A Supervised Method for Nonlinear Hyperspectral Unmixing
Abstract
1. Introduction
2. Methodology
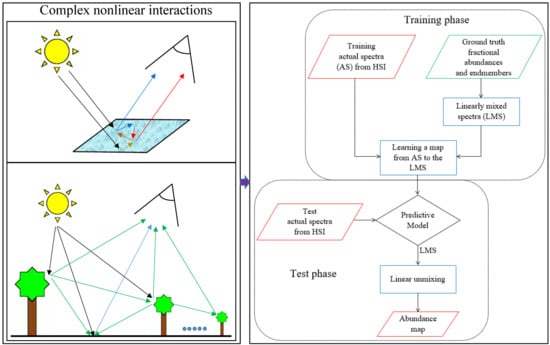
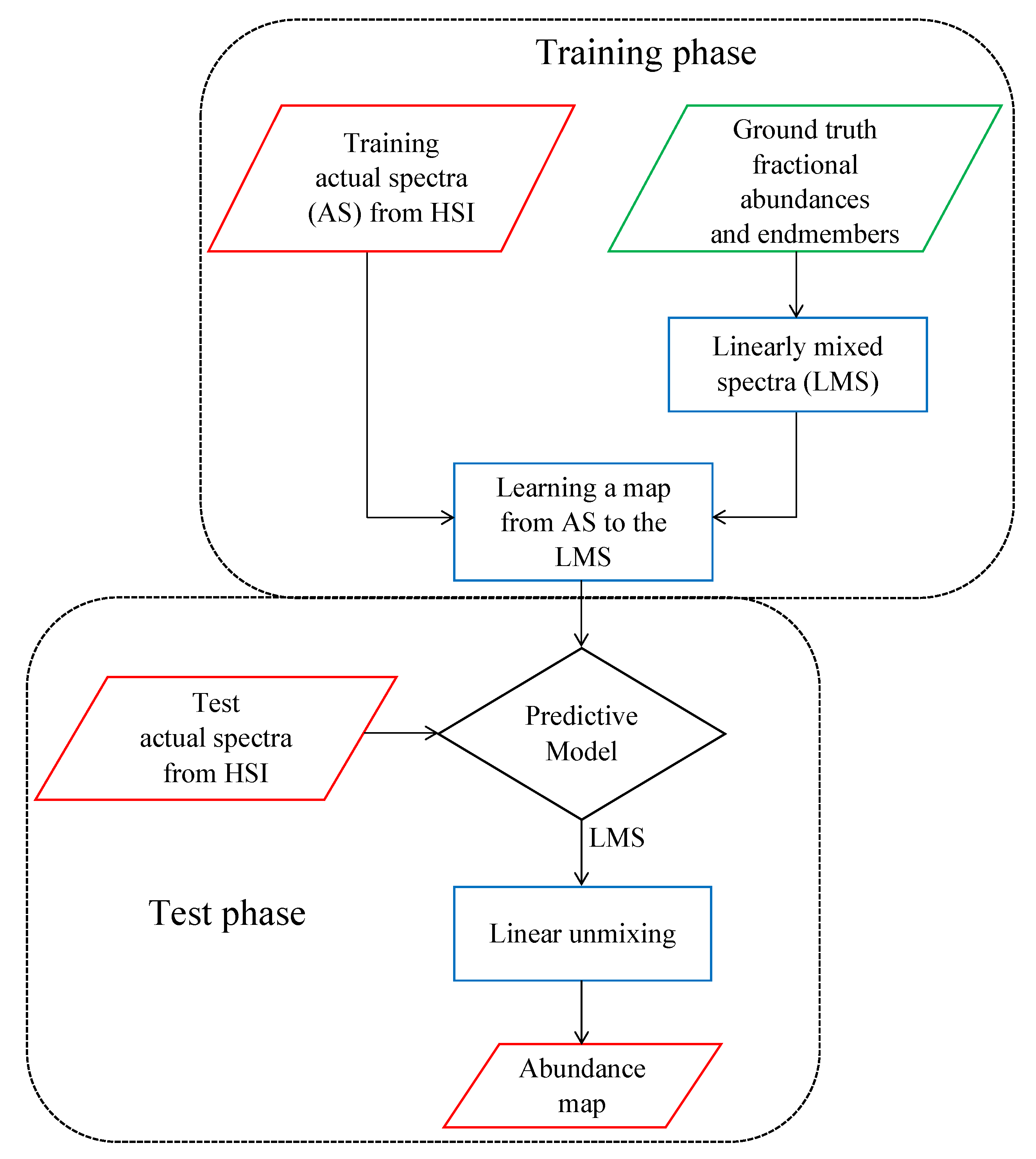
2.1. Generating Linearly Mixed Training Spectra
2.2. Mapping
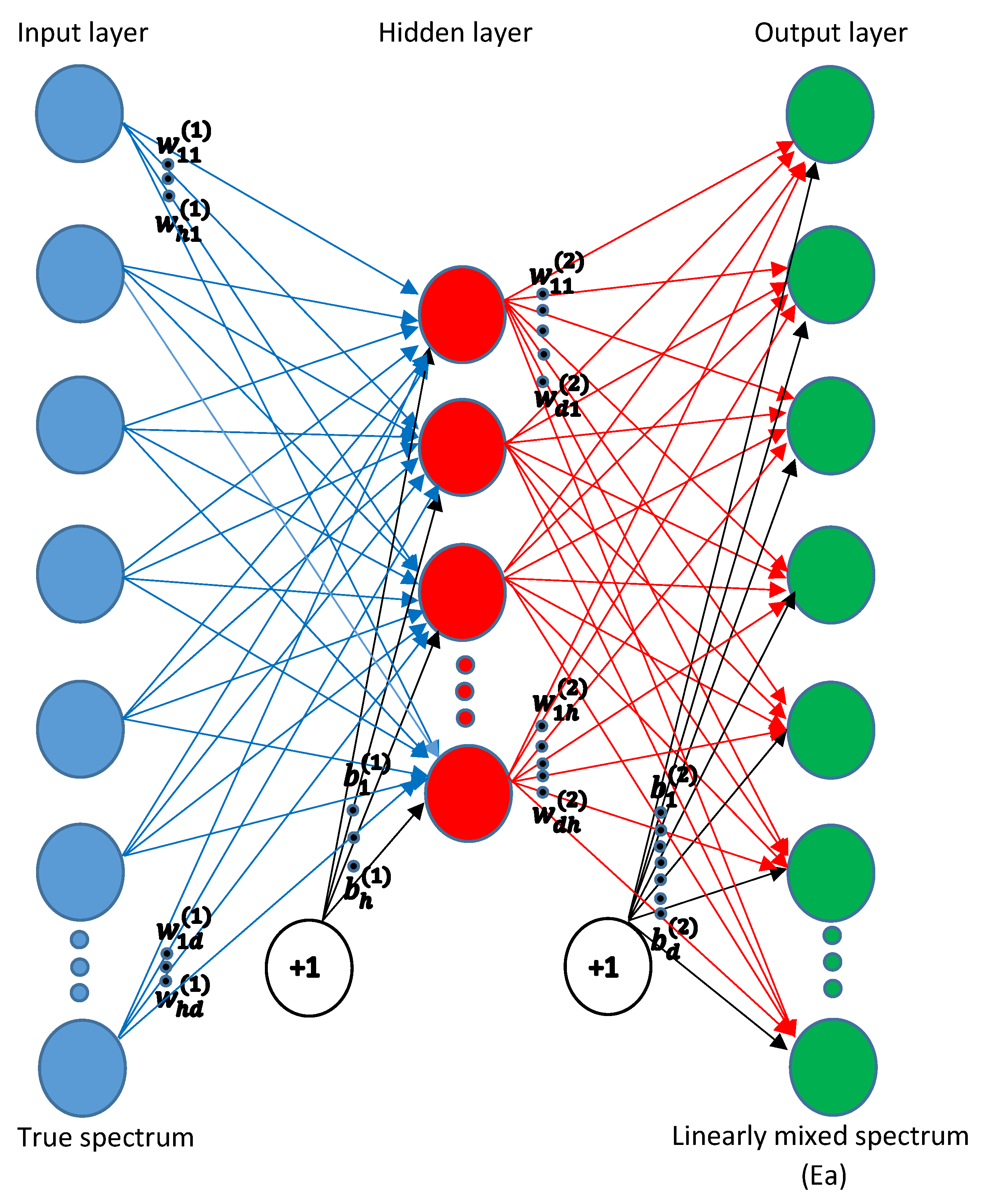
2.2.1. Mapping Using NN
2.2.2. Mapping Using KRR
2.2.3. Mapping Using GP
2.3. Linear Unmixing
3. Eperimental Data
3.1. Dataset 1: Simulated Mineral Dataset
3.2. Dataset 2: Ray Tracing Vegetation Dataset
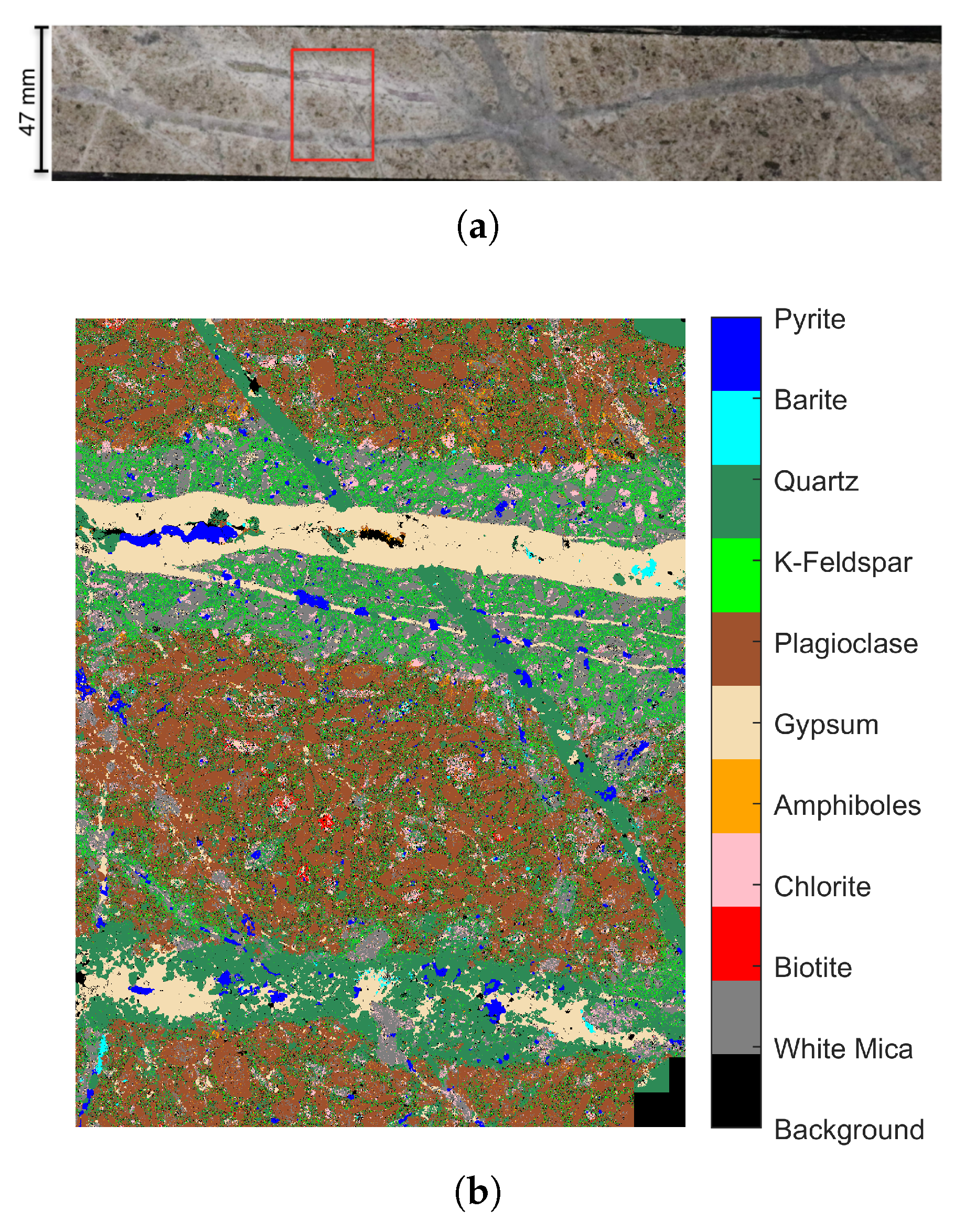
3.3. Dataset 3: Drill Core Hyperspectral Dataset
4. Experiments
4.1. Experimental Set-Up
4.1.1. Comparison to Unsupervised Unmixing Models
4.1.2. Comparison to Other Mapping Methods
4.1.3. Performance Measures
4.2. Experiments Using the Simulated Mineral Dataset
4.2.1. Comparison with Direct Mapping to the Fractional Abundances
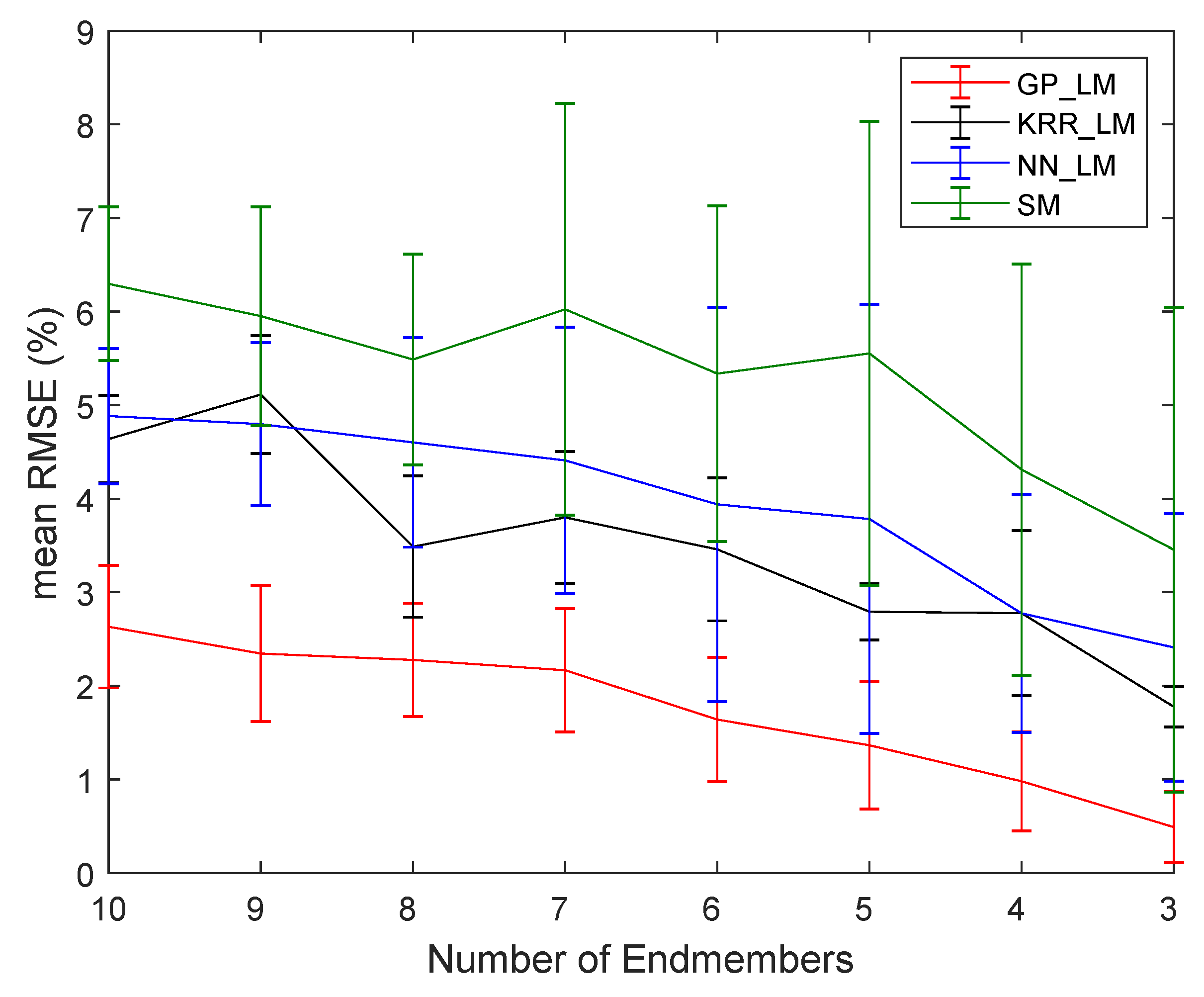
4.2.2. Robustness to Noise and the Number of Endmembers
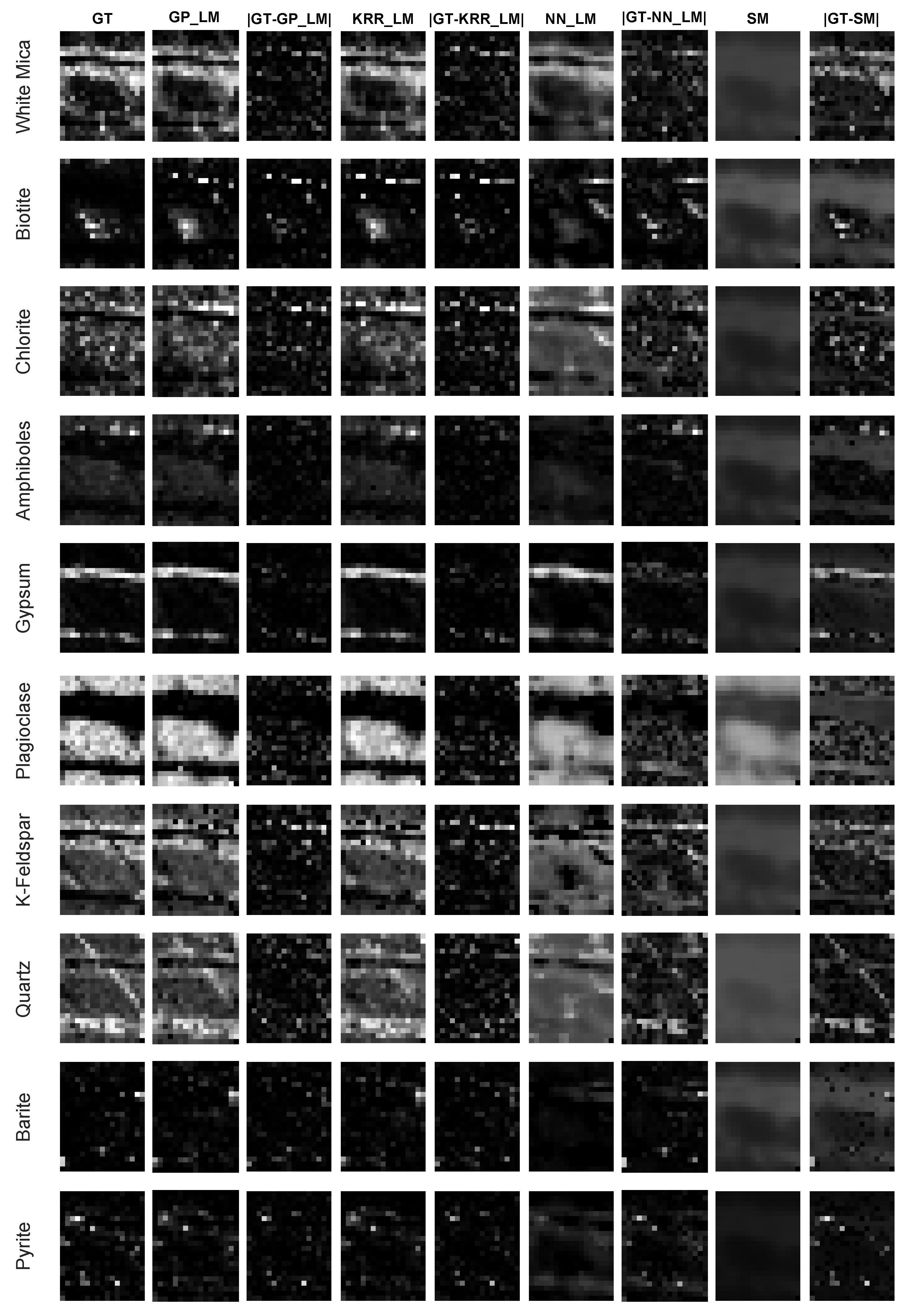
4.2.3. Accuracy of the Mapping
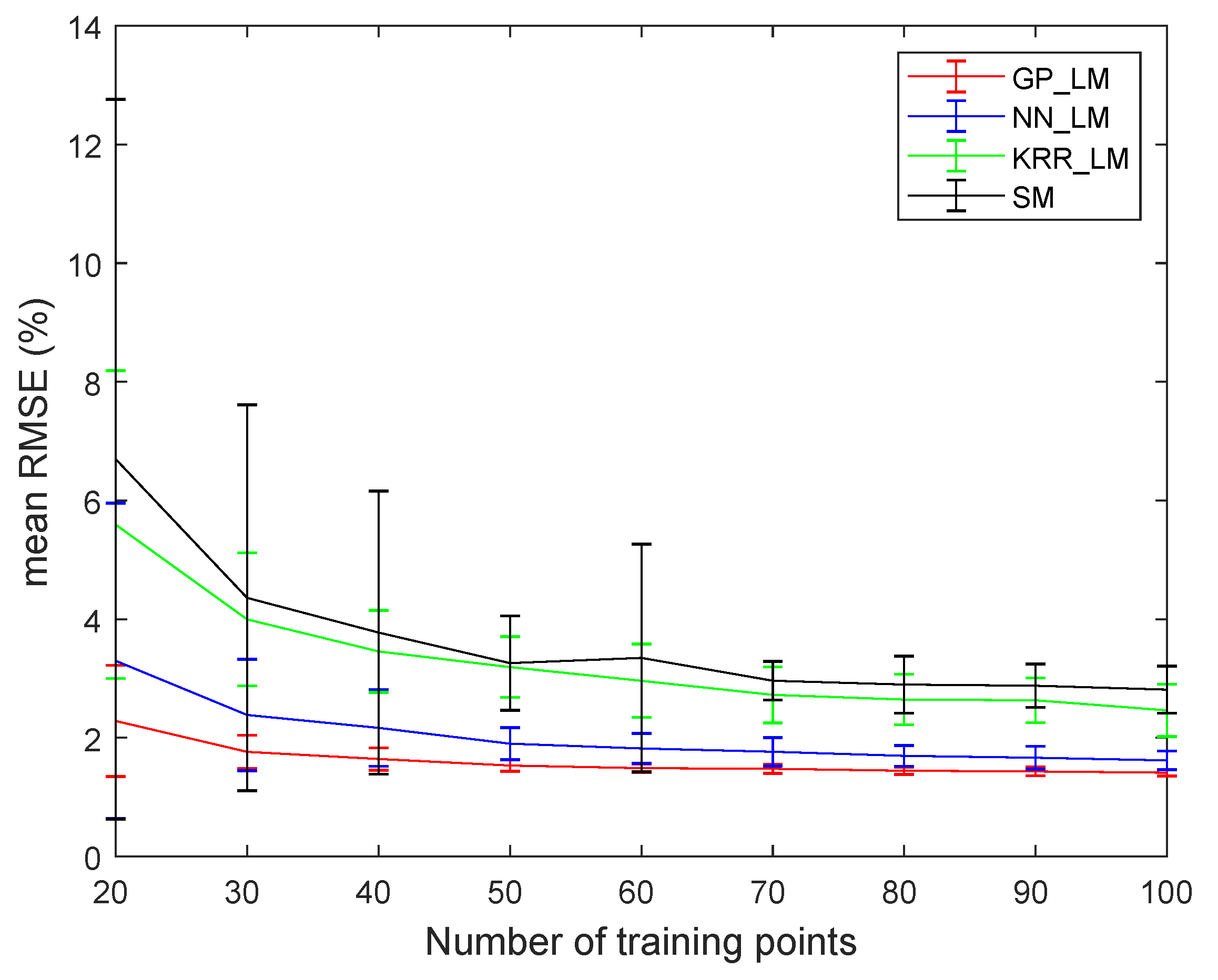
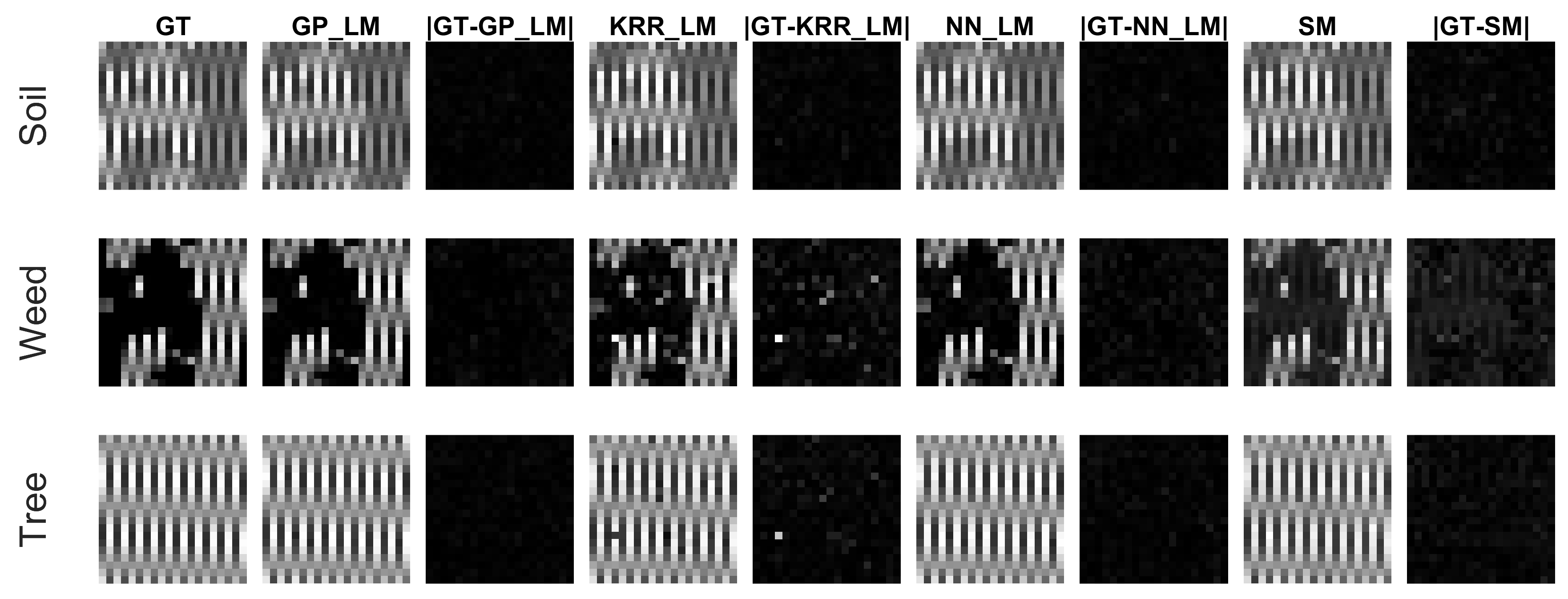
4.3. Experiments on the Ray Tracing Vegetation Dataset
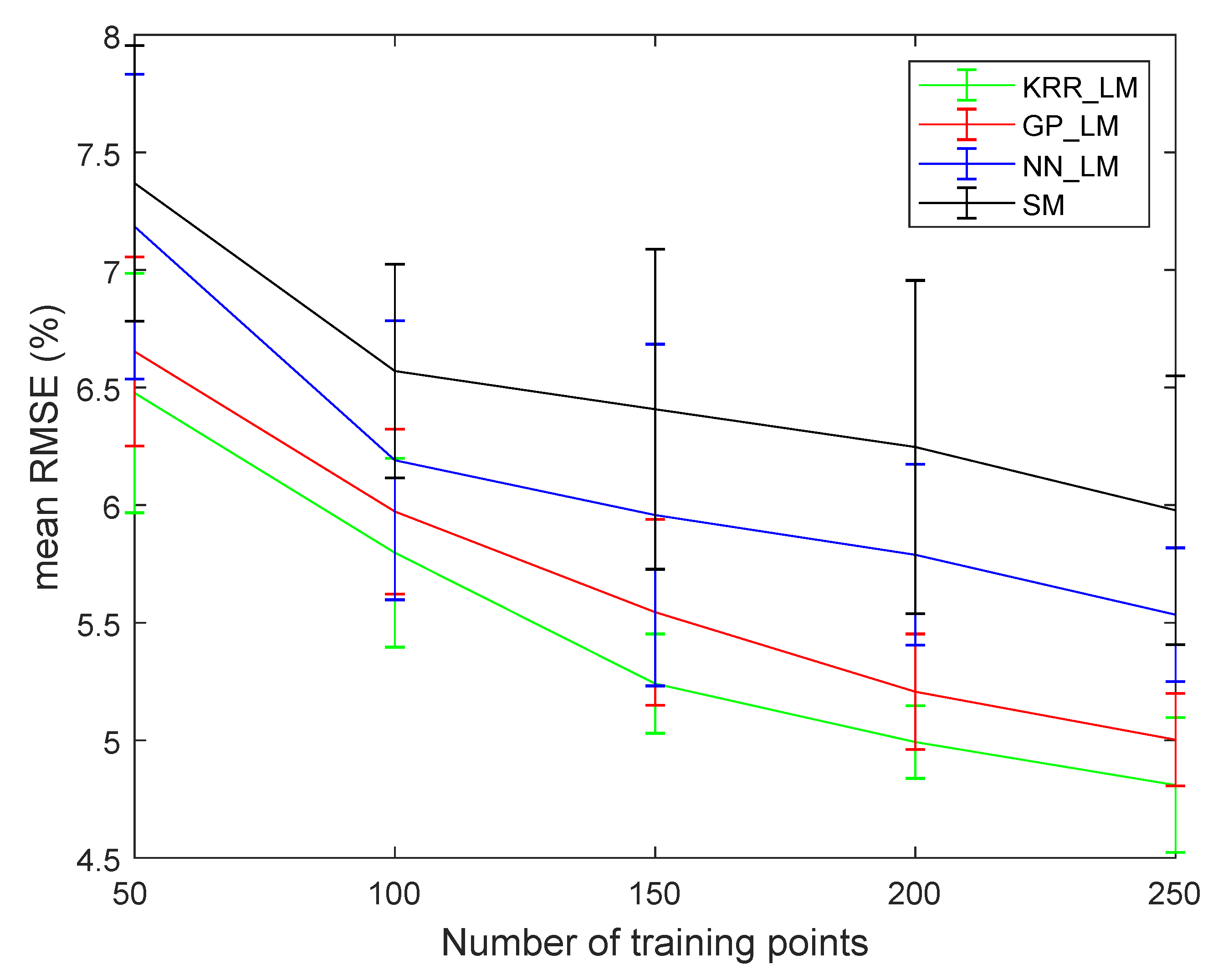
4.4. Experiments on the Drill Core Hyperspectral Dataset
5. Discussion
- The supervised methods all outperform the use of nonlinear spectral mixture models. This is of course partially due the fact that these methods make use of training data. However, the fact that they do not rely on a specific mixture model and the generic nature of these methods allows them to better account for nonlinearities and spectral variability in the data. Results show that the supervised methods can take nonlinearities of different nature simultaneously into account.
- The strategy of mapping onto the linear mixture model outperforms methods that directly map onto the fractional abundances. The main difference is that the proposed methodololgy inherently meets the nonnegativity and sum-to-one constraints. As opposed to the direct unconstrained mapping of the spectra to abundances, the estimated fractional abundances have a clear physical meaning and consequently the estimates are more accurate. Another difference with direct mapping is that our approach requires endmembers, while the direct mapping methods do not. A clear advantage is that in case no pure pixels are available in the data, endmember spectra from a spectral library can be applied. The mapping accounts for the spectral variability between these and the actual endmembers of the data.
- The proposed methodology is generic in the sense that a mapping can be learned to any model. However, learning a mapping to the linear model is favorable over learning mappings to nonlinear models. The higher the nonlinearity of the model, the higher the errors seem to be. The reason for this is that it is just easier to project onto a linear manifold, since a linear manifold can more easily be characterized by a limited number of training samples.
- The proposed methodology requires high quality ground truth data. Learning the mapping based on pure or linearly mixed spectra, as is done in the state of the art literature, will not improve results over the linear mixture model. High-quality ground truth data for spectral unmixing is hard to obtain, in particular in remote sensing applications. One way is to make use of a high-resolution multispectral image of the same scene (if available) to generate a groundcover classification map that can be used to generate ground truth fractional abundances for a low-resolution hyperspectral image. However, in the domain of mineralogy, it is more common to generate validation data with other characterization techniques, such as the MLA technique in the drill core example.
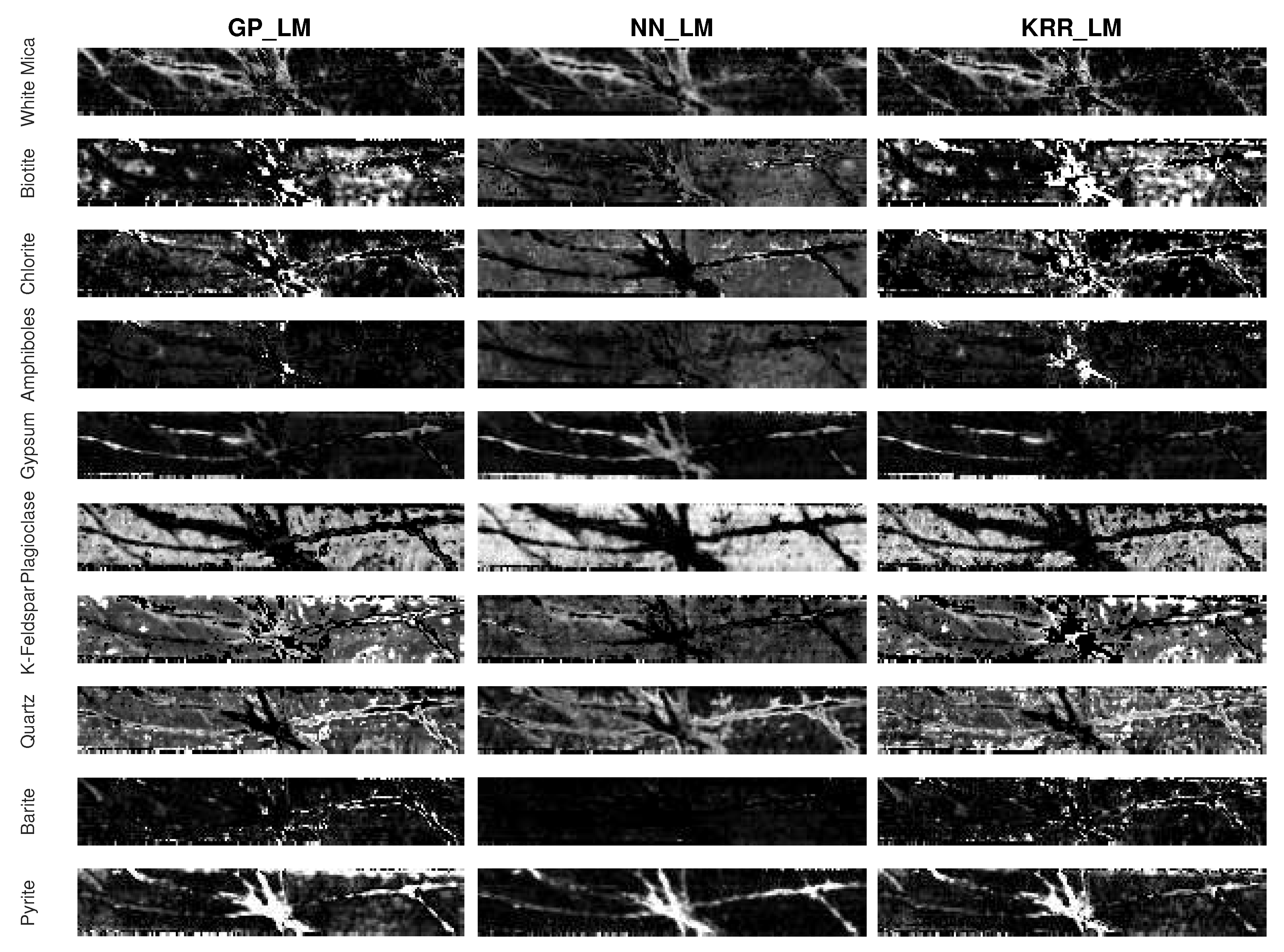
- An advantage of the proposed methodology is that any nonlinear regression method can be applied for learning the mapping. In this work, we compared three different ones. Gaussian processes generally seems to outperform kernel ridge regression and feedforward neural networks. Compared to kernel ridge regression, Gaussian processes contains more hyperparameters for a band-by-band adaptation to the nonlinearities. However, Gaussian processes can overfit the data in case the ground truth fractional abundance values are not very trustworthy. This can be observed in the obtained abundance maps of the entire drill core sample (Figure 13 and Figure 14). On the other hand, a neural network has better generalization properties, but its training can be computationally expensive.
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Keshava, N.; Mustard, J.F. Spectral unmixing. IEEE Signal Process. Mag. 2002, 19, 44–57. [Google Scholar] [CrossRef]
- Bioucas-Dias, J.M.; Plaza, A.; Dobigeon, N.; Parente, M.; Du, Q.; Gader, P.; Chanussot, J. Hyperspectral Unmixing Overview: Geometrical, Statistical, and Sparse Regression-Based Approaches. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2012, 5, 354–379. [Google Scholar] [CrossRef]
- Boardman, J.W. Geometric mixture analysis of imaging spectrometry data. In Proceedings of the Geoscience and Remote Sensing Symposium on Surface and Atmospheric Remote Sensing: Technologies, Data Analysis and Interpretation, Pasadena, CA, USA, 8–12 August 1994; Volume 4, pp. 2369–2371. [Google Scholar]
- Heylen, R.; Parente, M.; Gader, P. A Review of Nonlinear Hyperspectral Unmixing Methods. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2014, 7, 1844–1868. [Google Scholar] [CrossRef]
- Heylen, R.; Scheunders, P. A Multilinear Mixing Model for Nonlinear Spectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2016, 54, 240–251. [Google Scholar] [CrossRef]
- Marinoni, A.; Gamba, P. A Novel Approach for Efficient p-Linear Hyperspectral Unmixing. IEEE J. Sel. Top. Signal Process. 2015, 9, 1156–1168. [Google Scholar] [CrossRef]
- Marinoni, A.; Plaza, J.; Plaza, A.; Gamba, P. Nonlinear Hyperspectral Unmixing Using Nonlinearity Order Estimation and Polytope Decomposition. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2015, 8, 2644–2654. [Google Scholar] [CrossRef]
- Marinoni, A.; Plaza, A.; Gamba, P. Harmonic Mixture Modeling for Efficient Nonlinear Hyperspectral Unmixing. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2016, 9, 4247–4256. [Google Scholar] [CrossRef]
- Hapke, B.; Nelson, R.; Smythe, W. The Opposition Effect of the Moon: Coherent Backscatter and Shadow Hiding. Icarus 1998, 133, 89–97. [Google Scholar] [CrossRef]
- Hapke, B. Bidirectional reflectance spectroscopy: 1. Theory. J. Geophys. Res. 1981, 86, 3039–3054. [Google Scholar] [CrossRef]
- Broadwater, J.; Chellappa, R.; Banerjee, A.; Burlina, P. Kernel fully constrained least squares abundance estimates. In Proceedings of the 2007 IEEE International Geoscience and Remote Sensing Symposium, Barcelona, Spain, 23–28 July 2007; pp. 4041–4044. [Google Scholar]
- Broadwater, J.; Banerjee, A. A comparison of kernel functions for intimate mixture models. In Proceedings of the 2009 First Workshop on Hyperspectral Image and Signal Processing: Evolution in Remote Sensing, Grenoble, France, 26–28 August 2009; pp. 1–4. [Google Scholar]
- Miao, L.; Qi, H. Endmember Extraction From Highly Mixed Data Using Minimum Volume Constrained Nonnegative Matrix Factorization. IEEE Trans. Geosci. Remote. Sens. 2007, 45, 765–777. [Google Scholar] [CrossRef]
- Li, J.; Agathos, A.; Zaharie, D.; Bioucas-Dias, J.M.; Plaza, A.; Li, X. Minimum Volume Simplex Analysis: A Fast Algorithm for Linear Hyperspectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2015, 53, 5067–5082. [Google Scholar]
- Li, J.; Bioucas-Dias, J.M.; Plaza, A.; Liu, L. Robust Collaborative Nonnegative Matrix Factorization for Hyperspectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2016, 54, 6076–6090. [Google Scholar] [CrossRef]
- Fei, Z.; Honeine, P.; Kallas, M. Kernel nonnegative matrix factorization without the pre-image problem. In Proceedings of the 2014 IEEE International Workshop on Machine Learning for Signal Processing (MLSP), Reims, France, 21–24 September 2014; pp. 1–6. [Google Scholar]
- Guo, R.; Wang, W.; Qi, H. Hyperspectral image unmixing using autoencoder cascade. In Proceedings of the 2015 7th Workshop on Hyperspectral Image and Signal Processing: Evolution in Remote Sensing (WHISPERS), Tokyo, Japan, 2–5 June 2015; p. 4. [Google Scholar]
- Su, Y.; Marinoni, A.; Li, J.; Plaza, A.; Gamba, P. Nonnegative sparse autoencoder for robust endmember extraction from remotely sensed hyperspectral images. In Proceedings of the 2017 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Fort Worth, TX, USA, 23–28 July 2017; pp. 205–208. [Google Scholar]
- Qu, Y.; Qi, H. uDAS: An Untied Denoising Autoencoder With Sparsity for Spectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2019, 57, 1698–1712. [Google Scholar] [CrossRef]
- Ozkan, S.; Kaya, B.; Akar, G.B. EndNet: Sparse AutoEncoder Network for Endmember Extraction and Hyperspectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2019, 57, 482–496. [Google Scholar] [CrossRef]
- Palsson, B.; Sigurdsson, J.; Sveinsson, J.R.; Ulfarsson, M.O. Hyperspectral Unmixing Using a Neural Network Autoencoder. IEEE Access 2018, 6, 25646–25656. [Google Scholar] [CrossRef]
- Su, Y.; Marinoni, A.; Li, J.; Plaza, J.; Gamba, P. Stacked Nonnegative Sparse Autoencoders for Robust Hyperspectral Unmixing. IEEE Geosci. Remote. Sens. Lett. 2018, 15, 1427–1431. [Google Scholar] [CrossRef]
- Iordache, M.; Bioucas-Dias, J.M.; Plaza, A. Sparse Unmixing of Hyperspectral Data. IEEE Trans. Geosci. Remote. Sens. 2011, 49, 2014–2039. [Google Scholar] [CrossRef]
- Iordache, M.; Bioucas-Dias, J.M.; Plaza, A. Total Variation Spatial Regularization for Sparse Hyperspectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2012, 50, 4484–4502. [Google Scholar] [CrossRef]
- Iordache, M.; Bioucas-Dias, J.M.; Plaza, A. Collaborative Sparse Regression for Hyperspectral Unmixing. IEEE Trans. Geosci. Remote. Sens. 2014, 52, 341–354. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G.P.; Tits, L.; Coppin, P. Endmember variability in Spectral Mixture Analysis: A review. Remote. Sens. Environ. 2011, 115, 1603–1616. [Google Scholar] [CrossRef]
- Roberts, D.; Gardner, M.; Church, R.; Ustin, S.; Scheer, G.; Green, R. Mapping Chaparral in the Santa Monica Mountains Using Multiple Endmember Spectral Mixture Models. Remote. Sens. Environ. 1998, 65, 267–279. [Google Scholar] [CrossRef]
- Drumetz, L.; Chanussot, J.; Jutten, C. Variability of the endmembers in spectral unmixing: Recent advances. In Proceedings of the 2016 8th Workshop on Hyperspectral Image and Signal Processing: Evolution in Remote Sensing (WHISPERS), Los Angeles, CA, USA, 21–24 August 2016; p. 5. [Google Scholar]
- Foody, G. Relating the land-cover composition of mixed pixels to artificial neural network classification output. Photogr. Eng. Remote Sens. 1996, 62, 491–499. [Google Scholar]
- Foody, G.; Lucas, R.; Curran, P.; Honzak, M. Non-linear mixture modeling without end-members using an artificial neural network. Int. J. Remote Sens. 1997, 18, 937–953. [Google Scholar] [CrossRef]
- Atkinson, P.; Cutler, M.; Lewis, H. Mapping sub-pixel proportional land cover with AVHRR imagery. Int. J. Remote. Sens. 1997, 18, 917–935. [Google Scholar] [CrossRef]
- Okujeni, A.; van der Linden, S.; Tits, L.; Somers, B.; Hostert, P. Support vector regression and synthetically mixed training data for quantifying urban land cover. Remote. Sens. Environ. 2013, 137, 184–197. [Google Scholar] [CrossRef]
- Licciardi, G.; Frate, F.D. Pixel Unmixing in Hyperspectral Data by Means of Neural Networks. IEEE Trans. Geosci. Remote. Sens. 2011, 49, 4163–4172. [Google Scholar] [CrossRef]
- Uezato, T.; Murphy, R.J.; Melkumyan, A.; Chlingaryan, A. A Novel Spectral Unmixing Method Incorporating Spectral Variability Within Endmember Classes. IEEE Trans. Geosci. Remote. Sens. 2016, 54, 2812–2831. [Google Scholar] [CrossRef]
- Plaza, J.M.; Martínez, P.J.M.; Pérez, R.M.; Plaza, A.C.R. Nonlinear neural network mixture models for fractional abundance estimation in AVIRIS hyperspectral images. In Proceedings of the NASA Jet Propulsion Laboratory AVIRIS Airborne Earth Science Workshop, Pasadena, CA, USA, 31 March–2 April 2004; p. 12. [Google Scholar]
- Koirala, B.; Heylen, R.; Scheunders, P. A Neural Network Method for Nonlinear Hyperspectral Unmixing. In Proceedings of the 2018 IEEE International Geoscience and Remote Sensing Symposium, Valencia, Spain, 22–27 July 2018; pp. 4233–4236. [Google Scholar]
- Welling, M. Kernel Ridge Regression; Max Welling’s Classnotes in Machine Learning: Toronto, ON, Canada, 2013; p. 3. [Google Scholar]
- Shawe-Taylor, J.; Cristianini, N. Kernel Methods for Pattern Analysis; Cambridge University Press: Cambridge, UK, 2004. [Google Scholar]
- Rasmussen, C.; Williams, C. Gaussian Processes for Machine Learning; The MIT Press: New York, NY, USA, 2006. [Google Scholar]
- Marquardt, D.W. An algorithm for least-squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Exterkate, P. Model selection in kernel ridge regression. Comput. Stat. Data Anal. 2013, 68, 16. [Google Scholar] [CrossRef]
- Somers, B.; Tits, L.; Coppin, P. Quantifying Nonlinear Spectral Mixing in Vegetated Areas: Computer Simulation Model Validation and First Results. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2014, 7, 1956–1965. [Google Scholar] [CrossRef]
- Stuckens, J.; Somers, B.; Delalieux, S.; Verstraeten, W.; Coppin, P. The impact of common assumptions on canopy radiative transfer simulations: A case study in Citrus orchards. J. Quant. Spectrosc. Radiat. Transf. 2009, 110, 21. [Google Scholar] [CrossRef]
- Cecilia Contreras Acosta, I.; Khodadadzadeh, M.; Tusa, L.; Ghamisi, P.; Gloaguen, R. A Machine Learning Framework for Drill-Core Mineral Mapping Using Hyperspectral and High-Resolution Mineralogical Data Fusion. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2019, 1–14. [Google Scholar] [CrossRef]






| Model | Equation | Parameters |
|---|---|---|
| Linear | ||
| FM | ||
| PPNM | ||
| MLM | ||
| Hapke | : cosine incident angle | |
| : cosine reflectance angle |
| Method | GP_LM | GP | KRR_LM | KRR | NN_LM | NN | SVR | SM |
|---|---|---|---|---|---|---|---|---|
| training set 1 | ||||||||
| RMSE | 19.88 ± 0.62 | 40.89 ± 0.01 | 31.81 ± 1.71 | 40.89 ± 0.01 | 23.57 ± 1.97 | 36.57 ± 6.40 | 34.91 ± 9.48 | 33.68 ± 0.01 |
| NEFA | 0 | 48.32 ± 0.31 | 0 | 25.12 ± 0.27 | 0 | 22.34 ± 0.36 | 24.37 ± 0.62 | 0 |
| training set 2 | ||||||||
| RMSE | 3.05 ± 1.10 | 5.54 ± 1.31 | 4.05 ± 0.58 | 5.55 ± 0.95 | 4.15 ± 1.17 | 5.15 ± 0.80 | 7.10 ± 0.95 | 15.65 ± 5.88 |
| NEFA | 0 | 5.13 ± 1.97 | 0 | 4.66 ± 0.82 | 0 | 8.71 ± 2.40 | 8.96 ± 1.67 | 0 |
| Method | GP_LM | GP_Fan | GP_Hapke | KRR_LM | KRR_Fan | KRR_Hapke | NN_LM | NN_Fan | NN_Hapke |
|---|---|---|---|---|---|---|---|---|---|
| RMSE | 1.19 ± 0.72 | 1.44 ± 0.93 | 1.46 ± 0.74 | 3.04 ± 0.32 | 3.06 ± 0.35 | 3.61 ± 0.64 | 3.65 ± 0.59 | 4.12 ± 0.52 | 5.20 ± 1.25 |
| RE | 2.4 ± 1.98 | 5.8 ± 4.00 | 19 ± 18 | 6.25 ± 3.28 | 12 ± 2.14 | 21 ± 4.47 | 15 ± 7.37 | 29 ± 7.32 | 50 ± 22 |
| Endmember Method | GP_LM | KRR_LM | NN_LM | SM | LMM | Fan | PPNM | MLM | Hapke |
|---|---|---|---|---|---|---|---|---|---|
| Soil | 1.59 ± 0.16 | 2.39 ± 0.93 | 1.86 ± 0.34 | 3.07 ± 0.46 | 11.69 | 12.40 | 16.15 | 14.06 | 9.79 |
| Weed | 1.18 ± 0.18 | 3.77 ± 1.06 | 1.62 ± 0.40 | 3.86 ± 1.42 | 14.84 | 12.72 | 21.40 | 14.42 | 5.94 |
| Tree | 2.06 ± 0.21 | 3.95 ± 0.67 | 2.63 ± 0.46 | 3.74 ± 1.20 | 24.36 | 19.13 | 8.28 | 26.06 | 13.39 |
| Mineral Method | GT | KRR_LM | GP_LM | NN_LM | SM | LMM | Fan | PPNM | MLM | Hapke |
|---|---|---|---|---|---|---|---|---|---|---|
| White Mica | 14.14 | 13.40 ± 0.83 | 13.57 ± 0.79 | 13.61 ± 1.22 | 13.56 ± 1.32 | 9.01 | 3.82 | 6.93 | 9.63 | 6.91 |
| Biotite | 0.46 | 0.63 ± 0.12 | 0.62 ± 0.12 | 0.59 ± 0.20 | 1.04 ± 0.61 | 0 | 0 | 0 | 0 | 0 |
| Chlorite | 4.17 | 4.66 ± 0.35 | 4.56 ± 0.33 | 4.61 ± 0.85 | 3.64 ± 0.60 | 0 | 0 | 0 | 0 | 0 |
| Amphiboles | 1.43 | 1.37 ± 0.10 | 1.38 ± 0.09 | 1.43 ± 0.40 | 1.83 ± 0.46 | 0.24 | 0 | 6.46 | 0.81 | 0 |
| Gypsum | 10.93 | 11.21 ± 1.24 | 11.11 ± 1.19 | 11.08 ± 1.20 | 11.71 ± 1.61 | 32.32 | 31.31 | 44.26 | 31.66 | 42.54 |
| Plagioclase | 33.20 | 32.24 ± 1.58 | 32.74 ± 1.44 | 32.32 ± 2.69 | 33.72 ± 2.69 | 7.30 | 3.66 | 7.62 | 20.43 | 0 |
| K-Feldspar | 6.68 | 5.93 ± 0.30 | 5.98 ± 0.32 | 6.27 ± 1.28 | 5.58 ± 0.76 | 4.77 | 6.33 | 4.92 | 7.74 | 41.21 |
| Quartz | 26.85 | 28.15 ± 1.26 | 27.62 ± 1.12 | 27.19 ± 2.52 | 25.91 ± 1.70 | 0.43 | 0.14 | 20.64 | 3.79 | 0 |
| Barite | 0.33 | 0.46 ± 0.10 | 0.43 ± 0.09 | 0.45 ± 0.24 | 0.98 ± 0.59 | 0 | 0 | 3.40 | 0 | 0 |
| Pyrite | 1.80 | 1.95 ± 0.32 | 2.00 ± 0.34 | 2.46 ± 1.14 | 2.02 ± 0.44 | 45.93 | 54.73 | 5.76 | 25.93 | 9.34 |
| Method | time (s) | time (s) |
|---|---|---|
| KRR_LM | 0.89 | 20.55 |
| GP_LM | 43.84 | 840.62 |
| NN_LM | 105.15 | 7499.84 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koirala, B.; Khodadadzadeh, M.; Contreras, C.; Zahiri, Z.; Gloaguen, R.; Scheunders, P. A Supervised Method for Nonlinear Hyperspectral Unmixing. Remote Sens. 2019, 11, 2458. https://doi.org/10.3390/rs11202458
Koirala B, Khodadadzadeh M, Contreras C, Zahiri Z, Gloaguen R, Scheunders P. A Supervised Method for Nonlinear Hyperspectral Unmixing. Remote Sensing. 2019; 11(20):2458. https://doi.org/10.3390/rs11202458
Chicago/Turabian StyleKoirala, Bikram, Mahdi Khodadadzadeh, Cecilia Contreras, Zohreh Zahiri, Richard Gloaguen, and Paul Scheunders. 2019. "A Supervised Method for Nonlinear Hyperspectral Unmixing" Remote Sensing 11, no. 20: 2458. https://doi.org/10.3390/rs11202458
APA StyleKoirala, B., Khodadadzadeh, M., Contreras, C., Zahiri, Z., Gloaguen, R., & Scheunders, P. (2019). A Supervised Method for Nonlinear Hyperspectral Unmixing. Remote Sensing, 11(20), 2458. https://doi.org/10.3390/rs11202458