Imaging Spectroscopic Analysis of Biochemical Traits for Shrub Species in Great Basin, USA
Abstract
:1. Introduction
2. Materials and Data
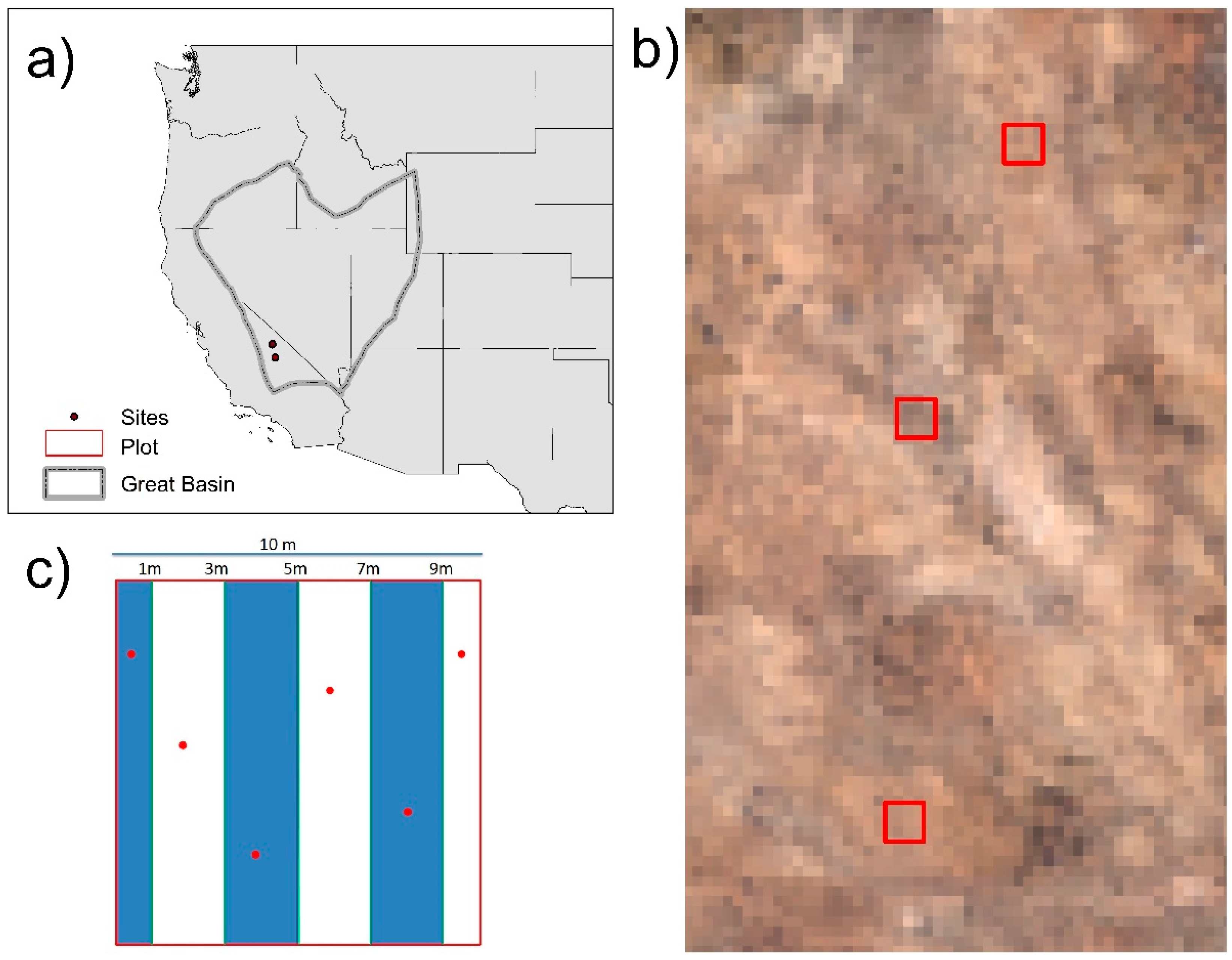
2.1. Stuty Sites
2.2. Field Data Collection
2.3. Biochemical Measurements
2.4. Remote Sensing Data
2.5. Data Analysis
3. Results
3.1. Variation in Biochemical Traits and Reflectance
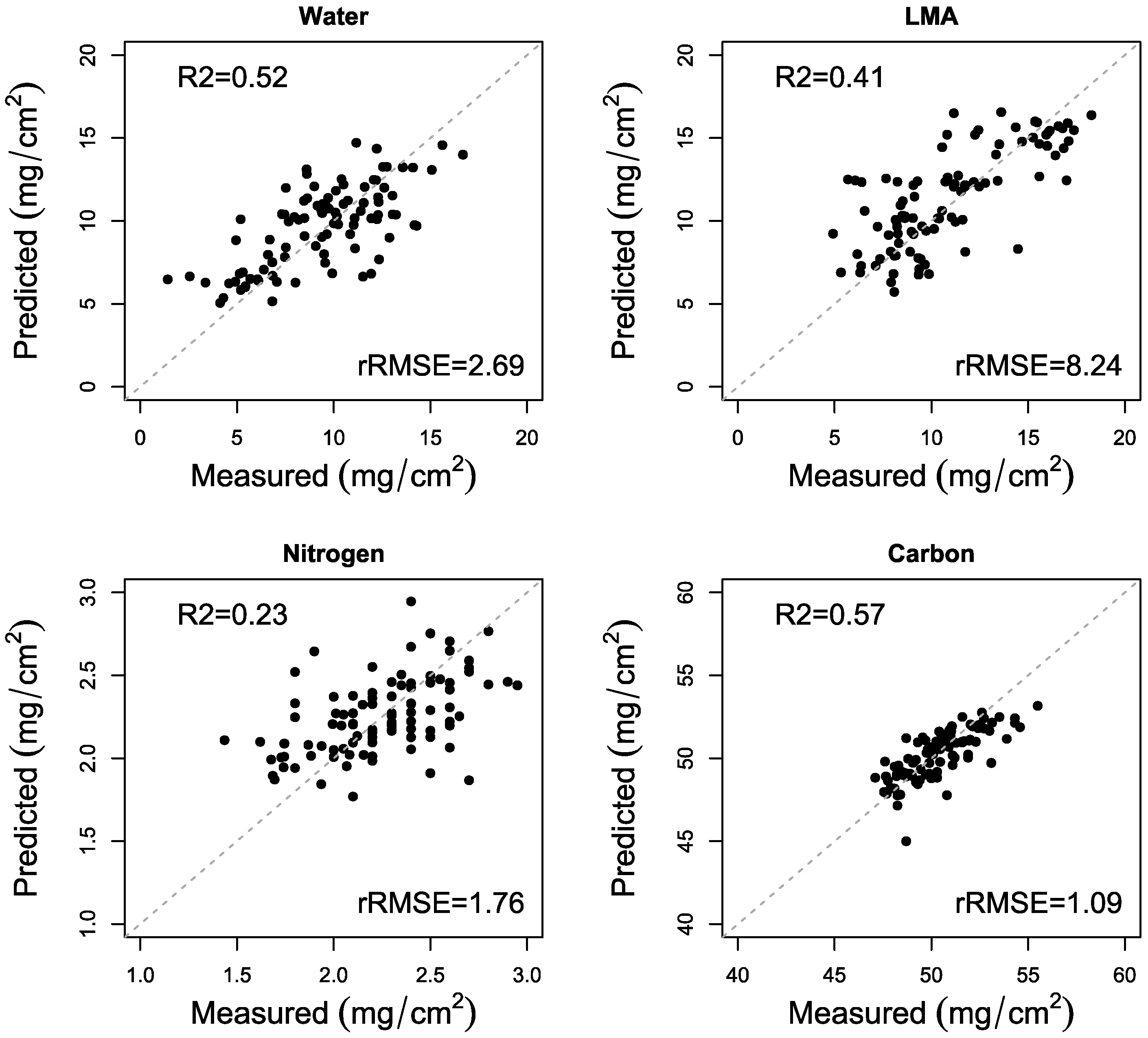
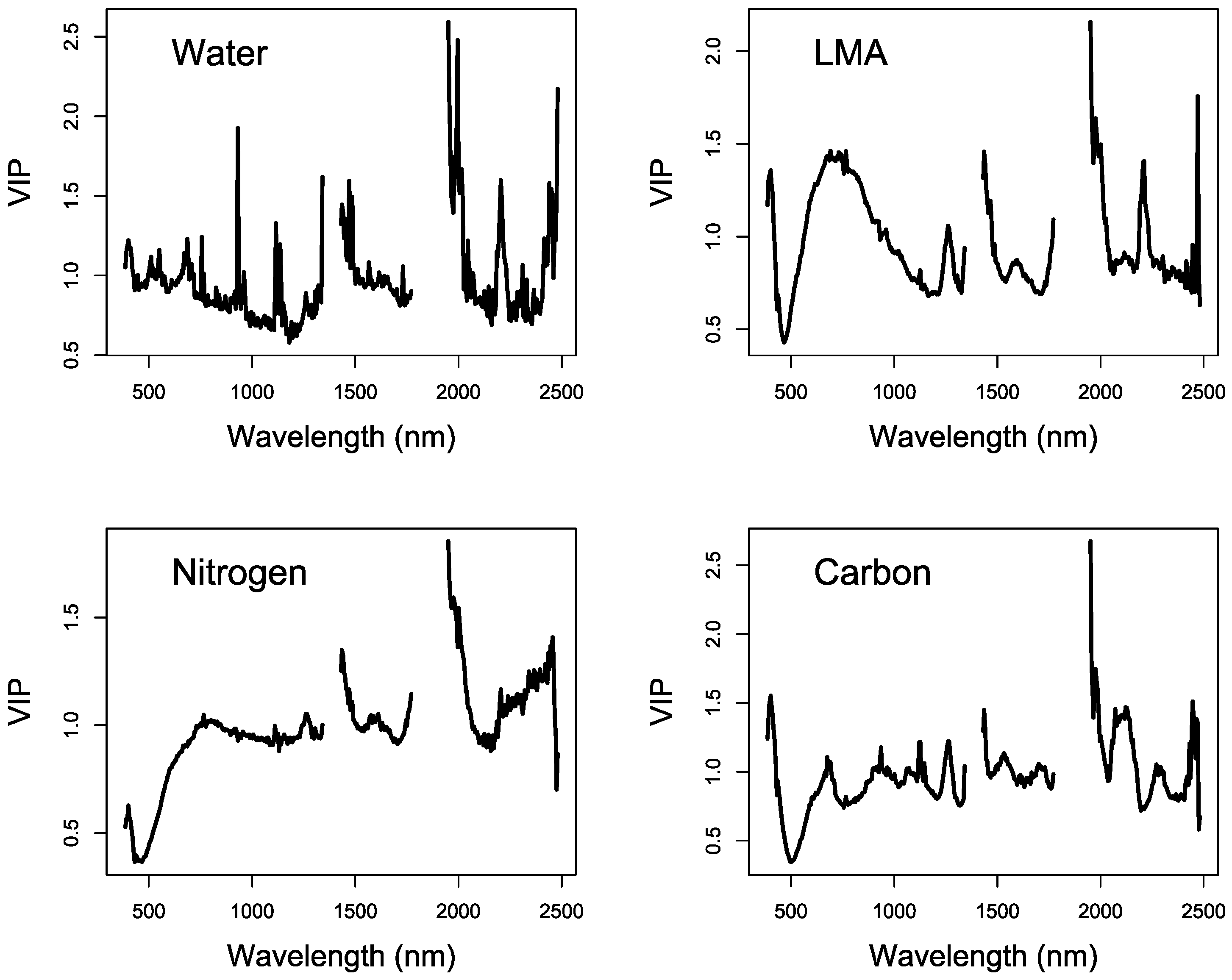
3.2. PLSR Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Melillo, J.M.; Aber, J.D.; Muratore, J.F. Nitrogen and Lignin Control of Hardwood Leaf Litter Decomposition Dynamics. Ecology 1982, 63, 621–626. [Google Scholar] [CrossRef] [Green Version]
- Shipley, B.; Vile, D.; Garnier, E.; Wright, I.J.; Poorter, H. Functional linkages between leaf traits and net photosynthetic rate: reconciling empirical and mechanistic models. Funct. Ecol. 2005, 19, 602–615. [Google Scholar] [CrossRef] [Green Version]
- Santiago, L.S.; Wright, S.J. Leaf functional traits of tropical forest plants in relation to growth form. Funct. Ecol. 2007, 21, 19–27. [Google Scholar] [CrossRef]
- Cornwell, W.K.; Cornelissen, J.H.C.; Amatangelo, K.; Dorrepaal, E.; Eviner, V.T.; Godoy, O.; Hobbie, S.E.; Hoorens, B.; Kurokawa, H.; Pérez-Harguindeguy, N.; et al. Plant species traits are the predominant control on litter decomposition rates within biomes worldwide. Ecol. Lett. 2008, 11, 1065–1071. [Google Scholar] [CrossRef] [PubMed]
- Wright, I.J.; Reich, P.B.; Westoby, M.; Ackerly, D.D.; Baruch, Z.; Bongers, F.; Cavender-Bares, J.; Chapin, T.; Cornelissen, J.H.C.; Diemer, M.; et al. The worldwide leaf economics spectrum. Nature 2004, 428, 821–827. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poorter, L. Leaf traits show different relationships with shade tolerance in moist versus dry tropical forests. New Phytol. 2009, 181, 890–900. [Google Scholar] [CrossRef] [PubMed]
- Niinemets, Ü. Photosynthesis and resource distribution through plant canopies. Plant Cell Environ. 2007, 30, 1052–1071. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goetz, S.J.; Prince, S.D. Remote sensing of net primary production in boreal forest stands. Agric. For. Meteorol. 1996, 78, 149–179. [Google Scholar] [CrossRef]
- Fourty, T.; Baret, F. Vegetation water and dry matter contents estimated from top-of-the-atmosphere reflectance data: A simulation study. Remote Sens. Environ. 1997, 61, 34–45. [Google Scholar] [CrossRef]
- Kramer, P.; Boyer, J. Water Relations of Plants and Soils; Academic Press: Cambridge, MA, USA, 1995; p. 495. [Google Scholar]
- Field, C.; Mooney, H. The photosynthesis-nitrogen relationship in wild plants. In on the Economy of Plant Form and Function; Givnish, T., Ed.; Cambridge University Press: Cambridge, UK, 1986; pp. 25–55. [Google Scholar]
- Reich, P.B.; Walters, M.B.; Ellsworth, D.S. Leaf Life-Span in Relation to Leaf, Plant, and Stand Characteristics among Diverse Ecosystems. Ecol. Monogr. 1992, 62, 365–392. [Google Scholar] [CrossRef] [Green Version]
- Kokaly, R.F.; Asner, G.P.; Ollinger, S.V.; Martin, M.E.; Wessman, C.A. Characterizing canopy biochemistry from imaging spectroscopy and its application to ecosystem studies. Remote Sens. Environ. 2009, 113, S78–S91. [Google Scholar] [CrossRef]
- Ustin, S.L.; Roberts, D.A.; Gamon, J.A.; Asner, G.P.; Green, R.O. Using imaging spectroscopy to study ecosystem processes and properties. BioScience 2004, 54, 523–534. [Google Scholar] [CrossRef]
- Asner, G.; Martin, R. Spectral and chemical analysis of tropical forests: Scaling from leaf to canopy levels. Remote Sens. Environ. 2008, 112, 3958–3970. [Google Scholar] [CrossRef]
- Asner, G.P.; Martin, R.E.; Knapp, D.E.; Tupayachi, R.; Anderson, C.B.; Sinca, F.; Vaughn, N.R.; Llactayo, W. Airborne laser-guided imaging spectroscopy to map forest trait diversity and guide conservation. Science 2017, 355, 385. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Serbin, S.P.; McNeil, B.E.; Kingdon, C.C.; Townsend, P.A. Imaging spectroscopy algorithms for mapping canopy foliar chemical and morphological traits and their uncertainties. Ecol. Appl. 2015, 25, 2180–2197. [Google Scholar] [CrossRef] [PubMed]
- Serbin, S.P.; Singh, A.; McNeil, B.E.; Kingdon, C.C.; Townsend, P.A. Spectroscopic determination of leaf morphological and biochemical traits for northern temperate and boreal tree species. Ecol. Appl. 2014, 24, 1651–1669. [Google Scholar] [CrossRef]
- Serrano, L.; Peñuelas, J.; Ustin, S.L. Remote sensing of nitrogen and lignin in Mediterranean vegetation from AVIRIS data: Decomposing biochemical from structural signals. Remote Sens. Environ. 2002, 81, 355–364. [Google Scholar] [CrossRef]
- Roberts, D.A.; Dennison, P.E.; Peterson, S.; Sweeney, S.; Rechel, J. Evaluation of Airborne Visible/Infrared Imaging Spectrometer (AVIRIS) and Moderate Resolution Imaging Spectrometer (MODIS) measures of live fuel moisture and fuel condition in a shrubland ecosystem in southern California. J. Geophys. Res. 2006, 111. [Google Scholar] [CrossRef]
- Roberts, D.A.; Ustin, S.L.; Ogunjemiyo, S.; Greenberg, J.; Bobrowski, S.Z.; Chen, J.; Hinckley, T.M. Spectral and structural measures of northwest forest vegetation at leaf to landscape scales. Ecosystems 2004, 7, 545–562. [Google Scholar] [CrossRef]
- Yang, X.; Tang, J.; Mustard, J.F.; Wu, J.; Zhao, K.; Serbin, S.; Lee, J.-E. Seasonal variability of multiple leaf traits captured by leaf spectroscopy at two temperate deciduous forests. Remote Sens. Environ. 2016, 179, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Asner, G.P.; Wessman, C.A.; Bateson, C.A.; Privette, J.L. Impact of tissue, canopy, and landscape factors on the Hyperspectral reflectance variability of arid ecosystems. Remote Sens. Environ. 2000, 74, 69–84. [Google Scholar] [CrossRef]
- Smith, M.O.; Ustin, S.L.; Adams, J.B.; Gillespie, A.R. Vegetation in deserts: I. A regional measure of abundance from multispectral images. Remote Sens. Environ. 1990, 31, 1–26. [Google Scholar] [CrossRef]
- Okin, G.S.; Roberts, D.A.; Murray, B.; Okin, W.J. Practical limits on hyperspectral vegetation discrimination in arid and semiarid environments. Remote Sens. Environ. 2001, 77, 212–225. [Google Scholar] [CrossRef]
- Ustin, S.L.; Smith, M.O.; Adams, J.B. A strategy for Developing Ecological Models Using Spectral Mixture Analysis. In Scaling Physiological Processes: Leaf to Globe; Ehlringer, J., Field, C., Eds.; Academic Press: Cambridge, MA, USA, 1993; pp. 339–357. [Google Scholar]
- Grossman, Y.L.; Ustin, S.L.; Jacquemoud, S.; Sanderson, E.W.; Schmuck, G.; Verdebout, J. Critique of stepwise multiple linear regression for the extraction of leaf biochemistry information from leaf reflectance data. Remote Sens. Environ. 1996, 56, 182–193. [Google Scholar] [CrossRef]
- Næsset, E.; Bollandsås, O.M.; Gobakken, T. Comparing regression methods in estimation of biophysical properties of forest stands from two different inventories using laser scanner data. Remote Sens. Environ. 2005, 94, 541–553. [Google Scholar] [CrossRef]
- Asner, G.P.; Martin, R.E.; Anderson, C.B.; Knapp, D.E. Quantifying forest canopy traits: Imaging spectroscopy versus field survey. Remote Sens. Environ. 2015, 158, 15–27. [Google Scholar] [CrossRef]
- Ustin, S.L.; Roberts, D.A.; Pinzón, J.; Jacquemoud, S.; Gardner, M.; Scheer, G.; Castañeda, C.M.; Palacios-Orueta, A. Estimating canopy water content of chaparral shrubs using optical methods. Remote Sens. Environ. 1998, 65, 280–291. [Google Scholar] [CrossRef]
- Serrano, L.; Ustin, S.L.; Roberts, D.A.; Gamon, J.A.; Peñuelas, J. Deriving water content of chaparral vegetation from AVIRIS data. Remote Sens. Environ. 2000, 74, 570–581. [Google Scholar] [CrossRef]
- Mirik, M.; Norland, J.E.; Crabtree, R.L.; Biondini, M.E. Hyperspectral One-Meter-Resolution Remote Sensing in Yellowstone National Park, Wyoming: I. Forage Nutritional Values. Rangel. Ecol. Manag. 2005, 58, 452–458. [Google Scholar] [CrossRef]
- Mitchell, J.J.; Glenn, N.F.; Sankey, T.T.; Derryberry, D.R.; Germino, M.J. Remote sensing of sagebrush canopy nitrogen. Remote Sens. Environ. 2012, 124, 217–223. [Google Scholar] [CrossRef]
- Reynolds, J.F.; Smith, D.M.S.; Lambin, E.F.; Turner, B.L.; Mortimore, M.; Batterbury, S.P.J.; Downing, T.E.; Dowlatabadi, H.; Fernandez, R.J.; Herrick, J.E.; et al. Global Desertification: Building a Science for Dryland Development. Science 2007, 316, 847–851. [Google Scholar] [CrossRef] [PubMed]
- Balch, J.K.; Bradley, B.A.; D’Antonio, C.M.; Gómez-Dans, J. Introduced annual grass increases regional fire activity across the arid western USA (1980–2009). Glob. Chang. Boil. 2013, 19, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Cook, E.R.; Woodhouse, C.A.; Eakin, C.M.; Meko, D.M.; Stahle, D.W. Long-Term Aridity Changes in the Western United States. Science 2004, 306, 1015–1018. [Google Scholar] [CrossRef] [PubMed]
- Asner, G.P.; Brodrick, P.G.; Anderson, C.B.; Vaughn, N.; Knapp, D.E.; Martin, R.E. Progressive forest canopy water loss during the 2012–2015 California drought. Proc. Natl. Acad. Sci. USA 2016, 113, E249–E255. [Google Scholar] [CrossRef] [PubMed]
- Xue, T.; Tang, G.; Sun, L.; Wu, Y.; Liu, Y.; Dou, Y. Long-term trends in precipitation and precipitation extremes and underlying mechanisms in the U.S. Great Basin during 1951–2013. J. Geophys. Res. Atmos. 2017, 122, 6152–6169. [Google Scholar] [CrossRef]
- Kampe, T.U.; Asner, G.P.; Green, R.O.; Eastwood, M.; Johnson, B.R.; Kuester, M. Advances in airborne remote sensing of ecosystem processes and properties—Toward high-quality measurement on a global scale. Proc. SPIE 2010, 7809, 7809–7813. [Google Scholar]
- Gao, B.-C.; Heidebrecht, K.B.; Goetz, A.F.H. Derivation of scaled surface reflectances from AVIRIS data. Remote Sens. Environ. 1993, 44, 165–178. [Google Scholar] [CrossRef]
- Martens, H.; Naes, T. Multivariate calibration by data compression. In Near-Infrared Technology in the Agriculture and Food Industries, 2nd ed.; Norris, W., Norris, K., Eds.; American Association of Cereal Chemists: St. Paul, MN, USA, 2001; pp. 59–100. [Google Scholar]
- Qi, Y.; Dennison, P.E.; Jolly, W.M.; Kropp, R.C.; Brewer, S.C. Spectroscopic analysis of seasonal changes in live fuel moisture content and leaf dry mass. Remote Sens. Environ. 2014, 150, 198–206. [Google Scholar] [CrossRef]
- Kennard, R.W.; Stone, L.A. Computer Aided Design of Experiments. Technometrics 1969, 11, 137–148. [Google Scholar] [CrossRef]
- Wold, S.; Sjöström, M.; Eriksson, L. PLS-regression: a basic tool of chemometrics. Chemom. Intell. Lab. Syst. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Mehmood, T.; Liland, K.H.; Snipen, L.; Sæbø, S. A review of variable selection methods in Partial Least Squares Regression. Chemom. Intell. Lab. Syst. 2012, 118, 62–69. [Google Scholar] [CrossRef]
- Dina, S.J.; Klikoff, L.G. Effect of Plant Moisture Stress on Carbohydrate and Nitrogen Content of Big Sagebrush. J. Range Manag. 1973, 26, 207–209. [Google Scholar] [CrossRef]
- Bolster, K.L.; Martin, M.E.; Aber, J.D. Determination of carbon fraction and nitrogen concentration in tree foliage by near infrared reflectances: a comparison of statistical methods. Can. J. For. Res. 1996, 26, 590–600. [Google Scholar] [CrossRef]
- Ourcival, J.M.; Joffre, R.; Rambal, S. Exploring the relationships between reflectance and anatomical and biochemical properties in Quercus ilex leaves. New Phytol. 1999, 143, 351–364. [Google Scholar] [CrossRef] [Green Version]
- Asner, G.P.; Martin, R.E.; Tupayachi, R.; Emerson, R.; Martinez, P.; Sinca, F.; Powell, G.V.N.; Wright, S.J.; Lugo, A.E. Taxonomy and remote sensing of leaf mass per area (LMA) in humid tropical forests. Ecol. Appl. 2011, 21, 85–98. [Google Scholar] [CrossRef] [PubMed]
- Serbin, S.P.; Dillaway, D.N.; Kruger, E.L.; Townsend, P.A. Leaf optical properties reflect variation in photosynthetic metabolism and its sensitivity to temperature. J. Exp. Bot. 2012, 63, 489–502. [Google Scholar] [CrossRef] [PubMed]
- Cheng, T.; Rivard, B.; Sánchez-Azofeifa, A.G.; Féret, J.-B.; Jacquemoud, S.; Ustin, S.L. Deriving leaf mass per area (LMA) from foliar reflectance across a variety of plant species using continuous wavelet analysis. ISPRS J. Photogramm. Remote Sens. 2014, 87, 28–38. [Google Scholar] [CrossRef]
- Kokaly, R.F.; Clark, R.N. Spectroscopic determination of leaf biochemistry using band-depth analysis of absorption features and stepwise multiple linear regression. Remote Sens. Environ. 1999, 67, 267–287. [Google Scholar] [CrossRef]
- Kokaly, R.F. Investigating a physical basis for spectroscopic estimates of leaf nitrogen concentration. Remote Sens. Environ. 2001, 75, 153–161. [Google Scholar] [CrossRef]
- Knyazikhin, Y.; Schull, M.A.; Stenberg, P.; Mõttus, M.; Rautiainen, M.; Yang, Y.; Marshak, A.; Latorre Carmona, P.; Kaufmann, R.K.; Lewis, P.; et al. Hyperspectral remote sensing of foliar nitrogen content. Proc. Natl. Acad. Sci. USA 2013, 110, E185–E192. [Google Scholar] [CrossRef] [PubMed]
- Asner, G.P.; Heidebrecht, K.B. Spectral unmixing of vegetation, soil and dry carbon cover in arid regions: Comparing multispectral and hyperspectral observations. Int. J. Remote Sens. 2002, 23, 3939–3958. [Google Scholar] [CrossRef]
- Huang, S.; Potter, C.; Crabtree, R.L.; Hager, S.; Gross, P. Fusing optical and radar data to estimate sagebrush, herbaceous, and bare ground cover in Yellowstone. Remote Sens. Environ. 2010, 114, 251–264. [Google Scholar] [CrossRef]
- McGwire, K.; Minor, T.; Fenstermaker, L. Hyperspectral Mixture Modeling for Quantifying Sparse Vegetation Cover in Arid Environments. Remote Sens. Environ. 2000, 72, 360–374. [Google Scholar] [CrossRef]
- Guerschman, J.P.; Hill, M.J.; Renzullo, L.J.; Barrett, D.J.; Marks, A.S.; Botha, E.J. Estimating fractional cover of photosynthetic vegetation, non-photosynthetic vegetation and bare soil in the Australian tropical savanna region upscaling the EO-1 Hyperion and MODIS sensors. Remote Sens. Environ. 2009, 113, 928–945. [Google Scholar] [CrossRef]
- Mundt, J.T.; Streutker, D.R.; Glenn, N.F. Mapping Sagebrush Distribution Using Fusion of Hyperspectral and Lidar Classifications. Photogramm. Eng. Remote Sens. 2006, 72, 47–54. [Google Scholar] [CrossRef]
- Mitchell, J.J.; Shrestha, R.; Spaete, L.P.; Glenn, N.F. Combining airborne hyperspectral and LiDAR data across local sites for upscaling shrubland structural information: Lessons for HyspIRI. Remote Sens. Environ. 2015, 167, 98–110. [Google Scholar] [CrossRef]

| Sagebrush | Bitterbrush | |||
|---|---|---|---|---|
| 2014 | 2015 | 2014 | 2015 | |
| LMA (mg/cm2) | 13.36 (3.24) | 8.67 (1.82) | 21.52 (3.92) | 18.22 (2.26) |
| Water (mg/cm2) | 9.24 (3.59) | 9.79 (2.49) | 14.73 (2.91) | 14.82 (2.38) |
| Nitrogen (%) | 2.09 (0.27) | 2.43 (0.28) | 1.49 (0.13) | 1.56 (0.16) |
| Carbon (%) | 51.56 (1.51) | 49 (1.27) | 50.27 (1.43) | 50.24 (1.12) |
| LAI | 1.91 (0.46) | 1.98 (0.49) | 1.99 (0.43) | 1.91 (0.59) |
| Major Width (m) | 1.68 (0.71) | 1.25 (0.46) | 1.47 (0.94) | 1.40 (0.57) |
| Minor Width (m) | 1.24 (0.54) | 1.13 (1.49) | 1.39 (1.33) | 1.17 (0.51) |
| Vegetation Cover | 0.34 (0.13) | 0.33 (0.17) | 0.14 (0.12) | 0.13 (0.1) |
| Water | LMA | Nitrogen | Carbon | |
|---|---|---|---|---|
| Water | 1.00 (1.00) | |||
| LMA | 0.60 * (−0.01) | 1.00 (1.00) | ||
| Nitrogen | 0.19 (−0.01) | −0.13 (0.08) | 1.00 (1.00) | |
| Carbon | −0.20 (−0.15) | −0.05 (0.31 *) | −0.03 (−0.14) | 1.00 (1.00) |
| Calibration | Validation | ||||
|---|---|---|---|---|---|
| R2 (S.D.) | rRMSE (S.D.) | R2 (S.D.) | rRMSE (S.D.) | ||
| Sagebrush (n = 194) | LMA | 0.66 (0.04) | 3.28 (0.39) | 0.52 (0.01) | 2.69 (0.01) |
| Water | 0.70 (0.03) | 3.98 (0.37) | 0.41 (0.06) | 8.24 (0.08) | |
| Nitrogen | 0.42 (0.04) | 1.57 (0.02) | 0.23 (0.05) | 1.76 (0.01) | |
| Carbon | 0.60 (0.03) | 1.10 (0.01) | 0.57 (0.02) | 1.09 (0.01) | |
| Bitterbrush (n = 74) | LMA | 0.86 (0.04) | 1.72 (0.11) | - | - |
| Water | 0.51 (0.06) | 2.04 (0.18) | 0.06 (0.17) | 2.34 (0.03) | |
| Nitrogen | 0.04 (0.04) | 1.49 (0.05) | - | - | |
| Carbon | 0.17 (0.06) | 1.09 (0.01) | 0.27 (0.06) | 1.07 (0.01) | |
| Calibration | Validation | ||||
|---|---|---|---|---|---|
| R2 (S.D.) | rRMSE (S.D.) | R2 (S.D.) | rRMSE (S.D.) | ||
| Sagebrush (n = 35) | LMA | 0.73 (0.09) | 1.31 (0.11) | 0.67 (0.06) | 1.32 (0.01) |
| Water | 0.67 (0.07) | 1.19 (0.11) | 0.1 (0.11) | 1.49 (0.02) | |
| Nitrogen | 0.66 (0.08) | 3.04 (0.24) | 0.52 (0.04) | 2.75 (0.01) | |
| Carbon | 0.7 (0.06) | 9.26 (1.16) | 0.38 (0.04) | 9.17 (0.01) | |
| Bitterbrush (n = 24) | LMA | 0.81 (0.05) | 2.07 (0.18) | 0.08 (0.05) | 1.94 (0.02) |
| Water | 0.81 (0.08) | 2.47 (0.58) | 0.68 (0.13) | 2.74 (0.01) | |
| Nitrogen | 0.12 (0.11) | 2.94 (0.83) | -- | -- | |
| Carbon | 0.8 (0.1) | 13.97 (2.26) | -- | -- | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qi, Y.; Ustin, S.L.; Glenn, N.F. Imaging Spectroscopic Analysis of Biochemical Traits for Shrub Species in Great Basin, USA. Remote Sens. 2018, 10, 1621. https://doi.org/10.3390/rs10101621
Qi Y, Ustin SL, Glenn NF. Imaging Spectroscopic Analysis of Biochemical Traits for Shrub Species in Great Basin, USA. Remote Sensing. 2018; 10(10):1621. https://doi.org/10.3390/rs10101621
Chicago/Turabian StyleQi, Yi, Susan L. Ustin, and Nancy F. Glenn. 2018. "Imaging Spectroscopic Analysis of Biochemical Traits for Shrub Species in Great Basin, USA" Remote Sensing 10, no. 10: 1621. https://doi.org/10.3390/rs10101621






