the influence of the sparse convolutional neural network improved by Mamba, the HDBSCAN clustering algorithm, and the W-KNN neighborhood search algorithm on the refined tree instance segmentation method of the local forest, this study conducts a comprehensive evaluation across seven local forest datasets, comparing our proposed Mamba-enhanced Sparse Convolutional Neural Network for point cloud instance segmentation with the conventional TreeLearn pipeline approach.
As demonstrated in
Table 4, our optimized method demonstrates superior performance over the TreeLearn pipeline across multiple benchmark datasets, as evidenced by both instance detection and instance segmentation evaluation metrics. Furthermore, the reported mean values and standard deviations from three independent trials corroborate the robustness of these findings. The consistently low standard deviations across most datasets (e.g., C, OE, and F1 scores typically exhibiting SD < 2.0%) indicate that the performance of our method is stable and not contingent upon a specific favorable initialization. This reproducibility strengthens the validity of the observed performance advantages. In addition, the experimental results based on the best overall result demonstrate robust performance across six local forest plots, achieving a callback rate (r) exceeding 90%, classification accuracy (p) above 84%, F-score surpassing 74%, and coverage (Cov) greater than 82%, confirming the model’s prominent instance segmentation capability. However, the best results from both the TreeLearn model and our optimized method exhibit higher omission errors (8–15 percentage points) in the TUWIEN and RMIT regions compared to the artificial tall tree local forest in the TreeLearn dataset, as well as the CULS, NIBIO, and SCION regions. The Completeness of RMIT instance detection evaluation does not even reach 85%. The best result on the TUWIEN dataset exhibits a significantly higher commission error (40%) compared to both the dense local forests in the TreeLearn and Merge-Forest benchmark datasets. This performance discrepancy can be attributed to two key factors: (1) the higher density of local forest data in these regions; (2) significant variations in tree height distributions and species composition between TUWIEN and Merge-Forest samples. However, the high mean commission error (M = 35.1%, SD = 6.6) also exhibits considerable variability, suggesting that the model’s performance on this challenging, dense forest is more sensitive to initial conditions compared to other datasets. Correspondingly, our model’s best results demonstrate salient performance on the TreeLearn dataset, CULS, NIBIO, and SCION regions. Notably, the best result achieves perfect segmentation (0% omission error) on the relatively sparse CULS data with well-defined tree structures, indicating complete semantic recognition of all point cloud instances. This perfect segmentation is consistently achievable, as reflected by the negligible standard deviation (M = 99.6%, SD = 0.6 for C) across runs. A comparison of the overall best results in
Table 4 also reveals that, compared with the baseline model, our model demonstrates significant improvements in segmentation accuracy for the low-canopy RMIT woodland dataset, achieving a 3.8 percentage point increase in F1-score and a 4.9 percentage point enhancement in precision compared to baseline methods. These results demonstrate the efficacy of our optimization model in addressing the challenging task of instance segmentation and accurate assignment within low-canopy woodland environments. The specific reasons can be referred to in
Section 4.5. Consistent improvements are observed in the best results across the CULS, NIBIO, TUWIEN, SCION, and Merge-Forest datasets, with our optimized model showing measurable gains (typically ± 1 percentage point) in both instance detection and instance segmentation evaluation metrics. While the best result on the TUWIEN dataset exhibits a slight precision reduction, it achieves a substantial 4 percentage points improvement in commission error and F-score. Similarly, while the best result from our network exhibits marginal decreases in callback rate (r) and coverage (Cov) on the TreeLearn dataset, it achieves substantial improvements in F1-score and precision—the two metrics that most critically reflect segmentation quality.
In contrast to baseline methods, our approach preserves all segmentation results without post-processing pruning across benchmark datasets. This strategy maximizes retention of ecological information within woodland areas, despite potential edge segmentation imperfections. The comparative analysis of means and standard deviations further reveals that our optimized model not only achieves higher average performance but also demonstrates comparable or reduced performance variance relative to the TreeLearn pipeline (e.g., lower SD in F1-score for RMIT and Merge-Forest). These results, obtained from experiments trained over 500–1000 rounds, indicate a consistent enhancement in both the accuracy and reliability of instance tree recognition, with particular improvements in edge forest feature extraction capabilities.
4.2.2. Visual Analysis of Experimental Results
To visualize its optimization by our model more intuitively,
Figure 7 combines the box plot and normal distribution curves to show the instance-level visualization results for all local forests. The visualization integrates boxplots, jitter points, and normal curves to comprehensively reflect the performance distribution: (1) boxplots represent the interquartile range (25th–75th percentiles) of metric values; (2) the horizontal line within each box denotes the median; (3) whiskers extend to the maximum and minimum values excluding outliers; (4) jitter points show the raw distribution of individual instance data, including outliers; (5) normal curves illustrate the overall distribution trend and probability density of each metric. The figure shows that the segmentation effect of forest inventory is negatively correlated with the number and density of trees in the local forest.
For individual tree segmentation within the same local forest, baseline results reveal fundamental challenges posed by intersecting branches and occlusion effects. Our optimized model demonstrates significant improvements in addressing these complex segmentation scenarios.
Figure 8 is the resulting visualization of the simulated forest point cloud in the FOR-instance dataset. Among these datasets, the trees in the CULS dataset are relatively sparse, and the stems and leaves are distinct. The species in the RMIT dataset exhibit the lowest height. The Merge-Forest and TUWIEN datasets show obvious height differences and densely interleaved trees. However, the SCION and NIBIO datasets have more regular tree planting and better overall segmentation results. From the model comparison results in
Figure 7, the denser the box plots and normal curves of the corresponding indicators, the better the overall segmentation effect of the instances. Similarly, the fewer the outliers of the curves, the more accurate the segmented instances. In summary, as shown in the individual tree segmentation results, our optimization method demonstrates significant reductions in errors (or convergence) across the CULS, NIBIO, RMIT, Merge-Forest, and TreeLearn datasets. This indicates that our model effectively addresses the original network’s limitations in recognizing challenging forest scenes, including low-canopy height (RMIT), dense vegetation (SCION, TreeLearn), and structurally complex environments (Merge-Forest). Notably, our methodology enhances the original TreeLearn framework through significant improvements to its offset prediction component. The baseline model defines a “tree base” with a height of 3 m, as detailed in the original paper [
38]. This means that its pipeline has a considerable error in the identification of classes, such as RMIT low shrubs, resulting in no reference for the experiment. It has been proven that the model uses the offset prediction of
Section 3.3, which is more suitable for different complex scenes and improves the accuracy of instance segmentation.
4.3. Benchmark Comparison with Existing Methods
Table 5 extends the comparative analysis by distinguishing between Ours (Raw data) and Ours (Processed data) on the FOR-instance dataset. This subdivision enables a two-fold analysis: benchmarking our core model against state-of-the-art methods via raw data, and quantifying the pipeline’s impact through raw/processed data comparisons. It is important to note that, due to limitations in the network evaluation methodology and the constraints of open-source access, we are only able to compare with the corresponding network on select parameters.
On the TreeLearn dataset, SegmentAnyTree has a slight lead in evaluation metrics such as Completeness on the TreeLearn dataset. However, the 2% improvement in F1-score indicates that our model has greater flexibility and better overall performance in instance detection or segmentation. This trade-off is attributed to the dataset’s dense European beech stands, where overlapping canopies and ambiguous inter-tree boundaries pose inherent challenges. Our method’s advantage in instance delineation stems from the multi-dimensional attention mechanism of Mamba, which effectively models long-range feature dependencies in 3D point clouds, overcoming the limitations of conventional sparse CNNs in capturing complex canopy structures. Similarly, as previously noted, the 0.5 percentage point completeness drop versus SegmentAnyTree is exacerbated by the irregular crown shapes of mature beech trees. Many of these trees have sparse outer foliage and fragmented canopy edges, where the network’s attention mechanism fails to prioritize sparse boundary points. Unlike SegmentAnyTree, which may use more aggressive boundary expansion, our method’s focus on dense core features leads to the under-segmentation of these marginal regions, highlighting a trade-off between precision in core instance delineation and completeness in boundary regions.
The raw data results on FOR-instance reflect the intrinsic capability of the Mamba-Enhanced Sparse CNN to handle unprocessed forest point clouds. Across all datasets, the model demonstrates competitive or leading performance in key metrics, with notable strengths and limitations. On CULS, it matches the completeness and omission error of the top competing methods while exceeding ITS-Net in F1-score by 2.4 points. This highlights the effectiveness of our optimized HDBSCAN and W-KNN clustering in refining instance assignments for well-structured coniferous forests. On the relatively dense coniferous forest NIBIO dataset, it surpasses ITS-Net by over 3.4 points in completeness, with an F1-score improvement of 4.4 points. It is shown that our network effectively captures the complex structure of coniferous forests, achieving exceptional accuracy in individual tree segmentation. For the high-resolution and informative TUWIEN plots, the model addresses severe under-segmentation issues of existing methods, with precision improved by 3.8 points over HFC and the F1-score up by 10.3 points. This breakthrough comes from Mamba’s ability to capture multi-scale spatial features, combined with our robust clustering strategy. On the low-canopy RMIT dataset, our model achieves superior completeness, exhibiting a detection rate 16.3% higher than SegmentAnyTree and 16.5% higher than ITS-Net. This improvement correlates with the inclusion of our offset prediction module, which was specifically designed to mitigate the prevalent under-segmentation challenges in low-canopy vegetation.
However, the SCION dataset consists of non-native pure coniferous temperate forests with uniform tree spacing and regular canopy structures. Our raw data method achieves 100% completeness, 0% omission error, but a lower F1-score (83.3%) compared to ITF-Net (92.7%). This 9.4 percentage point F1-score drop is a notable limitation caused by the network’s over-complexity for highly structured datasets. ITS-Net outperforms rule-based methods in the SCION dataset by leveraging its dual-coordinate positional embedding and global-view feature encoding modules, which capture nuanced structural variations and contextual relationships even in forests with uniform spacing and regular canopies. In contrast, our method’s Mamba-enhanced feature extraction and clustering pipeline over-extracts features, leading to minor over-segmentation of uniform canopies (e.g., misclassifying small foliage variations as separate instances) and thus reducing the F1-score. This limitation reflects the network’s inability to adapt to overly structured scenarios where simpler rule-based methods outperform complex deep learning approaches.
The results of Ours(Processed data) in
Table 5 represent the performance of our synthetic data pipeline (
Section 3.1) in test set fusion evaluation. This pipeline was specifically designed to address the FOR-instance dataset’s limitations, including geographically imbalanced samples, insufficient instance counts, and a lack of block coherence. Importantly, these processed data results validate the pipeline’s effectiveness in mitigating limitations and enhancing model generalization. The purpose of our pipeline, as an explicit data-augmentation method, is to make the FOR-instance dataset expandable and aligned. It provides an integrated evaluation of both the data-augmentation procedure and the test set. In most forests, the results are equivalent to those on the original dataset (in CULS and NIBIO, the metrics are slightly lower than the raw-data results).
However, compared with the raw data, there are still some differences in the processed data results. On the RMIT dataset, our processing pipeline yielded the most substantial performance gains, elevating the F1-score and recall by 6.8 and 6.6 percentage points, respectively. This improvement can be attributed to the pipeline’s effective noise suppression, which enhances the distinction between low-canopy trees and understory shrubs. On SCION, processed data leads to a substantial 13.6 percentage point F1-score improvement, which, to some extent, addresses the raw data’s performance drop versus HFC. The synthetic pipeline generates more uniformly structured tree instances with consistent spacing and canopy shapes. This reduces over-segmentation and false positives, aligning the network’s performance with the simplicity of the dataset. In contrast to previous findings, on TUWIEN, processed data shows an F1-score decline of 18.3 percentage points and a significant increase in commission error (37.5%), but this is a deliberate and meaningful trade-off. The synthetic pipeline simulates more complex, dense mixed-species stands with extreme canopy overlap—scenarios that are underrepresented in the original dataset. While this leads to an increase in false positives during the test set fusion evaluation, this trade-off is instrumental in enhancing the model’s generalization capability to dense forest environments beyond the scope of the benchmark.
4.4. Ablation Experiment
To systematically evaluate the contribution of individual model components to instance segmentation performance, we conduct comprehensive ablation studies across three representative forest datasets: the low-canopy RMIT dataset
Table 6(a), the dense canopy crossing TUWIEN dataset
Table 6(b), and the hybrid heterogeneous Merge-Forest dataset
Table 6c. These datasets collectively represent the spectrum of segmentation challenges in real-world forestry applications. As shown in
Table 6, the experimental results reveal distinct component contributions under different forest structural conditions.
Improved offset prediction method: Our offset prediction module (Method IV, V, VI) consistently delivers the most significant performance improvements across all three forest types. In the low-canopy RMIT environment, Method IV boosts the F1-score by over 26 points and precision by the same margin, establishing it as the most effective single-component enhancement for scenarios where conventional height-based approaches falter. For the dense, overlapping canopies of TUWIEN, the module raises completeness substantially from 72.7% to 88.3%, though the concurrent rise in commission error highlights a limitation in highly entangled scenes where spatial cues alone can be ambiguous. Most notably, in the heterogeneous Merge-Forest, Method IV attains the highest completeness and lowest omission error among all single-module variants, while also leading in both F1-score and precision. This outperforms the Mamba-only and clustering-only configurations, which exhibit either lower completeness or higher error rates. The results collectively affirm that the proposed offset mechanism provides a robust and universally applicable spatial anchor, adept at capturing tree centers across diverse forest structures from sparse low-canopy to dense mixed stands. This demonstrates that the component’s lower-level architecture and optimized configuration significantly enhance segmentation performance, effectively resolving a critical challenge in current research: accurate segmentation of low-canopy forest datasets.
Clustering post-processing method: The HDBSCAN + W-KNN clustering component (Method II, III, VI, VII) demonstrates complementary strengths that vary markedly with forest density. In the dense TUWIEN dataset, the standalone clustering module increases completeness but at a severe cost: commission error skyrockets from a baseline of 3.7% to 57.8%, causing the F1-score to plummet by nearly 30 points. This reveals a critical weakness in applying density-based clustering alone to overlapping canopies, where it tends to erroneously merge adjacent crowns. However, when paired with offset prediction (Method VI), the synergy proves transformative across all datasets. On the low-canopy RMIT, this combination achieves an F1-score of 89.1% and precision of 84.4%, significantly outperforming offset prediction alone. Similarly, in the complex Merge-Forest environment, the pairing delivers well-balanced metrics with 91.3% completeness and 74.9% F1-score. These results underscore that while clustering alone can be detrimental in dense conditions, it becomes a powerful refinement tool when guided by accurate spatial anchors, enabling precise point reassignment and boundary delineation. The improvement stems from the component’s ability to leverage reliable reference data, effectively addressing segmentation challenges caused by structural complexities, such as uneven canopy heights and branch occlusion.
Mamba-Enhanced module: The Mamba module (Method I, III, V, VII) provides a distinct form of enhancement centered on global context integration rather than spatial precision. on the low-canopy RMIT dataset, its most consistent contribution is in recall improvement, with values consistently exceeding 94% in configurations that include Mamba. This demonstrates its core strength in minimizing under-segmentation by capturing long-range dependencies among tree structures. In the challenging TUWIEN environment, Mamba alone achieves 80.3% completeness while keeping commission error at a moderate level, outperforming the clustering module in this dense setting. When integrated with offset prediction (Method V), the combination excels at capturing complex forest patterns, achieving 90.2% completeness on TUWIEN and the highest completeness of 92.0% on Merge-Forest among all configurations. However, this comes with elevated commission error in some cases, revealing that while Mamba excels at finding tree structures, it benefits from additional constraints for precise boundary definition. The module’s value is particularly evident in the full model (Method VII), which maintains excellent completeness (84.3% on RMIT, 90.8% on TUWIEN, 92.6% on Merge-Forest) while controlling error rates through complementary component integration. This confirms that the Mamba module effectively addresses the fundamental limitation of conventional sparse CNNs in modeling long-range dependencies, enabling the network to maintain structural coherence across fragmented point clouds and complex canopy arrangements where local receptive fields prove insufficient.
Component synergy and dataset-specific adaptations: The full model (Method VII) achieves optimal or near-optimal balance across all datasets, but the relative component contributions reveal important patterns. For low-canopy forests like RMIT, the combination of offset prediction and clustering (Method VI) nearly matches the full model’s performance, suggesting that precise localization and point assignment are paramount in these environments. In contrast, for dense overlapping canopies like TUWIEN, all three components are essential to achieve both high completeness (90.8%) and manageable error rates. The heterogeneous Merge-Forest presents a more complex picture, where different component combinations excel at different metrics, with the full model providing the most balanced overall performance. This analysis demonstrates that our modular architecture enables adaptive optimization based on forest characteristics while maintaining robust performance across diverse conditions, offering practical flexibility for real-world forest inventory applications.
4.5. Computational Efficiency and Memory Usage Analysis
To evaluate the practical applicability of our method,
Table 7 compares its computational efficiency and memory usage against the TreeLearn baseline across all seven forest datasets. As summarized in
Table 7, our method generally achieves lower average CPU memory utilization, for example, 17.8 percent versus 21.7 percent on the TreeLearn dataset, and 13.6 percent versus 14.0 percent on CULS, while maintaining comparable peak GPU memory consumption. This reduction in average memory footprint can be attributed to the algorithmic design of HDBSCAN employed in our pipeline. Unlike the conventional DBSCAN (Density-based spatial clustering of applications with noise) used in the baseline, which requires a global density threshold and may perform redundant neighborhood queries in sparse regions, HDBSCAN adopts a hierarchical, fixed-scale approach. This design inherently constrains memory consumption by avoiding the dynamic expansion of search radii and the associated overhead of maintaining large adjacency matrices.
Although the fixed neighborhood scale of HDBSCAN leads to more serial processing and thus longer runtime, for example, 29.3 versus 26.5 min on TreeLearn and 116.4 versus 101.2 min on SCION, it provides finer-grained and less memory-intensive clustering. This is especially beneficial in dense point clouds where excessive parallelism can cause memory spikes. Notably, on the densest NIBIO and SCION plots, our method maintains stable GPU memory usage while slightly increasing computation time, reflecting a trade-off between segmentation precision and processing speed. Overall, the results confirm that our approach offers a favorable balance of memory efficiency and robustness, suitable for real-world forest inventory where hardware resources may be constrained.
4.6. Noise Robustness Evaluation
To further assess the practical applicability of our model in real-world scanning environments, we conduct a noise robustness evaluation, with the results illustrated in
Figure 9. The RMIT test set, characterized by low canopy and a complex, disorganized structure, was selected for this experiment due to its high sensitivity to scanning artifacts and point cloud imperfections, making it an ideal benchmark for evaluating model stability under adverse conditions.
We simulate two common types of data corruption: point perturbation and point deletion. For perturbation, independent Gaussian noise is added to each point in the point cloud set , such that , where is a zero-mean Gaussian noise vector with covariance matrix (I denotes the identity matrix, and is the noise standard deviation). Here, is set proportionally to the bounding box diagonal of the input cloud to maintain scale consistency. For deletion, we perform Bernoulli sampling on each point with a retention probability , where r is the point deletion ratio.
The results in Figure demonstrate distinct characteristics of our model’s robustness. First, under Gaussian perturbation, our method maintains precision above 80% even at a 50% noise level, whereas TreeLearn drops below 75%. This resilience highlights the effectiveness of our offset prediction mechanism, which reinforces spatial consistency by guiding points toward tree centers, thereby mitigating the effect of positional jitter. Second, the model shows greater sensitivity to point deletion than to perturbation: under 50% deletion, precision declines more sharply. This suggests that while our architecture can effectively smooth local noise, the removal of structural points directly impacts the density cues critical for HDBSCAN clustering and offset calculation. Nevertheless, across all noise levels in both settings, our model consistently surpasses TreeLearn, with the performance gap widening as noise increases. Even under an extreme 90% point deletion rate that completely prevented TreeLearn from splitting instances, our method still maintained a segmentation accuracy of 23.9%. This demonstrates that our Mamba-enhanced feature extraction preserves robust global dependencies even under substantial data loss, offering a clear advantage for processing incomplete or highly corrupted real-world scans.
4.7. Parameter Sensitivity Analysis
We conduct a systematic sensitivity analysis on the W-KNN neighborhood size K. Four representative datasets are evaluated: low-canopy RMIT, structured plantation CULS, dense deciduous TUWIEN, and mixed-species Merge-Forest, covering key structural variations encountered in operational forest inventories.
As shown in
Table 8, segmentation precision remains stable across a wide range of
K values (5–30), demonstrating the robustness of the clustering framework. In structured plantations (CULS), performance is nearly invariant, indicating minimal sensitivity to neighborhood size in uniform canopies. Conversely, in complex low-canopy (RMIT) and dense deciduous (TUWIEN) environments, precision improves with moderate
K value (peaking at
–20), as larger neighborhoods provide sufficient local context for distinguishing adjacent trees in overlapping vegetation. Beyond
, further increases yield diminishing returns while linearly increasing computational cost.
Based on this empirical analysis, we select as the optimal balance between accuracy and efficiency. This value provides adequate contextual information for robust clustering across diverse forest structures, while avoiding the computational overhead and potential over-smoothing associated with excessively large neighborhoods.
4.8. Comparative Analysis of Forest Structural Metrics Between Raw and Processed Data
To verify that our simulated forest point cloud synthesis pipeline (
Section 3.1) preserves critical tree structural characteristics, while also assessing potential differences in forest complexity, we conducted a quantitative comparison between raw and processed data. Specifically, we focused on three key structural metrics: the trunk-to-canopy height ratio for instances, and two plot-level forest complexity measures: canopy overlap rate and foliage height diversity (FHD). Each metric was purposefully designed based on established principles in forest remote sensing literature to reflect meaningful ecological differences in forest structure.
For each instance, we compute a robust trunk-to-canopy height ratio. Let
and
denote the sets of vertical coordinates for stem and canopy points, respectively. The trunk height
is defined as
, and the canopy height
as
, where
denotes the
k-th percentile. The ratio
is then calculated. This percentile-based approach, following [
60], minimizes the influence of outlier points in the point cloud.
The canopy overlap rate quantifies horizontal canopy complexity by measuring the intermingling of adjacent tree canopies. For a given plot, let
represent the set of canopy points with their corresponding tree ID
t. These points are projected onto a horizontal grid. The grid cell size
r is determined adaptively based on the local 2D point density
(points per m
2) following the empirical approach in [
61], where higher density areas use finer grids to capture detailed structure while sparser areas use coarser grids to maintain spatial coherence. Specifically, we set
m for
pts/m
2,
m for
pts/m
2, and
m otherwise, balancing resolution and computational efficiency as established in prior studies. A grid cell is considered valid for analysis only if it contains a number of points exceeding a minimum threshold that scales with
to maintain statistical reliability. The overlap rate is then computed as
, where
is the number of valid grid cells containing points from two or more unique tree IDs, and
is the total number of valid grid cells.
Foliage height diversity (FHD) assesses the vertical heterogeneity of the canopy. For each plot, we first estimate the ground elevation as the 5
th percentile of points classified as ground. The relative height above ground for each canopy point is calculated as
, where
z is the point’s absolute height, and
is the ground elevation estimate. To exclude extreme outlier heights, we define the effective maximum relative height as
, the 95th percentile of all relative heights. The relative height range from 0 to
is divided into
equally sized vertical bins. Each canopy point is assigned to a bin based on its
h value, producing a histogram of point counts
across bins. For bins with
, we compute the relative frequency
. The FHD is then calculated using the Shannon entropy formula:
, normalized by the maximum possible entropy
where
is the number of non-empty bins. The approach is adapted from methods used in LiDAR-based canopy structure analysis [
62].
The results in
Table 9 demonstrate that, for most forests (CULS, NIBIO, RMIT, SCION), the trunk-canopy height ratio remains largely consistent between raw and processed data. This indicates that our synthetic pipeline successfully preserves this critical vertical structural characteristic of individual trees, validating its capability to maintain foundational tree architecture.
A subtle but consistent trend across most datasets is a slight increase in the canopy overlap rate for the processed data. This aligns with the design objective of our synthesis pipeline, which inherently tends to integrate and densify canopy regions during the fusion of multiple tree instances and the simulation of plot-level interactions. The process thus amplifies horizontal canopy intermingling compared to the often more discretely represented original point clouds.
The behavior of foliage height diversity (FHD) is more variable. For some forests like RMIT and SCION, the processed data shows a marginal decrease in FHD. This can be attributed to the nature of their original stands, which may have had relatively concentrated or uniform canopy layers. The synthesis process, while adding complexity, might have preferentially amplified the dominant vertical strata (e.g., the main canopy), slightly reducing the evenness of the vertical point distribution and thus the entropy-based FHD metric.
The TUWIEN dataset presents a notable and deliberate exception, with significant increases across all three metrics. The higher trunk-to-canopy ratio primarily results from the synthesis amplifying the influence of larger trees present in the original data. While the raw TUWIEN point cloud contains trees of highly varying sizes, the processed data accentuates the structural contribution of taller specimens. Simultaneously, the processed data exhibits a substantially elevated canopy overlap rate and FHD. This intentional increase in density and complexity, while making the segmentation task more difficult (as reflected in the commission error decline noted in
Section 4.3, serves the crucial purpose of enhancing model generalization for real-world, dense forest environments not well-captured by conventional datasets.
5. Discussion
5.1. Instance Segmentation Performance in Challenging Forest Scenarios
To further verify the influence of the Mamba-Enhanced Sparse CNN on individual tree instance segmentation in local forests, we use visualizations of simulated local forest dataset slices from the TreeLearn dataset and FOR-instance as examples.
In
Figure 10, the comparison of instance segmentation results between our model and the baseline model is shown. To demonstrate our model’s improvements, the experiments select representative samples from all benchmark datasets containing challenging forest scenarios: edge regions, dense canopies, and areas with frequent tree crown intersections. Due to the lack of context information and the dense intersections between tree points, these slices are located at the edge of the forest, which easily leads to point misassignment. Both our optimized model and the TreeLearn model have some errors in the segmentation of tree instances along the edges. Overall, our model is more robust and can correctly segment all tree instances in each dataset.
Specifically, on the TreeLearn dataset, which has the widest woodland area and the densest trees, our model only has marginal outlier errors on instances 1, 3, and 4. In contrast, the baseline suffers from pervasive adjacent instance confusion: extraneous stems/branches are misassigned to six instances, with massive cross-instance outliers (except instance 1), and instances 5/6 are heavily contaminated by structures from other trees. This gap may stem from the limited global feature modeling capability of conventional sparse CNNs. To mitigate this issue, our model leverages the selective state-space modeling principle of the Mamba architecture, thereby improving its capability for dynamic long-range dependency tracking.
On the sparsest and most regular CULS dataset, our model has a minimal number of assignment errors on instances 1, 4, and 5, where instance 5 incorrectly assigns the bottom trunk part from instance 4. However, in addition to the assignment error for instance 5, the baseline model incorrectly segments two distinct trees as four separate instances: instances 1, 7, and instances 6, 8 should be treated as two separate ones. This highlights the baseline’s difficulty in preserving 3D structural coherence. To mitigate this challenge, our model employs the multi-dimensional feature coupling inherent in the Mamba architecture to better align vertical trunk and horizontal canopy features.
On the densely planted NIBIO dataset, the optimized model only has significant outliers on instances 1, 3, and 4, while the baseline model incorrectly divides one instance of one tree into two (instances 6, 7). Compared with the optimized model, the outliers in instances 1 to 4 are more serious. Instance 1 even has large blocks of outliers that belong to instance 2.
On the height-heterogeneous TUWIEN dataset, our model has minor misallocation on instances 1, 3, 4, 6, while the baseline has three critical failures: instance misassignment, target misidentification, and extreme over-segmentation (one tree split into four instances: 4/7/8/9). These results validate the capability of our Mamba-based model to adapt to height-induced feature variations. This addresses a key shortcoming of sparse CNNs, which are limited by their local receptive fields. These results suggest that the architecture of Mamba may help adapt to height-induced feature variations, an area where sparse CNNs, due to their local receptive fields, often show limitations. Furthermore, our integration of HDBSCAN for hierarchical density clustering and W-KNN for post-processing enhances performance in high-density forests. This approach thereby addresses the limitations of fixed cluster sizes in traditional DBSCAN and KNN, which can cause over-clustering (tree merging) or under-clustering (tree separation).
The low-canopy RMIT dataset presents a significant challenge for tree structural feature extraction. Our model incurs only minor misassignment and missing-point errors on instances 2 and 6. However, experimental results reveal that the baseline model has significant performance limitations on this dataset. The segmentation results exhibit two distinct error types: first, the complete absence of correct instance partitioning; second, the misclassification of three low-canopy trees as a single instance (instance 3), along with the erroneous division of one tree into two separate instances (instances 4 and 5, 6 and 7). This notable performance difference suggests that our offset mechanism is particularly effective for low-rise trees. Furthermore, the improved results can be attributed to a combination of factors: The long-range feature extraction enabled by Mamba enhances context understanding, while the subsequent HDBSCAN clustering and W-KNN refinement collectively contribute to more rational point cloud allocation, especially at object boundaries.
On the SCION dataset, where the trees are arranged regularly and neatly, our model only has assignment errors on structures of instances 2 and 3. The baseline persists in exhibiting redundant instance assignments, particularly between instance pairs (2, 7), (5, 9), and (6, 8), indicating unresolved challenges in instance differentiation.
In summary, our Mamba-Enhanced Sparse Convolutional Neural Network demonstrates superior performance in edge-region segmentation in local forest environments, showing particular effectiveness in capturing instance-level features from context-deficient point cloud data. In the comparison of instance segmentation results for closely intertwined trees, which are common in local forests, our optimized model can clearly reduce or even solve incorrect instance assignments. This means that the model introduces a multi-dimensional attention mechanism through Mamba, which can capture and process multi-dimensional point cloud information at the same time. The designed offset mechanism contributes to improved segmentation across various forest environments, with notable gains in low-canopy regions. Within our framework, its synergy with HDBSCAN and W-KNN specifically targets key challenges, including overlapping tree crowns and segmentation errors arising from significant height variations.
5.2. Spatial Directional Accuracy of Instance Segmentation
Figure 11 compares the segmentation accuracy of our model and the TreeLearn baseline along the horizontal (x, y) and vertical (z) axes. It shows that for forest edge point clouds, the horizontal accuracy exhibits a progressive decreasing trend for both networks—a universal challenge due to context deficiency at edges. However, our Mamba-enhanced sparse convolutional neural network demonstrates diversified accuracy improvements in both vertical (z-axis) and horizontal (x/y-axis) directions across the seven datasets. In general, our model outperforms the baseline in vertical and horizontal evaluations for all datasets except TUWIEN, with notable gains on CULS, RMIT, Merge-Forest, and NIBIO datasets.
Specifically, in the vertical direction, for the TreeLearn dataset, our model maintains comparable superior overall precision to the baseline while delivering superior vertical feature extraction performance. This improvement can be attributed to the selective state-space model (SSM) mechanism in Mamba, which excels at capturing long-range vertical dependencies. The capability is critical for capturing continuous trunk structures from base to canopy.
The Merge-Forest dataset reveals a distinctive performance trend. Our network far exceeds the baseline in early stages but gradually converges to it from the L3 position onward, ultimately sustaining a leading edge. This pattern may suggest that the dynamic long-range dependency of Mamba tracking gradually accumulates discriminative vertical features as slice depth increases, overcoming the baseline’s inherent limitation of local receptive fields.
On the CULS dataset, the proposed network demonstrates superior and more stable accuracy in the vertical dimension compared to the baseline. This improved vertical consistency indicates a stronger capability to delineate individual tree instances across varying heights, thus contributing to more robust and stable overall segmentation outcomes.
NIBIO dataset experiments show that our network exhibits noticeably less accuracy fluctuation and a marginal overall advantage over the baseline. This stability gain likely stems from the synergy between Mamba’s multi-dimensional feature coupling and hierarchical clustering of HDBSCAN, which enhances vertical feature modeling robustness in densely planted forest scenarios.
On the TUWIEN dataset, where tree heights exhibit significant heterogeneity, our network demonstrates a consistent smoother and more stable accuracy across all spatial positions. This result indicates that the architecture of Mamba may be better equipped to handle height-induced feature variations, a known limitation for baseline models with local receptive fields.
Consistent vertical accuracy improvements are also observed for our method on the RMIT dataset, a low-canopy scenario where weak trunk features pose major segmentation hurdles. These gains may stem from the combined effect of the long-distance feature extraction of Mamba and the designed offset mechanism, both of which boost the detection of subtle vertical features in sparse environments.
Finally, the SCION dataset brings competitive performance to the enhanced network as it exhibits slightly higher accuracy than the baseline in most spatial locations. This indicates that in regularly arranged dense artificial forests, our model better guarantees the segmentation accuracy.
In the horizontal direction, our optimized method outperforms the baseline overall, except for TUWIEN, which is basically flat. This suggests that our model’s integration of the multi-dimensional feature coupling of Mamba and W-KNN post-processing helps refine horizontal canopy boundary delineation by enhancing cross-dimensional feature coherence. However, the inferior performance on TUWIEN implies that our current network still has room for improvement in horizontal spatial information capture and feature extraction. Although the combined algorithm (M3Unet with HDBSCAN and W-KNN) alleviates some existing segmentation limitations, it does not establish a decisive advantage in handling highly complex forest landscapes, especially when available training data are limited.
5.3. Implications and Limitations
Table 10 systematically contrasts our model with three state-of-the-art approaches (2024–2026), highlighting core differences in terms of advantages and limitations. The key implication of our method lies in resolving the long-standing challenge of balancing global feature modeling and local detail capture in forest point cloud segmentation. This distinguishes it from prior work: ITS-Net focuses on sensor-agnostic feature learning, SegmentAnyTree on cross-platform adaptability, and RsegNet on rubber species-specific optimization.
Unlike ITS-Net reliance on global-view feature encoding and dual-coordinate positional embedding to handle geometric variations, our model leverages Mamba’s SSM to capture long-range feature dependencies, effectively addressing crown overlap and ambiguous boundaries—issues that the memory-enhanced versions of ITS-Net and ScoreNet still struggle to fully resolve in highly overlapping canopies. Our segmentation offset mechanism, tailored for diverse forest scenes, mitigates the performance degradation observed in ITS-Net in sparse point-density environments (e.g., RMIT with 498 pts/m2), where ITS-Net exhibits lower detection rates and higher omission errors due to under-segmentation in low-density canopies. Additionally, our global-local feature fusion strategy enhances adaptability to structural heterogeneity (mixed coniferous/deciduous forests), whereas ITS-Net faces higher commission errors in complex canopy scenarios.
In contrast to SegmentAnyTree focus on sensor-agnostic adaptation, our model prioritizes segmentation accuracy in complex structural environments over cross-platform scalability. While SegmentAnyTree excels in computational efficiency for large-scale scenes, it suffers from over-segmentation on irregular crowns and insufficient discrimination in overlap regions. The limitations of our Mamba-enhanced feature extraction are mitigated by modeling fine-grained spatial relationships. While SegmentAnyTree achieves its versatility via random subsampling augmentation that simulates sparse ALS data from dense ULS/MLS inputs, our model prioritizes segmentation precision in complex forest structures (low-canopy, dense crown overlap) over cross-platform flexibility, which we will consider in the future.
The primary strength of RsegNet lies in specialized rubber tree segmentation via cosine similarity and dual-channel clustering, but its narrow applicability and poor generalization to mixed-species forests contrast with our model’s robustness across diverse forest types. Our approach avoids species-specific constraints, making it suitable for plantation, natural, and mixed forests, while RsegNet employs a specialized dynamic clustering approach tailored to rubber tree plantations, which limits its generalizability to other forest types.
Our model faces two main limitations: restricted cross-platform transferability and performance degradation in highly complex forest landscapes with scarce training data. The cross-platform limitation arises as the model learns platform-specific feature distributions during training. To address this, future work will focus on developing domain-adaptive training strategies that enable model transfer across different LiDAR platforms (e.g., ALS, MLS, TLS) with minimal labeled data, and exploring self-supervised pre-training on multi-platform point cloud datasets to learn platform-invariant representations. These approaches aim to eliminate retraining requirements and enhance model adaptability to diverse data sources. Meanwhile, performance in complex landscapes suffers because the framework relies on sufficient data to learn intricate feature patterns. Mitigating this will involve augmenting training data through more advanced synthetic generation techniques and optimizing the segmentation pipeline with adaptive clustering parameters. These improvements aim to enhance the model’s practicality while maintaining its performance in heterogeneous environments.
In summary, our core contribution is a versatile segmentation framework that balances global dependency modeling and local feature capture, outperforming existing methods in complex, heterogeneous forest scenes. While ITS-Net, SegmentAnyTree, and RsegNet excel in specific scenarios (platform-agnostic generalization, cross-platform scalability, species-specific segmentation), our model offers a more comprehensive solution for diverse real-world forest inventory needs.
6. Conclusions
This study proposes an individual tree segmentation method for local forest point clouds based on the Mamba-Enhanced Sparse Convolutional Neural Network, which provides an innovative approach for the field of forest resource inventory. The research demonstrates that integrating the Mamba-Enhanced Sparse Convolutional Neural Network with novel offset prediction and clustering methodologies significantly enhances the 3D U-Net architecture, improving its capacity to extract and process high-dimensional point cloud features.
Additionally, we develop a synthetic point cloud generation pipeline to address key limitations in the FOR-instance dataset, enhancing both data diversity and quality for improved model training. After completing the individual tree segmentation process with the improved offset prediction, the HDBSCAN clustering algorithm and the W-KNN neighborhood search algorithm are used to achieve fine instance segmentation of the local forest point cloud. Quantitative evaluations confirm consistent performance improvements across all benchmark datasets. Across multiple forest plots including CULS, NIBIO, RMIT, and SCION, the model achieves prediction precision exceeding 85% while maintaining coverage (Cov) above 80%. Compared with the traditional sparse convolutional neural network model, our model has significant advantages in both instance detection and segmentation evaluation metrics, especially in the RMIT dataset, which has the lowest canopy and is relatively dense, with increases of 3.8 and 4.9 percentage points in F1-score and accuracy, respectively. Our method achieves robust instance segmentation performance across all seven local forest plots, with key metrics exceeding benchmark thresholds: detection rate (r > 90%), precision (p > 84%), F-score (>74%), and coverage (Cov > 82%). Visual analysis further verifies the effectiveness of the model. When handling low-canopy, dense, and complex forest scenes, the model can effectively reduce or eliminate recognition confusion present in the original network, and edge-region segmentation is more reasonable, allowing the model to better capture instance-level information.
While our framework fuses global and local features effectively, it still faces challenges in highly complex forest landscapes when training data is scarce and exhibits limited cross-platform transferability. Future research can be carried out by developing domain-adaptive learning methods to enhance cross-platform generalization, augmenting training data via advanced synthetic generation techniques, and optimizing the pipeline’s adaptive clustering mechanisms to further improve the model’s performance in edge cases and practical deployment. In general, for forest resource inventory management in limited woodland or local forests in different regions, this study provides advanced techniques and groundbreaking approaches for the automation and refinement of forest resource inventory.