Classification of Expansive Grassland Species in Different Growth Stages Based on Hyperspectral and LiDAR Data
Abstract
1. Introduction
Objectives
- compare different times of airborne data acquisition depending on the growing phase of analysed species, to point out the most optimal time of proper species detection,
- collate different datasets containing spectral data and additional different vegetation with high layers to choose the most optimal dataset to detect these species.
2. Data and Methods
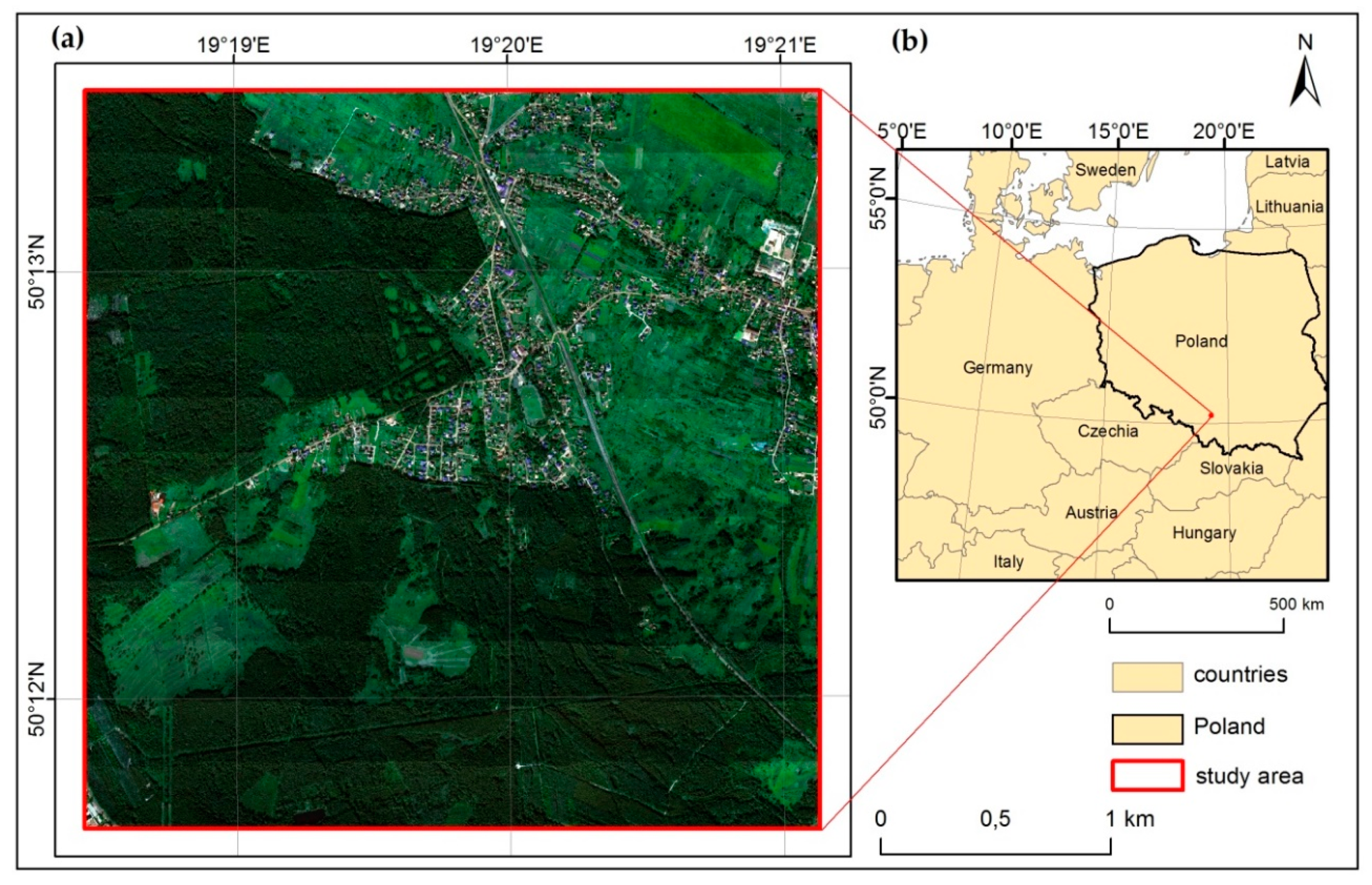
2.1. Study Area and Object of the Study
2.2. Remote Sensing Data
2.3. Reference Botanical Data
2.4. Random Forest Classification and Accuracy Assessment
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hooftman, D.A.P.; Bullock, J.M. Mapping to inform conservation: A case study of changes in semi-natural habitats and their connectivity over 70 years. Biol. Conserv. 2012, 145, 30–38. [Google Scholar] [CrossRef]
- Wesche, K.; Krause, B.; Culmsee, H.; Leuschner, C. Fifty years of change in Central European grassland vegetation: Large losses in species-richness and animal-pollinated plants. Biol. Conserv. 2012, 150, 76–85. [Google Scholar] [CrossRef]
- Krause, B.; Culmsee, H. The significance of habitat continuity and current management on the compositional and functional diversity of grasslands in the uplands of Lower Saxony, Germany. Flora 2013, 208, 299–311. [Google Scholar] [CrossRef]
- Rebele, F.; Lehmann, C. Biological Flora of Central Europe: Calamagrostis epigejos (L.) Roth. Flora 2001, 196, 325–344. [Google Scholar] [CrossRef]
- Hejcman, M.; Češková, M.; Pavlů, V. Control of Molinia caerulea by cutting management on sub-alpine grassland. Flora Morphol. Distrib. Funct. Ecol. Plants 2010, 205, 577–582. [Google Scholar] [CrossRef]
- Holub, P.; Tůma, I.; Záhora, J.; Fiala, K. Different nutrient use strategies of expansive grasses Calamagrostis epigejos and Arrhenatherum elatius. Biologia 2012, 67, 673–680. [Google Scholar] [CrossRef]
- Pruchniewicz, D.; Donath, T.W.; Otte, A.; Żołnierz, L.; Eckstein, R.L. Effect of expansive species on seed rain and soil seed bank of mountain mesic meadows. Tuexenia 2016, 36, 81–96. [Google Scholar]
- Dorigo, W.; Lucieer, A.; Podobnikar, T.; Čarni, A. Mapping invasive Fallopia japonica by combined spectral, spatial, and temporal analysis of digital orthophotos. Int. J. Appl. Earth Obs. Geoinf. 2012, 19, 185–195. [Google Scholar] [CrossRef]
- Bradley, B.A. Remote detection of invasive plants: A review of spectral, textural and phenological approaches. Biol. Invasions 2014, 16, 1411–1425. [Google Scholar] [CrossRef]
- Müllerová, J.; Brůna, J.; Bartaloš, T.; Dvořák, P.; Vítková, M.; Pyšek, P. Timing Is Important: Unmanned Aircraft vs. Satellite Imagery in Plant Invasion Monitoring. Front. Plant Sci. 2017, 8, 887. [Google Scholar] [CrossRef]
- Huang, C.Y.; Asner, G.P. Applications of remote sensing to alien invasive plant studies. Sensors 2009, 9, 4869–4889. [Google Scholar] [CrossRef] [PubMed]
- Zlinszky, A.; Kania, A. Will it blend? Visualization and accuracy evaluation of high-resolution fuzzy vegetation maps. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2016, XLI-B2, 335–343. [Google Scholar] [CrossRef]
- Gao, B.C. NDWI—A normalized difference water index for remote sensing of vegetation liquid water from space. Remote Sens. Environ. 1996, 58, 257–266. [Google Scholar] [CrossRef]
- Goward, S.N.; Markham, B.; Dye, D.G.; Dulaney, W.; Yang, J. Normalized difference vegetation index measurements from the Advanced Very High Resolution Radiometer. Remote Sens. Environ. 1991, 35, 257–277. [Google Scholar] [CrossRef]
- Wang, L.; Qu, J.J. NMDI: A normalized multi-band drought index for monitoring soil and vegetation moisture with satellite remote sensing. Geoph. Res. Lett. 2007, 34. [Google Scholar] [CrossRef]
- Kycko, M.; Zagajewski, B.; Lavender, S.; Romanowska, E.; Zwijacz–Kozica, M. The Impact of Tourist Traffic on the Condition and Cell Structures of Alpine Swards. Remote Sens. 2018, 10, 220. [Google Scholar] [CrossRef]
- Zagajewski, B.; Tømmervik, H.; Bjerke, J.W.; Raczko, E.; Bochenek, Z.; Kłos, A.; Jarocińska, A.; Lavender, S.; Ziółkowski, D. Intraspecific Differences in Spectral Reflectance Curves as Indicators of Reduced Vitality in High–Arctic Plants. Remote Sens. 2017, 9, 1289. [Google Scholar] [CrossRef]
- Hamada, Y.; Stow, D.A.; Coulter, L.L.; Jafolla, J.C.; Hendricks, L.W. Detecting Tamarisk species (Tamarix spp.) in riparian habitats of Southern California using high spatial resolution hyperspectral imagery. Remote Sens. Environ. 2007, 109, 237–248. [Google Scholar] [CrossRef]
- Marcinkowska–Ochtyra, A.; Zagajewski, B.; Ochtyra, A.; Jarocińska, A.; Wojtuń, B.; Rogass, C.; Mielke, C.; Lavender, S. Subalpine and alpine vegetation classification based on hyperspectral APEX and simulated EnMAP images. Int. J. Remote Sens. 2017, 38, 1839–1864. [Google Scholar] [CrossRef]
- Marcinkowska–Ochtyra, A.; Zagajewski, B.; Raczko, E.; Ochtyra, A.; Jarocińska, A. Classification of High-Mountain Vegetation Communities within a Diverse Giant Mountains Ecosystem Using Airborne APEX Hyperspectral Imagery. Remote Sens. 2018, 10, 570. [Google Scholar] [CrossRef]
- Jensen, J.R. Biophysical remote sensing. Ann. Assoc. Am. Geogr. 1983, 73, 111–132. [Google Scholar] [CrossRef]
- Carrão, H.; Gonçalves, P.; Caetano, M. Contribution of multispectral and multitemporal information from MODIS images to land cover classification. Remote Sens. Environ. 2008, 112, 986–997. [Google Scholar] [CrossRef]
- Schneider, J.; Grosse, G.; Wagner, D. Land cover classification of tundra environments in the Arctic Lena Delta based on Landsat 7 ETM+ data and its application for upscaling of methane emissions. Remote Sens. Environ. 2009, 113, 380–391. [Google Scholar] [CrossRef]
- Golenia, M.; Zagajewski, B.; Ochtyra, A.; Hościło, A. Semiautomatic land cover mapping according to the 2nd level of the CORINE Land Cover legend. Pol. Cartog. Rev. 2015, 47, 203–212. [Google Scholar] [CrossRef]
- Riaño, D.; Chuvieco, E.; Salas, J.; Aguado, I. Assessment of different topographic corrections in Landsat–TM data for mapping vegetation types. IEEE Trans. Geosci. Remote Sens. 2003, 41, 1056–1061. [Google Scholar] [CrossRef]
- Jędrych, M.; Zagajewski, B.; Marcinkowska–Ochtyra, A. Application of Sentinel–2 and EnMAP new satellite data to the mapping of alpine vegetation of the Karkonosze Mountains. Pol. Cartog. Rev. 2017, 49, 107–119. [Google Scholar] [CrossRef]
- Chan, J.C.W.; Spanhove, T.; Ma, J.; Borre, J.V.; Paelinckx, D.; Canters, F. Natura 2000 Habitat Identification and Conservation Status Assessment with Superresolution Enhanced Hyperspectral (CHRIS/Proba) Imagery. In Proceedings of the GEOBIA 2010 Geographic Object-Based Image Analysis, Ghent, Belgium, 29 June–2 July 2010. [Google Scholar]
- Goetz, A.F.H.; Vane, G.; Solomon, J.E.; Rock, B.N. Imaging spectroscopy for earth remote sensing. Science 1985, 228, 1147–1153. [Google Scholar] [CrossRef]
- Hotelling, H. Analysis of a complex of statistical variables into principal components. J. Educ. Psychol. 1933, 24, 417–441. [Google Scholar] [CrossRef]
- Green, A.A.; Berman, M.; Switzer, P.; Craig, M.D. A transformation for ordering multispectral data in terms of image quality with implications for noise removal. IEEE Trans. Geosci. Remote Sens. 1988, 26, 65–74. [Google Scholar] [CrossRef]
- Xie, Y.; Sha, Z.; Yu, M. Remote sensing imagery in vegetation mapping: A review. J. Plant Ecol. 2008, 1, 9–23. [Google Scholar] [CrossRef]
- Geerken, R.; Zaitchik, B.; Evans, J.P. Classifying rangeland vegetation type and coverage from NDVI time series using Fourier Filtered Cycle Similarity. Int. J. Remote Sens. 2005, 26, 5535–5554. [Google Scholar] [CrossRef]
- Schuster, C.; Schmidt, T.; Conrad, C.; Kleinschmit, B.; Förster, M. Grassland habitat mapping by intra-annual time series analysis–Comparison of RapidEye and TerraSAR-X satellite data. Int. J. Appl. Earth Obs. Geoinf. 2015, 34, 25–34. [Google Scholar] [CrossRef]
- Zhang, C.; Xie, Z. Object-based vegetation mapping in the Kissimmee River watershed using HyMap data and machine learning techniques. Wetlands 2013, 33, 233–244. [Google Scholar] [CrossRef]
- Burai, P.; Deák, B.; Valkó, O.; Tomor, T. Classification of herbaceous vegetation using airborne hyperspectral imagery. Remote Sens. 2015, 7, 2046–2066. [Google Scholar] [CrossRef]
- Lawrence, R.L.; Wood, S.D.; Sheley, R.L. Mapping invasive plants using hyperspectral imagery and Breiman Cutler classifications (RandomForest). Remote Sens. Environ. 2006, 100, 356–362. [Google Scholar] [CrossRef]
- Cutler, D.R.; Edwards, T.C., Jr.; Beard, K.H.; Cutler, A.; Hess, K.T.; Gibson, J.; Lawler, J.J. Random forests for classification in ecology. Ecology 2007, 88, 2783–2792. [Google Scholar] [CrossRef] [PubMed]
- Naidoo, L.; Cho, M.A.; Mathieu, R.; Asner, G. Classification of savanna tree species, in the Greater Kruger National Park region, by integrating hyperspectral and LiDAR data in a Random Forest data mining environment. ISPRS J. Photogramm. Remote Sens. 2012, 69, 167–179. [Google Scholar] [CrossRef]
- Ghosh, A.; Fassnacht, F.E.; Joshi, P.K.; Koch, B. A framework for mapping tree species combining hyperspectral and LiDAR data: Role of selected classifiers and sensor across three spatial scales. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 49–63. [Google Scholar] [CrossRef]
- Raczko, E.; Zagajewski, B. Comparison of Support Vector Machine, Random Forest and Neural Network Classifiers for Tree Species Classification on Airborne Hyperspectral APEX images. Eur. J. Remote Sens. 2017, 50, 144–154. [Google Scholar] [CrossRef]
- Ustin, S.L.; DiPietro, D.; Olmstead, K.; Underwood, E.; Scheer, G.J. Hyperspectral Remote Sensing for Invasive Species Detection and Mapping. In Proceedings of the 2002 IEEE International Geoscience and Remote Sensing Symposium, Toronto, ON, Canada, 24–28 June 2002. [Google Scholar]
- Pengra, B.W.; Johnston, C.A.; Loveland, T.R. Mapping an invasive plant, Phragmites australis, in coastal wetlands using the EO–1 Hyperion hyperspectral sensor. Remote Sens. Environ. 2007, 108, 74–81. [Google Scholar] [CrossRef]
- Asner, G.P.; Jones, M.O.; Martin, R.E.; Knapp, D.E.; Hughes, R.F. Remote sensing of native and invasive species in Hawaiian forests. Remote Sens. Environ. 2008, 112, 1912–1926. [Google Scholar] [CrossRef]
- Házi, J.; Bartha, S.; Szentes, S.; Wichmann, B.; Penksza, K. Seminatural grassland management by mowing of Calamagrostis epigejos in Hungary. Hung. Acad. Sci. 2011, 145, 699–707. [Google Scholar] [CrossRef]
- Jacquemyn, H.; Brys, R.; Neubert, M.G. Fire increases invasive spread of Molinia caerulea mainly through changes in demographic parameters. Ecol. Appl. 2005, 15, 2097–2108. [Google Scholar] [CrossRef]
- Mücher, C.A.; Kooistra, L.; Vermeulen, M.; Borre, J.V.; Haest, B.; Haveman, R. Quantifying structure of Natura 2000 heathland habitats using spectral mixture analysis and segmentation techniques on hyperspectral imagery. Ecol. Ind. 2013, 33, 71–81. [Google Scholar] [CrossRef]
- Haest, B.; Borre, J.V.; Spanhove, T.; Thoonen, G.; Delalieux, S.; Kooistra, L.; Mücher, C.A.; Paelinckx, D.; Scheunders, P.; Kempeneers, P. Habitat mapping and quality assessment of NATURA 2000 heathland using airborne imaging spectroscopy. Remote Sens. 2017, 9. [Google Scholar] [CrossRef]
- Schmidt, J.; Fassnacht, F.E.; Neff, C.; Lausch, A.; Kleinschmit, B.; Förster, M.; Schmidtlein, S. Adapting a Natura 2000 field guideline for a remote sensing-based assessment of heathland conservation status. Int. J. Appl. Earth Obs. Geoinf. 2017, 60, 61–71. [Google Scholar] [CrossRef]
- Ali, I.; Cawkwell, F.; Dwyer, E.; Barrett, B.; Green, S. Satellite remote sensing of grasslands: From observation to management—A review. J. Plant Ecol. 2016, 9, 649–671. [Google Scholar] [CrossRef]
- Evangelista, P.H.; Stohlgren, T.J.; Morisette, J.T.; Kumar, S. Mapping invasive tamarisk (Tamarix): A comparison of single–scene and time–series analyses of remotely sensed data. Remote Sens. 2009, 1, 519–533. [Google Scholar] [CrossRef]
- Somers, B.; Asner, G.P. Hyperspectral time series analysis of native and invasive species in Hawaiian rainforests. Remote Sens. 2009, 4. [Google Scholar] [CrossRef]
- Lopes, M.; Fauvel, M.; Girard, S.; Sheeren, D. Object-based classification of grasslands from high resolution satellite image time series using gaussian mean map kernels. Remote Sens. 2017, 9, 688. [Google Scholar] [CrossRef]
- Voss, M.; Sugumaran, R. Seasonal effect on tree species classification in an urban environment using hyperspectral data, LiDAR, and an object–oriented approach. Sensors 2008, 8, 3020–3036. [Google Scholar] [CrossRef] [PubMed]
- Zlinszky, A.; Schroiff, A.; Kania, A.; Deák, B.; Mücke, W.; Vári, Á.; Székely, B.; Pfeifer, N. Categorizing grassland vegetation with full–waveform airborne laser scanning: A feasibility study for detecting Natura 2000 habitat types. Remote Sens. 2014, 6, 8056–8087. [Google Scholar] [CrossRef]
- Rapinel, S.; Rossignol, N.; Hubert-Moy, L.; Bouzillé, J.B.; Bonis, A. Mapping grassland plant communities using a fuzzy approach to address floristic and spectral uncertainty. Appl. Veg. Sci. 2018, 21, 678–693. [Google Scholar] [CrossRef]
- Kopeć, D.; Michalska-Hejduk, D.; Sławik, S.; Berezowski, T.; Borowski, M.; Rosadziński, S.; Chormański, J. Application of multisensoral remote sensing data in the mapping of alkaline fens Natura 2000 habitat. Ecol. Indic. 2016, 70, 196–208. [Google Scholar] [CrossRef]
- Dalponte, M.; Bruzzone, L.; Gianelle, D. Tree species classification in the Southern Alps based on the fusion of very high geometrical resolution multispectral/hyperspectral images and LiDAR data. Remote Sens. Environ. 2012, 123, 258–270. [Google Scholar] [CrossRef]
- Hollaus, M.; Mücke, W.; Höfle, B.; Dorigo, W.; Pfeifer, N.; Wagner, W.; Bauerhansl, C.; Regner, B. Tree Species Classification Based on Full-Waveform Airborne Laser Scanning Data. In Silvilaser 2009; Texas A & M University: College Station, TX, USA, 2009. [Google Scholar]
- Chance, C.M.; Coops, N.C.; Plowright, A.A.; Tooke, T.R.; Christen, A.; Aven, N. Invasive shrub mapping in an urban environment from hyperspectral and LiDAR–derived attributes. Front. Plant Sci. 2016, 7, 1528. [Google Scholar] [CrossRef]
- Geerling, G.W.; Labrador-Garcia, M.; Clevers, J.G.P.W.; Ragas, A.M.J.; Smits, A.J.M. Classification of floodplain vegetation by data fusion of spectral (CASI) and LiDAR data. Int. J. Remote Sens. 2007, 28, 4263–4284. [Google Scholar] [CrossRef]
- Zlinszky, A.; Mücke, W.; Lehner, H.; Briese, C.; Pfeifer, N. Categorizing wetland vegetation by Airborne Laser Scanning on Lake Balaton and Kis-Balaton, Hungary. Remote Sens. 2012, 4, 1617–1650. [Google Scholar] [CrossRef]
- Hellesen, T.; Matikainen, L. An object-based approach for mapping shrub and tree cover on grassland habitats by use of LiDAR and CIR orthoimages. Remote Sens. 2013, 5, 558–583. [Google Scholar] [CrossRef]
- Ward, R.D.; Burnside, N.G.; Joyce, C.B.; Sepp, K. The use of medium point density LiDAR elevation data to determine plant community types in Baltic coastal wetlands. Ecol. Indic. 2013, 33, 96–104. [Google Scholar] [CrossRef]
- Debes, C.; Merentitis, A.; Heremans, R.; Hahn, J.; Frangiadakis, N.; van Kasteren, T.; Wenzhi, L.; Bellens, R.; Pizurica, A.; Gautama, S.; et al. Hyperspectral and LiDAR data fusion: Outcome of the 2013 GRSS data fusion contest. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 2405–2418. [Google Scholar] [CrossRef]
- Luo, S.; Wang, C.; Xi, X.; Zeng, H.; Li, D.; Xia, S.; Wang, P. Fusion of airborne discrete-return LiDAR and hyperspectral data for land cover classification. Remote Sens. 2016, 8, 3. [Google Scholar] [CrossRef]
- Buck, O.; Millán, V.E.G.; Klink, A.; Pakzad, K. Using information layers for mapping grassland habitat distribution at local to regional scales. Int. J. Appl. Earth Obs. Geoinf. 2015, 37, 83–89. [Google Scholar] [CrossRef]
- Reese, H.; Nyström, M.; Nordkvist, K.; Olsson, H. Combining airborne laser scanning data and optical satellite data for classification of alpine vegetation. Int. J. Appl. Earth Obs. Geoinf. 2014, 27, 81–90. [Google Scholar] [CrossRef]
- Chan, J.C.W.; Beckers, P.; Spanhove, T.; Borre, J.V. An evaluation of ensemble classifiers for mapping Natura 2000 heathland in Belgium using spaceborne angular hyperspectral (CHRIS/Proba) imagery. Int. J. Appl. Earth Obs. Geoinf. 2012, 18, 13–22. [Google Scholar] [CrossRef]
- Fassnacht, F.E.; Latifi, H.; Ghosh, A.; Joshi, P.K.; Koch, B. Assessing the potential of hyperspectral imagery to map bark beetle–induced tree mortality. Remote Sens. 2014, 140, 533–548. [Google Scholar] [CrossRef]
- Pyšek, P.; Richardson, D.M.; Rejmánek, M.; Webster, G.L.; Williamson, M.; Kirschner, J. Alien Plants in Checklists and Floras: Towards Better Communication between Taxonomists and Ecologists. Taxon 2004, 53, 131. [Google Scholar] [CrossRef]
- Mróz, W. Monitoring of Natural Habitats. Methodological Guide; GIOŚ: Warszawa, Poland, 2013. [Google Scholar]
- Cope, T.A.; Gray, A.J. Grasses of the British Isles; Botanical Society of the British Isles: Harpenden, UK, 2009; Volume 13, p. 612. [Google Scholar]
- Rutkowski, L. Klucz do Oznaczania Roślin Naczyniowych Polski Niżowej; PWN: Warszawa, Poland, 1998; p. 112. [Google Scholar]
- Hubbard, C.E. Grasses, A guide to Their Structure, Identification, Uses, and Distribution in the British Isles, 3rd ed.; Penguin Books Ltd.: Harmondsworth, UK, 1984. [Google Scholar]
- Březina, S.; Koubek, T.; Münzbergová, Z.; Herben, T. Ecological benefits of integration of Calamagrostis epigejos ramets under field conditions. Flora 2006, 201, 461–467. [Google Scholar] [CrossRef]
- Sedláková, I.; Fiala, K. Ecological problems of degradation of alluvial meadows due to expanding Calamagrostis epigejos. Ekologia 2001, 20 (Suppl. 3), 226–233. [Google Scholar]
- Somodi, I.; Virágh, K.; Podani, J. The effect of the expansion of the clonal grass Calamagrostis epigejos on the species turnover of a semi–arid grassland. Appl. Veg. Sci. 2008, 11, 187–192. [Google Scholar] [CrossRef]
- Schläpfer, D.; Richter, R. Geo-atmospheric processing of airborne imaging spectrometry data. Part 1, Parametric orthorectification. Int. J. Remote Sens. 2002, 23, 2609–2630. [Google Scholar] [CrossRef]
- Richter, R.; Schläpfer, D. Geo-Atmospheric Processing of Wide FOV Airborne Imaging Pectrometry Data. In Remote Sensing for Environmental Monitoring, GIS Applications, and Geology; Ehlers, M., Ed.; Society of Photo Optical: Bellingham, WA, USA, 2002; Volume 4545, pp. 264–273. [Google Scholar]
- Tokarska-Guzik, B.; Błońska, A. (University of Silesia, Katowice, Poland); Jarocińska, A. (University of Warsaw, Warsaw, Poland). Personal Communication, 2017.
- ENVI. User’s Guide; RSI Research Systems: Boulder, CO, USA, 2004. [Google Scholar]
- RIEGL Laser Measurement Systems GmbH. 2015. Available online: http://www.riegl.com/uploads/tx_pxpriegldownloads/11_Datasheet_RiProcess_2016-09-16_01.pdf (accessed on 8 December 2018).
- Mallet, C.; Bretar, F. Full–waveform topographic lidar: State–of–the–art. ISPRS J. Photogramm. Remote Sens. 2009, 64, 1–16. [Google Scholar] [CrossRef]
- RIEGL Laser Measurement Systems GmbH. 2015. Available online: http://www.riegl.com/uploads/tx_pxpriegldownloads/11_Datasheet_RiANALYZE_2015-01-22.pdf (accessed on 8 December 2018).
- Terrasolid TerraScan User’s Guide. Available online: http://www.terrasolid.com/download/tscan.pdf (accessed on 12 August 2018).
- Pfeifer, N.; Mandlburger, G.; Otepka, J.; Karel, W. OPALS—A framework for Airborne Laser Scanning data analysis. Comput. Environ. Urban Syst. 2014, 45, 125–136. [Google Scholar] [CrossRef]
- Conrad, O.; Bechtel, B.; Bock, M.; Dietrich, H.; Fischer, E.; Gerlitz, L.; Wehberg, J.; Wichmann, V.; Böhner, J. System for Automated Geoscientific Analyses (SAGA) v. 2.1.4. Geosci. Model Dev. 2015, 8, 1991–2007. [Google Scholar] [CrossRef]
- Maaten, L.V.D.; Hinton, G. Visualizing data using t–SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Congalton, R.G. A review of assessing the accuracy of classifications of remotely sensed data. Remote Sens. Environ. 1991, 37, 35–46. [Google Scholar] [CrossRef]
- Manel, S.; Williams, H.C.; Ormerod, S.J. Evaluating presence–absence models in ecology: The need to account for prevalence. J. App. Ecol. 2002, 38, 921–931. [Google Scholar] [CrossRef]
- Rijsbergen, C.J.V. Information Retrieval; Butterworths: London, UK, 1979. [Google Scholar]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 159–174. [Google Scholar] [CrossRef]
- Marcinkowska-Ochtyra, A. Classification of Molinia caerulea and Calamagrostis epigejos Expansive Species Based on LiDAR Vegetation Structure Data. Unpublished work.
- Tokarska-Guzik, B. (University of Silesia, Katowice, Poland); Raczko, E. (University of Warsaw, Warsaw, Poland). Personal Communication, 2017.
- Zagajewski, B. Assessment of neural networks and Imaging Spectroscopy for vegetation classification of the High Tatras. Teledetekcja Środowiska 2010, 43, 1–113. [Google Scholar]



| Characteristics | Scanner | |
|---|---|---|
| VNIR | SWIR | |
| Spatial pixels | 1800 | 384 |
| Minimum wavelength [nm] | 416 | 954 |
| Maximum wavelength [nm] | 995 | 2510 |
| Spectral sampling [nm] | 3.26 | 5.45 |
| No. of bands | 182 (163 1) | 288 |
| Radiometric resolution [bit] | 16 | 16 |
| Field of view (FOV) [°] | 17–34 | 16–32 |
| Instantaneous field of view (IFOV) [°] | 0.01–0.04 | 0.04–0.08 |
| Characteristics | Riegl LMS-Q680i |
|---|---|
| Wavelength [nm] | 1550 |
| Pulse duration [ns] | <4 |
| Avg. flight altitude [m] | 500 |
| Ground footprint size [cm] | 25 |
| Average point density [p/m2] 1 | 8 |
| Average point spacing [m] 2 | 0.34 |
| Full waveform registered | Yes (Riegl FWF) |
| Number of echoes for each pulse 3 | 7 |
| Scenario No. | Dataset |
|---|---|
| sc01 | MOSAIC |
| sc02 | MNF |
| sc03 | MNF+CHM |
| sc04 | MNF+VIS |
| sc05 | MNF+DISC |
| sc06 | MNF+DISC+FWF 1 |
| sc07 | MNF+TOPO |
| sc08 | MNF+FWF1 |
| sc09 | MNF+CHM+VIS+DISC+FWF1+TOPO |
| Species | Polygons | June 2017 | August 2017 | September 2016 1 |
|---|---|---|---|---|
| C. epigejos | species from field measurements | 234 | 256 | 245 |
| background from field measurements | 662 | 679 | 614 | |
| species selected 2 | 222 | 241 | 237 | |
| background selected 2 | 643 | 660 | 614 | |
| M. caerulea | species from field measurements | 195 | 198 | 197 |
| background from field measurements | 656 | 737 | 654 | |
| species selected 2 | 174 | 183 | 177 | |
| background selected 2 | 649 | 728 | 649 |
| sc. No. | Class | June | August | September | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| UA | PA | F1 | UA | PA | F1 | UA | PA | F1 | ||
| sc01 | M. caerulea | 0.88 | 0.61 | 0.72 | 0.89 | 0.79 | 0.84 | 0.85 | 0.67 | 0.75 |
| background | 0.93 | 0.98 | 0.96 | 0.95 | 0.98 | 0.97 | 0.94 | 0.98 | 0.96 | |
| sc02 | Molinia caerulea | 0.88 | 0.79 | 0.83 | 0.88 | 0.77 | 0.82 | 0.91 | 0.81 | 0.86 |
| background | 0.96 | 0.98 | 0.97 | 0.95 | 0.98 | 0.96 | 0.97 | 0.99 | 0.98 | |
| sc03 | M. caerulea | 0.92 | 0.75 | 0.82 | 0.89 | 0.83 | 0.86 | 0.87 | 0.8 | 0.84 |
| background | 0.95 | 0.99 | 0.97 | 0.96 | 0.98 | 0.97 | 0.96 | 0.98 | 0.97 | |
| sc04 | M. caerulea | 0.92 | 0.72 | 0.81 | 0.9 | 0.83 | 0.86 | 0.93 | 0.81 | 0.87 |
| background | 0.95 | 0.99 | 0.97 | 0.96 | 0.98 | 0.97 | 0.97 | 0.99 | 0.98 | |
| sc05 | M. caerulea | 0.92 | 0.73 | 0.81 | 0.94 | 0.79 | 0.86 | 0.93 | 0.82 | 0.87 |
| background | 0.95 | 0.99 | 0.97 | 0.96 | 0.99 | 0.97 | 0.97 | 0.99 | 0.98 | |
| sc06 | M. caerulea | 0.94 | 0.72 | 0.82 | 0.91 | 0.82 | 0.86 | 0.85 | 0.84 | 0.84 |
| background | 0.95 | 0.99 | 0.97 | 0.96 | 0.98 | 0.97 | - | - | - | |
| sc07 | M. caerulea | 0.9 | 0.68 | 0.78 | 0.92 | 0.82 | 0.87 | - | - | - |
| background | 0.94 | 0.99 | 0.96 | 0.96 | 0.99 | 0.97 | 0.96 | 0.99 | 0.98 | |
| sc08 | M. caerulea | 0.95 | 0.8 | 0.87 | 0.93 | 0.79 | 0.86 | - | - | - |
| background | 0.96 | 0.99 | 0.98 | 0.96 | 0.99 | 0.97 | - | - | - | |
| sc09 | M. caerulea | 0.96 | 0.75 | 0.84 | 0.94 | 0.85 | 0.89 | 0.92 | 0.85 | 0.88 |
| background | 0.95 | 0.99 | 0.97 | 0.97 | 0.99 | 0.98 | 0.97 | 0.98 | 0.98 | |
| sc. No. | Class | June | August | September | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| UA | PA | F1 | UA | PA | F1 | UA | PA | F1 | ||
| sc01 | C. epigejos | 0.64 | 0.46 | 0.54 | 0.64 | 0.57 | 0.6 | 0.67 | 0.51 | 0.58 |
| background | 0.85 | 0.92 | 0.89 | 0.87 | 0.9 | 0.89 | 0.86 | 0.93 | 0.89 | |
| sc02 | C. epigejos | 0.78 | 0.47 | 0.56 | 0.73 | 0.53 | 0.61 | 0.81 | 0.64 | 0.72 |
| background | 0.85 | 0.96 | 0.9 | 0.87 | 0.94 | 0.91 | 0.9 | 0.96 | 0.93 | |
| sc03 | C. epigejos | 0.74 | 0.57 | 0.64 | 0.76 | 0.57 | 0.65 | 0.81 | 0.65 | 0.72 |
| background | 0.88 | 0.94 | 0.91 | 0.88 | 0.95 | 0.91 | 0.9 | 0.95 | 0.92 | |
| sc04 | C. epigejos | 0.72 | 0.47 | 0.57 | 0.71 | 0.63 | 0.67 | 0.78 | 0.53 | 0.63 |
| background | 0.86 | 0.95 | 0.9 | 0.89 | 0.92 | 0.91 | 0.87 | 0.96 | 0.91 | |
| sc05 | C. epigejos | 0.75 | 0.45 | 0.57 | 0.79 | 0.61 | 0.69 | 0.88 | 0.63 | 0.73 |
| background | 0.85 | 0.95 | 0.9 | 0.89 | 0.95 | 0.92 | 0.9 | 0.97 | 0.93 | |
| sc06 | C. epigejos | 0.78 | 0.57 | 0.66 | 0.84 | 0.56 | 0.67 | - | - | - |
| background | 0.88 | 0.95 | 0.91 | 0.88 | 0.97 | 0.92 | - | - | - | |
| sc07 | C. epigejos | 0.81 | 0.41 | 0.55 | 0.82 | 0.54 | 0.65 | 0.87 | 0.54 | 0.67 |
| background | 0.84 | 0.97 | 0.9 | 0.86 | 0.96 | 0.91 | 0.87 | 0.97 | 0.92 | |
| sc08 | C. epigejos | 0.8 | 0.47 | 0.6 | 0.79 | 0.52 | 0.63 | - | - | - |
| background | 0.86 | 0.97 | 0.91 | 0.86 | 0.96 | 0.91 | - | - | - | |
| sc09 | C. epigejos | 0.83 | 0.56 | 0.67 | 0.75 | 0.66 | 0.7 | 0.82 | 0.62 | 0.7 |
| background | 0.88 | 0.97 | 0.92 | 0.91 | 0.94 | 0.92 | 0.89 | 0.96 | 0.92 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marcinkowska-Ochtyra, A.; Jarocińska, A.; Bzdęga, K.; Tokarska-Guzik, B. Classification of Expansive Grassland Species in Different Growth Stages Based on Hyperspectral and LiDAR Data. Remote Sens. 2018, 10, 2019. https://doi.org/10.3390/rs10122019
Marcinkowska-Ochtyra A, Jarocińska A, Bzdęga K, Tokarska-Guzik B. Classification of Expansive Grassland Species in Different Growth Stages Based on Hyperspectral and LiDAR Data. Remote Sensing. 2018; 10(12):2019. https://doi.org/10.3390/rs10122019
Chicago/Turabian StyleMarcinkowska-Ochtyra, Adriana, Anna Jarocińska, Katarzyna Bzdęga, and Barbara Tokarska-Guzik. 2018. "Classification of Expansive Grassland Species in Different Growth Stages Based on Hyperspectral and LiDAR Data" Remote Sensing 10, no. 12: 2019. https://doi.org/10.3390/rs10122019
APA StyleMarcinkowska-Ochtyra, A., Jarocińska, A., Bzdęga, K., & Tokarska-Guzik, B. (2018). Classification of Expansive Grassland Species in Different Growth Stages Based on Hyperspectral and LiDAR Data. Remote Sensing, 10(12), 2019. https://doi.org/10.3390/rs10122019