1. Introduction
Forest biomass is a fundamental parameter for evaluating ecosystem functioning and resource sustainability [
1]. It is also essential for quantifying forest carbon sequestration, providing baseline information for carbon cycle and storage assessments [
2]. Reliable estimation of aboveground biomass (AGB), which includes stems, branches, foliage, and canopy, is critical at regional scales because it determines overall biomass and carbon dynamics [
3]. Such assessments are vital for addressing global climate change and advancing carbon neutrality and peaking goals [
4].
Remote sensing provides the primary means for large-scale AGB estimation due to its ability to provide spatially continuous observations. Among existing techniques, passive optical data is widely applied. However, optical sensors are limited by their poor penetration of dense canopies, which reduces sensitivity to AGB variation and leads to early saturation of biomass models [
5,
6]. Light Detection and Ranging (Lidar) alleviates this issue through direct canopy profiling, but spaceborne missions remain scarce, and airborne campaigns are costly, restricting their large-scale applicability [
7]. By contrast, Synthetic Aperture Radar (SAR) offers distinct advantages through weather-independent operation and enhanced canopy penetration [
8].
The backscattered signal originates from interactions with multiple layers within the forest, which include leaves, branches, trunks, and the ground surface, thereby facilitating the estimation of vertical structural properties. This makes SAR particularly well-suited for both total and component-level AGB estimation at regional scales. Early SAR studies based on single-frequency, single-polarization data showed limited performance in AGB estimation [
9]. Advances in multi-frequency and multi-polarization SAR systems have since enabled more detailed characterization of forest structure. Shortwave bands (X- and C-band) interact primarily with the upper canopy layers, including leaves and fine twigs [
10], while longwave bands (L-, P-band) penetrate deeper into canopies, capturing larger branches, trunks, and even ground–trunk interactions [
11,
12,
13]. The saturation levels are influenced by factors such as frequency, polarization, and forest structural complexity [
14]. Among these, L-band achieves an effective balance between penetration and structural sensitivity and is widely regarded as the most reliable for biomass retrieval [
15], whereas C-band remains useful for characterizing canopy biomass [
16]. Combining multiple frequencies and polarizations has proven particularly effective in improving AGB estimation accuracy and raising saturation thresholds [
17,
18].
Advancements in SAR technology, transitioning from single-parameter to multi-dimensional configuration acquisitions, have significantly improved the capacity for AGB estimation [
19]. Among SAR features, backscatter coefficients are widely used for AGB estimation. In addition to backscatter coefficients, which are influenced by soil moisture, roughness, and other confounding factors [
20,
21]. Polarimetric decomposition provides improved sensitivity to scattering mechanisms and forest structure [
22]. Interferometric coherence information is strongly correlated with forest height, a critical auxiliary variable for biomass estimation. Previous studies integrating backscatter with interferometric coherence have reported improved accuracy and raised saturation points [
23]. Texture parameters, which characterize canopy heterogeneity, have also demonstrated potential when combined with other SAR features [
24,
25]. However, current research has focused on limited combinations, and few have fully integrated all four SAR dimensions (backscatter, polarimetric decomposition, interferometric coherence, and texture) for AGB modeling. Research specifically targeting biomass components using dual-frequency SAR data also remains scarce [
26].
With the increasing dimensionality of SAR features, appropriate feature selection and modeling strategies have become critical for reliable biomass estimation. Early approaches relied on simple empirical relationships, such as linear or exponential models [
27,
28], but these methods cannot adequately capture the nonlinear relationships between AGB and multi-dimensional SAR features. Multivariate regression improved estimation performance [
29], but suffers from redundancy and overfitting when handling large feature sets. Semi-empirical models [
30] and machine learning approaches have been widely applied in multi-frequency SAR-based AGB estimation, where the latter have demonstrated strong predictive potential [
31,
32,
33]. Random Forest (RF), in particular, has proven effective in handling high-dimensional datasets through ensemble learning and feature importance ranking. Genetic Algorithms (GA) provide complementary advantages by optimizing feature selection and parameter tuning [
4,
34], and their integration with RF has demonstrated significant improvements in high-dimensional AGB estimation [
35,
36].
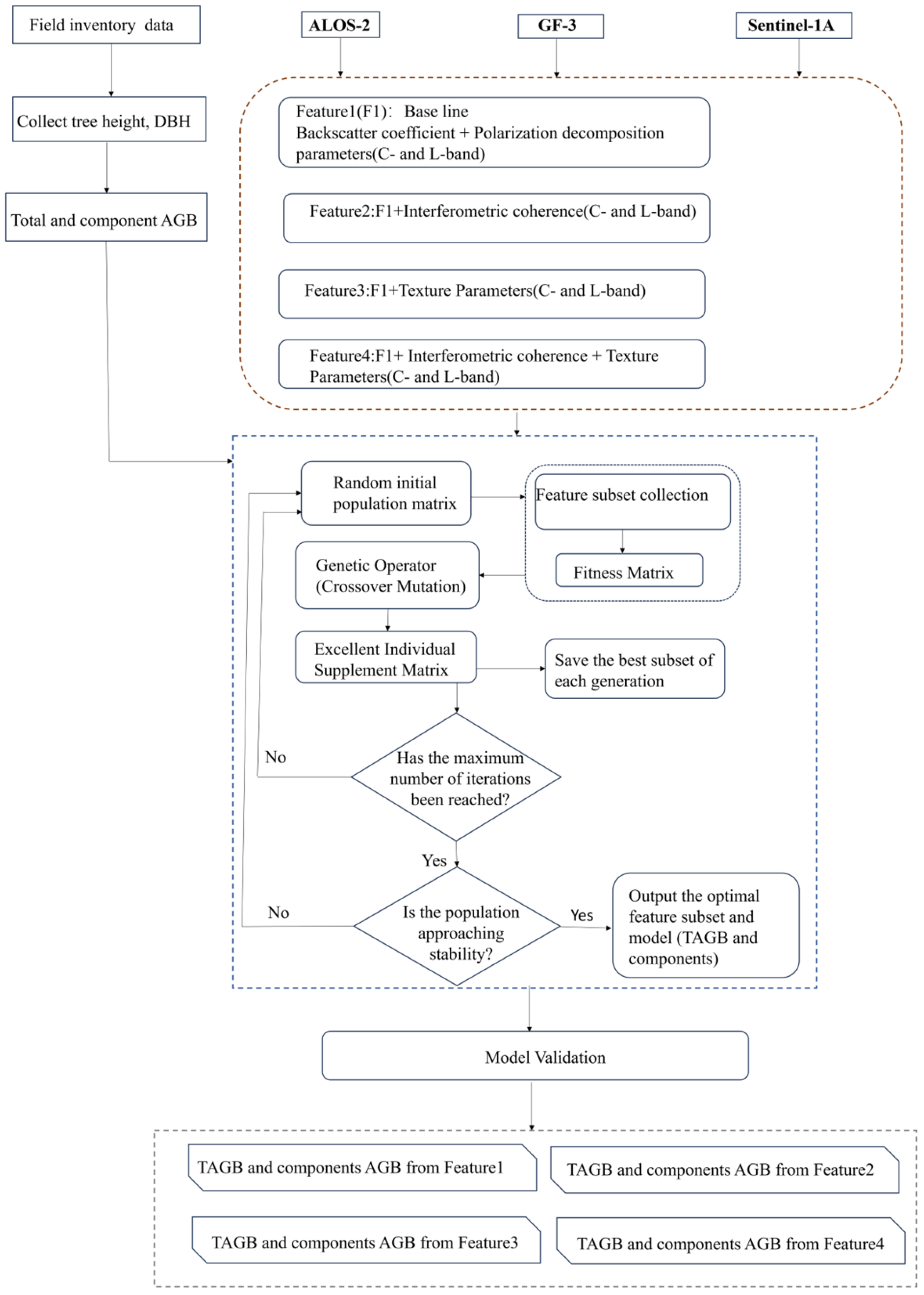
Therefore, this study integrates dual-frequency SAR observations with interferometric coherence and texture features and adopts the Random Forest model optimized using an Adaptive Genetic Algorithm (RF–AGA) modeling framework to exploit complementary sensitivities while addressing challenges associated with feature dimensionality, redundancy, and parameter tuning. In particular, deeper L-band penetration is expected to enhance sensitivity to coarse woody elements, thereby contributing more strongly to trunk and total AGB retrieval, whereas C-band measurements are anticipated to be more responsive to canopy-related biomass. Interferometric coherence, reflecting vertical structure information, is expected to further strengthen trunk and total biomass estimation when combined with backscatter-based predictors. Texture features derived from SAR intensity are also assumed to mitigate biomass saturation and improve estimation in forests with dense crowns or fine-scale structural variability. Moreover, the integration of Random Forest with an Adaptive Genetic Algorithm is designed to improve model robustness and predictive accuracy by efficiently filtering redundant variables and optimizing parameter configurations.
5. Discussion
5.1. Mechanisms of SAR Parameter Responses to Biomass Components
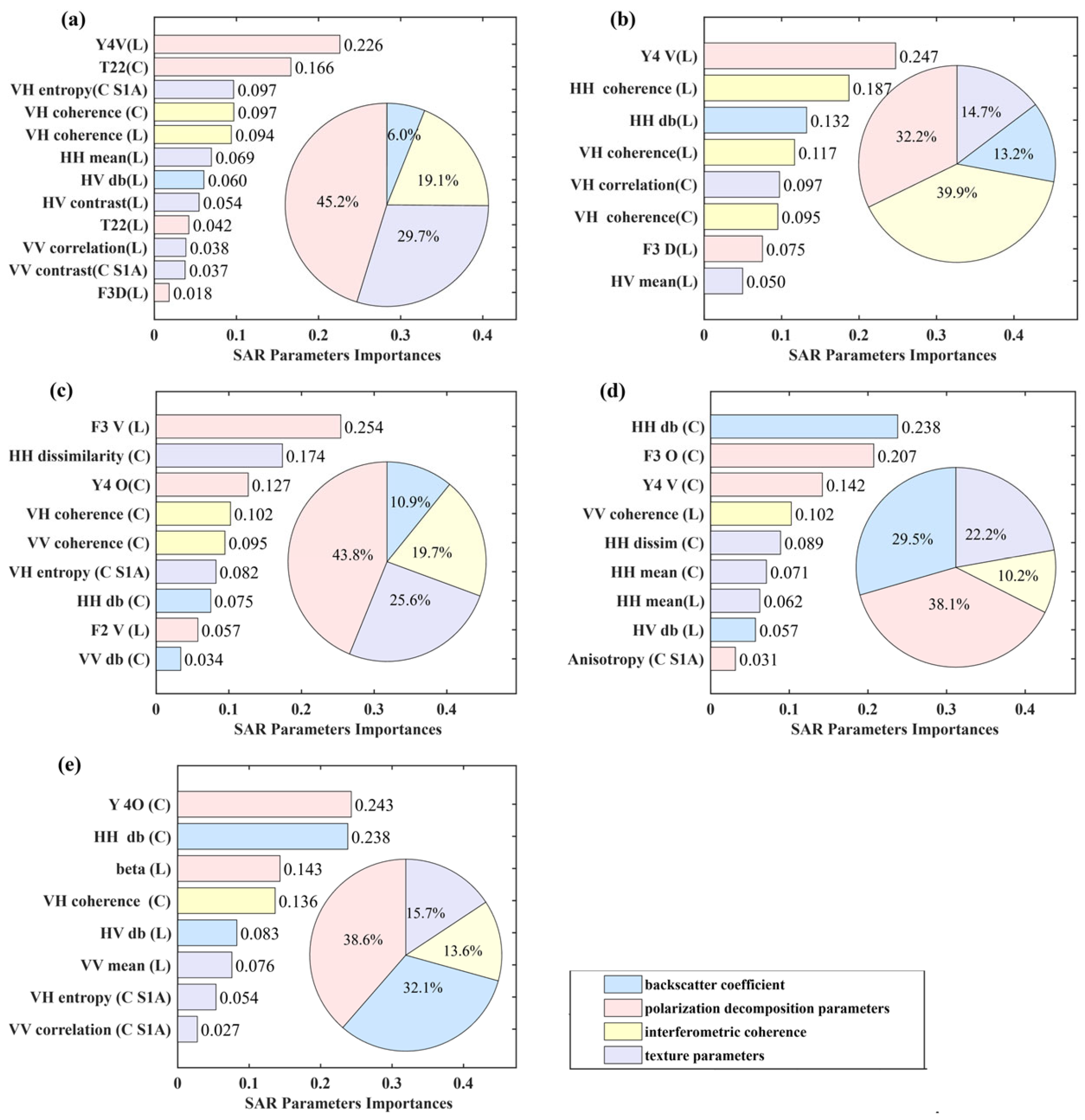
This study systematically elucidates the differential response mechanisms of multi-dimensional SAR parameters to TAGB and its components (trunk, canopy, branch, and leaves). Significant variations in parameter importance across biomass models reflect the distinct physical structures of forest components and their unique interactions with SAR data. These differences were evident not only in parameter rankings but also in component-specific dependencies and the contrasting contributions of C- and L-band parameters. Polarimetric decomposition parameters dominated across most models, though their influence varied by component. For TAGB and trunk biomass, the L-band volume scattering component (Y4V, 22.6–24.7%) was most important, linked to interactions with branches deep in the canopy or ground-canopy scattering. For trunk biomass, the L-band double-bounce component (Y4D) validated the trunk-ground dihedral reflection mechanism [
59]. By contrast, canopy and leaf components exhibited unique scattering responses, with C-band surface scattering and L-band volume scattering emerging as key parameters. These results differ from [
4,
56], where volume scattering was less dominant in low-biomass settings, likely due to higher biomass density and structural complexity in our study area. Differences in backscatter coefficients further corroborated these mechanisms. The cross-polarization (VH) backscatter coefficients in the TAGB model underscore its sensitivity to vertical vegetation structure [
25], while L-band HH polarization backscatter coefficients in the trunk model emphasized co-polarized backscatter interactions with trunk geometry, and canopy biomass; however, they showed more prominence for L- and C-band co-polarization (HH), corroborating findings by [
15].
Texture features, representing spatial heterogeneity, ranked second in importance after polarimetric decomposition for TAGB and canopy models. C-band VH correlation was effective for trunk geometry, while C-band HH contrast and dissimilarity were crucial for canopy biomass estimation. Interferometric coherence was critical for trunk biomass, with L-band HH coherence and C-band VH coherence contributing 39.9%. Although coherence ranked third for TAGB and canopy models, the joint inclusion of L- and C-band VH coherence in the TAGB model highlights their complementary roles. This synergy reflects the combined contributions of trunks and canopy to biomass estimation and underscores the potential of interferometric coherence for improving SAR-based AGB modeling [
60,
61].
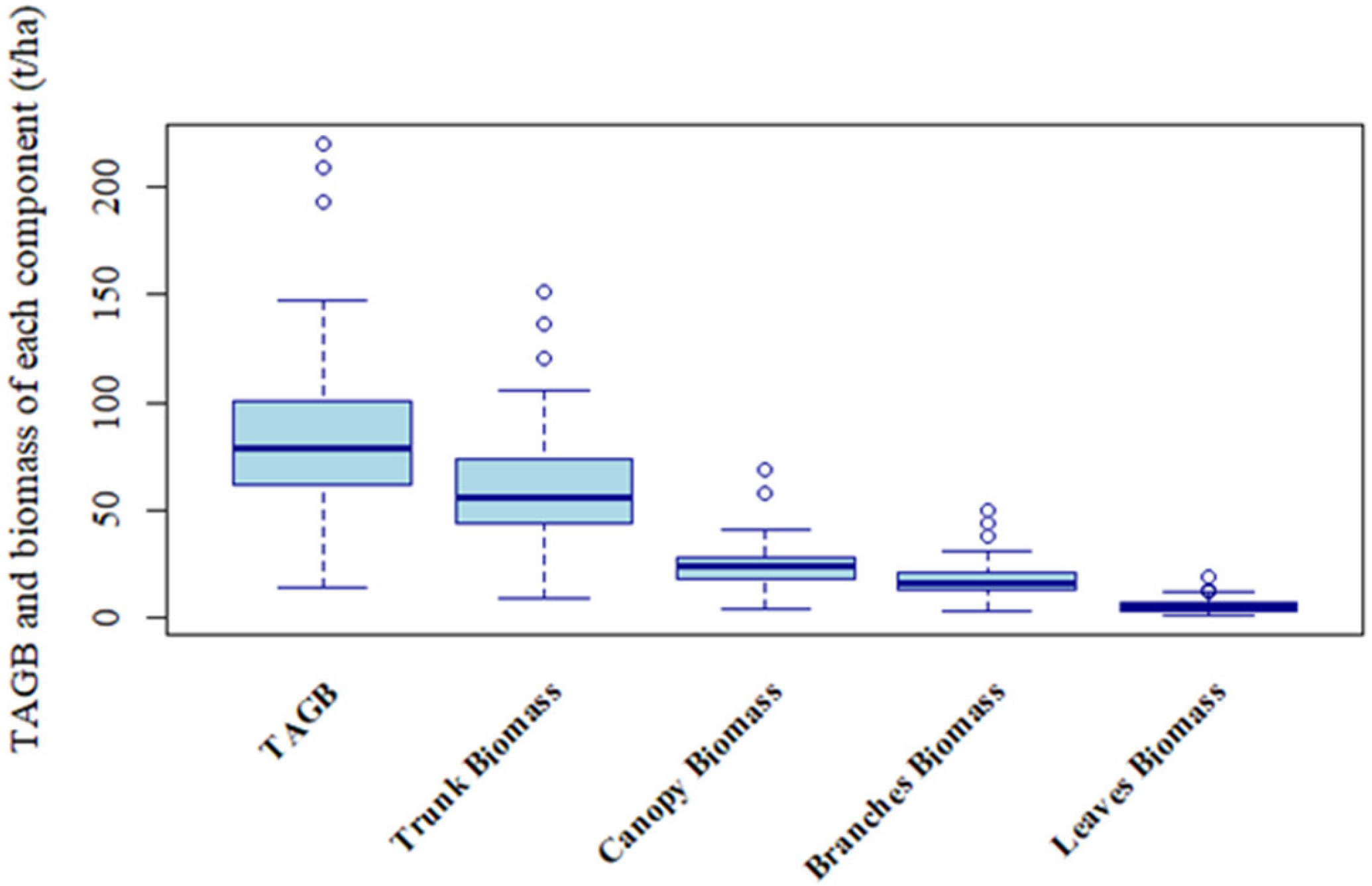
Ref. [
56] achieved optimal TAGB estimation (
R2 = 0.562) in low-biomass forests (<120 t/ha) using C-band data, while [
4] reported improved accuracy (
R2 = 0.639) at Genhe with a combined C- and L-band approach. Our results, however, demonstrate clear L-band dominance in high-biomass forests (<219.9 t/ha). These differences reflect distinct scattering mechanisms: C-band effectively interacts with fine canopy elements in sparse stands, enhancing sensitivity to leaf biomass [
26,
56], whereas L-band penetrates dense canopies to capture trunk and branch structures, making it indispensable for TAGB and trunk estimation. These findings underscore the need to tailor SAR band selection to forest structure. For carbon monitoring, the C-band is more suitable for young, low-biomass stands, while the L-band provides more reliable assessments in mature, high-biomass forests.
5.2. Multidimensional Parameter Fusion for Biomass Estimation
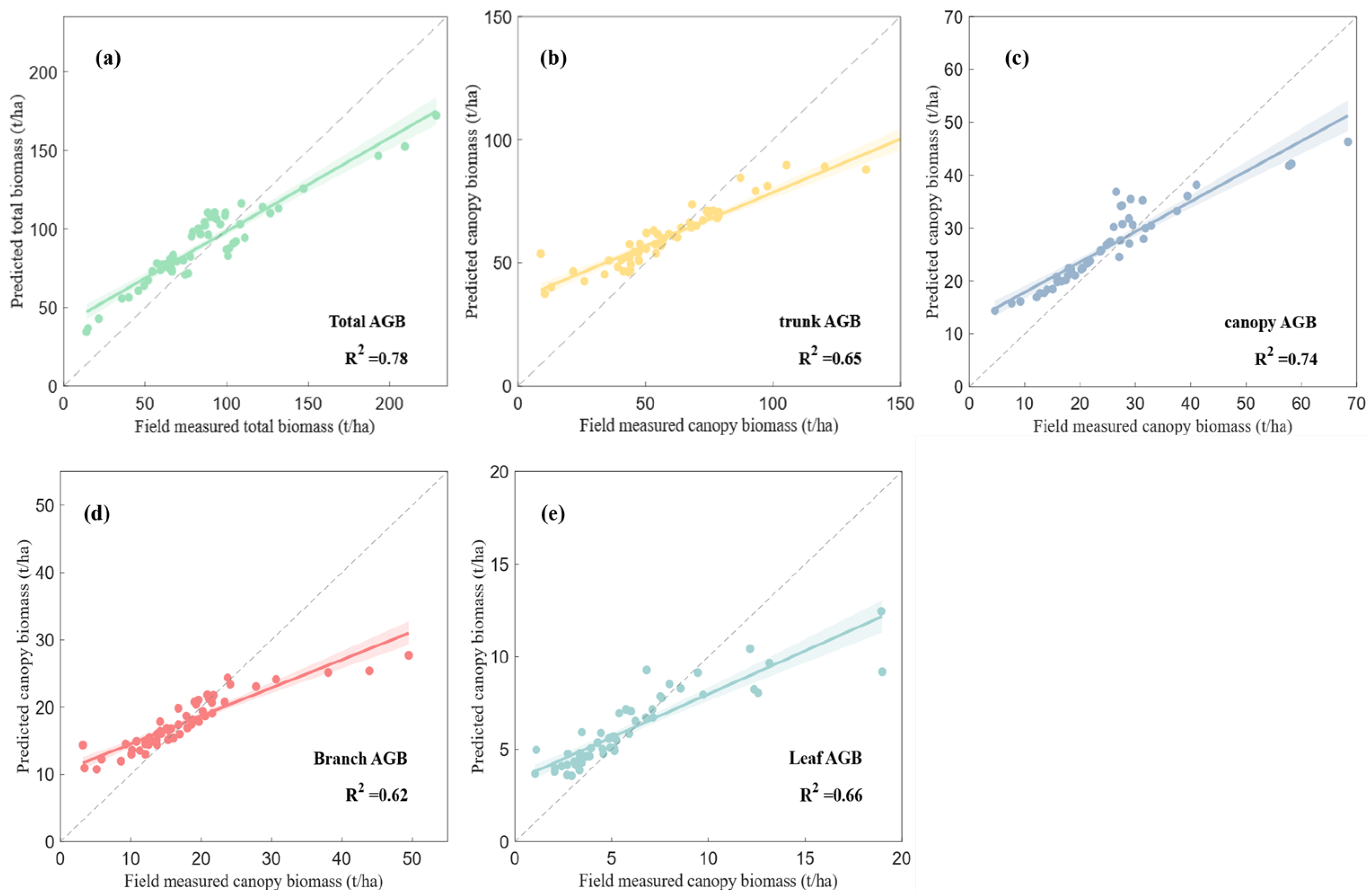
The comparative evaluation of feature combination strategies demonstrated that integrating multi-dimensional SAR parameters substantially improves the estimation of forest biomass components. The baseline two-dimensional model provided moderate performance (TAGB R2 = 0.64), but accuracy was constrained by signal saturation at higher biomass levels.
The inclusion of interferometric coherence (Strategy 2) enhanced estimation of TAGB and trunk biomass, improving
R2 by 13% and reducing RMSE by12% (M7). Ref. [
61] reported TAGB (
R2 = 0.645) and stem biomass (
R2 = 0.614) using X-band PolSAR and InSAR regression. In contrast, our integration of C- and L-band Strategy 2 with RF-AGA achieved higher accuracy, with TAGB (
R2 = 0.69) and trunk biomass (
R2 = 0.68). These results demonstrate that dual-frequency data fusion combined with ensemble learning offers clear advantages and underscores the potential of multi-dimensional SAR data for enhancing biomass retrieval.
The inclusion of texture features significantly improved canopy biomass estimation. Compared with models using only backscatter and polarimetric decomposition, texture increased
R2 by 28%. By quantifying canopy spatial heterogeneity, texture reduced signal saturation in high-biomass regions [
60] and enhanced component-level predictions. These findings align with global evidence demonstrating that texture mitigates random backscatter heterogeneity [
25]. Studies in subtropical forests [
62] and tropical forests [
24] confirm its value. For instance, ref. [
24] reported a 23% accuracy improvement when adding texture to an L-band polarimetric model. Overall, texture extends SAR-derived structural information beyond traditional backscatter, enabling more accurate biomass retrieval across ecosystems.
Full four-dimensional feature fusion (Strategy 4), integrating backscatter, polarimetric decomposition, interferometric coherence, and texture, achieved superior performance for all components. The R2 for TAGB estimation improved by 27% over the two-dimensional baselines. Models reflected the complementary roles of coherence (vertical structure) and texture (canopy heterogeneity).
Compared with previous multi-frequency studies, our results demonstrate clear advantages. Refs. [
63,
64] reported
R2 = 0.71 for AGB using X- and P-band data with polarimetric and interferometric features, whereas our GA-RF–based four-dimensional C- and L-band fusion achieved substantially higher accuracy (
R2 = 0.88). This improvement reflects the RF-AGA algorithm’s capacity to optimize hyperparameters and feature subsets while integrating coherence and texture, thereby expanding the feature space and capturing both structural and spatial complexity.
These findings confirm the effectiveness of multi-dimensional SAR parameter fusion for biomass estimation. Interferometric coherence enhances trunk and TAGB estimation by providing sensitivity to vertical structure, while texture improves canopy and fine-component estimation by capturing heterogeneity. Their combined use with backscatter and polarimetric decomposition mitigates signal saturation and delivers performance gains beyond previous approaches. Overall, this strategy provides a robust framework for regional-scale forest carbon monitoring and highlights the importance of exploiting complementary SAR features to address biomass saturation and component-level variability.
5.3. Limitations and Future Directions
Despite the encouraging performance of the proposed RF–AGA framework, several limitations should be acknowledged. First, the field sample size remains relatively small (n = 55 plots) in relation to the high-dimensional feature space. Although additional analyses, including nonparametric bootstrap resampling of LOOCV errors and targeted evaluations at higher biomass levels, were conducted to assess the stability of model performance, uncertainty associated with limited sample support cannot be fully eliminated. Expanding field measurements, particularly in high-biomass conditions, would help to further improve the reliability of parameter estimation and error characterization.
Second, model validation at the plot scale relied primarily on leave-one-out cross-validation. While a preliminary Global Moran’s I test did not indicate statistically significant spatial autocorrelation (Moran’s I = 0.032, p = 0.293), the limited number and spatial dispersion of plots may reduce the sensitivity of this test. To partially address this issue, an independent nationwide forest AGB product was employed for cross-validation within the study area. The consistency between plot-based validation results and those derived from the AIRI dataset suggests that the RF–AGA framework exhibits reasonable transferability at the regional scale. Nevertheless, future studies should adopt spatially explicit validation strategies, such as block-based cross-validation, in conjunction with both global and local spatial autocorrelation diagnostics, to more rigorously evaluate spatial dependence effects.
Third, the use of multi-sensor SAR data acquired at different times introduces potential temporal inconsistencies. Although coherence-related features were consistently selected by RF–AGA as important predictors for trunk biomass, the present analysis does not explicitly resolve the underlying physical scattering mechanisms. These effects may be associated with the temporal stability of woody components, differential decorrelation behavior, or sensor-specific acquisition characteristics. Further investigation using denser SAR time series and physically based scattering models would be necessary to clarify these mechanisms.
In summary, while the above limitations should be considered when interpreting the results, the combination of bootstrap-based uncertainty assessment, saturation-oriented diagnostic analyses, and independent dataset-based cross-validation provides converging evidence supporting the applicability of the proposed framework within the scope of the current study.