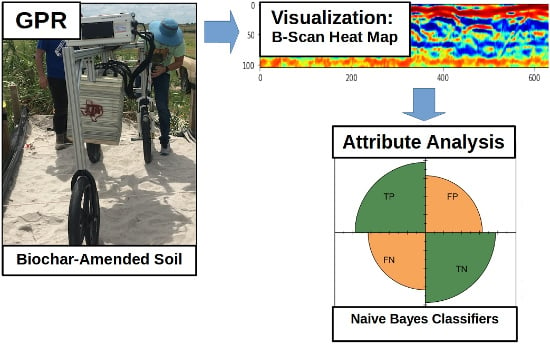
Quantification of Soil Organic Carbon in Biochar-Amended Soil Using Ground Penetrating Radar (GPR)
Abstract
1. Introduction
2. Materials and Methods
2.1. Material Preparation and Treatments
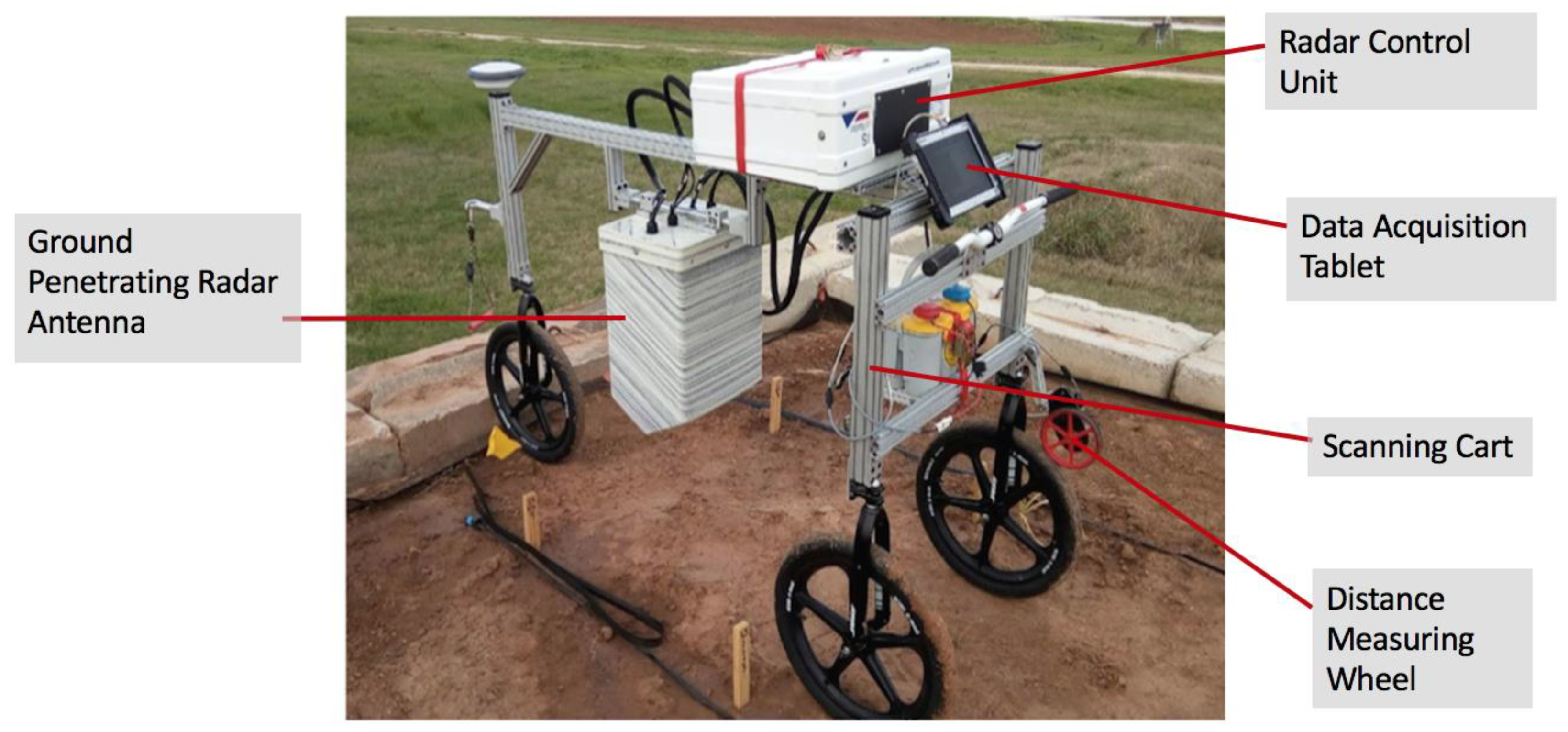
2.2. Data Collection
2.3. Data Processing and Analyses
2.4. Statistical Analysis
3. Results
3.1. Moisture Levels
3.2. Treatment Differences
3.3. Naïve Bayes Predictive Model
3.4. Wavelet Analysis
4. Discussion
Limitation of This Study
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Amundson, R.; Berhe, A.A.; Hopmans, J.W.; Olson, C.; Sztein, A.E.; Sparks, D.L. Soil and Human Security in the 21st Century. Science 2015, 348. [Google Scholar] [CrossRef]
- Montanarella, L.; Pennock, D.J.; McKenzie, N.; Badraoui, M.; Chude, V.; Baptista, I.; Mamo, T.; Yemefack, M.; Aulakh, M.S.; Yagi, K.; et al. World’s soils are under threat. Soil 2016, 2, 79–82. [Google Scholar] [CrossRef]
- Scharpenseel, H.W.; Becker-Heidmann, P. Sustainable Land Use in the Light of Resilience/Elasticity to Soil Organic Matter Functions. In Soil Resilience and Sustainable Land Use, 2nd ed.; Greenland, D.J., Szabolcs, I., Eds.; CAB International: Wallingford, Oxon, UK, 1994; pp. 249–262. [Google Scholar]
- Lal, R. World Cropland Soils as a Source or Sink for Atmospheric Carbon. Adv. Agron. 2004, 71, 145–191. [Google Scholar] [CrossRef]
- Sanderman, J.; Hengl, T.; Fiske, G.J. Soil Carbon Debt of 12,000 Years of Human Land Use. Proc. Natl. Acad. Sci. USA 2017, 114, 9575–9580. [Google Scholar] [CrossRef] [PubMed]
- Bruun, E.W.; Ambus, P.; Egsgaard, H.; Hauggaard-Nielsen, H. Effects of slow and fast pyrolysis biochar on soil C and N turnover dynamics. Soil Biol. Biochem. 2012, 46, 73–79. [Google Scholar] [CrossRef]
- Fowles, M. Black carbon sequestration as an alternative to bioenergy. Biomass Bioenergy 2007, 31, 426–432. [Google Scholar] [CrossRef]
- Woolf, D.; Amonette, J.E.; Street-Perrott, F.A.; Lehmann, J.; Joseph, S. Sustainable Biochar to Mitigate Global Climate Change. Nat. Commun. 2010, 1, 56. [Google Scholar] [CrossRef]
- Clough, T.J.; Bertram, J.E.; Ray, J.L.; Condron, L.M.; O’Callaghan, M.; Sherlock, R.R.; Wells, N.S. Unweathered wood biochar impact on nitrous oxide emissions from a bovine-urine-amended pasture soil. Soil Sci. Soc. Am. J. 2010, 74, 852–860. [Google Scholar] [CrossRef]
- Van Zwieten, L.; Kimber, S.; Morris, S.; Downie, A.; Berger, E.; Rust, J.; Scheer, C. Influence of biochars on flux of N2O and CO2 from Ferrosol. Soil Res. 2010, 48, 555–568. [Google Scholar] [CrossRef]
- Shackley, S.; Hammond, J.; Gaunt, J.; Ibarrola, R. The Feasibility and Costs of Biochar Deployment in the UK. Carbon Manag. 2011, 2, 335–356. [Google Scholar] [CrossRef]
- Schumacher, B.A. Methods for the Determination of Total Organic Carbon (TOC) in Soils and Sediments; U.S. Environmental Protection Agency: Washington, DC, USA, 2002.
- Hammes, K.; Schmidt, M.W.; Smernik, R.J.; Currie, L.A.; Ball, W.P.; Nguyen, T.H.; Louchouarn, P.; Houel, S.; Gustafsson, Ö.; Elmquist, M.; et al. Comparison of quantification methods to measure fire-derived (black/elemental) carbon in soils and sediments using reference materials from soil, water, sediment and the atmosphere. Glob. Biogeochem. Cycles 2007, 21. [Google Scholar] [CrossRef]
- McClellan, M.; Comas, X.; Benscoter, B.; Hinkle, R.; Sumner, D. Estimating Belowground Carbon Stocks in Isolated Wetlands of the Northern Everglades Watershed, Central Florida, Using Ground Penetrating Radar and Aerial Imagery. J. Geophys. Res. Biogeosci. 2017, 122, 2804–2816. [Google Scholar] [CrossRef]
- El-Mahallawy, M.S.; Hashim, M. Material classification of underground utilities from GPR images using DCT-based SVM approach. IEEE Geosci. Remote Sens. Lett. 2013, 10, 1542–1546. [Google Scholar] [CrossRef]
- Anbazhagan, P.; Dixit, P.N.; Bharatha, T.P. Identification of type and degree of railway ballast fouling using ground coupled GPR antennas. J. Appl. Geophys. 2016, 126, 183–190. [Google Scholar] [CrossRef]
- Nicolotti, G.; Socco, L.V.; Martinis, R.; Godio, A.; Sambuelli, L. Application and comparison of three tomographic techniques for detection of decay in trees. J. Arboric. 2003, 29, 66–78. [Google Scholar]
- Cassidy, N.J. Electrical and magnetic properties of rocks, soil sand fluids. In Ground Penetrating Radar Theory and Applications; Jol, H.M., Ed.; Elsevier: Amsterdam, The Netherlands, 2008; pp. 41–67. [Google Scholar]
- Le Gall, A.; Ciarletti, V.; Berthelier, J.J.; Reineix, A.; Guiffaut, C.; Ney, R.; Dolon, F.; Bonaime, S. An imaging HF GPR using stationary antennas: Experimental validation over the Antarctic ice sheet. IEEE Trans. Geosci. Remote Sens. 2008, 46, 3975–3986. [Google Scholar] [CrossRef]
- Dogan, M.; Turhan-Sayan, G. Preprocessing of A-Scan GPR Data Based on Energy Features. In Proceedings of the Conference on Detection and Sensing of Mines, Explosive Objects, and Obscured Targets XXI, Baltimore, MD, USA, 17–21 April 2016; Volume 9823, p. 98231E. [Google Scholar] [CrossRef]
- Delgado, A.; Novo, A.; Hays, D.B. Data Acquisition Methodologies Utilizing Ground Penetrating Radar for Cassava (Manihot Esculenta Crantz) Root Architecture. Geoscience 2019, 9. [Google Scholar] [CrossRef]
- Zhang, X.; Derival, M.; Albrecht, U.; Ampatzidis, Y. Evaluation of a Ground Penetrating Radar to Map the Root Architecture of HLB-Infected Citrus Trees. Agronomy 2019, 9. [Google Scholar] [CrossRef]
- Olhoeft, G.R. Electrical properties from 10−3 Hz to 10+9 Hz—Physics and chemistry. In Proceedings of the American Institute of Physics Conference, Anaheim, CA, USA, 15–17 October 1987; pp. 281–298. [Google Scholar]
- Zheng, H.; He, X.; Li, L.; Zhang, W.; Xia, Y.; Su, Y.; Qiu, H.; Liu, S.; Chen, X. Modified Method for Estimating Organic Carbon Density in Discontinuous Karst Soil Using Ground-Penetrating Radar and Geostatistics. J. Mt. Sci. 2015, 12, 1229–1240. [Google Scholar] [CrossRef]
- Slater, L.; Hanson, P.J.; Kolka, R.K.; Parsekian, A.D.; Nolan, J.; Sebesteyen, S.D.; Ntarlagiannis, D. Uncertainty in Peat Volume and Soil Carbon Estimated Using Ground-Penetrating Radar and Probing. Soil Sci. Soc. Am. J. 2012, 76, 1911–1918. [Google Scholar] [CrossRef]
- Loisel, J.; Yu, Z.; Parsekian, A.; Nolan, J.; Slater, L. Quantifying Landscape Morphology Influence on Peatland Lateral Expansion Using Ground-Penetrating Radar (GPR) and Peat Core Analysis. J. Geophys. Res. Biogeosci. 2013, 118, 373–384. [Google Scholar] [CrossRef]
- Jol, H.M.; Smith, D.G. Ground Penetrating Radar Surveys of Peatlands for Oilfield Pipelines in Canada. J. Appl. Geophys. 1995, 34, 109–123. [Google Scholar] [CrossRef]
- Iryanti, M.; Nugraha, H.D.; Setiawan, T.; Bijaksana, S. Mapping Peat Morphology in Sag Pond with Ground Penetrating Radar. AIP Conf. Proc. 2013, 1554, 265–268. [Google Scholar] [CrossRef]
- Darusman, T.; Kristiyono, A.; Warren, M.; Slater, L.; Kolka, R.; Sudiana, N.; Terry, N.; Comas, X.; Nurjaman, D. Imaging Tropical Peatlands in Indonesia Using Ground-Penetrating Radar (GPR) and Electrical Resistivity Imaging (ERI): Implications for Carbon Stock Estimates and Peat Soil Characterization. Biogeosciences 2015, 12, 2995–3007. [Google Scholar] [CrossRef]
- Morris, I.; Glisic, B. GPR Attribute Analysis for Material Property Identification. In Proceedings of the 2017 9th International Workshop on Advanced Ground Penetrating Radar (IWAGPR 2017), Edinburgh, UK, 28–30 June 2017; pp. 1–5. [Google Scholar] [CrossRef]
- Zhao, W.; Forte, E.; Pipan, M.; Tian, G. Ground Penetrating Radar (GPR) Attribute Analysis for Archaeological Prospection. J. Appl. Geophys. 2013, 97, 107–117. [Google Scholar] [CrossRef]
- McCallum, A.; Nigam, K. A comparison of event models for naive bayes text classification. In Proceedings of the AAAI-98 Workshop on Learning for Text Categorization, Madison, WI, USA, 26–27 July 1998; Volume 52, pp. 41–48. [Google Scholar]
- Rennie, J.D.; Shih, L.; Teevan, J.; Karger, D.R. Tackling the poor assumptions of naive bayes text classifiers. In Proceedings of the 20th International conference on machine learning (icml-03), Washington, DC, USA, 21–24 August 2003; pp. 616–623. [Google Scholar]
- Pang, B.; Lee, L.; Vaithyanathan, S. Thumbs up? sentiment classification using machine learning techniques. In Proceedings of the ACL-02 Conference on Empirical Methods in Natural Language Processing; Association for Computational Linguistics: Stroudsburg, PA, USA, 2002; Volume 10, pp. 79–86. [Google Scholar]
- Rish, I. An empirical study of the naive Bayes classifier. In Proceedings of the IJCAI 2001 Workshop on Empirical Methods in Artificial Intelligence, Seattle, WA, USA, 4–6 August 2001; Volume 3, pp. 41–46. [Google Scholar]
- Lee, D.T.; Yamamoto, A. Wavelet analysis: Theory and applications. Hewlett Packard J. 1994, 45, 44. [Google Scholar]
- Javadi, M.; Ghasemzadeh, H. Wavelet analysis for ground penetrating radar applications: A case study. J. Geophys. Eng. 2017, 14, 1189–1202. [Google Scholar] [CrossRef]
- Heil, C.E.; Walnut, D.F. Continuous and discrete wavelet transforms. SIAM Rev. 1989, 31, 628–666. [Google Scholar] [CrossRef]
- Steelman, C.M.; Endres, A.L. Comparison of petrophysical relationships for soil moisture estimation using GPR ground waves. Vadose Zone J. 2011, 10, 270–285. [Google Scholar] [CrossRef]
- Benedetto, A.; Tosti, F.; Ortuani, B.; Giudici, M.; Mele, M. Mapping the spatial variation of soil moisture at the large scale using GPR for pavement applications. Near Surf. Geophys. 2015, 13, 269–278. [Google Scholar] [CrossRef]
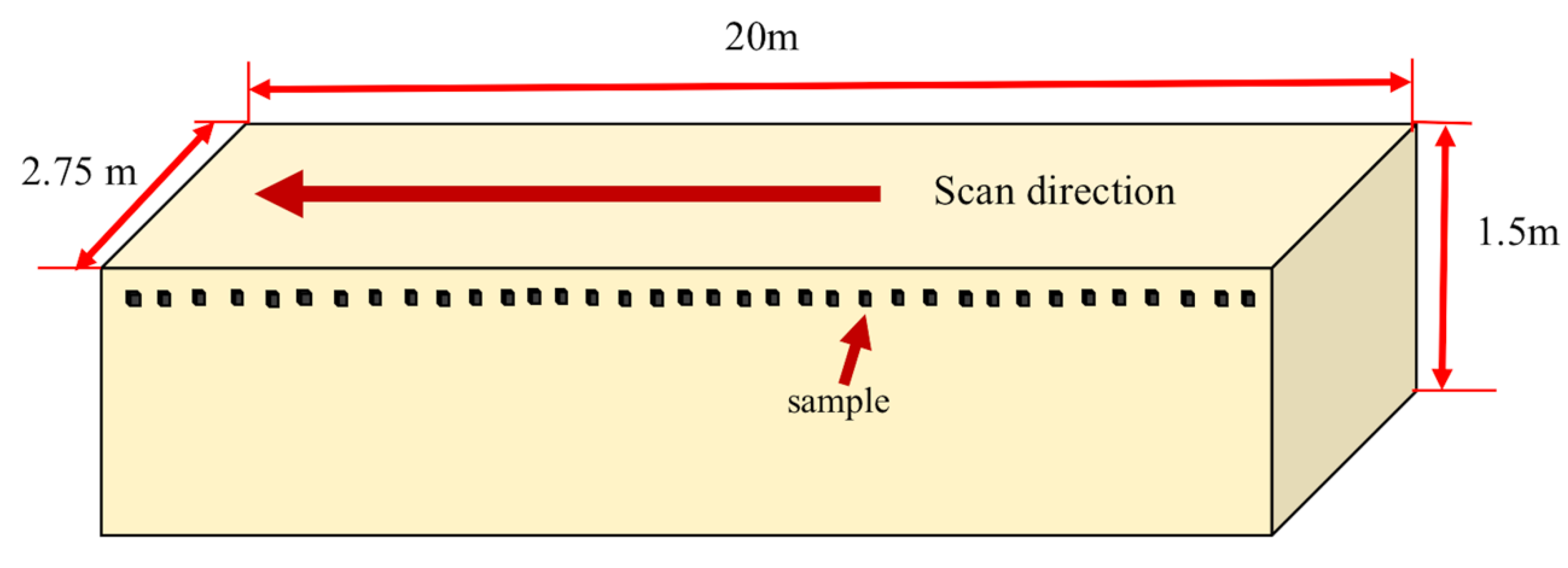
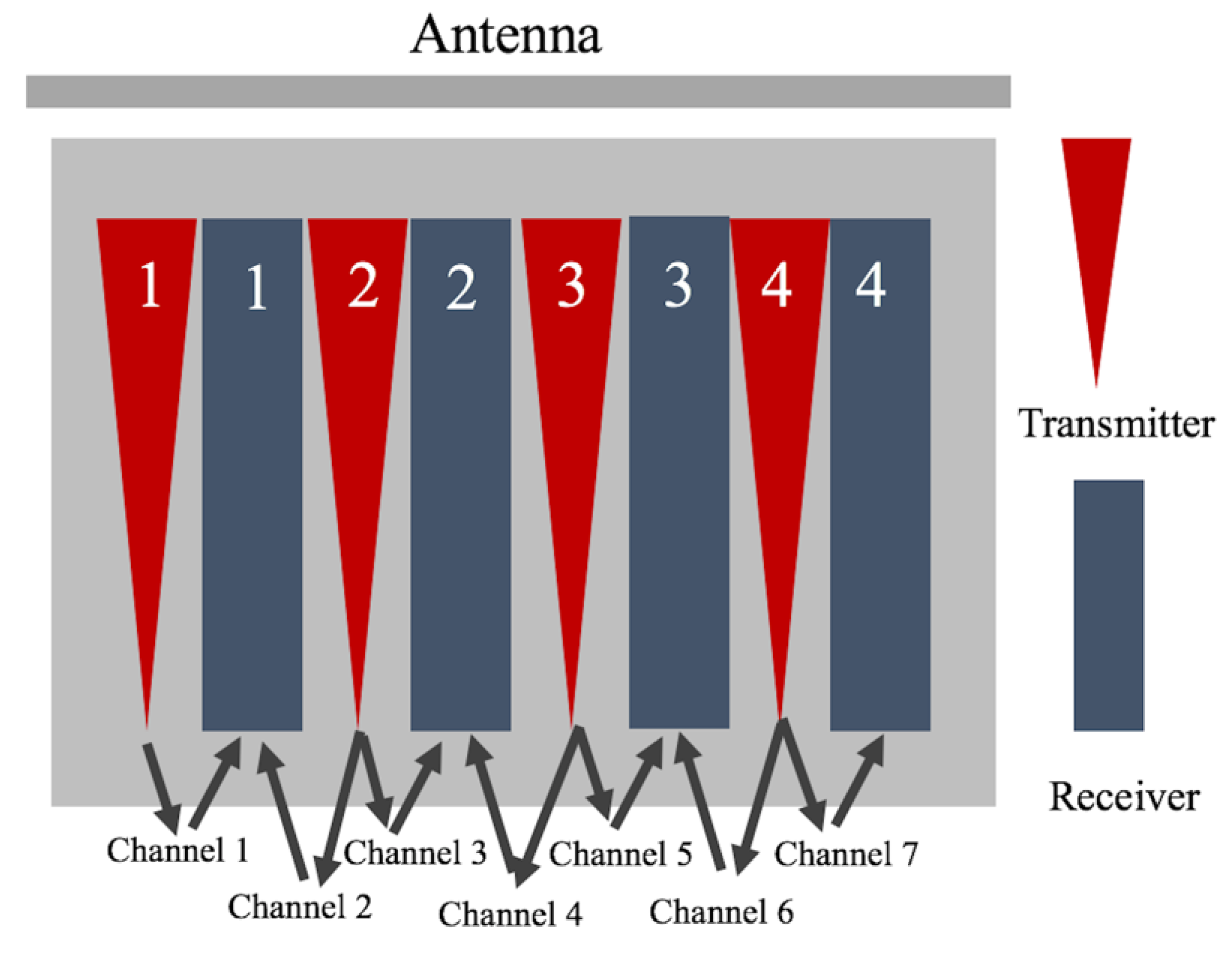

| Treatment Number | Treatment | Carbon Content by Weight Per Sample (g) | Carbon Content by Percent Per Sample (%) |
|---|---|---|---|
| 1 | 0% Biochar | 0 | 0 |
| 2 | 2% Biochar | 7.5012 | 1.0840 |
| 3 | 4% Biochar | 13.6150 | 2.1680 |
| 4 | 6% Biochar | 18.7640 | 3.2520 |
| 5 | 8% Biochar | 24.0214 | 4.3360 |
| 6 | 10% Biochar | 28.9428 | 5.4200 |
| 7 | 50% Biochar | 67.7500 | 27.1000 |
| 8 | 100% Biochar | 86.7200 | 54.2000 |
| 9 | 50% Graphite | 250 | 50 |
| 10 | 100% Graphite | 430 | 100 |
| 11 | 50% Activated Carbon | 200 | 40 |
| 12 | 100% Activated Carbon | 256 | 80 |
| Attributes | Spearman’s Correlation Coefficient | p-Value |
|---|---|---|
| Maximum amplitude | 0.1930 | <2.20 × 10−16 *** |
| Intensity | 0.1627 | 1.456 × 10−14 *** |
| Area | 0.0874 | 3.912 × 10−5 *** |
| Energy | 0.2278 | <2.20 × 10−16 *** |
| Attributes | Pearson Correlation Coefficient | p-Value |
|---|---|---|
| Maximum amplitude | 0.2822 | 3.706 × 10−12 *** |
| Intensity | 0.2821 | 3.803 × 10−12 *** |
| Area | 0.3501 | <2.20 × 10−16 *** |
| Energy | −0.4138 | <2.20 × 10−16 *** |
| Attributes | Pearson Correlation Coefficient | p-Value |
|---|---|---|
| Maximum amplitude | −0.2316 | 6.911 × 10−15 *** |
| Intensity | −0.2114 | 1.338 × 10−12 *** |
| Area | −0.2408 | 5.252 × 10−16 *** |
| Energy | 0.1678 | 1.378 × 10−7 *** |
| Reference | |||
|---|---|---|---|
| Prediction | Biochar | Non-Biochar | |
| Biochar | 64 | 42 | |
| Non-Biochar | 12 | 18 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, X.; Foster, T.; Baldi, H.; Dobreva, I.; Burson, B.; Hays, D.; Tabien, R.; Jessup, R. Quantification of Soil Organic Carbon in Biochar-Amended Soil Using Ground Penetrating Radar (GPR). Remote Sens. 2019, 11, 2874. https://doi.org/10.3390/rs11232874
Shen X, Foster T, Baldi H, Dobreva I, Burson B, Hays D, Tabien R, Jessup R. Quantification of Soil Organic Carbon in Biochar-Amended Soil Using Ground Penetrating Radar (GPR). Remote Sensing. 2019; 11(23):2874. https://doi.org/10.3390/rs11232874
Chicago/Turabian StyleShen, Xiaoqing, Tyler Foster, Heather Baldi, Iliyana Dobreva, Byron Burson, Dirk Hays, Rodante Tabien, and Russell Jessup. 2019. "Quantification of Soil Organic Carbon in Biochar-Amended Soil Using Ground Penetrating Radar (GPR)" Remote Sensing 11, no. 23: 2874. https://doi.org/10.3390/rs11232874
APA StyleShen, X., Foster, T., Baldi, H., Dobreva, I., Burson, B., Hays, D., Tabien, R., & Jessup, R. (2019). Quantification of Soil Organic Carbon in Biochar-Amended Soil Using Ground Penetrating Radar (GPR). Remote Sensing, 11(23), 2874. https://doi.org/10.3390/rs11232874