1. Introduction
Phytoplankton account for approximately half of global primary production via photosynthesis [
1] and form the base of the marine food web. Intracellular pigments of phytoplankton, composed of chlorophylls (a, b and c), carotenoids (carotenes and xanthophylls) and phycobiliproteins (phycoerythrin, phycocyanin and allophycocyanin) [
2], play a vital role in photoprotection and the light-driven part of photosynthesis. Chlorophyll-b, -c and photosynthetic carotenoids (PSC), such as fucoxanthin (Fuco), act as antenna pigments that transfer the light energy to chlorophyll-a in the photosynthetic reaction centers of photosystems, assisting in light harvesting for photosynthesis. In cyanobacteria, red algae, and cryptophytes, phycobiliproteins are the major light-harvesting pigments [
3]. Chlorophyll-a is crucial in converting the received light energy to chemically bonded energy. The carotenoids not involved in photosynthesis are photoprotective (PPC). In particular, some xanthophylls such as violaxanthin (Viola), zeaxanthin (Zea), diadinoxanthin (Diadino) and diatoxanthin (Diato) are involved in the xanthophyll cycle, one of the most important photoprotective mechanisms that drives the non-radiative dissipation of the excess light energy to prevent photoinhibition [
4,
5]. Therefore, their relative abundance can be used as a tracer of photoacclimation processes [
5].
In the context of global climate change, knowledge of the distributions of phytoplankton pigments is useful to understand the impacts of the changing environment on primary productivity [
6], phytoplankton diversity and community composition through appropriate analysis, for example, CHEMTAX [
7] and diagnostic pigment analysis [
8]. In remote sensing applications, phytoplankton pigment databases have been extensively used to develop, validate, or refine bio-optical algorithms for estimating phytoplankton biomass (often estimated using total chlorophyll-a (TChl-a) concentration) and functional types (via diagnostic pigment analysis) based on both cell size (micro-, nano- and pico-phytoplankton) and biogeochemical functions (e.g., calcification, silicification, dimethyl sulphide production and nitrogen fixation) [
9] and references therein. These data sets are mainly based on high-performance liquid chromatography (HPLC) analysis of discrete water samples. This technique enables the accurate quantification of 25–50 pigments in a single analysis [
10]. However, it requires highly trained personnel, intensive labor and time, expensive and complex analysis, and is limited by the sampling frequency, spatial coverage and additional issues related to discrete sampling such as sample handling, storage and transportation. While HPLC pigment analysis remains indispensable, it is necessary to explore methods that enable easier access to pigment data at higher spatial-temporal resolution.
Because optical measurements are currently the only means of collecting synoptic scale information on upper ocean particles (e.g., operational open-ocean satellite ocean color provides data daily with pixel size down to 300 m by 300 m), attempts have been made to quantify the concentrations of various phytoplankton pigments from these measurements (e.g., absorption or reflectance spectra). Optical methods take advantage of the distinctive absorption characteristics of different pigments and various approaches are applied, such as the decomposition of spectra into Gaussian functions, e.g., [
11], spectral reconstruction, e.g., [
12], derivative analysis, e.g., [
13], partial least squares regression, e.g., [
14], multiple linear regression [
15], reflectance band ratio, e.g., [
16,
17], principal component analysis, e.g., [
18] and artificial neural networks [
19,
20].
The Gaussian decomposition method decomposes phytoplankton absorption spectra (
) into Gaussian functions and correlates the amplitudes of the Gaussian functions with the concentrations of major pigment groups. The amplitude of each Gaussian function is assumed to represent the magnitude of the absorption coefficient of a specific pigment or pigment group at the Gaussian peak wavelength, based on known pigment absorption properties determined in laboratory analyses. This method simultaneously retrieves the concentrations of chlorophyll-a, chlorophyll-b, chlorophyll-c and carotenoids [
11,
21,
22,
23] or of chlorophyll-a and phycocyanin [
24,
25]. However, the retrieval accuracy is generally limited by the variations in pigment package effect of field samples. Nevertheless, the Gaussian absorption coefficients of specific pigment groups were recently incorporated into the reconstruction of hyper- and multi-spectral remote sensing reflectance, allowing the robust estimation of the concentrations of TChl-a, total chlorophyll-b (TChl-b), the combination of chlorophyll-c1 and -c2 (Chl-c1/2) and PPC globally [
23] as well as of phycocyanin in cyanobacteria bloom waters [
24,
25] from remote sensing reflectance data.
The spectral reconstruction method assumes that
can be reconstructed from the linear combination of pigment-specific absorption coefficients multiplied by corresponding pigment concentrations [
26]. Moisan et al. [
27,
28] applied matrix inversion analysis to the reconstruction model and successfully estimated the concentrations of a series of pigments directly from
. This technique involves a first inversion of the observed pigment concentrations that derives pigment-specific absorption spectra and a second inversion of these derived pigment-specific absorption spectra that solves for pigment concentrations. Four methods that solve least squares problems, i.e., singular value decomposition (SVD) [
29], non-negative least squares (NNLS) [
30] and two nonlinear least squares minimization schemes based on the Levenberg–Marquardt algorithm [
31,
32] were compared for the two inversions. They found that when the first inversion was carried out with SVD and the second one with NNLS, the inverse modeling technique yielded the most accurate pigment estimates. However, the retrieval accuracy is affected by the level of correlation between pigment concentrations, the contribution of a specific pigment to the spectral
, pigment package effect, the missing absorption components by the pigments that exist in the samples but are not obtained by standard HPLC (e.g., mycosporine-like amino acids and phycobiliproteins) [
27,
28], and the number of spectral bands of
used in the inversion model [
27,
28,
33]. Overall, the SVD-NNLS method achieved simultaneous statistically significant retrievals of TChl-a, total chlorophyll-c (TChl-c),
-carotene (
-Caro), Fuco, Viola, Diadino and peridinin (Peri) in U.S. east coast waters [
27,
28]. It was recently applied to
modeled from MODIS-Aqua TChl-a data for northeastern U.S. waters, yielding maps of the concentrations of ten pigments [
34]. Similar approaches were successful in infering phytoplankton size classes globally [
35,
36] and taxonomic groups in the Chukchi and Bering Seas [
33] from absorption data.
Derivative analysis of absorption spectra separates the secondary absorption peaks and shoulders contributed by phytoplankton pigments within the overlapping absorption regions [
37]. Bidigare et al. [
13] found that the fourth derivative maxima of particulate absorption spectra (
) provided strong linear relationships with chlorophylls (a, b and c) concentrations in Sargasso Sea. However, this method failed to estimate carotenoid concentrations because of the similarity of their spectral properties, the broad spectral absorption and relatively rounded absorption peaks that are less accessible to derivative analysis.
Principal component analysis (a.k.a. empirical orthogonal function analysis) derives several dominant modes (known as “principal components”) of the spectra that mainly account for the variability in spectral shape and relates them to pigment concentrations. Bracher et al. [
18] performed this analysis on both hyperspectral and multispectral remote sensing reflectance data and retrieved the concentrations of TChl-a, monovinyl-chlorophyll-a, PPC, PSC, Chl-c1/2, 19
-butanoyloxyfucoxanthin (But), 19
-hexanoyloxyfucoxanthin (Hex), Zea, phycoerythrin and the sum of
- and
-Caro from the linear combinations of the principal components in the Atlantic Ocean. This method is, however, only applicable to the pigments that have been identified in most collocated samples. It failed to retrieve the pigments that are mostly absent or below detection limit. Similarly, Soja–Woźniak et al. [
38] applied this analysis on both hyperspectral and multispectral remote sensing reflectance data and successfully retrieved TChl-a, phycocyanin and phycoerythrin in the Gulf of Gdansk.
An artificial neural network relates spectra to pigment data with a nonlinear model that self-adjusts the model parameters (i.e., weight matrix) for the best fit. Bricaud et al. [
19] developed a multilayer perceptron using a global data set and obtained estimations of the concentrations of TChl-a, TChl-b, TChl-c, PSC and PPC, with TChl-a and TChl-b being the most accurate and poorest estimates, respectively. The main limitation of this method lies in the biological variability embedded in the training data set.
More recently, there has been an increased use of in situ hyperspectral optical sensors to obtain pigment data from continuous optical measurements, e.g., [
22]. In-line and autonomous measurements by new miniature sensors deployed on various platforms (e.g., profiling floats, autonomous surface water vehicles) have substantially increased the sampling frequency and spatial coverage of measurements. The shipboard underway spectrophotometry considerably facilitates the acquisition of
with unprecedented spatial resolution. It utilizes an AC-S hyperspectral spectrophotometer (or the 9-wavelength resolved AC-9) (Sea-Bird Scientific, Philomath, OR, USA) operated in flow-through mode and derives
by differencing the bulk seawater absorption measurements from temporally adjacent 0.2-μm filtered water sample measurements, e.g., [
22,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48]. It has provided surface TChl-a data along cruise tracks via the empirical relationships between the spectrophotometry derived
and HPLC measured TChl-a concentrations [
39,
40,
45,
46,
47,
48]. Furthermore, Gaussian decomposition has been performed by Chase et al. [
22] to retrieve major pigment groups from a globally extensive underway AC-S derived
data set. Here we use a data set obtained with a similar underway system to compare and contrast two different methods to obtain information on the underlying pigments.
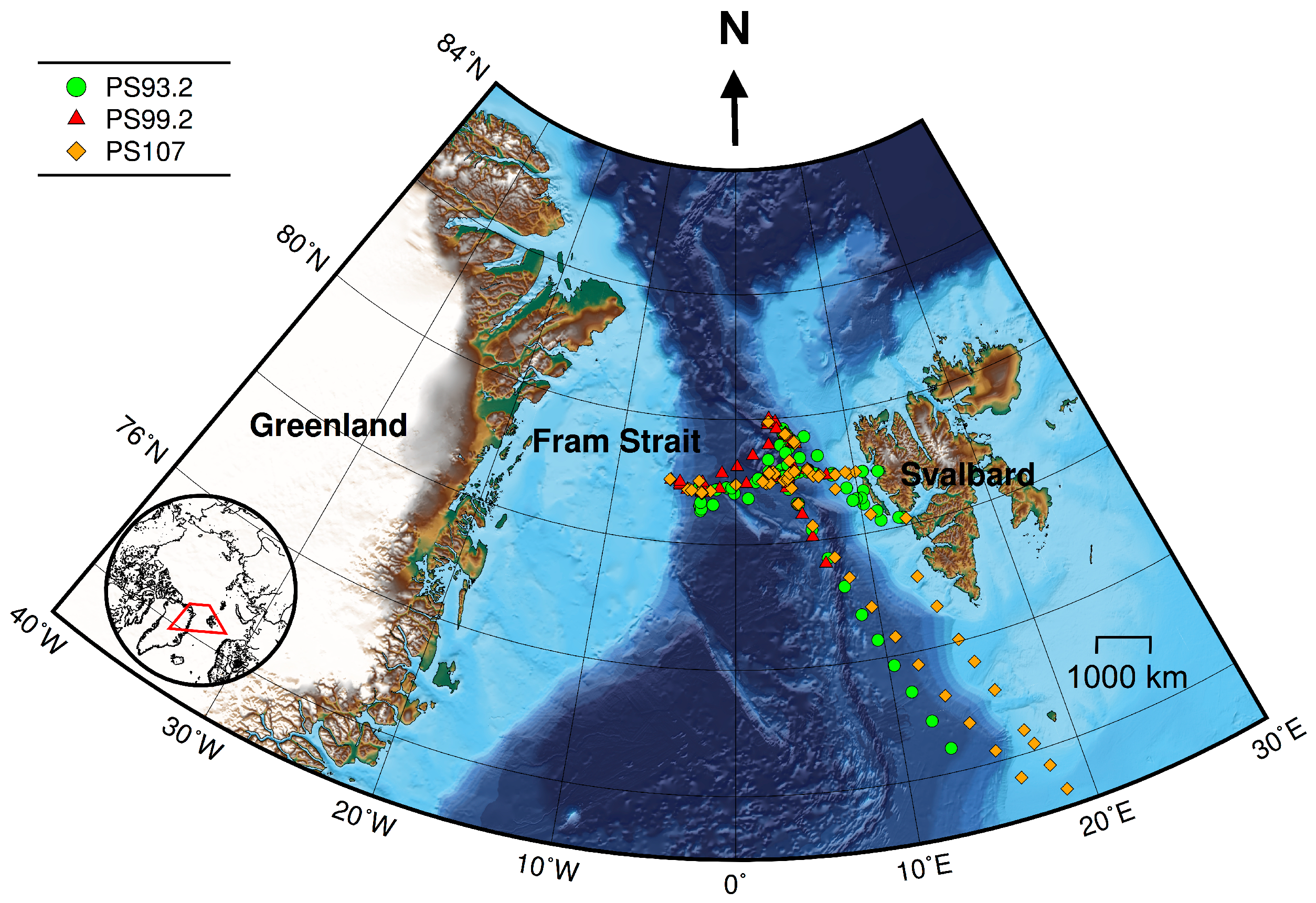
The Fram Strait, the region between Svalbard and Greenland, provides the only deep connection between the North Atlantic and Arctic Oceans (
Figure 1). It is of great importance to the climate in the Arctic region, as it accounts for 75% of the mass exchange and 90% of the heat exchange between the Arctic Ocean and the rest of the world’s ocean [
49]. In recent decades, the Fram Strait has undergone a significant warming, high variability of Atlantic water inflow [
50] and an overall increase of sea ice area export [
51,
52,
53,
54,
55]. This impacts phytoplankton biomass, community composition and distribution by altering light and nutrient regimes. The seasonal cycle of phytoplankton biomass has been significantly enhanced in the shallow upper water layers since 2008 [
56]. Phytoplankton distributions reflect the dominant local physical processes [
56,
57]. A significant increase in summertime chlorophyll-a concentration in the eastern Fram Strait was observed, whereas on the western side there were minor changes [
56]. Furthermore, a shift of dominant phytoplankton assemblages from diatoms (mainly
Thalassiosira spp.,
Chaetoceros spp. and
Fragilariopsis spp.) towards coccolithophores (mainly
Emiliania huxleyi) and more recently,
Phaeocystis spp. (mainly
Phaeocystis pouchetii) and other small pico- and nanoflagellates during summer months was suggested [
56,
58,
59], which can strongly affect the functioning and stability of marine food webs [
60,
61]. The studies of phytoplankton community composition in this region are mainly based on discrete water samples or moored sediment traps. Because of the inherent limitations of these methods, the observations are scarce. Furthermore, it remains difficult to obtain information on phytoplankton community composition via satellite due to the poor spatial-temporal coverage of ocean color data in this region, e.g., [
57] and the lack of assessment of the applicability of satellite algorithms determining the phytoplankton community structure for this region. Additionally, algorithms applicable to other waters for quantifying phytoplankton community structure or pigment composition from in situ optical measurements have not been assessed yet in this region.
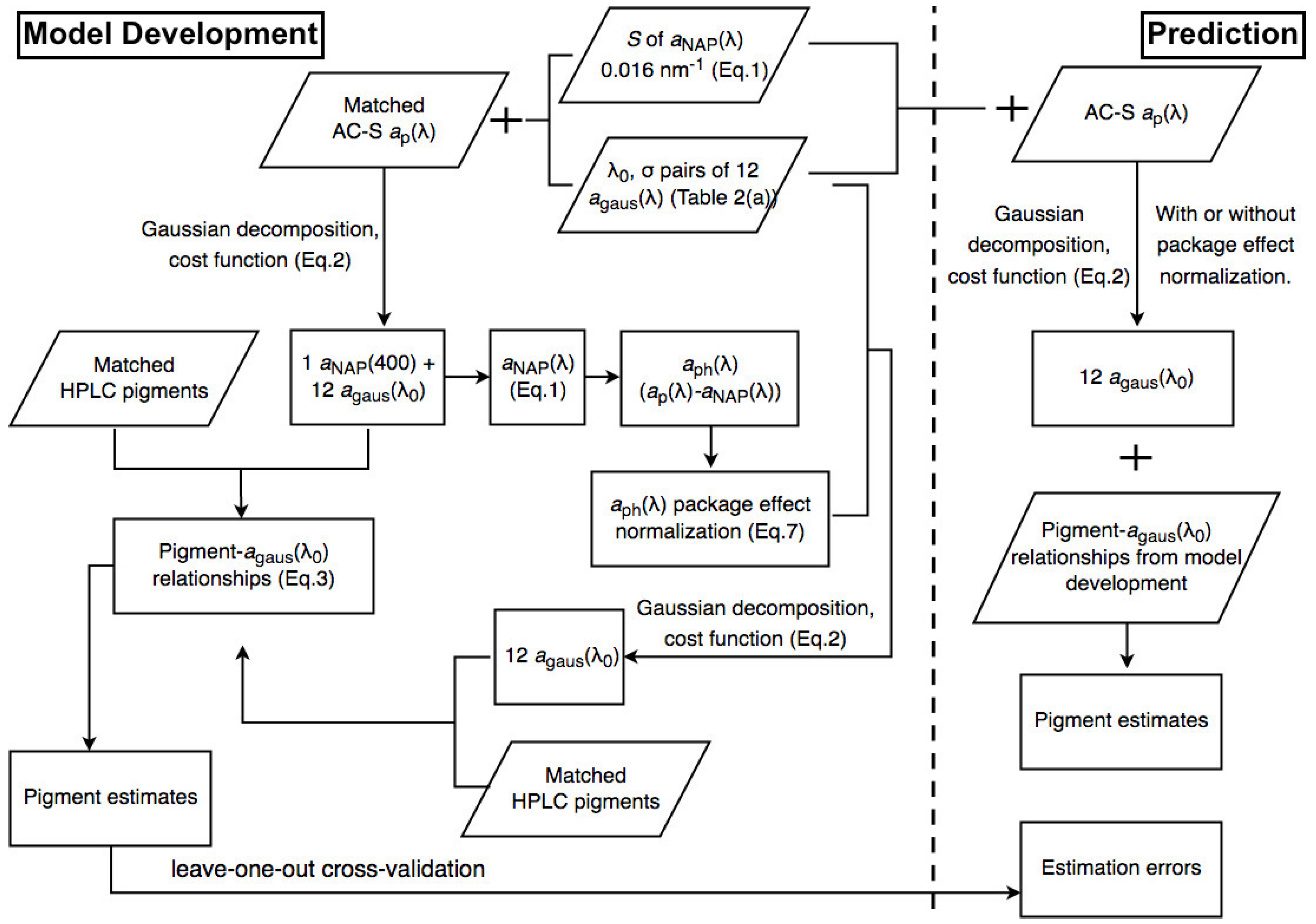
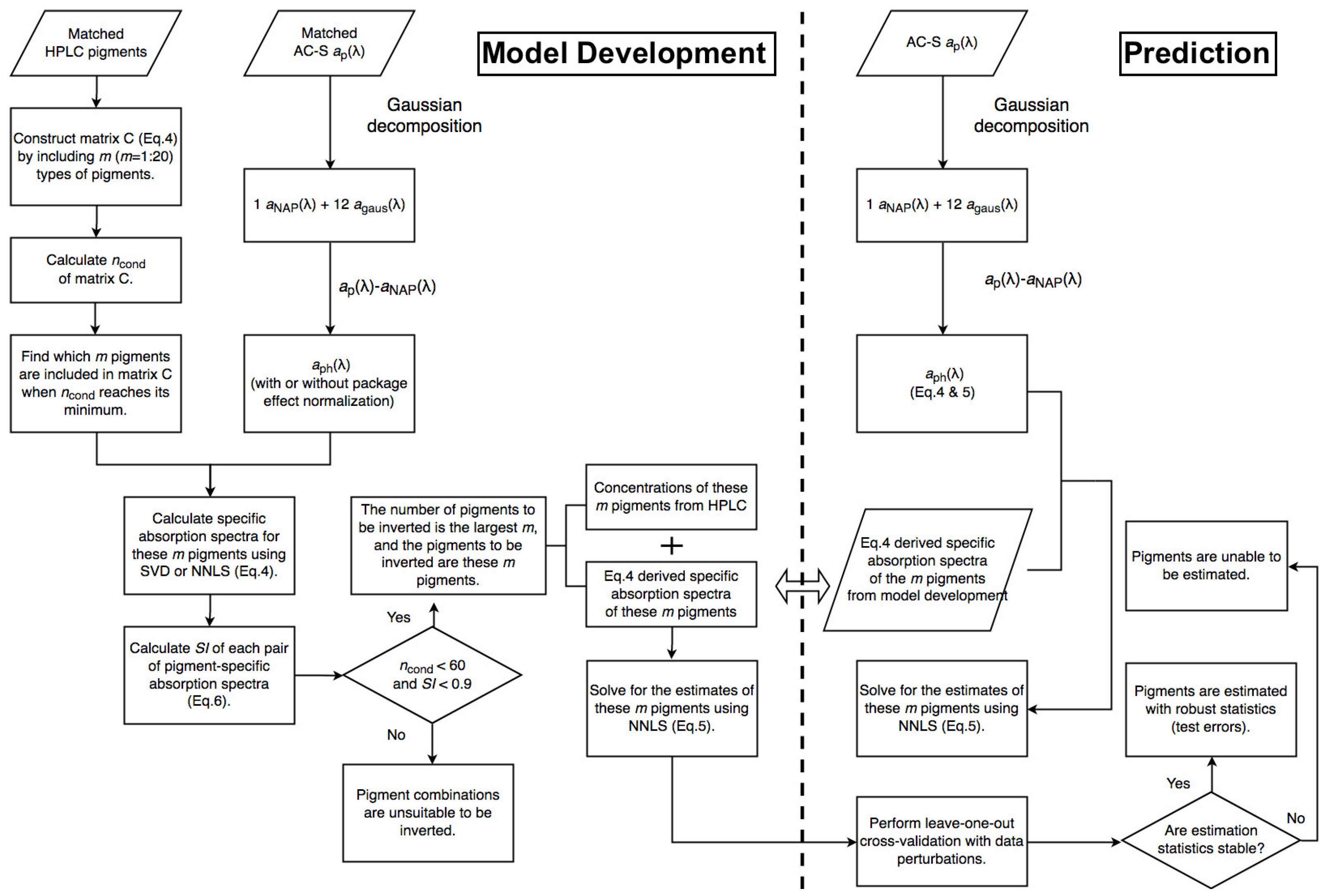
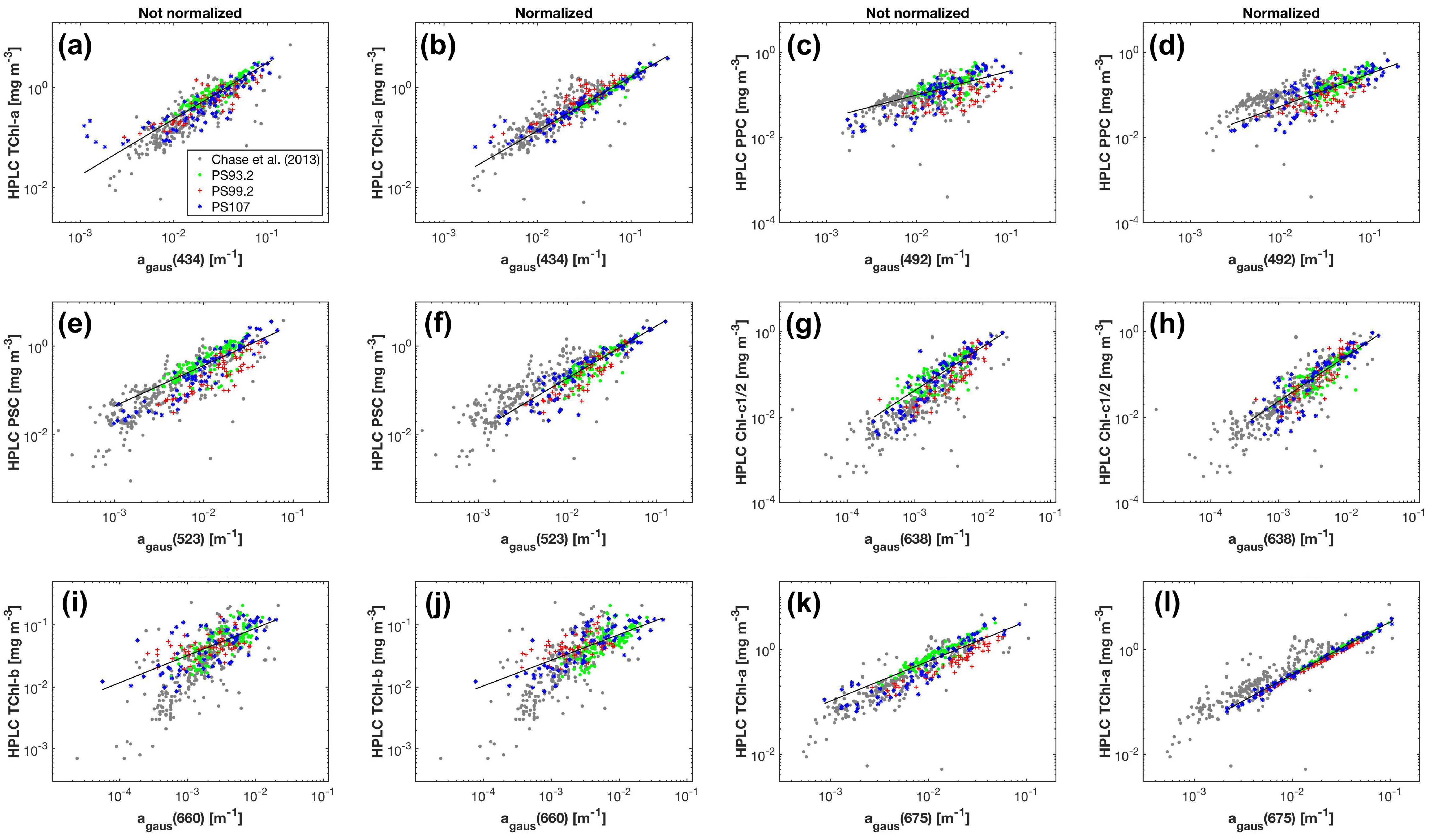
The Fram Strait cruises PS93.2, PS99.2 and PS107 on R/V Polarstern collected a comprehensive in situ bio-optical data set and offer a unique opportunity for bio-optical modeling. In particular, underway spectrophotometry was applied during all three cruises. To obtain the information of individual phytoplankton pigments or pigment groups (e.g., PSC and PPC) from underway spectrophotometry, here, we compare and optimize the performances of two pigment retrieval approaches, Gaussian decomposition [
22] and the matrix inversion technique [
27,
28], find the potential number and types of pigments that can be retrieved, and assess the applicability of the two approaches to the Fram Strait and its vicinity.
5. Conclusions
We demonstrated the retrieval of high spatially resolved phytoplankton pigment concentrations in the Fram Strait and its vicinity from underway hyperspectral
(400–700 nm, ~3.5 nm wavelength resolution) by the application of Gaussian decomposition [
22] and the matrix inversion technique [
27]. Gaussian decomposition enables robust predictions of Gauss-5 pigments (MPE 21–34%). Improved retrieval accuracy was obtained by normalizing the
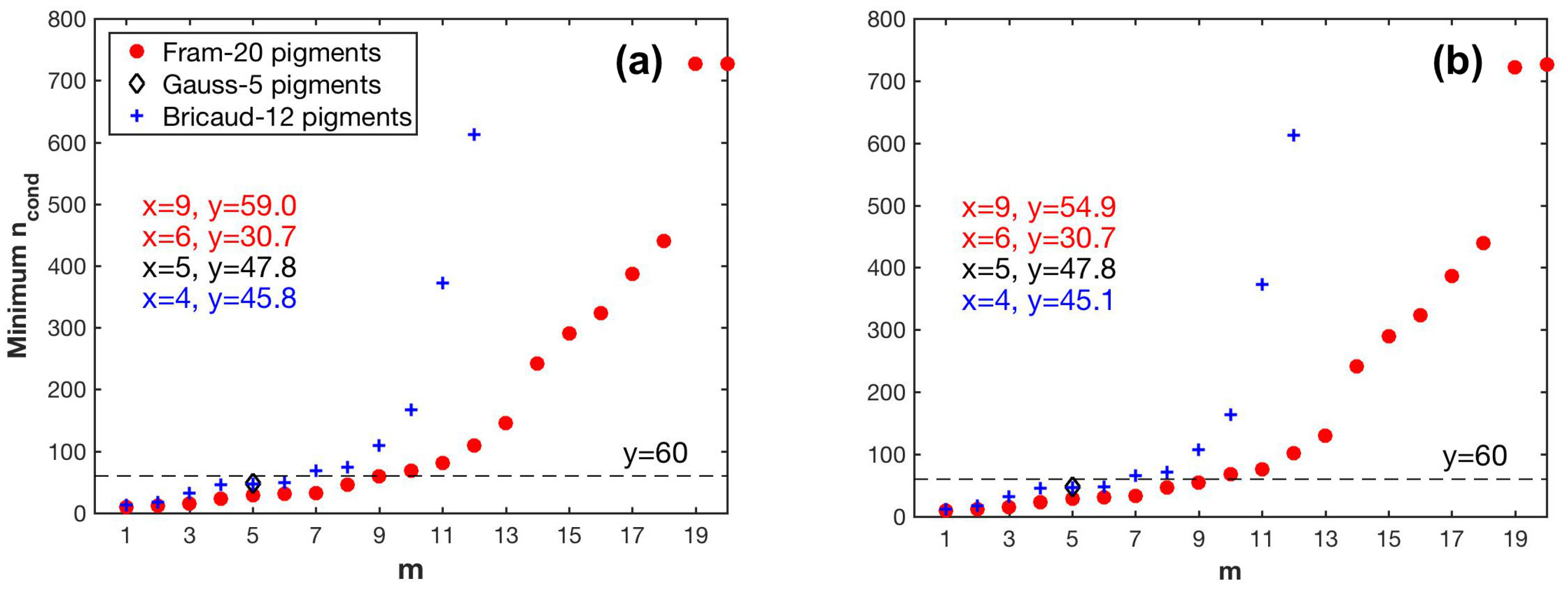
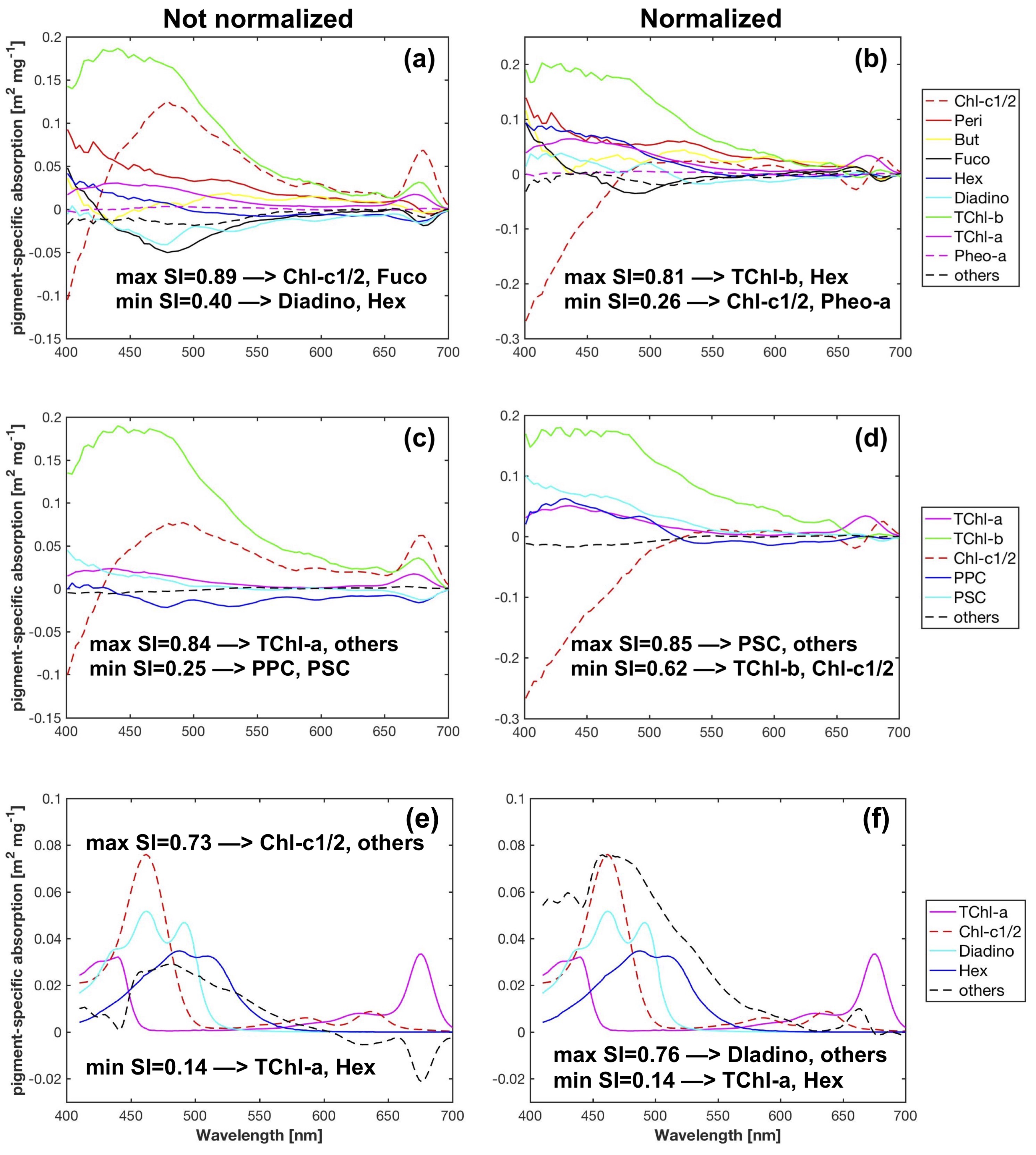
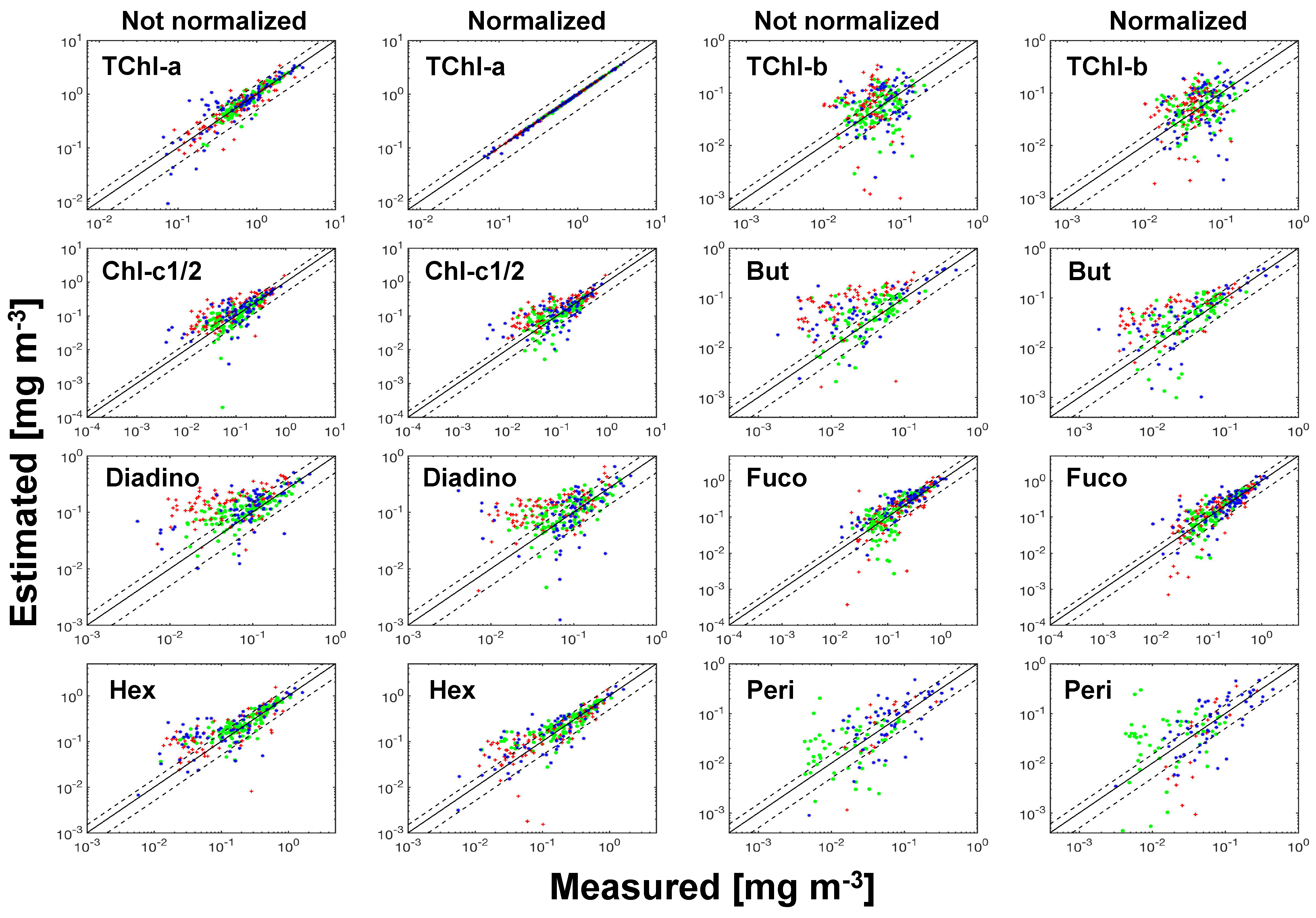
spectra with the pigment package effect factor at 675 nm. For the matrix inversion technique, although SVD cannot guarantee the derivation of non-negative pigment-specific absorption spectra, it generates more accurate pigment estimates compared to the NNLS derived spectra or the measured spectra from pigments in solution. To minimize the effect of the ill-conditioned matrices on pigment retrieval accuracy, we propose an innovative approach in selecting the pigments to be determined based on the combined use of data perturbations and leave-one-out cross-validation to generate robust pigment estimation statistics. Considering the overall pigment retrieval accuracy, SVD-NNLS-9 performed best among the three SVD-NNLS methods. The SVD-NNLS-9 method enables the robust estimations of six pigments (MPE 16-65%), i.e., TChl-a, TChl-b, Chl-c1/2, Diadino, Fuco and Hex, and two more being less accuraely estimated (MPE 67–76%), i.e., But and Peri, with the application of the package effect normalization. Gaussian decomposition outperforms SVD-NNLS-5
in retrieving the TChl-b, Chl-c1/2, PPC and PSC, while both methods show similar capability in estimating TChl-a.
The matrix inversion technique has the advantage of retrieving the concentrations of several specific carotenoids, which is currently not accomplished by Gaussian decomposition, derivative analysis [
13], partial least squares regression [
14], and multiple linear regression [
15]. However, its performance is sensitive to input errors when the input matrix is to some extent ill-conditioned. Therefore, sensitivity analysis such as the one based on data perturbations used in our study is always needed when assessing the performance of the matrix inversion technique in retrieving phytoplankton pigments or pigment related parameters. Future studies using methods such as principle component analysis and artificial neural network may show promise to obtain not only chlorophylls but also different types of carotenoids in our study area.
In addition to the number of pigments, the number of spectral bands used for pigment retrieval also significantly influence the performance of the matrix inversion technique. Compared with the results using hyperspectral , the number of pigments able to be retrieved by SVD-NNLS-9 was reduced to four, i.e., TChl-a, TChl-b, Chl-c1/2 and Hex, with increased estimation errors, especially for Chl-c1/2 and Hex, when using multispectral (at ten MODIS bands). This suggests the advantage of using hyperspectral data for increasing the accuracy of phytoplankton pigment retrievals. It follows that inverted from hyperspectral remote sensing reflectance measured by in situ or satellite radiometry has a greater potential for the application of Gaussian decomposition and the matrix inversion technique than multispectral radiometric measurements.
To apply Gaussian decomposition or the matrix inversion technique to a study area, prior knowledge of concurrent AC-S derived and HPLC pigment concentrations in this region is necessary to derive either the regional -pigment concentration relationship or the regional pigment-specific absorption spectra. With this knowledge, we apply both approaches to underway AC-S measurements in times when no HPLC data is available. Given that proxy-relation may change in the future, it is imperative to always collect some HPLC data to validate that derived relations or coefficients are still consistent.
The application of the two methods to our data obtain in three Fram Strait expeditions enables the derivation of pigment data sets along the cruise tracks. Future work could build upon these results, by deriving phytoplankton functional types based on retrieved marker pigments from hyperspectral phytoplankton absorption as well as hyperspectral remote sensing reflectance data. Such a high resolution data set will strengthen the study of phytoplankton dynamics in responses to environmental variables in the context of climate change.