Author Contributions
Conceptualization, M.P., M.S., and H.N.; Methodology, M.S. and M.P.; Software, M.P.; Validation, M.P., M.S., and H.N.; Formal Analysis, M.P.; Investigation, M.P.; Resources, H.N.; Data Curation, M.S.; Writing—Original Draft Preparation, M.S., M.P., and H.N.; Writing—Review and Editing, H.N.; Visualization, M.P. and M.S.; Supervision, H.N.; Project Administration, H.N. and M.P.; Funding Acquisition, H.N.
Figure 1.
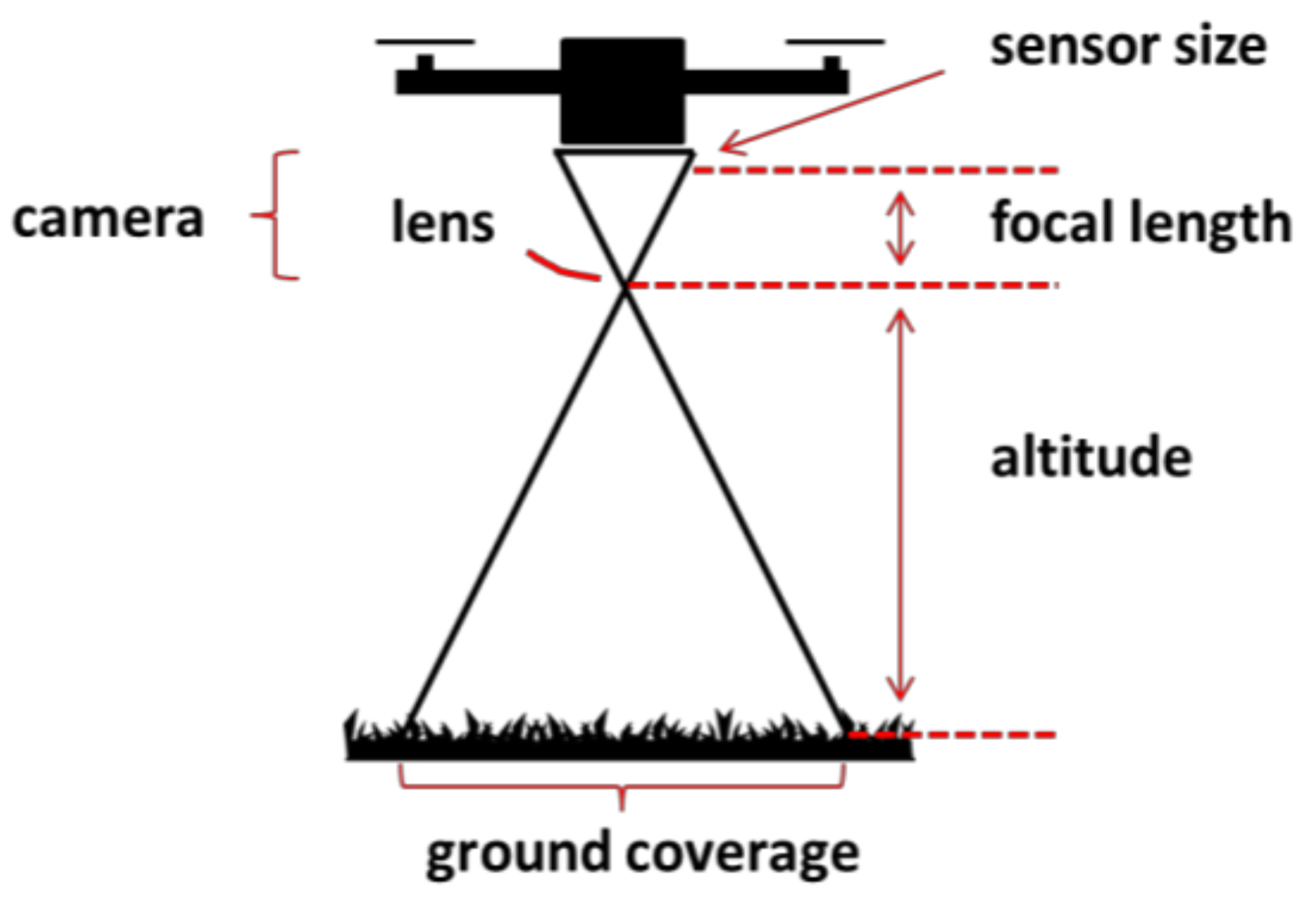
Capturing high resolution images at more than 100 waypoints (+) from Nadir. At a flight altitude of 5 m the ground sample distance (GSD) was 0.4 mm with a lens having a focal length of 60 mm and a camera equipped with an APS-C sensor.
Figure 1.
Capturing high resolution images at more than 100 waypoints (+) from Nadir. At a flight altitude of 5 m the ground sample distance (GSD) was 0.4 mm with a lens having a focal length of 60 mm and a camera equipped with an APS-C sensor.
Figure 2.
Test field of approx. 6.2 ha, unmanned aircraft system (UAS) flight routes (FL), image positions and allocation of images in test and training data sets. Coordinates shown are in UTM zone 32 N, ETRS89 (EPSG code: 5972).
Figure 2.
Test field of approx. 6.2 ha, unmanned aircraft system (UAS) flight routes (FL), image positions and allocation of images in test and training data sets. Coordinates shown are in UTM zone 32 N, ETRS89 (EPSG code: 5972).
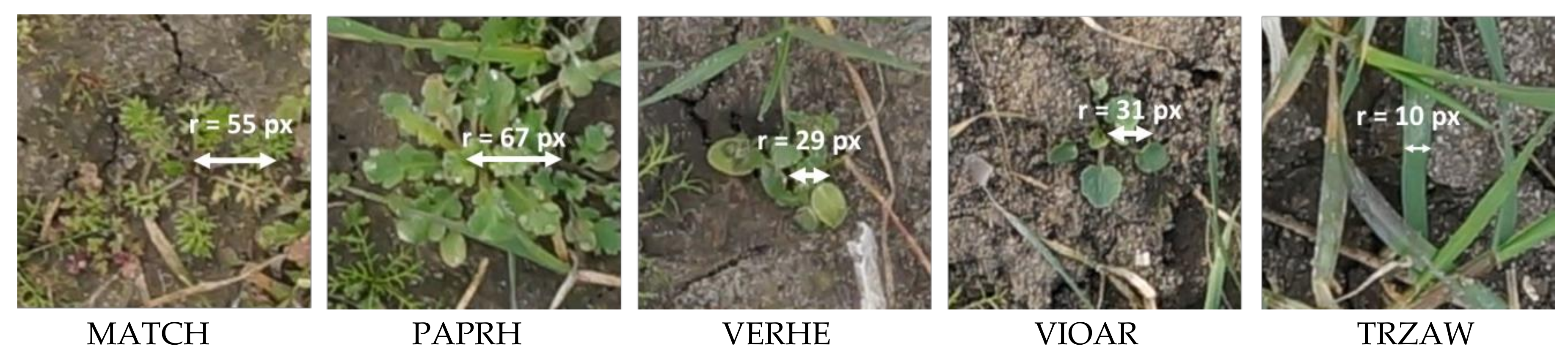
Figure 3.
Examples of sub-images extracted from the UAS imagery showing the different weed species and wheat plant. The average dimension (in pixel) was manually estimated.
Figure 3.
Examples of sub-images extracted from the UAS imagery showing the different weed species and wheat plant. The average dimension (in pixel) was manually estimated.
Figure 4.
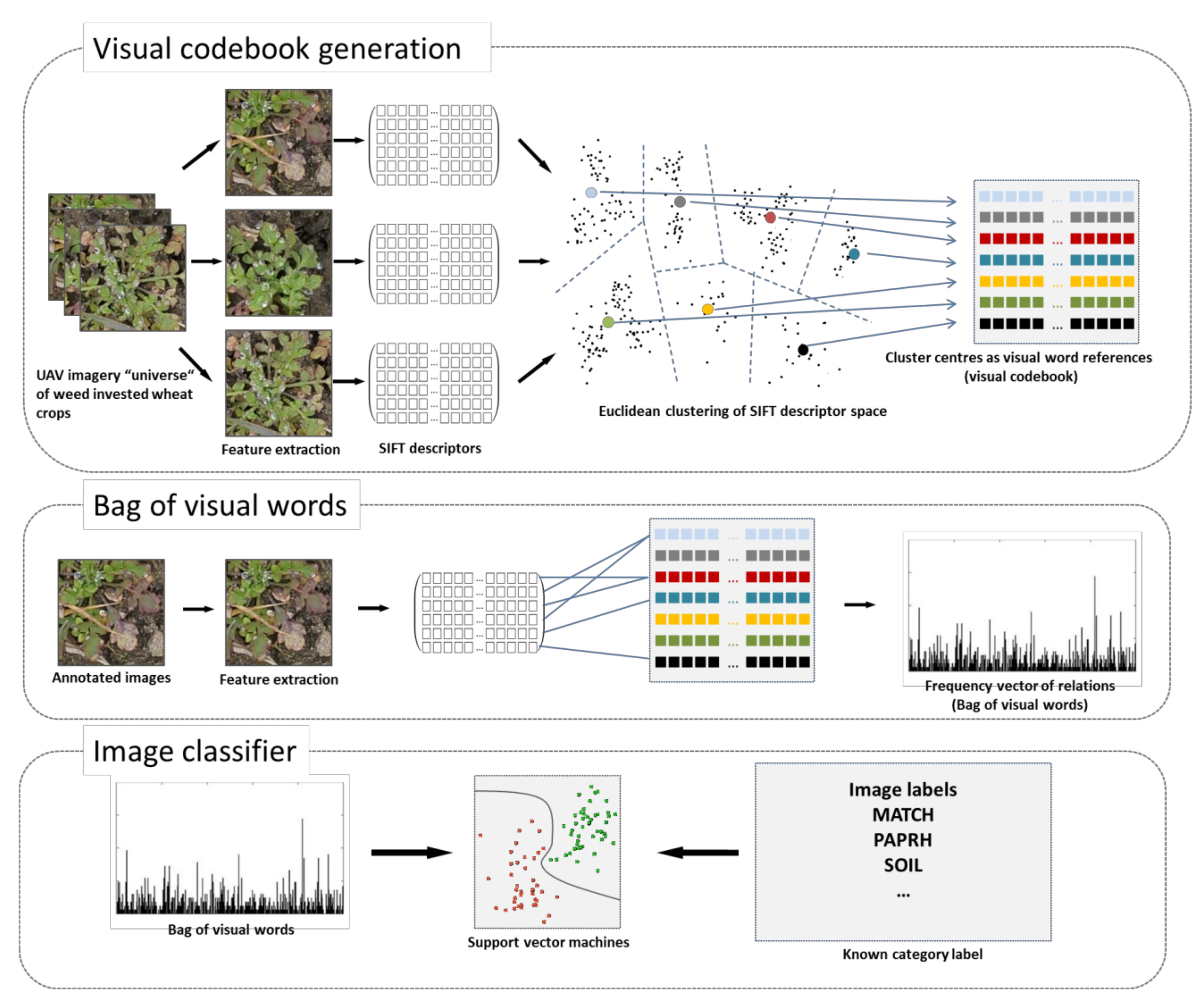
Bag of Visual Words (BoVW) framework: To train an image classifier using the BoVW concept, three steps need to be performed: First, a visual dictionary must be generated from many images. These images may be related to a specific theme, such as unspecific images from plants. From these images, the most relevant information will be extracted using key point extractors and descriptors. Depending on the type of the key point extractor, it finds key points at edges, corners, homogeneous regions, or blobs in the images, and the key point descriptor describes the immediate local neighborhood of the key points invariant to rotation or scale. The key point descriptions are collected over the entire image set and are generalized to a smaller set of code words by finding the centroids with a vector quantization method, such as k-means, to generate a visual dictionary. The second step involves a new labelled set of images. With the same key point descriptor as the visual dictionary that was built, the features of the new image will be extracted and described. The new key point descriptions are then referenced to the code words of the visual dictionary based on its similarity. The frequencies of the key point allocations are recorded over the code words to generate the BoVW vector. In the last step, the labelled BoVW vectors are used to train the image classifier using in our case a support vector machine (SVM). In case of a prediction given a new image, a new BoVW vector must be produced in relation to the codebook, and using the calibrated image classifier, the new BoVW vector will be decided to belong to a specific category, such as a specific weed species.
Figure 4.
Bag of Visual Words (BoVW) framework: To train an image classifier using the BoVW concept, three steps need to be performed: First, a visual dictionary must be generated from many images. These images may be related to a specific theme, such as unspecific images from plants. From these images, the most relevant information will be extracted using key point extractors and descriptors. Depending on the type of the key point extractor, it finds key points at edges, corners, homogeneous regions, or blobs in the images, and the key point descriptor describes the immediate local neighborhood of the key points invariant to rotation or scale. The key point descriptions are collected over the entire image set and are generalized to a smaller set of code words by finding the centroids with a vector quantization method, such as k-means, to generate a visual dictionary. The second step involves a new labelled set of images. With the same key point descriptor as the visual dictionary that was built, the features of the new image will be extracted and described. The new key point descriptions are then referenced to the code words of the visual dictionary based on its similarity. The frequencies of the key point allocations are recorded over the code words to generate the BoVW vector. In the last step, the labelled BoVW vectors are used to train the image classifier using in our case a support vector machine (SVM). In case of a prediction given a new image, a new BoVW vector must be produced in relation to the codebook, and using the calibrated image classifier, the new BoVW vector will be decided to belong to a specific category, such as a specific weed species.
![Remotesensing 10 01530 g004 Remotesensing 10 01530 g004]()
Figure 5.
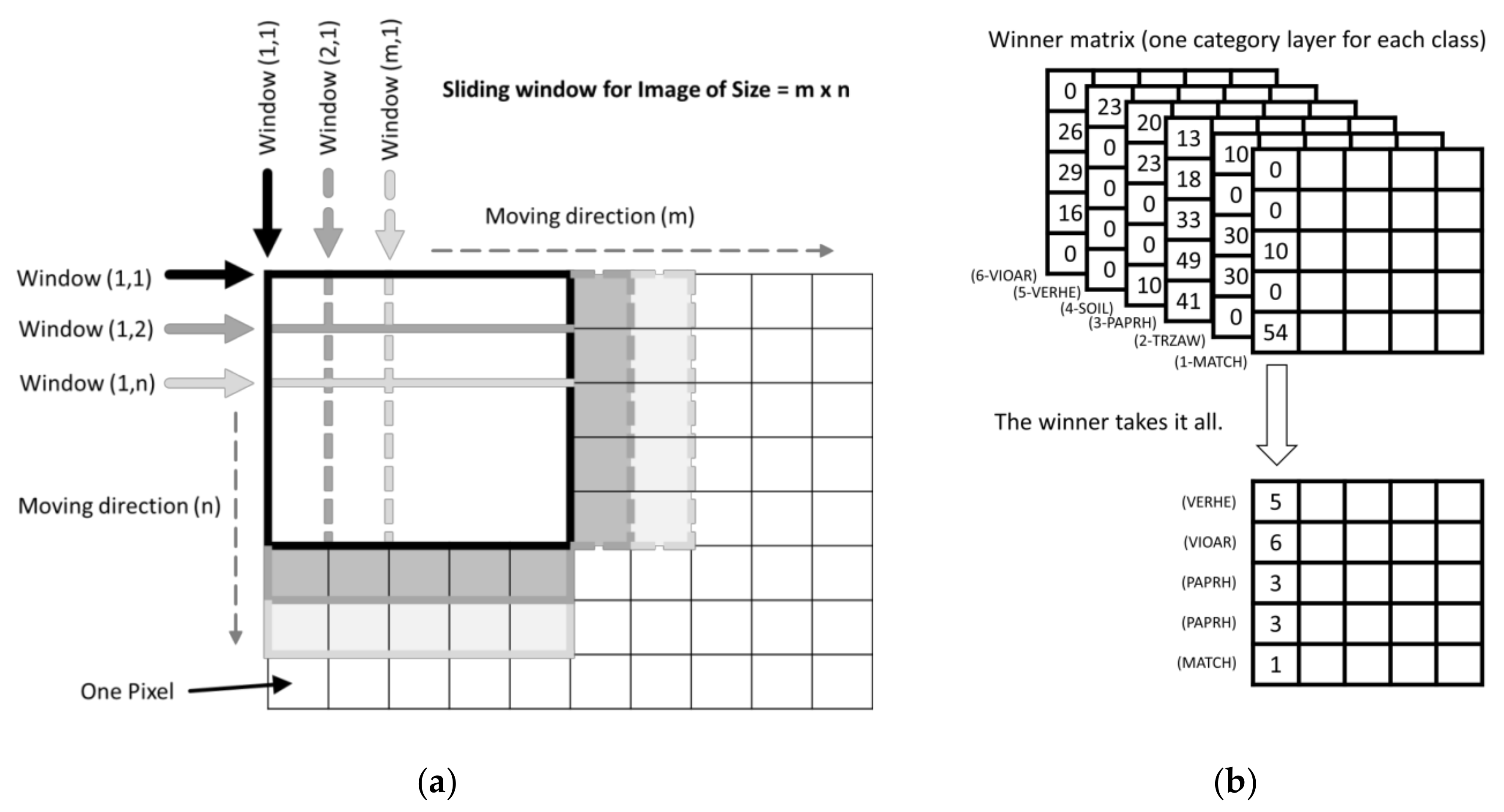
(a) Principle of the sliding window approach used; (b) One layer for each category was pre-allocated with the size m × n of the original image.
Figure 5.
(a) Principle of the sliding window approach used; (b) One layer for each category was pre-allocated with the size m × n of the original image.
Figure 6.
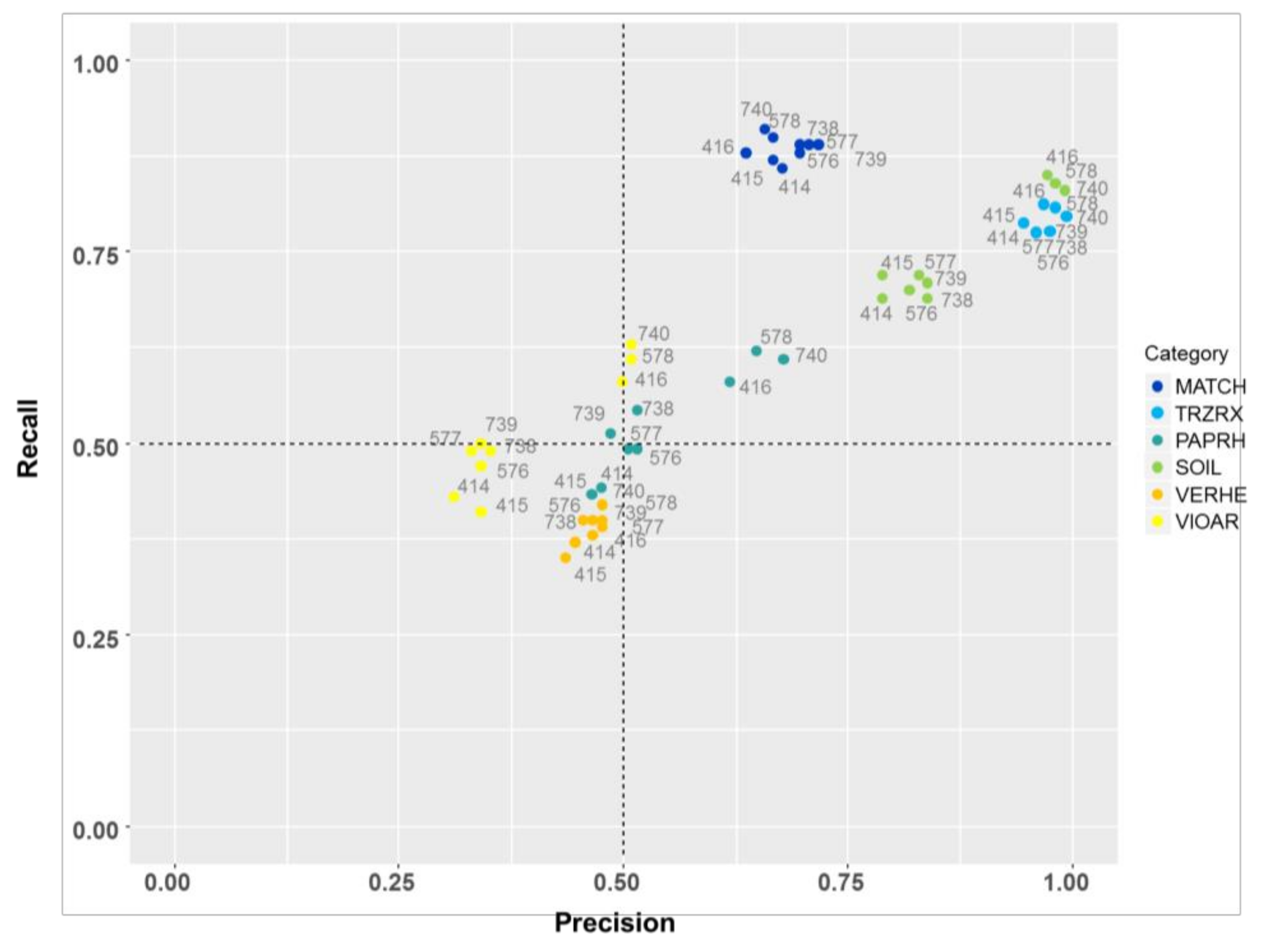
Precision-recall-plots of the different BoVW models shown in
Table 4 for the classes (a) MATCH, (b) TRZAW, (c) PAPRH, (d) SOIL, (e) VERHE, and (f) VIOAR.
Figure 6.
Precision-recall-plots of the different BoVW models shown in
Table 4 for the classes (a) MATCH, (b) TRZAW, (c) PAPRH, (d) SOIL, (e) VERHE, and (f) VIOAR.
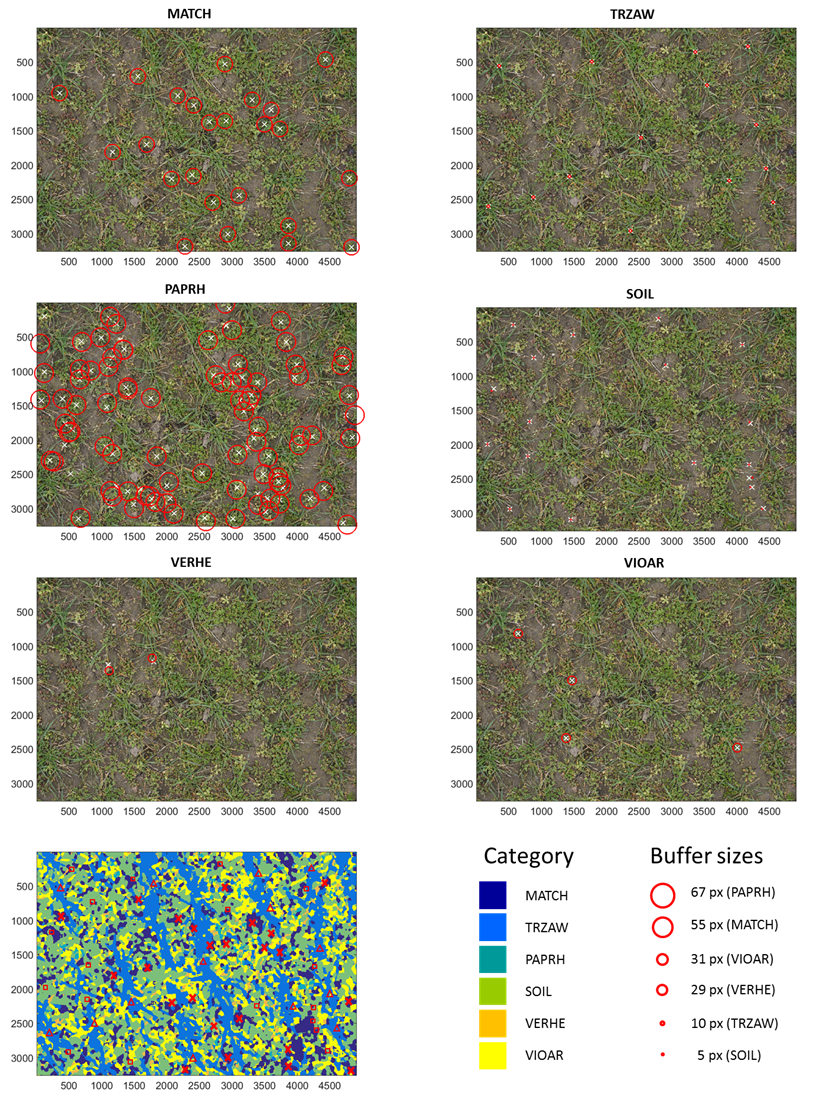
Figure 7.
Weed classification results of the model 578 shown for the UAS image taken at field position 42. The sliding window has a block size of 50 px and a shift of 5 px. A category map (bottom left) draws the spatial distribution of weeds, winter wheat, and soil. Ground truthing points for MATCH (x), TRZAW (∆), and soil (□) illustrate the accuracies.
Figure 7.
Weed classification results of the model 578 shown for the UAS image taken at field position 42. The sliding window has a block size of 50 px and a shift of 5 px. A category map (bottom left) draws the spatial distribution of weeds, winter wheat, and soil. Ground truthing points for MATCH (x), TRZAW (∆), and soil (□) illustrate the accuracies.
Table 1.
Summary of categories from weed species and crop plants, their EPPO-Codes, plant development stage, and specific plant dimensions. Category #4 comprises images from soil only and is not shown here.
Table 1.
Summary of categories from weed species and crop plants, their EPPO-Codes, plant development stage, and specific plant dimensions. Category #4 comprises images from soil only and is not shown here.
| Species | # | EPPO-Code | BBCH | Avg. Radius (px) |
|---|
| Matricaria recutita L. (Wild Chamomille) | 1 | MATCH | 16–18 | 55 |
| Papaver rhoeas L. (Common Poppy) | 3 | PAPRH | 17–19 | 67 |
| Veronica hederifolia L. (Ivyleaf Speedwell) | 5 | VERHE | 21 | 29 |
| Viola arvensis M. (Field Pansy) | 6 | VIOAR | 15 | 31 |
| Triticum aestivum L. (Winter Wheat) | 2 | TRZAW | 23 | 10 |
Table 2.
Allocation of annotated sub-images into train set and test set for wheat, soil, and the four weed species.
Table 2.
Allocation of annotated sub-images into train set and test set for wheat, soil, and the four weed species.
| n | MATCH | VERHE | PAPRH | VIOAR | TRZAW | SOIL | Total |
|---|
| Train Set | 6397 | 1728 | 1598 | 1701 | 2164 | 4077 | 17,665 |
| Test Set | 2086 | 891 | 711 | 1107 | 1038 | 1954 | 7787 |
| ∑ | 8483 | 2619 | 2309 | 2808 | 3202 | 6031 | 25,452 |
Table 3.
Permuted repetition of input parameters for generating n = 486 BoVW-models.
Table 3.
Permuted repetition of input parameters for generating n = 486 BoVW-models.
| Input | Variation |
|---|
| Vocabulary size | {200…500…1000} |
| Color space | {gray…rgb…hsv} |
| Number of spatial bins | {2…[2 4]…[2 4 8]} |
| Quantizer | {KDTREE…VQ} |
| SVM.C | {1…5…10} |
| SVM solver | {sdca… sgd…liblinear} |
Table 4.
Precision and recall values of the model testing with an independent image set. K-dimensional binary tree (KDTREE) was used as quantizer and Dual Coordinate Ascent solver (SDCA) as the support vector machines (SVM) solver.
Table 4.
Precision and recall values of the model testing with an independent image set. K-dimensional binary tree (KDTREE) was used as quantizer and Dual Coordinate Ascent solver (SDCA) as the support vector machines (SVM) solver.
| ID | Words | Color | Precision in % | Accuracy Overall in % |
|---|
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
|---|
| 414 | 200 | gray | 67.16 | 47.12 | 44.56 | 31.71 | 95.28 | 79.43 | 64.53 |
| 415 | 200 | rgb | 66.35 | 45.85 | 44.33 | 34.60 | 94.99 | 79.38 | 64.53 |
| 416 | 200 | hsv | 63.14 | 61.74 | 47.14 | 49.59 | 97.21 | 97.44 | 72.40 |
| 576 | 500 | gray | 68.60 | 50.63 | 45.79 | 34.96 | 96.15 | 82.24 | 66.66 |
| 577 | 500 | rgb | 69.99 | 50.07 | 48.04 | 34.42 | 96.15 | 82.60 | 67.25 |
| 578 | 500 | hsv | 65.87 | 64.98 | 47.92 | 51.31 | 97.98 | 98.31 | 74.09 |
| 738 | 1000 | gray | 68.55 | 51.34 | 47.03 | 35.59 | 96.53 | 83.98 | 67.43 |
| 739 | 1000 | rgb | 71.28 | 48.24 | 47.92 | 35.32 | 96.63 | 83.57 | 67.86 |
| 740 | 1000 | hsv | 65.44 | 67.51 | 47.59 | 50.68 | 98.65 | 98.52 | 74.21 |
| | Recall in % | |
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
| 414 | 200 | gray | 85.64 | 44.02 | 37.07 | 42.60 | 78.62 | 69.38 | 64.53 |
| 415 | 200 | rgb | 87.43 | 42.78 | 35.49 | 41.18 | 79.45 | 71.87 | 64.53 |
| 416 | 200 | hsv | 88.15 | 57.92 | 37.94 | 58.03 | 80.66 | 85.34 | 72.40 |
| 576 | 500 | gray | 87.52 | 49.18 | 39.92 | 46.80 | 78.34 | 69.96 | 66.66 |
| 577 | 500 | rgb | 88.65 | 49.24 | 38.77 | 48.78 | 78.27 | 71.51 | 67.25 |
| 578 | 500 | hsv | 90.28 | 61.68 | 41.78 | 60.62 | 80.71 | 83.63 | 74.09 |
| 738 | 1000 | gray | 88.65 | 54.15 | 40.33 | 48.88 | 78.22 | 69.12 | 67.43 |
| 739 | 1000 | rgb | 88.72 | 51.12 | 40.44 | 49.68 | 77.51 | 70.91 | 67.86 |
| 740 | 1000 | hsv | 91.18 | 60.76 | 42.27 | 62.75 | 79.75 | 83.01 | 74.21 |
Table 5.
Confusion matrix for the low resolution prediction (200/20) of model 578 on the UAS field scenes. The accuracy overall was 48%.
Table 5.
Confusion matrix for the low resolution prediction (200/20) of model 578 on the UAS field scenes. The accuracy overall was 48%.
| Actual/Predicted | 1 | 2 | 3 | 4 | 5 | 6 | Total | Recall |
|---|
| 1 | MATCH | 921 | 50 | 52 | 11 | 323 | 1149 | 2506 | 79.06 |
| 2 | PAPRH | 51 | 309 | 8 | 5 | 103 | 235 | 711 | 77.44 |
| 3 | VERHE | 16 | 16 | 51 | 7 | 244 | 567 | 901 | 28.65 |
| 4 | VIOAR | 135 | 18 | 49 | 58 | 291 | 1025 | 1576 | 69.05 |
| 5 | TRZAW | 9 | 2 | 10 | 2 | 913 | 102 | 1038 | 48.18 |
| 6 | SOIL | 33 | 4 | 8 | 1 | 21 | 1887 | 1954 | 38.01 |
| Total | 1165 | 399 | 178 | 84 | 1895 | 4965 | 8686 | |
| Precision | 36.75 | 43.46 | 5.66 | 3.68 | 87.96 | 96.57 | | |
Table 6.
Confusion matrix for the high resolution prediction (50/5) of model 578 on the UAS field scenes. The accuracy overall was 67%.
Table 6.
Confusion matrix for the high resolution prediction (50/5) of model 578 on the UAS field scenes. The accuracy overall was 67%.
| Actual/Predicted | 1 | 2 | 3 | 4 | 5 | 6 | Total | Recall |
|---|
| 1 | MATCH | 2185 | 0 | 27 | 34 | 155 | 105 | 2506 | 68.47 |
| 2 | PAPRH | 281 | 89 | 35 | 113 | 112 | 81 | 711 | 71.20 |
| 3 | VERHE | 252 | 19 | 106 | 211 | 141 | 172 | 901 | 34.53 |
| 4 | VIOAR | 356 | 16 | 122 | 743 | 209 | 130 | 1576 | 66.10 |
| 5 | TRZAW | 17 | 1 | 16 | 16 | 936 | 52 | 1038 | 59.77 |
| 6 | SOIL | 100 | 0 | 1 | 7 | 13 | 1833 | 1954 | 77.24 |
| Total | 3191 | 125 | 307 | 1124 | 1566 | 2373 | 8686 | |
| Precision | 87.19 | 12.52 | 11.76 | 47.14 | 90.17 | 93.81 | | |
Table 7.
Confusion matrix for the low resolution prediction (200/20) of model 578 on the UAS field scenes. When allowing a ground truthing tolerance considering plant dimensions, the accuracy overall was 80%.
Table 7.
Confusion matrix for the low resolution prediction (200/20) of model 578 on the UAS field scenes. When allowing a ground truthing tolerance considering plant dimensions, the accuracy overall was 80%.
| Actual/Predicted | 1 | 2 | 3 | 4 | 5 | 6 | Total | Recall |
|---|
| 1 | MATCH | 1998 | 18 | 20 | 3 | 112 | 355 | 2506 | 94.47 |
| 2 | PAPRH | 16 | 560 | 4 | 0 | 43 | 88 | 711 | 96.05 |
| 3 | VERHE | 6 | 2 | 709 | 2 | 48 | 134 | 901 | 92.08 |
| 4 | VIOAR | 71 | 1 | 26 | 769 | 185 | 524 | 1576 | 99.23 |
| 5 | TRZAW | 2 | 0 | 5 | 0 | 980 | 51 | 1038 | 70.86 |
| 6 | SOIL | 22 | 2 | 6 | 1 | 15 | 1908 | 1954 | 62.35 |
| Total | 2115 | 583 | 770 | 775 | 1383 | 3060 | 8686 | |
| Precision | 79.73 | 78.76 | 78.69 | 48.79 | 94.41 | 97.65 | | |
Table 8.
Confusion matrix for the high resolution prediction (50/5) of model 578 on the UAS field scenes. When allowing a ground truthing tolerance considering plant dimensions, the accuracy overall was 95%.
Table 8.
Confusion matrix for the high resolution prediction (50/5) of model 578 on the UAS field scenes. When allowing a ground truthing tolerance considering plant dimensions, the accuracy overall was 95%.
| Actual/Predicted | 1 | 2 | 3 | 4 | 5 | 6 | Total | Recall |
|---|
| 1 | MATCH | 2478 | 0 | 2 | 0 | 18 | 8 | 2506 | 92.50 |
| 2 | PAPRH | 62 | 573 | 8 | 23 | 30 | 15 | 711 | 98.79 |
| 3 | VERHE | 14 | 2 | 864 | 11 | 3 | 7 | 901 | 95.47 |
| 4 | VIOAR | 94 | 5 | 29 | 1359 | 53 | 36 | 1576 | 97.42 |
| 5 | TRZAW | 1 | 0 | 2 | 1 | 1026 | 8 | 1038 | 90.48 |
| 6 | SOIL | 30 | 0 | 0 | 1 | 4 | 1919 | 1954 | 96.29 |
| Total | 2679 | 580 | 905 | 1395 | 1134 | 1993 | 8686 | |
| Precision | 98.88 | 80.59 | 95.89 | 86.23 | 98.84 | 98.21 | | |
Table 9.
Correlation between precision, recall, altitude (ALT), and window filtering (block size, BS, and shift), where n is the number of ground truthing points from manual annotations of the images.
Table 9.
Correlation between precision, recall, altitude (ALT), and window filtering (block size, BS, and shift), where n is the number of ground truthing points from manual annotations of the images.
| ID | BS [px] | Shift [px] | ALT [m] | n | Precision in % | Accuracy Overall in % |
|---|
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
|---|
| 578 | 200 | 20 | 1.0–2.5 | 1177 | 58.55 | 89.08 | 0.00 | 24.14 | 93.84 | 97.77 | 73.24 |
| 578 | 200 | 20 | 2.5–4.0 | 1275 | 41.30 | 33.33 | 0.00 | 3.77 | 89.06 | 99.46 | 55.53 |
| 578 | 200 | 20 | 4.0–5.5 | 4144 | 37.39 | 28.87 | 2.88 | 0.83 | 89.46 | 93.14 | 40.15 |
| 578 | 200 | 20 | 5.5–6.0 | 2090 | 14.73 | 0.00 | 9.23 | 0.29 | 82.64 | 98.01 | 43.30 |
| 578 | 50 | 5 | 1.0–2.5 | 1177 | 83.22 | 13.79 | 0.00 | 87.93 | 82.88 | 86.59 | 73.15 |
| 578 | 50 | 5 | 2.5–4.0 | 1275 | 84.42 | 0.00 | 1.79 | 78.77 | 93.75 | 94.07 | 80.24 |
| 578 | 50 | 5 | 4.0–5.5 | 4144 | 88.40 | 12.26 | 10.86 | 37.87 | 94.09 | 93.59 | 63.88 |
| 578 | 50 | 5 | 5.5–6.0 | 2090 | 88.60 | 0.00 | 15.38 | 29.86 | 86.50 | 98.56 | 65.12 |
| | Recall in % | |
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
| 578 | 200 | 20 | 1.0–2.5 | 1177 | 88.12 | 81.15 | 0.00 | 82.35 | 74.86 | 63.99 | 73.24 |
| 578 | 200 | 20 | 2.5–4.0 | 1275 | 92.44 | 10.00 | 0.00 | 57.14 | 64.53 | 45.78 | 55.53 |
| 578 | 200 | 20 | 4.0–5.5 | 4144 | 73.31 | 77.27 | 10.00 | 46.67 | 37.22 | 28.49 | 40.15 |
| 578 | 200 | 20 | 5.5–6.0 | 2090 | 78.48 | NA | 54.55 | 25.00 | 50.20 | 38.29 | 43.30 |
| 578 | 50 | 5 | 1.0–2.5 | 1177 | 73.55 | 92.31 | 0.00 | 67.40 | 60.80 | 84.47 | 73.15 |
| 578 | 50 | 5 | 2.5–4.0 | 1275 | 87.37 | 0.00 | 16.67 | 69.29 | 69.77 | 90.41 | 80.24 |
| 578 | 50 | 5 | 4.0–5.5 | 4144 | 65.57 | 76.47 | 22.52 | 66.67 | 53.90 | 72.43 | 63.88 |
| 578 | 50 | 5 | 5.5–6.0 | 2090 | 62.90 | 0.00 | 53.85 | 58.52 | 62.56 | 72.51 | 65.12 |
Table 10.
Correlation between precision, recall, altitude (ALT), and window filtering (block size, BS, and shift), with buffered accuracies (tolerance threshold was chosen according to the average plant size of each species).
Table 10.
Correlation between precision, recall, altitude (ALT), and window filtering (block size, BS, and shift), with buffered accuracies (tolerance threshold was chosen according to the average plant size of each species).
| ID | BS [px] | Shift [px] | ALT [m] | n | Precision | Accuracy Overall in % |
|---|
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
|---|
| 578 | 200 | 20 | 1.0–2.5 | 1177 | 92.11 | 99.43 | 85.71 | 100.00 | 98.63 | 100.00 | 97.45 |
| 578 | 200 | 20 | 2.5–4.0 | 1275 | 91.69 | 100.00 | 83.04 | 66.98 | 95.31 | 99.46 | 89.65 |
| 578 | 200 | 20 | 4.0–5.5 | 4144 | 77.44 | 71.89 | 67.73 | 35.15 | 94.34 | 94.93 | 71.79 |
| 578 | 200 | 20 | 5.5–6.0 | 2090 | 67.46 | 75.00 | 84.84 | 45.22 | 91.96 | 98.19 | 79.38 |
| 578 | 50 | 5 | 1.0–2.5 | 1177 | 98.68 | 87.36 | 95.24 | 100.00 | 99.32 | 97.21 | 96.77 |
| 578 | 50 | 5 | 2.5–4.0 | 1275 | 100.00 | 100.00 | 99.11 | 95.28 | 98.44 | 99.46 | 98.75 |
| 578 | 50 | 5 | 4.0–5.5 | 4144 | 98.42 | 78.11 | 93.61 | 84.14 | 99.23 | 97.32 | 92.45 |
| 578 | 50 | 5 | 5.5–6.0 | 2090 | 99.52 | 100.00 | 96.70 | 78.84 | 98.39 | 99.10 | 95.22 |
| | Recall | |
| MATCH | PAPRH | VERHE | VIOAR | TRZAW | SOIL |
| 578 | 200 | 20 | 1.0–2.5 | 1177 | 100.00 | 98.30 | 94.74 | 100.00 | 97.96 | 93.96 | 97.45 |
| 578 | 200 | 20 | 2.5–4.0 | 1275 | 98.06 | 100.00 | 98.94 | 98.61 | 86.73 | 79.70 | 89.65 |
| 578 | 200 | 20 | 4.0–5.5 | 4144 | 91.69 | 95.01 | 85.48 | 98.67 | 56.64 | 46.60 | 71.79 |
| 578 | 200 | 20 | 5.5–6.0 | 2090 | 95.95 | 100.00 | 94.38 | 100.00 | 75.86 | 64.08 | 79.38 |
| 578 | 50 | 5 | 1.0–2.5 | 1177 | 96.15 | 100.00 | 86.96 | 95.60 | 94.16 | 98.31 | 96.77 |
| 578 | 50 | 5 | 2.5–4.0 | 1275 | 99.23 | 75.00 | 98.23 | 99.02 | 96.92 | 99.46 | 98.75 |
| 578 | 50 | 5 | 4.0–5.5 | 4144 | 90.04 | 99.04 | 93.02 | 96.73 | 84.10 | 94.50 | 92.45 |
| 578 | 50 | 5 | 5.5–6.0 | 2090 | 92.49 | 66.67 | 96.92 | 99.27 | 93.87 | 95.15 | 95.22 |