Evaluating Forest Aboveground Biomass Products by Incorporating Spatial Representativeness Analysis
Abstract
1. Introduction
2. Materials and Methods
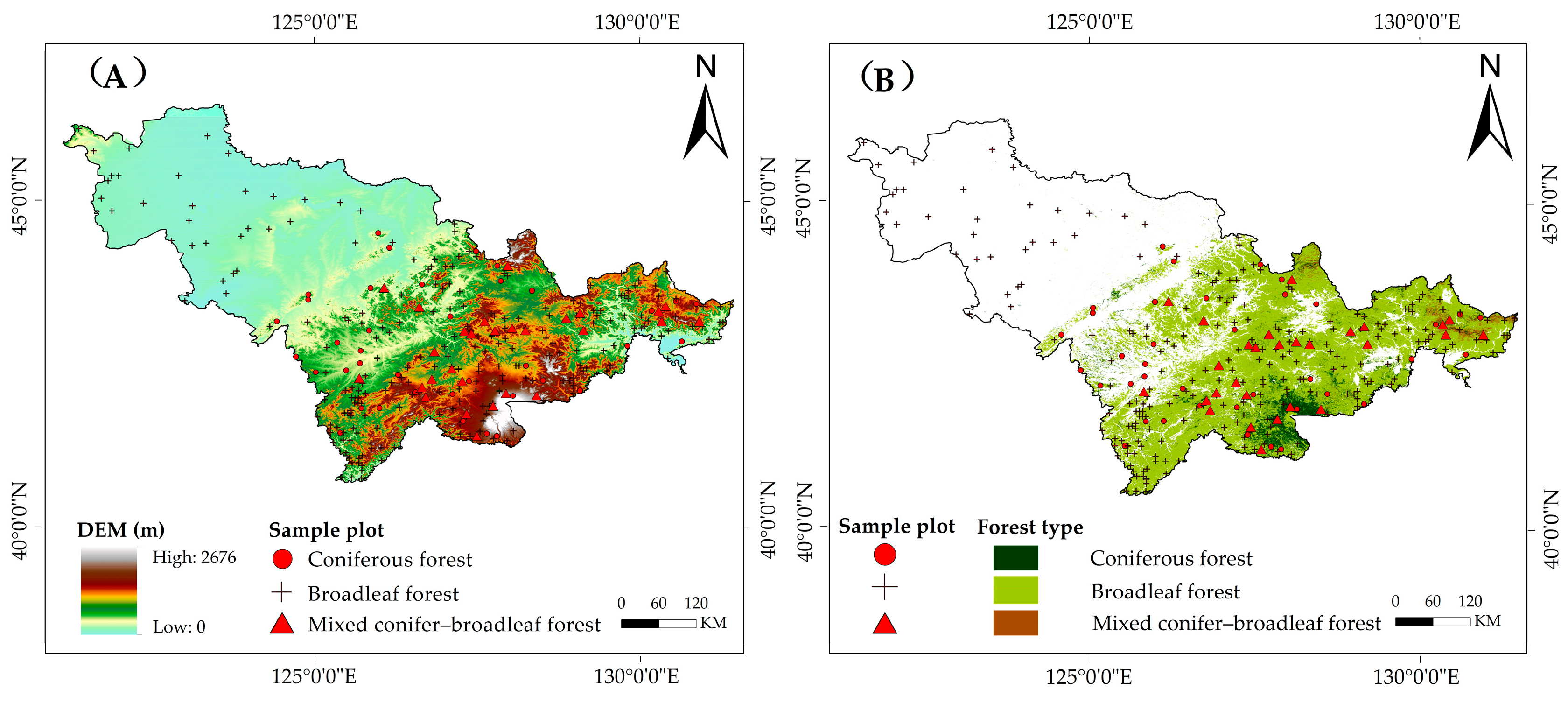
2.1. Study Area and Sample Plots
2.2. AGB Products
2.3. Auxiliary Datasets
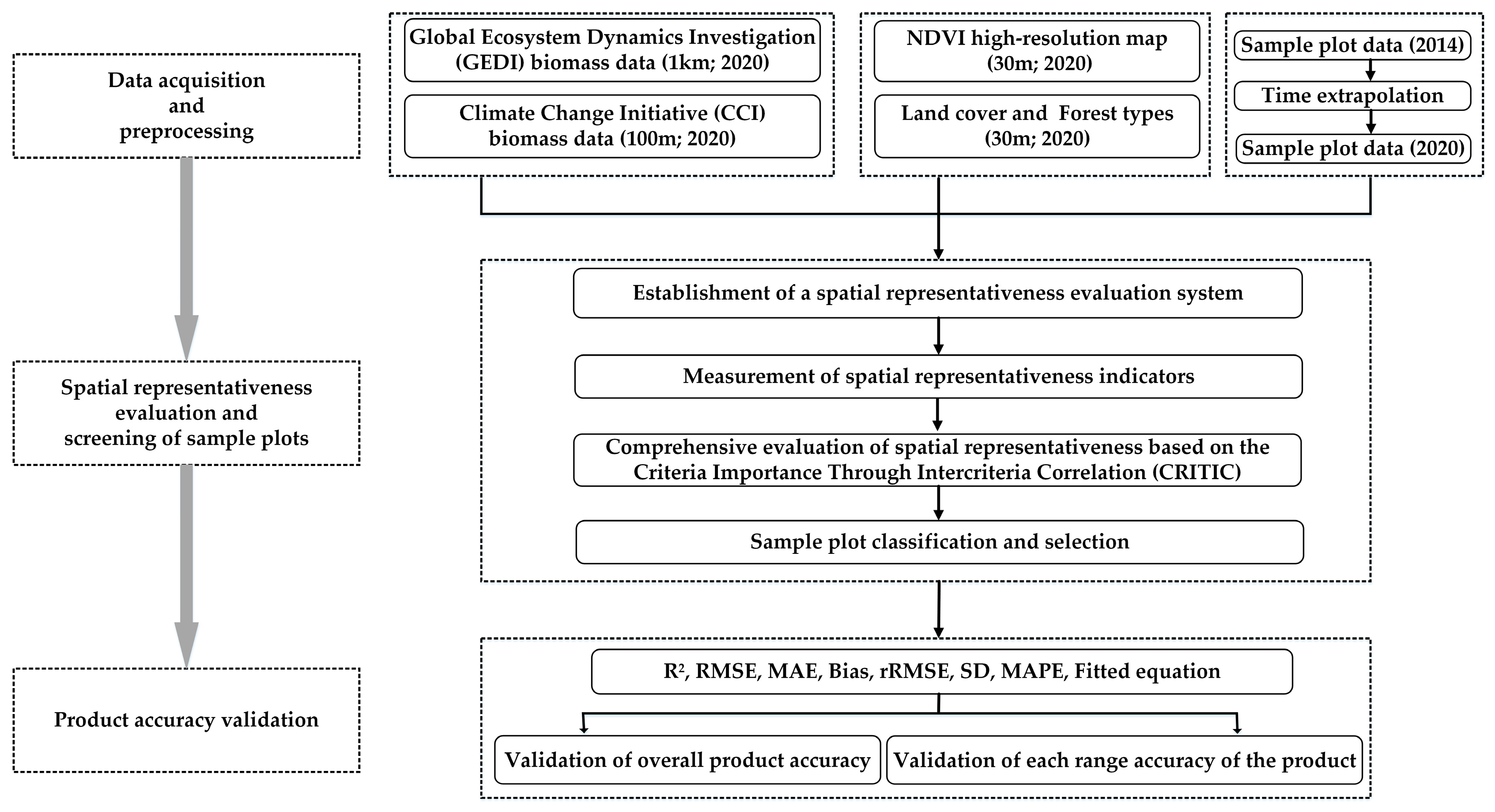
2.4. Methodological Framework
2.5. Spatial Representativeness Analysis of Sample Plots
2.5.1. Extracting Spatial Representativeness Indicators
2.5.2. Comprehensive Evaluation of Spatial Representativeness
2.6. AGB Product Evaluation
3. Results
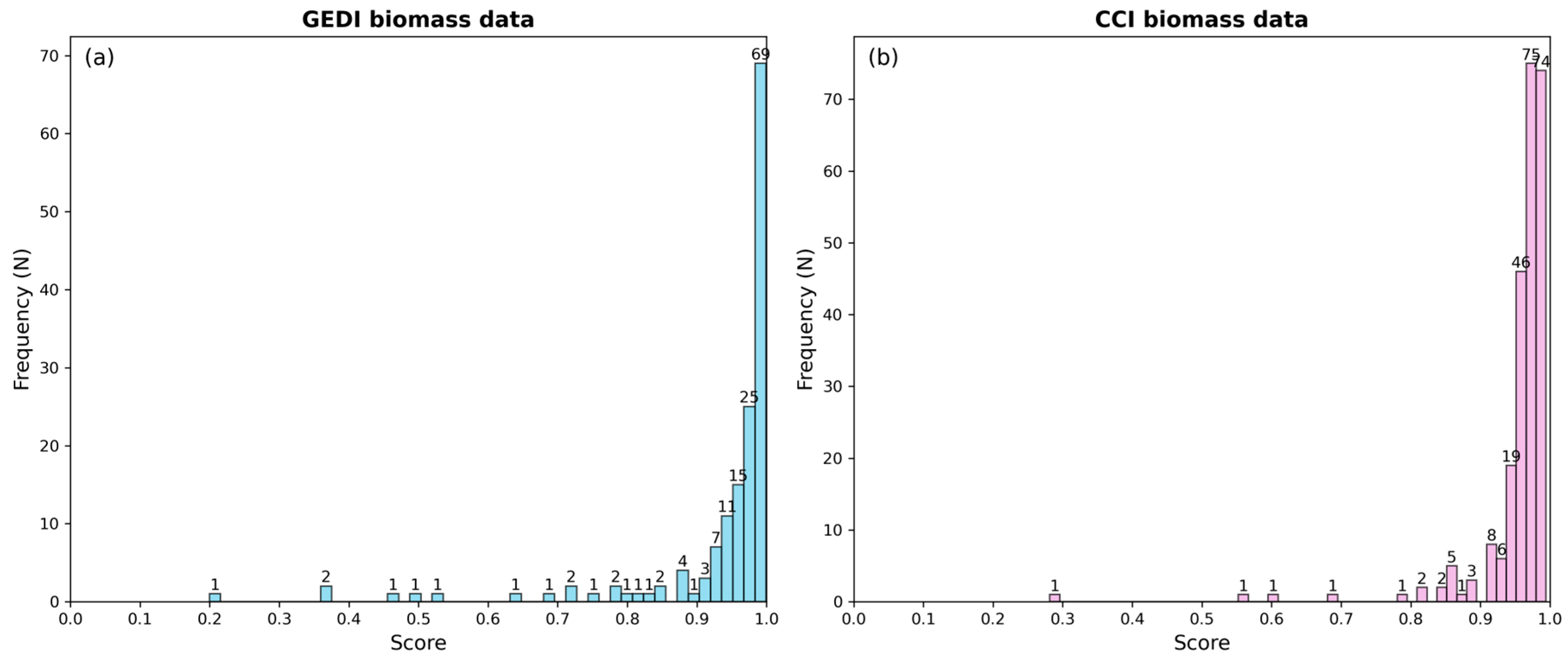
3.1. Sample Plot Selection
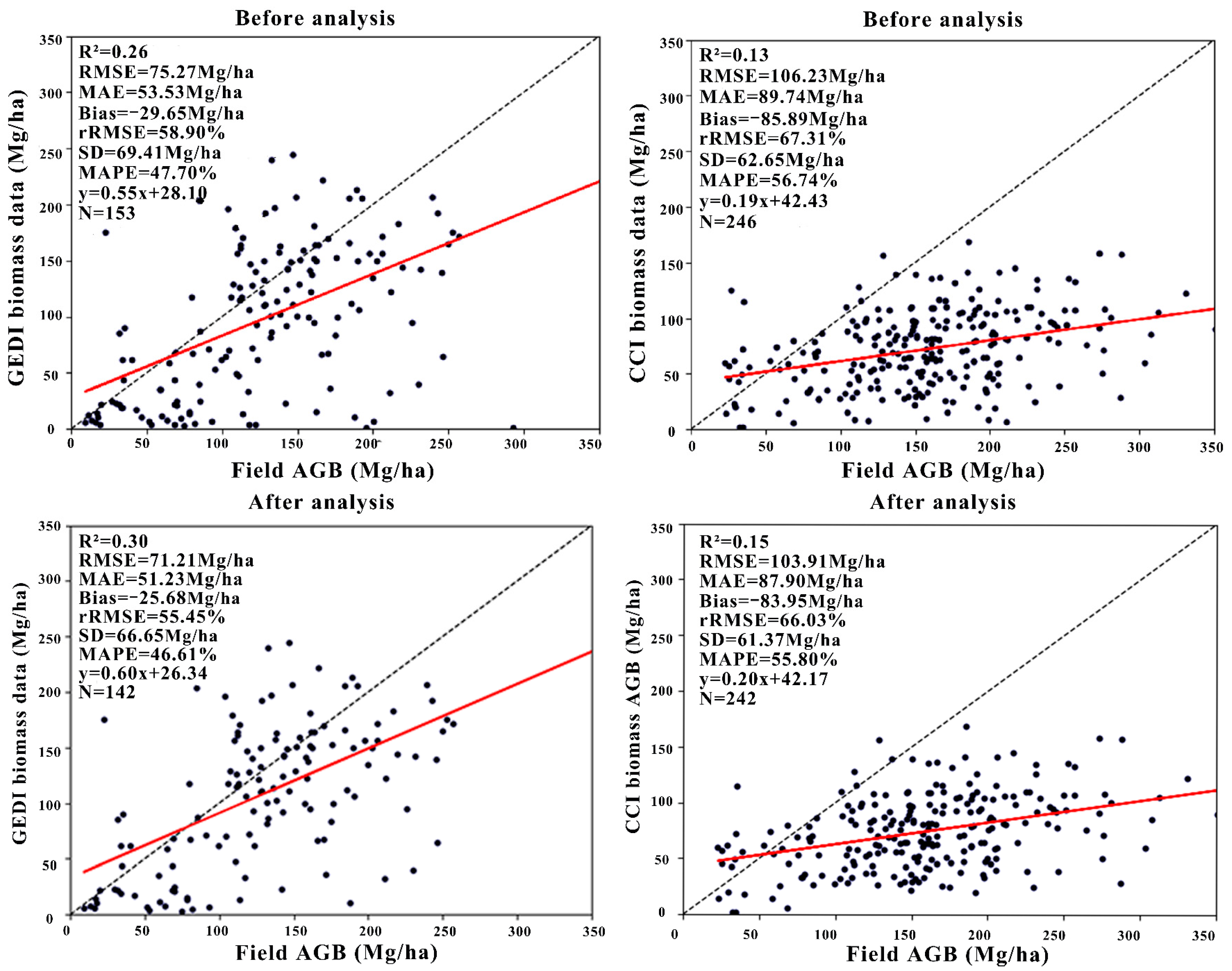
3.2. Overall Accuracy of AGB Product Evaluation
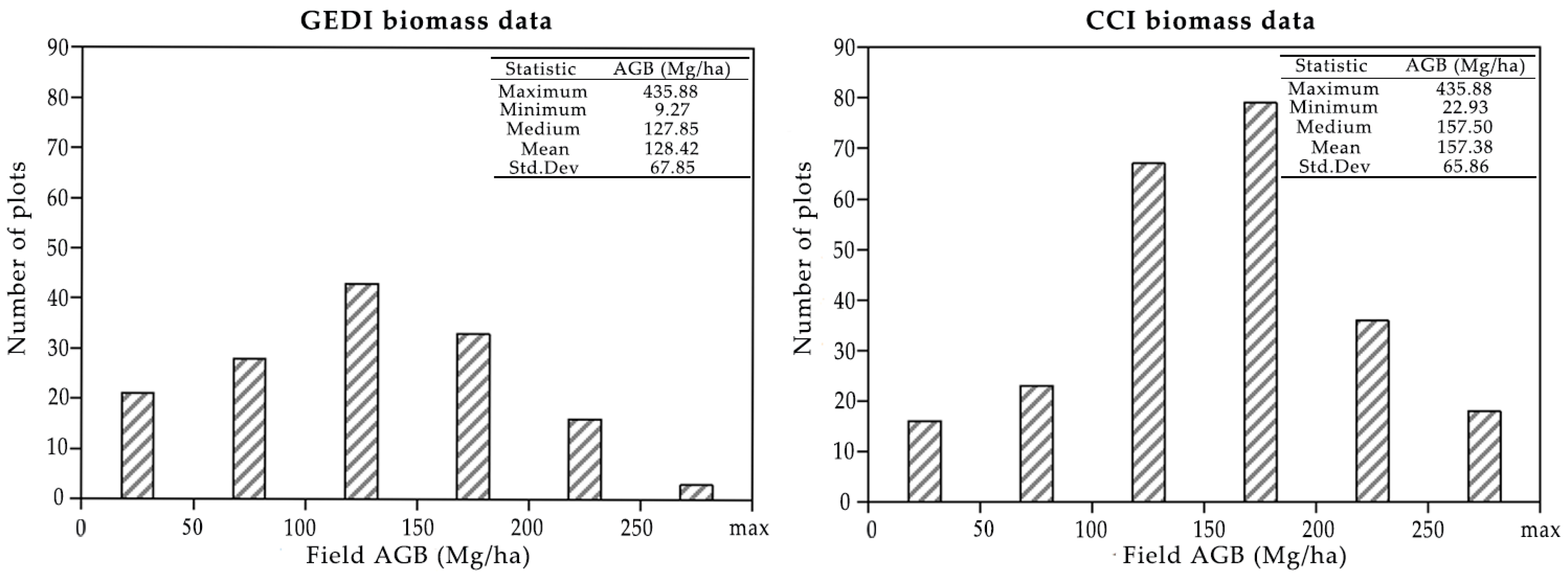
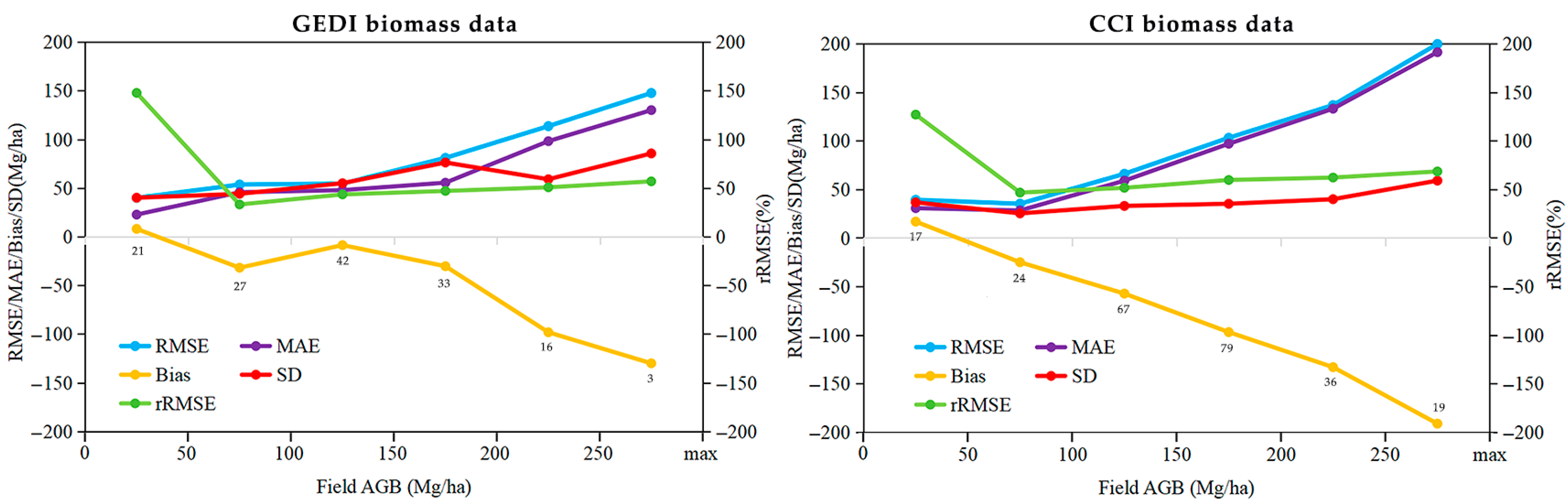
3.3. AGB Product Evaluation Grouped by Biomass Ranges
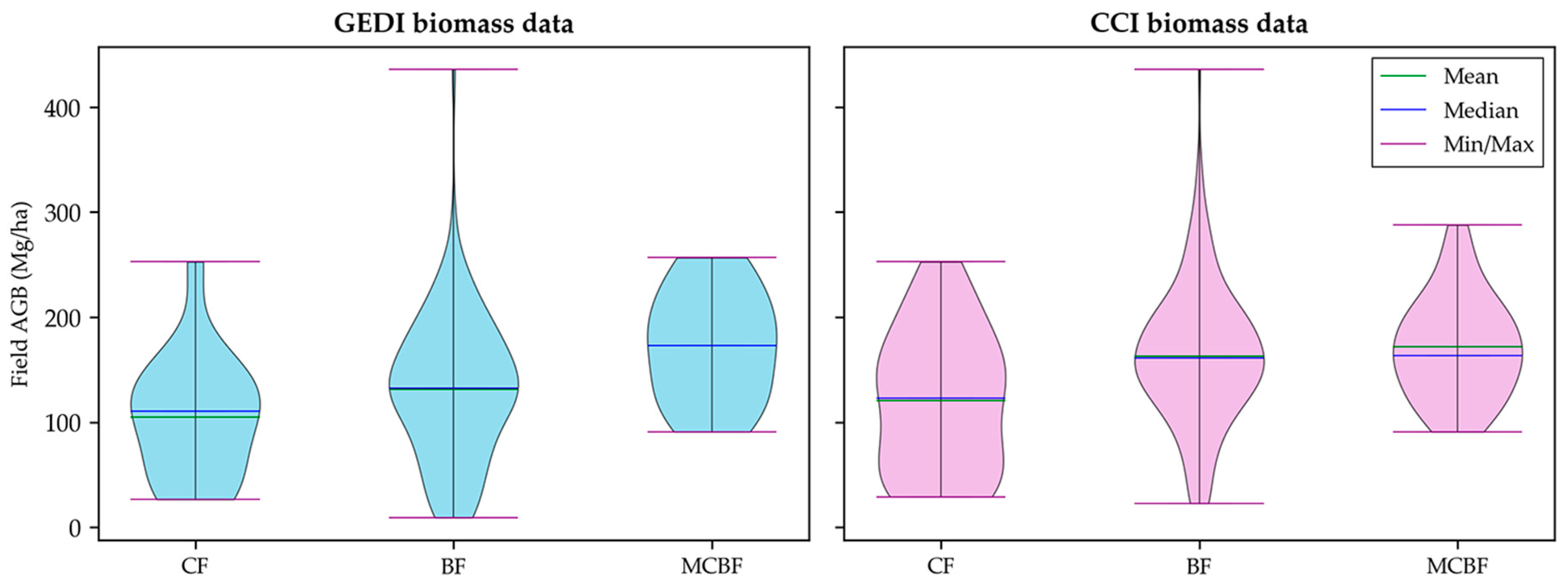
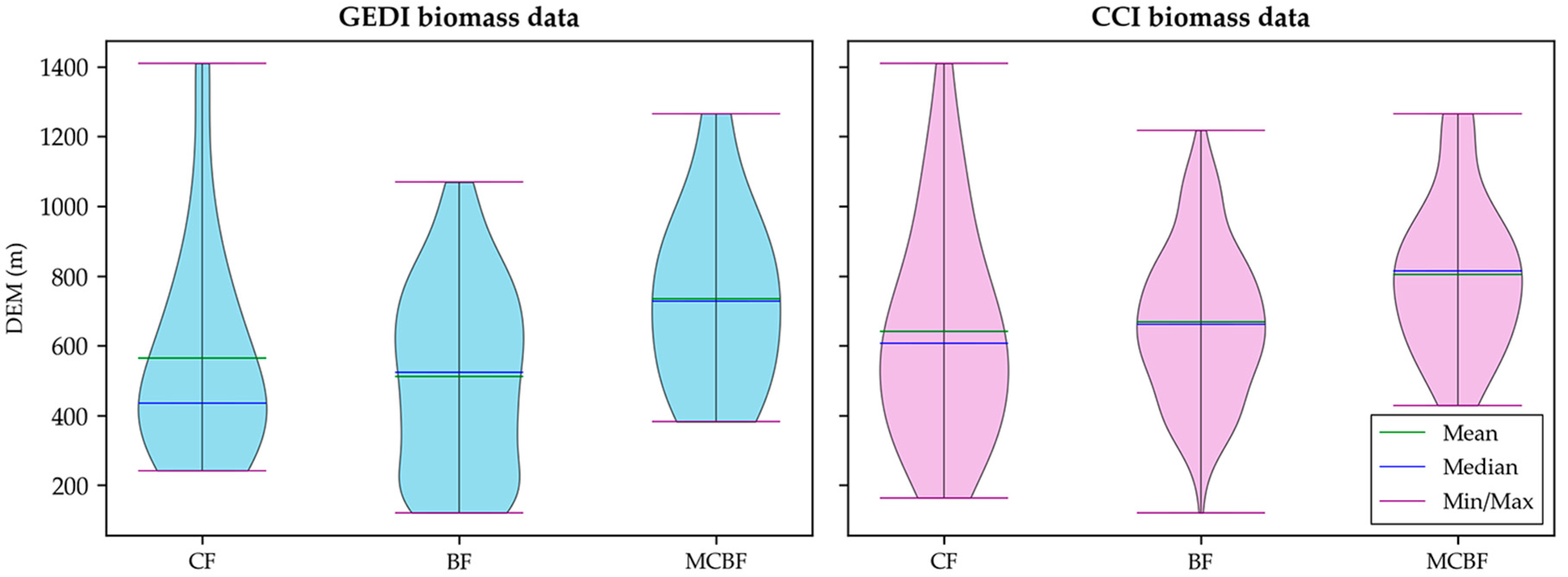
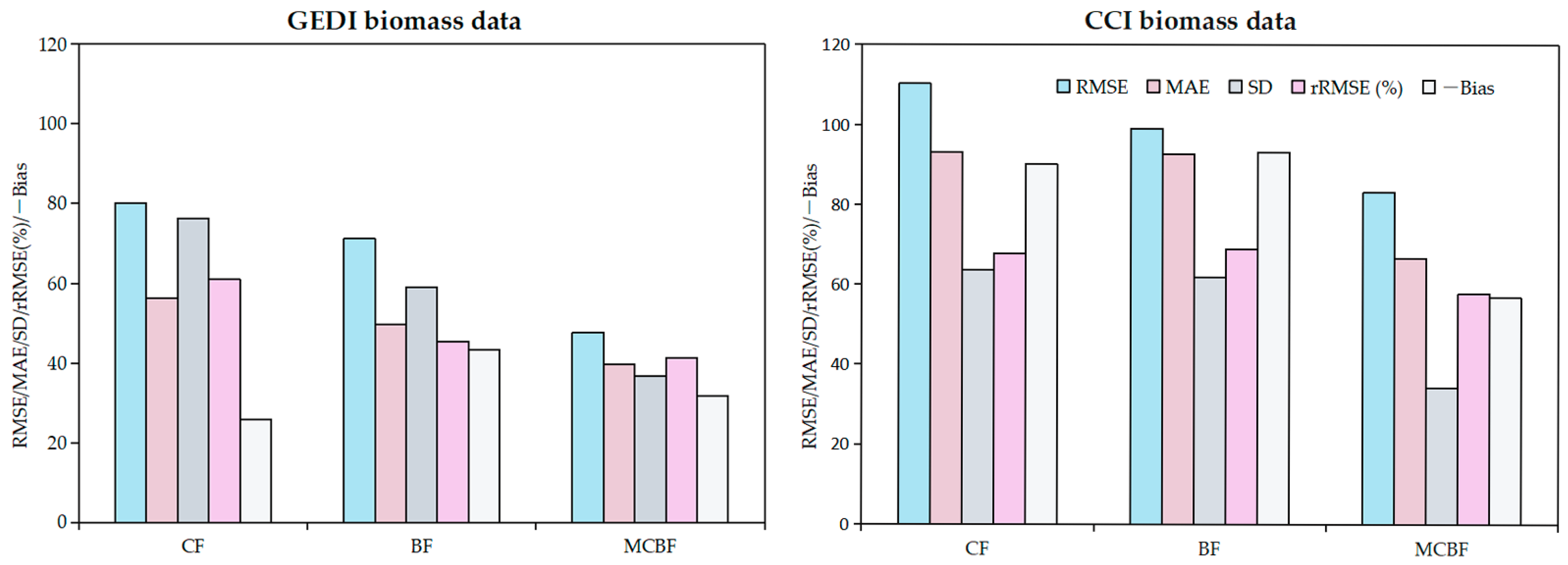
3.4. AGB Product Evaluation Grouped by Forest Types
4. Discussion
4.1. Effects of Field Data on Evaluation and Production of AGB Products
4.2. Bias of AGB Products
4.3. Adaption of Innovation and Applicability
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Pan, Y.; Birdsey, R.A.; Fang, J.; Houghton, R.; Kauppi, P.E.; Kurz, W.A.; Phillips, O.L.; Shvidenko, A.; Lewis, S.L.; Canadell, J.G.; et al. A large and persistent carbon sink in the world’s forests. Science 2011, 333, 988–993. [Google Scholar] [CrossRef]
- Aili, A.; Zhang, Y.; Lin, T.; Xu, H.; Bakayisire, F.; Waheed, A.; Zhang, Q. Ecological benefits of artificial vegetation restoration on local climate condition in abandoned mining area. Ecol. Indic. 2024, 169, 112964. [Google Scholar] [CrossRef]
- Schwärzel, K.; Zhang, L.; Montanarella, L.; Wang, Y.; Sun, G. How afforestation affects the water cycle in drylands: A process-based comparative analysis. Global Change Biol. 2019, 26, 944–959. [Google Scholar] [CrossRef]
- Obata, A.; Yoshida, T.; Hiura, T. Estimation of stand biomass and species-specific biomass in Japanese northern mixed forests in 1920–1930s: Understanding environmental factors affecting carbon sequestration before recent climate change. Ecol. Indic. 2023, 154, 110495. [Google Scholar] [CrossRef]
- Zheng, D.; Rademacher, J.; Chen, J.; Crow, T.; Bresee, M.; Le Moine, J.; Ryu, S.-R. Estimating aboveground biomass using Landsat 7 ETM+ data across a managed landscape in northern Wisconsin, USA. Remote Sens. Environ. 2004, 93, 402–411. [Google Scholar] [CrossRef]
- Avitabile, V.; Herold, M.; Heuvelink, G.B.M.; Lewis, S.L.; Phillips, O.L.; Asner, G.P.; Armston, J.; Ashton, P.S.; Banin, L.; Bayol, N.; et al. An integrated pan-tropical biomass map using multiple reference datasets. Global Change Biol. 2016, 22, 1406–1420. [Google Scholar] [CrossRef] [PubMed]
- Dubayah, R.O.; Armston, J.; Healey, S.P.; Yang, Z.; Patterson, P.L.; Saarela, S.; Stahl, G.; Duncanson, L.; Kellner, J.R. GEDI L4B Gridded Aboveground Biomass Density; Version 2.1; ORNL DAAC: Oak Ridge, TN, USA, 2023. Available online: https://www.earthdata.nasa.gov/data/catalog/ornl-cloud-gedi-l4b-gridded-biomass-v2-1-2299-2.1 (accessed on 22 July 2025).
- Kellndorfer, J.; Kirsch, K.; Fiske, G.; Bishop, J.; LaPoint, L.; Hoppus, M.; Westfall, J. NACP Aboveground Biomass and Carbon Baseline Data; V.2 (NBCD 2000); ORNL DAAC: Oak Ridge, TN, USA, 2000. Available online: https://www.earthdata.nasa.gov/data/catalog/ornl-cloud-nbcd2000-v2-1161-2 (accessed on 22 July 2025).
- Su, Y.; Guo, Q.; Xue, B.; Hu, T.; Alvarez, O.; Tao, S.; Fang, J. Spatial distribution of forest aboveground biomass in China: Estimation through combination of spaceborne lidar, optical imagery, and forest inventory data. Remote Sens. Environ. 2016, 173, 187–199. [Google Scholar] [CrossRef]
- Chen, C.; He, Y.; Zhang, J.; Xu, D.; Han, D.; Liao, Y.; Luo, L.; Teng, C.; Yin, T. Estimation of aboveground biomass for Pinus densata using multi-source time series in Shangri-La considering seasonal effects. Forests 2023, 14, 1747. [Google Scholar] [CrossRef]
- Huang, T.; Ou, G.; Wu, Y.; Zhang, X.; Liu, Z.; Xu, H.; Xu, X.; Wang, Z.; Xu, C. Estimating the aboveground biomass of various forest types with high heterogeneity at the provincial scale based on multi-source data. Remote Sens. 2023, 15, 3550. [Google Scholar] [CrossRef]
- Luo, Y.; Wang, X.; Ouyang, Z.; Lu, F.; Feng, L.; Tao, J. A review of biomass equations for China’s tree species. Earth Syst. Sci. Data 2020, 12, 21–40. [Google Scholar] [CrossRef]
- Schwartz, M.; Ciais, P.; De Truchis, A.; Chave, J.; Ottlé, C.; Vega, C.; Wigneron, J.-P.; Nicolas, M.; Jouaber, S.; Liu, S.; et al. FORMS: Forest Multiple Source height, wood volume, and biomass maps in France at 10 to 30 m resolution based on Sentinel-1, Sentinel-2, and Global Ecosystem Dynamics Investigation (GEDI) data with a deep learning approach. Earth Syst. Sci. Data 2023, 15, 4927–4945. [Google Scholar] [CrossRef]
- Silva, C.A.; Duncanson, L.; Hancock, S.; Neuenschwander, A.; Thomas, N.; Hofton, M.; Fatoyinbo, L.; Simard, M.; Marshak, C.Z.; Armston, J.; et al. Fusing simulated GEDI, ICESat-2 and NISAR data for regional aboveground biomass mapping. Remote Sens. Environ. 2021, 253, 112234. [Google Scholar] [CrossRef]
- Zhang, Y.; Liang, S. Fusion of multiple gridded biomass datasets for generating a global forest aboveground biomass map. Remote Sens. 2020, 12, 2559. [Google Scholar] [CrossRef]
- Li, Y.; Li, M.; Li, C.; Liu, Z. Forest aboveground biomass estimation using Landsat 8 and Sentinel-1A data with machine learning algorithms. Sci. Rep. 2020, 10, 9952. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Liu, C.; Wang, X.; Zhou, X.; Gong, P. Integration of multi-resource remotely sensed data and allometric models for forest aboveground biomass estimation in China. Remote Sens. Environ. 2019, 221, 225–234. [Google Scholar] [CrossRef]
- Nandy, S.; Srinet, R.; Padalia, H. Mapping Forest Height and Aboveground Biomass by Integrating ICESat-2, Sentinel-1 and Sentinel-2 Data Using Random Forest Algorithm in Northwest Himalayan Foothills of India. Geophys. Res. Lett. 2021, 48, e2021GL093799. [Google Scholar] [CrossRef]
- Guerra-Hernández, J.; Narine, L.L.; Pascual, A.; Gonzalez-Ferreiro, E.; Botequim, B.; Malambo, L.; Neuenschwander, A.; Popescu, S.C.; Godinho, S. Aboveground biomass mapping by integrating ICESat-2, SENTINEL-1, SENTINEL-2, ALOS2/PALSAR2, and topographic information in Mediterranean forests. Gisci. Remote Sens. 2022, 59, 1509–1533. [Google Scholar] [CrossRef]
- Shendryk, Y. Fusing GEDI with earth observation data for large area aboveground biomass mapping. Int. J. Appl. Earth Obs. 2022, 115, 103108. [Google Scholar] [CrossRef]
- Hunka, N.; Santoro, M.; Armston, J.; Dubayah, R.; McRoberts, R.E.; Næsset, E.; Quegan, S.; Urbazaev, M.; Pascual, A.; May, P.B. On the NASA GEDI and ESA CCI biomass maps: Aligning for uptake in the UNFCCC global stocktake. Environ. Res. Lett. 2023, 18, 124042. [Google Scholar] [CrossRef]
- Healey, S.P.; Patterson, P.L.; Armston, J. Algorithm Theoretical Basis Document (ATBD) for GEDI Level-4B (L4B) Gridded Aboveground Biomass Density, Version 1.0. ORNL Distributed Active Archive Center. Available online: https://daac.ornl.gov/daacdata/gedi/GEDI_L4B_Gridded_Biomass/comp/GEDI_L4B_ATBD_v1.0.pdf (accessed on 12 March 2024).
- Santoro, M.; Cartus, O. Algorithm Theoretical Basis Document (ATBD), Year 4, Version 4.0. European Space Agency. Available online: https://climate.esa.int/media/documents/D2_2_Algorithm_Theoretical_Basis_Document_ATBD_V4.0_20230317.pdf (accessed on 20 April 2023).
- Avitabile, V.; Camia, A. An assessment of forest biomass maps in Europe using harmonized national statistics and inventory plots. Forest Ecol. Manag. 2018, 409, 489–498. [Google Scholar] [CrossRef]
- Zhang, Y.; Liang, S.; Yang, L. A review of regional and global gridded forest biomass datasets. Remote Sens. 2019, 11, 2744. [Google Scholar] [CrossRef]
- Araza, A.; De Bruin, S.; Herold, M.; Quegan, S.; Labriere, N.; Rodriguez-Veiga, P.; Avitabile, V.; Santoro, M.; Mitchard, E.T.A.; Ryan, C.M.; et al. A comprehensive framework for assessing the accuracy and uncertainty of global above-ground biomass maps. Remote Sens. Environ. 2022, 272, 112917. [Google Scholar] [CrossRef]
- Breidenbach, J.; Granhus, A.; Hylen, G.; Eriksen, R.; Astrup, R. A century of National Forest Inventory in Norway-informing past, present, and future decisions. For. Ecosyst. 2020, 7, 46. [Google Scholar] [CrossRef]
- Knott, J.; Liknes, G.C.; Giebink, C.L.; Oh, S.; Domke, G.M.; McRoberts, R.E.; Quirino, V.F.; Walters, B.F. Effects of outliers on remote sensing-assisted forest biomass estimation: A case study from the United States national forest inventory. Methods Ecol. Evol. 2023, 14, 1587–1602. [Google Scholar] [CrossRef]
- Réjou-Méchain, M.; Muller-Landau, H.C.; Detto, M.; Thomas, S.C.; Le Toan, T.; Saatchi, S.S.; Barreto-Silva, J.S.; Bourg, N.A.; Bunyavejchewin, S.; Butt, N.; et al. Local spatial structure of forest biomass and its consequences for remote sensing of carbon stocks. Biogeosciences 2014, 11, 6827–6840. [Google Scholar] [CrossRef]
- Zeng, W.; Tomppo, E.; Healey, S.P.; Gadow, K.V. The national forest inventory in China: History- results -international context. Forest Ecosyst. 2015, 2, 23. [Google Scholar] [CrossRef]
- Yu, Y.; Pan, Y.; Yang, X.; Fan, W. Spatial scale effect and correction of forest aboveground biomass estimation using remote sensing. Remote Sens. 2022, 14, 2828. [Google Scholar] [CrossRef]
- Nappo, C.J.; Caneill, J.Y.; Furman, R.W.; Gifford, F.A.; Kaimal, J.C.; Kramer, M.L.; Lockhart, T.J.; Pendergast, M.M.; Pielke, R.A.; Randerson, D.; et al. The workshop on the representativeness of meteorological observations, June 1981, Boulder, Colo. Bull. Am. Meteorol. Soc. 1982, 63, 761–764. Available online: https://www.jstor.org/stable/26222836 (accessed on 18 August 2025).
- Ma, J.; Zhou, J.; Liu, S.; Göttsche, F.M.; Zhang, X.; Wang, S.; Li, M. Continuous evaluation of the spatial representativeness of land surface temperature validation sites. Remote Sens. Environ. 2021, 265, 112669. [Google Scholar] [CrossRef]
- Rossini, M.; Celesti, M.; Bramati, G.; Migliavacca, M.; Cogliati, S.; Rascher, U.; Colombo, R. Evaluation of the spatial representativeness of in situ SIF observations for the validation of medium-resolution satellite SIF products. Remote Sens. 2022, 14, 5107. [Google Scholar] [CrossRef]
- Wang, H.; Yakir, D.; Rotenberg, E. Assessing the effectiveness of a central flux tower in representing the spatial variations in gross primary productivity in a semi-arid pine forest. Agr. Forest Meteorol. 2023, 333, 109415. [Google Scholar] [CrossRef]
- Liu, S.; Su, H.; Tian, J.; Wang, W. An analysis of spatial representativeness of air temperature monitoring stations. Theor. Appl. Climatol. 2017, 132, 857–865. [Google Scholar] [CrossRef]
- Schmid, H.P.; Lloyd, C.R. Spatial representativeness and the location bias of flux footprints over inhomogeneous areas. Agr. Forest Meteorol. 1999, 93, 195–209. [Google Scholar] [CrossRef]
- Gong, H.; Wang, Y.; Wang, G.; Gao, Y.; Li, G.; Kuang, Z.; Zhuo, X.; Bi, J.; Wang, P.; Wang, W.; et al. Quantifying the spatial representativeness of carbon flux footprints of a grassland ecosystem in the semi-arid region. J. Geophys. Res. Atmos. 2023, 128, e2022JD038269. [Google Scholar] [CrossRef]
- Román, M.O.; Schaaf, C.B.; Woodcock, C.E.; Strahler, A.H.; Yang, X.; Braswell, R.H.; Curtis, P.S.; Davis, K.J.; Dragoni, D.; Goulden, M.L.; et al. The MODIS (Collection V005) BRDF/albedo product: Assessment of spatial representativeness over forested landscapes. Remote Sens. Environ. 2009, 113, 2476–2498. [Google Scholar] [CrossRef]
- Wang, Y.; Zheng, Z. Spatial Representativeness Analysis for Snow Depth Measurements of Meteorological Stations in Northeast China. J. Hydrometeorol. 2020, 21, 791–805. [Google Scholar] [CrossRef]
- Lesht, B.M.; Barbiero, R.P.; Warren, G.J. Using satellite observations to assess the spatial representativeness of the GLNPO water quality monitoring program. J. Great Lakes Res. 2018, 44, 547–562. [Google Scholar] [CrossRef]
- Liu, Q.; Shan, Y.; Shu, L.; Sun, P.; Du, S. Spatial and temporal distribution of forest fire frequency and forest area burnt in Jilin Province, Northeast China. J. Forestry Res. 2018, 29, 1233–1239. [Google Scholar] [CrossRef]
- Zhang, X.; Lei, Y. Comparison of annual individual-tree growth models based on variable rate and constant rate methods. Forest Res. 2009, 22, 824–828. [Google Scholar] [CrossRef]
- Li, H.; Lei, Y. Estimation and Evaluation of Forest Biomass Carbon Storage in China; China Forestry Publishing House: Beijing, China, 2010. [Google Scholar]
- Li, H.; Fa, L. Height-diameter model for major tree species in China using the classified height method (in Chinese). Sci. Silvae Sin. 2011, 47, 83–90. [Google Scholar]
- GB/T 43648-2024; Tree Biomass Models and Related Parameters to Carbon Accounting for Major Tree Species. China Standards Press: Beijing, China, 2024.
- Buckley, S. NASADEM: Creating a New NASA Digital Elevation Model and Associated Products. Available online: https://earthdata.nasa.gov/esds/competitive-programs/measures/nasadem (accessed on 18 July 2025).
- Zhang, X.; Liu, L.; Chen, X.; Gao, Y.; Xie, S.; Mi, J. GLC_FCS30: Global land-cover product with fine classification system at 30 m using time-series Landsat imagery. Earth Syst. Sci. Data 2021, 13, 2753–2776. [Google Scholar] [CrossRef]
- Santoro, M.; Cartus, O. ESA Biomass Climate Change Initiative (Biomass_cci): Global Datasets of Forest Above-Ground Biomass for the Years 2010, 2017, 2018, 2019 and 2020. NERC EDS Centre for Environmental Data Analysis. Available online: https://climate.esa.int/en/data (accessed on 1 July 2024).
- Dong, J.; Zhou, Y.; You, N.; Chen, C. A 30-m Annual Maximum NDVI Dataset in China from 2000 to 2022. National Ecosystem Science Data Center, China. Available online: https://cstr.cn/15732.11.nesdc.ecodb.rs.2021.012 (accessed on 22 July 2025).
- Gorelick, N.; Hancher, M.; Dixon, M.; Ilyushchenko, S.; Thau, D.; Moore, R. Google Earth Engine: Planetary-scale geospatial analysis for everyone. Remote Sens. Environ. 2017, 202, 18–27. [Google Scholar] [CrossRef]
- Yang, J.; Huang, X. The 30 m annual land cover datasets and its dynamics in China from 1990 to 2019. Earth Syst. Sci. Data 2021, 13, 3907–3925. [Google Scholar] [CrossRef]
- Hakuba, M.Z.; Folini, D.; Sanchez-Lorenzo, A.; Wild, M. Spatial representativeness of ground-based solar radiation measurements. J. Geophys. Res. Atmos. 2013, 118, 8585–8597. [Google Scholar] [CrossRef]
- Diakoulaki, D.; Mavrotas, G.; Papayannakis, L. Determining objective weights in multiple criteria problems: The CRITIC method. Comput. Ops. Res. 1995, 22, 763–770. [Google Scholar] [CrossRef]
- Fang, H.; Zhang, Y.; Wei, S.; Li, W.; Ye, Y.; Sun, T.; Liu, W. Validation of global moderate resolution leaf area index (LAI) products over croplands in northeastern China. Remote Sens. Environ. 2019, 233, 111377. [Google Scholar] [CrossRef]
- Cochran, W.G. Sampling Techniques, 3rd ed.; Wiley: New York, NY, USA, 1977. [Google Scholar]
- Food and Agriculture Organization of the United Nations (FAO). Global Forest Resources Assessment 2010: Main Report; FAO: Rome, Italy, 2010. [Google Scholar]
- National Forestry and Grassland Administration (NFGA). Technical Guidelines for the Ninth National Forest Inventory of China; NFGA: Beijing, China, 2018. [Google Scholar]
- Kennedy, R.E.; Yang, Z.; Cohen, W.B. Detecting trends in forest disturbance and recovery using yearly Landsat time series: 1. LandTrendr-temporal segmentation algorithms. Remote Sens. Environ. 2010, 114, 2897–2910. [Google Scholar] [CrossRef]
- Huang, C.; Goward, S.N.; Schleeweis, K.; Thomas, N.; Masek, J.G.; Zhu, Z. Dynamics of national forests assessed using the Landsat record: Case studies in eastern U.S. Remote Sens. Environ. 2009, 113, 1430–1442. [Google Scholar] [CrossRef]
- Ji, W.; Zhongke, F.; Hanyue, Z.; Yuan, W. Assessment of Carbon sink potential of arbor forests based on DBH growth rate model for standing trees. J. Agric. Sci. Technol. 2024, 26, 99–109. [Google Scholar] [CrossRef]
- Santoro, M.; Cartus, O.; Quegan, S.; Kay, H.; Lucas, R.M.; Araza, A.; Herold, M.; Labrière, N.; Chave, J.; Rosenqvist, Å.; et al. Design and performance of the climate change initiative biomass global retrieval algorithm. Sci. Remote Sens. 2024, 10, 100169. [Google Scholar] [CrossRef]
- Kellner, J.R.; Armston, J.; Duncanson, L. Algorithm theoretical basis document for GEDI footprint aboveground biomass density. ESS 2023, 10, e2022EA002516. [Google Scholar] [CrossRef]
- Jia, D.; Wang, C.; Hakkenberg, C.R.; Numata, I.; Elmore, A.J.; Cochrane, M.A. Accuracy evaluation and effect factor analysis of GEDI aboveground biomass product for temperate forests in the conterminous United States. GISci. Remote Sens. 2024, 61, 2292374. [Google Scholar] [CrossRef]
- Pascual, A.; Guerra-Hernández, J.; Armston, J.; Minor, D.M.; Duncanson, L.I.; May, P.B.; Kellner, J.R.; Dubayah, R. Assessing the performance of NASA’s GEDI L4A footprint aboveground biomass density models using National Forest Inventory and airborne laser scanning data in Mediterranean forest ecosystems. Forest Ecol. Manag. 2023, 538, 120975. [Google Scholar] [CrossRef]
- Ehlers, D.; Wang, C.; Coulston, J.; Zhang, Y.; Pavelsky, T.; Frankenberg, E.; Woodcock, C.; Song, C. Mapping forest aboveground biomass using multisource remotely sensed data. Remote Sens. 2022, 14, 1115. [Google Scholar] [CrossRef]
- Gupta, R.; Sharma, L.K. Aboveground biomass prediction by fusing GEDI footprints with optical and SAR data using the random forest in the mixed tropical forest, India. In Proceedings of the IGARSS 2022-2022 IEEE International Geoscience and Remote Sensing Symposium, Kuala Lumpur, Malaysia, 17–22 July 2022; pp. 5460–5463. [Google Scholar] [CrossRef]
- Wang, X.; Liu, C.; Lv, G.; Xu, J.; Cui, G. Integrating multi-source remote sensing to assess forest aboveground biomass in the Khingan mountains of north-eastern China using machine-learning algorithms. Remote Sens. 2022, 14, 1039. [Google Scholar] [CrossRef]
- Banda, F.; Giudici, D.; Le Toan, T.; Mariotti d’Alessandro, M.; Papathanassiou, K.; Quegan, S.; Riembaur, G.; Scipal, K.; Soja, M.; Tebaldini, S.; et al. The BIOMASS Level 2 Prototype Processor: Design and experimental results of above-ground biomass estimation. Remote Sens. 2020, 12, 985. [Google Scholar] [CrossRef]
- Wang, M.; Wang, Q.; Wang, Z.; Gao, Y.; Wang, J.; Cui, C.; Li, Y.; Ding, Z.; Wang, K.; Xu, C.; et al. Unlocking aerobatic potential of quadcopters: Autonomous freestyle flight generation and execution. Sci. Robot. 2025, 10, eadp9905. [Google Scholar] [CrossRef]
- Zhou, X.; Wen, X.; Wang, Z.; Gao, Y.; Li, H.; Wang, Q.; Yang, T.; Lu, H.; Cao, Y.; Xu, C.; et al. Swarm of micro flying robots in the wild. Sci. Robot. 2022, 7, eabm5954. [Google Scholar] [CrossRef]
- Yang, S.; Xing, Y.; Xing, T.; Deng, H.; Xi, Z. Multisensors fusion slam-aided forest plot mapping with backpack Dual-LiDAR System. IEEE Jstars. 2024, 17, 16051–16070. [Google Scholar] [CrossRef]
- Liu, H.; Xu, G.; Liu, B.; Li, Y.; Yang, S.; Tang, J.; Pan, K.; Xing, Y. A real time LiDAR-Visual-Inertial object level semantic SLAM for forest environments. Isprs J. Photogramm. 2025, 219, 71–90. [Google Scholar] [CrossRef]
- Jagadish, B.; Behera, M.D.; Prakash, A.J.; Paramanik, S.; Ghosh, S.M.; Patnaik, C.; Das, A. Indicating saturation limits of multi-sensor satellite data in estimating aboveground biomass of a mangrove forest. J. Indian Soc. Remote. 2024, 52, 2483–2500. [Google Scholar] [CrossRef]
- Wang, Q.; Putri, N.A.; Gan, Y.; Song, G. Combining both spectral and textural indices for alleviating saturation problem in forest LAI estimation using Sentinel-2 data. Geocarto Int. 2022, 37, 10511–10531. [Google Scholar] [CrossRef]
- Tian, Z.; Fan, J.; Yu, T.; de Leon, N.; Kaeppler, S.M.; Zhang, Z. Mitigating NDVI saturation in imagery of dense and healthy vegetation. Isprs J. Photogramm. 2025, 227, 234–250. [Google Scholar] [CrossRef]
- Gao, S.; Zhong, R.; Yan, K.; Ma, X.; Chen, X.; Pu, J.; Gao, S.; Qi, J.; Yin, G.; Myneni, R.B. Evaluating the saturation effect of vegetation indices in forests using 3D radiative transfer simulations and satellite observations. Remote Sens. Environ. 2023, 295, 113665. [Google Scholar] [CrossRef]
| Product | Weight (%) | ||
|---|---|---|---|
| RMAD | RSSE | PVTP | |
| GEDI biomass data | 13.95 | 13.01 | 73.04 |
| CCI biomass data | 16.44 | 15.09 | 68.47 |
| Product | Score | |
|---|---|---|
| Non-Representative | Representative | |
| GEDI biomass data | <0.75 | ≥0.75 |
| CCI biomass data | <0.70 | ≥0.70 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Wang, X.; Ji, P.; Li, H.; Wei, S.; Peng, D. Evaluating Forest Aboveground Biomass Products by Incorporating Spatial Representativeness Analysis. Remote Sens. 2025, 17, 2898. https://doi.org/10.3390/rs17162898
Wang Y, Wang X, Ji P, Li H, Wei S, Peng D. Evaluating Forest Aboveground Biomass Products by Incorporating Spatial Representativeness Analysis. Remote Sensing. 2025; 17(16):2898. https://doi.org/10.3390/rs17162898
Chicago/Turabian StyleWang, Yin, Xiaohui Wang, Ping Ji, Haikui Li, Shengrong Wei, and Daoli Peng. 2025. "Evaluating Forest Aboveground Biomass Products by Incorporating Spatial Representativeness Analysis" Remote Sensing 17, no. 16: 2898. https://doi.org/10.3390/rs17162898
APA StyleWang, Y., Wang, X., Ji, P., Li, H., Wei, S., & Peng, D. (2025). Evaluating Forest Aboveground Biomass Products by Incorporating Spatial Representativeness Analysis. Remote Sensing, 17(16), 2898. https://doi.org/10.3390/rs17162898