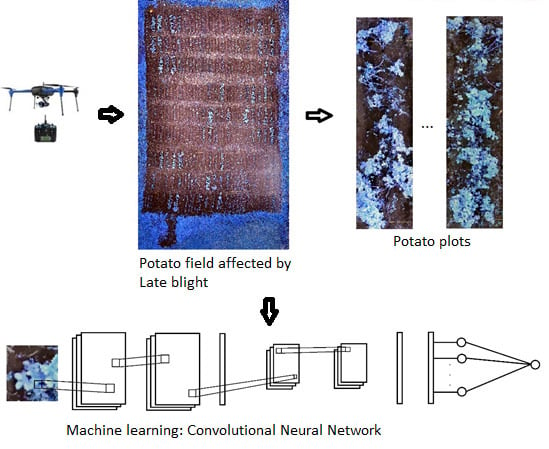
Evaluating Late Blight Severity in Potato Crops Using Unmanned Aerial Vehicles and Machine Learning Algorithms
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Design
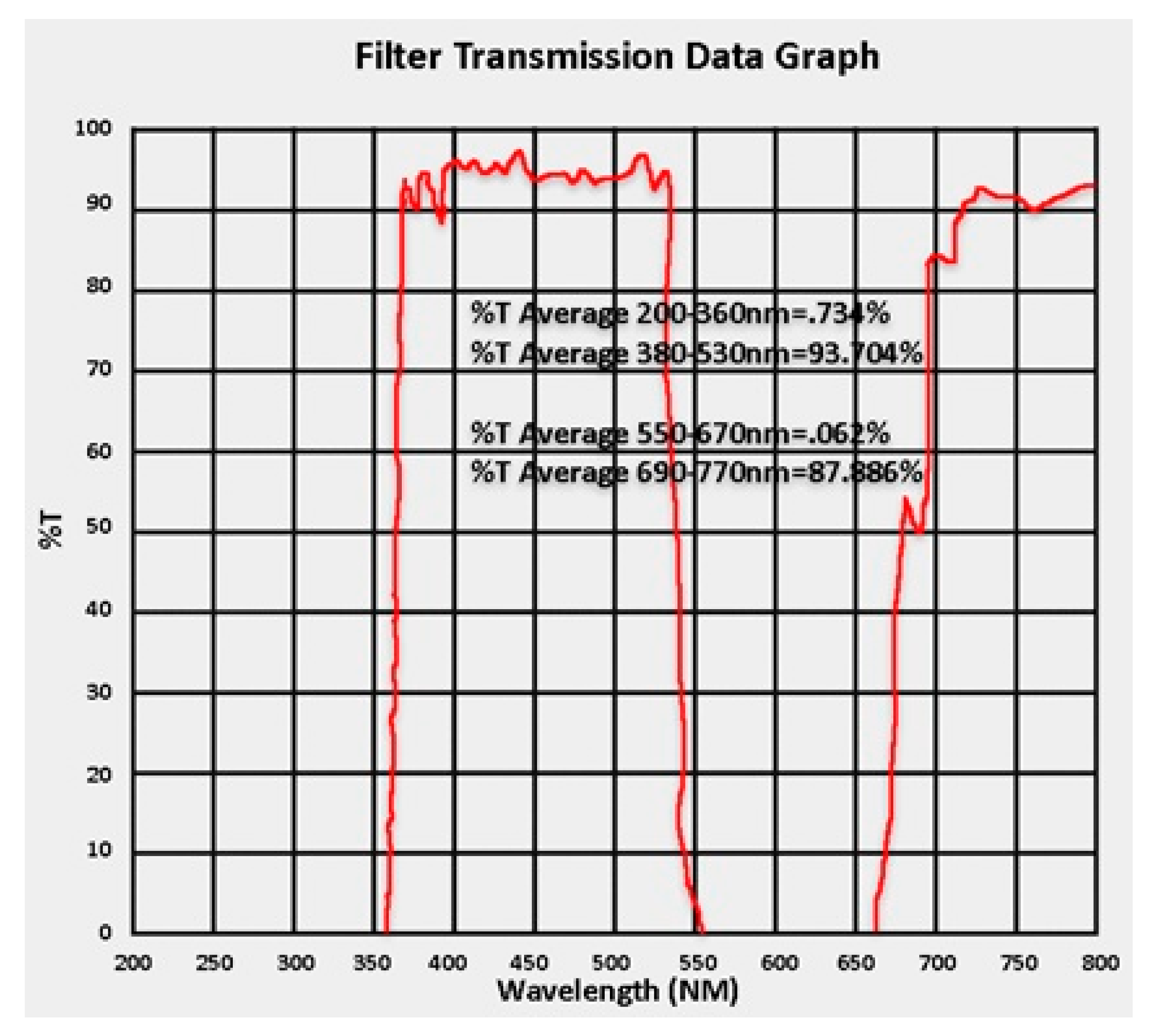
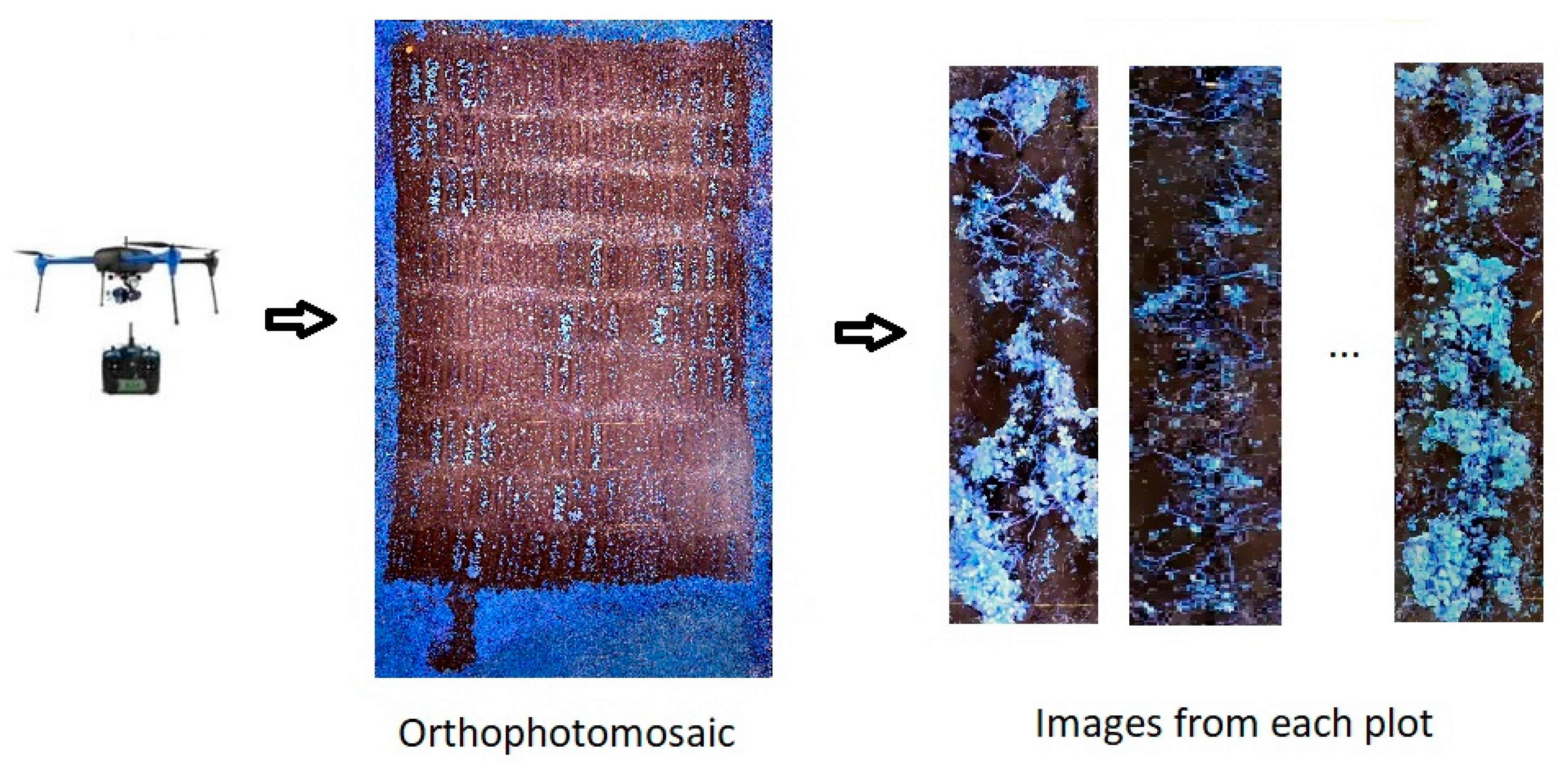
2.2. Image Acquisition and Processing
2.3. Ground Truth
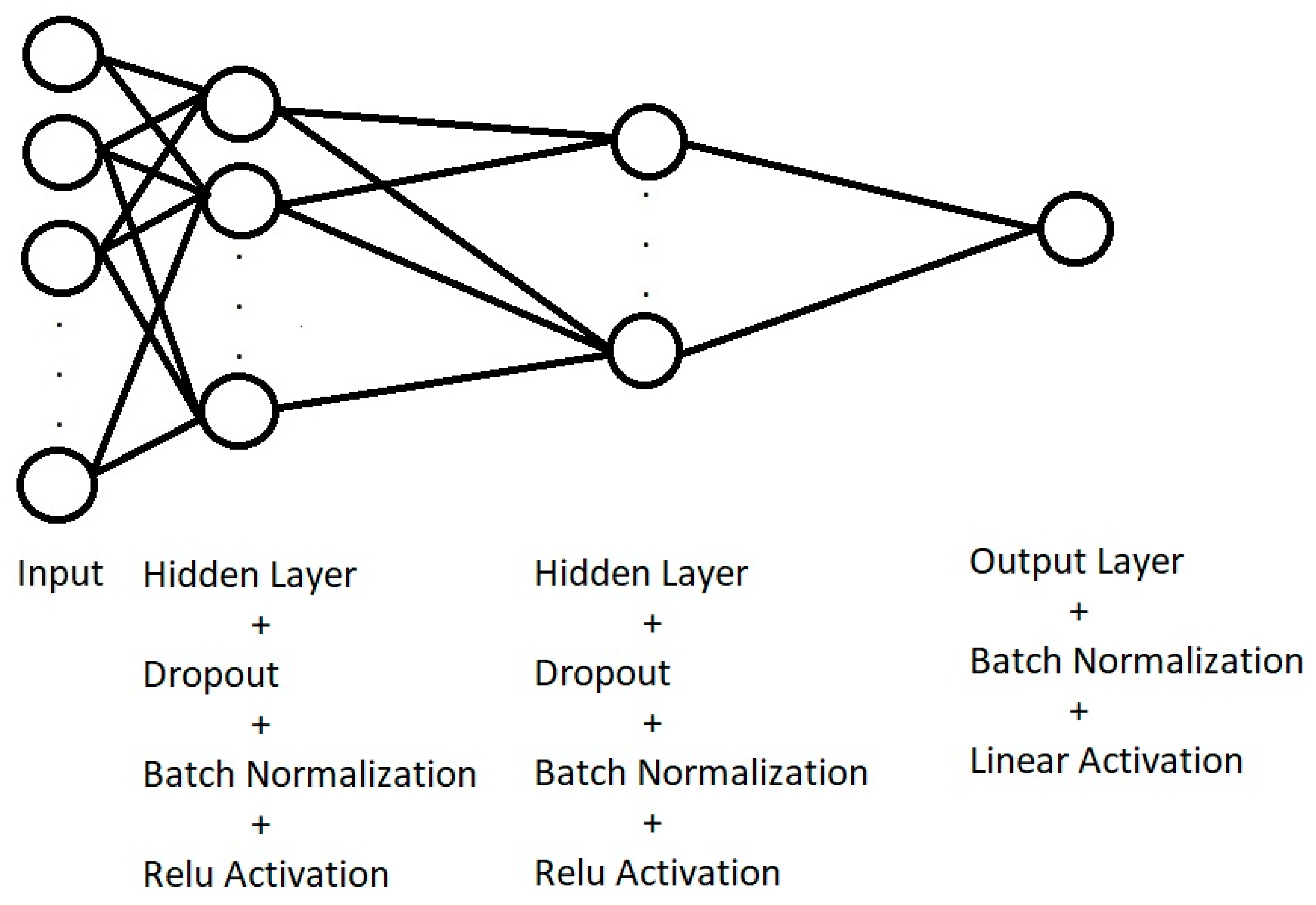
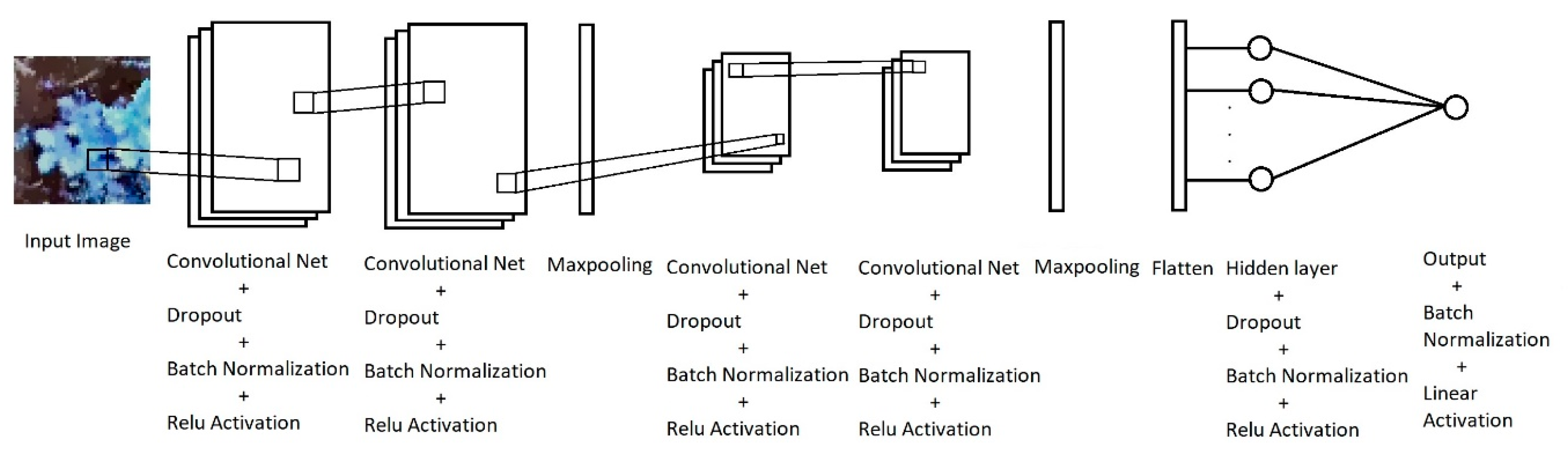
2.4. Machine Learning Algorithms
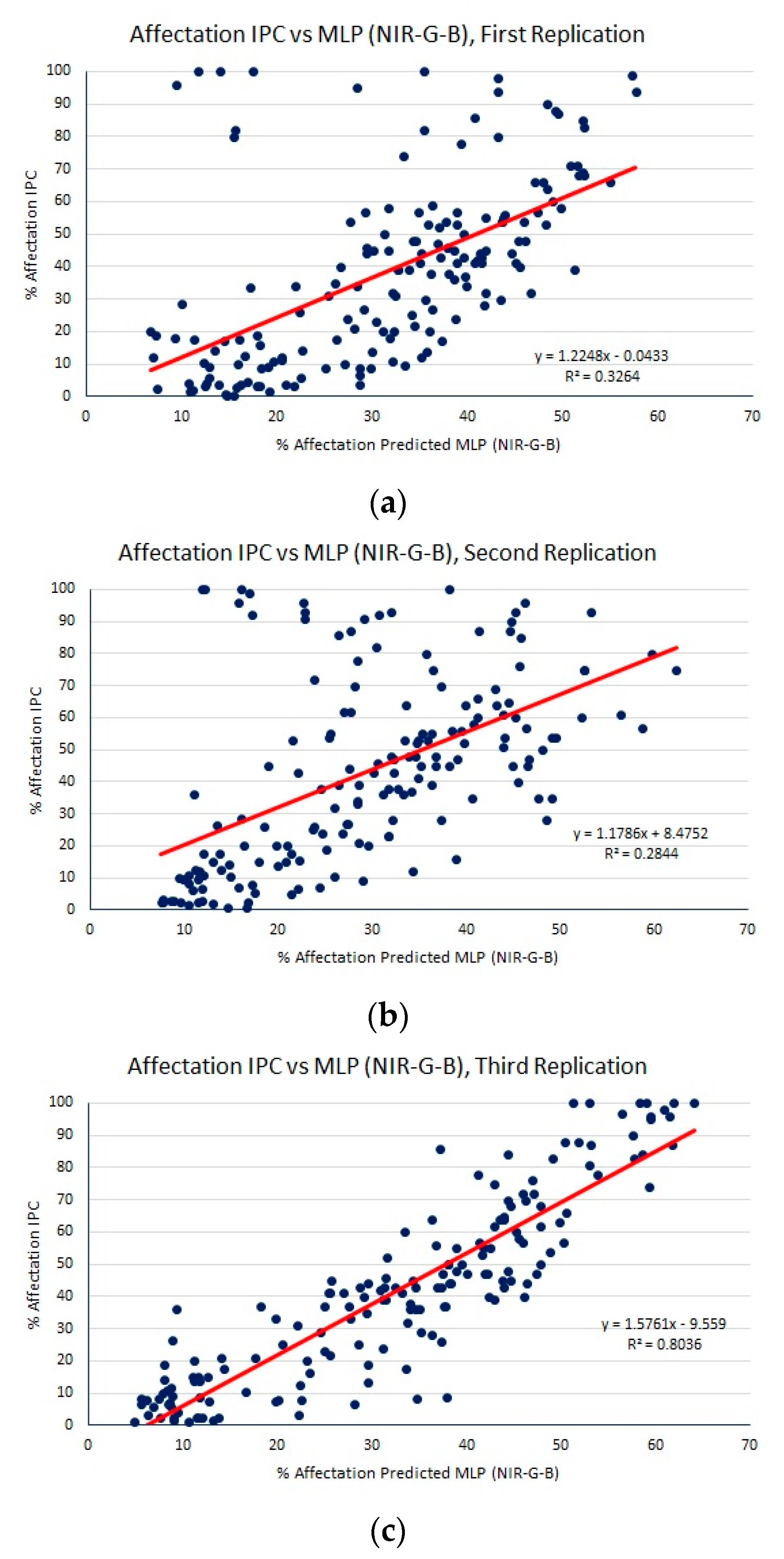
- The spectral differences between green and blue bands and between NIR and green bands. Hence, we obtain a dataset of samples of size 50 × 40 × 2.
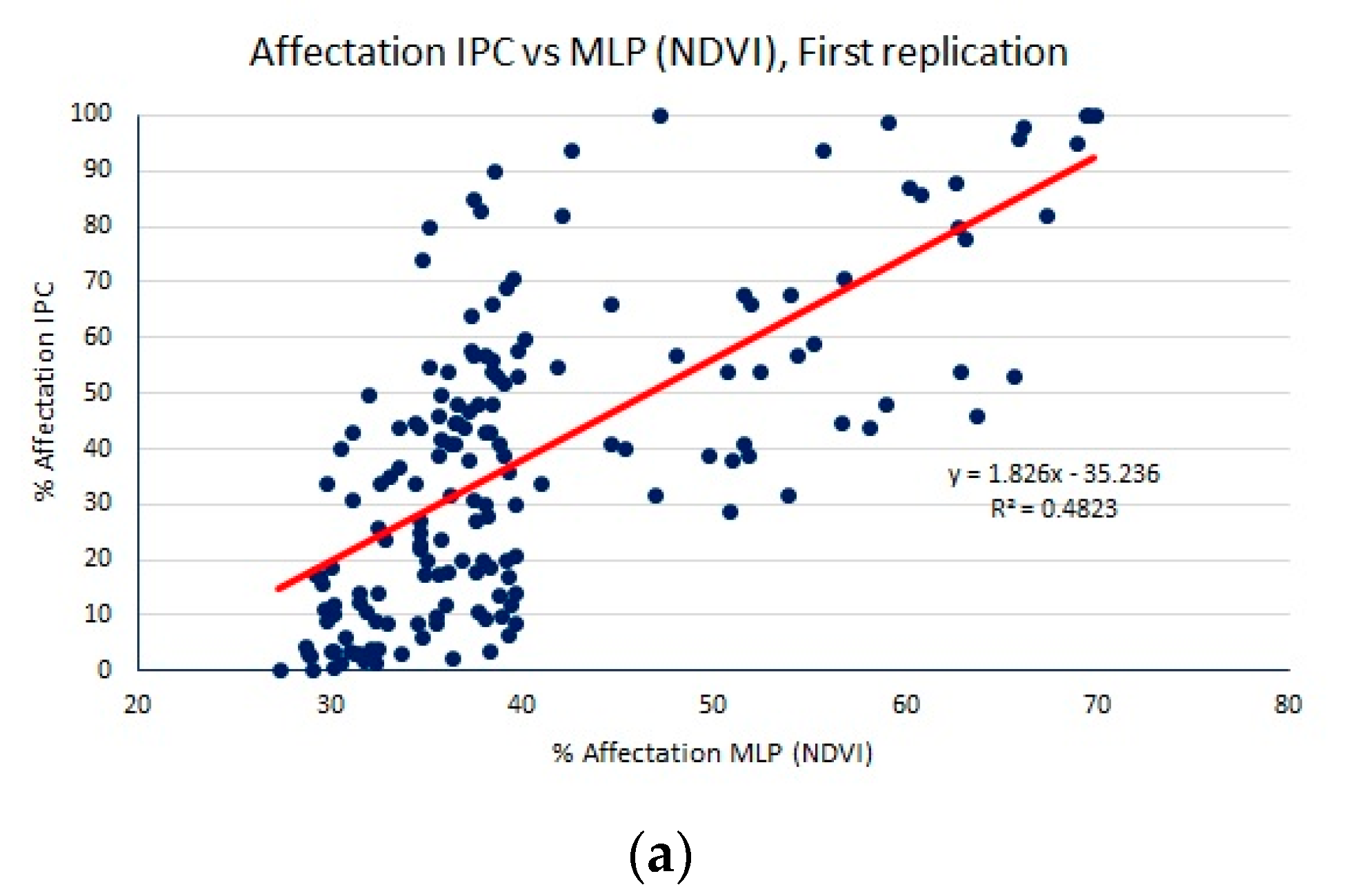
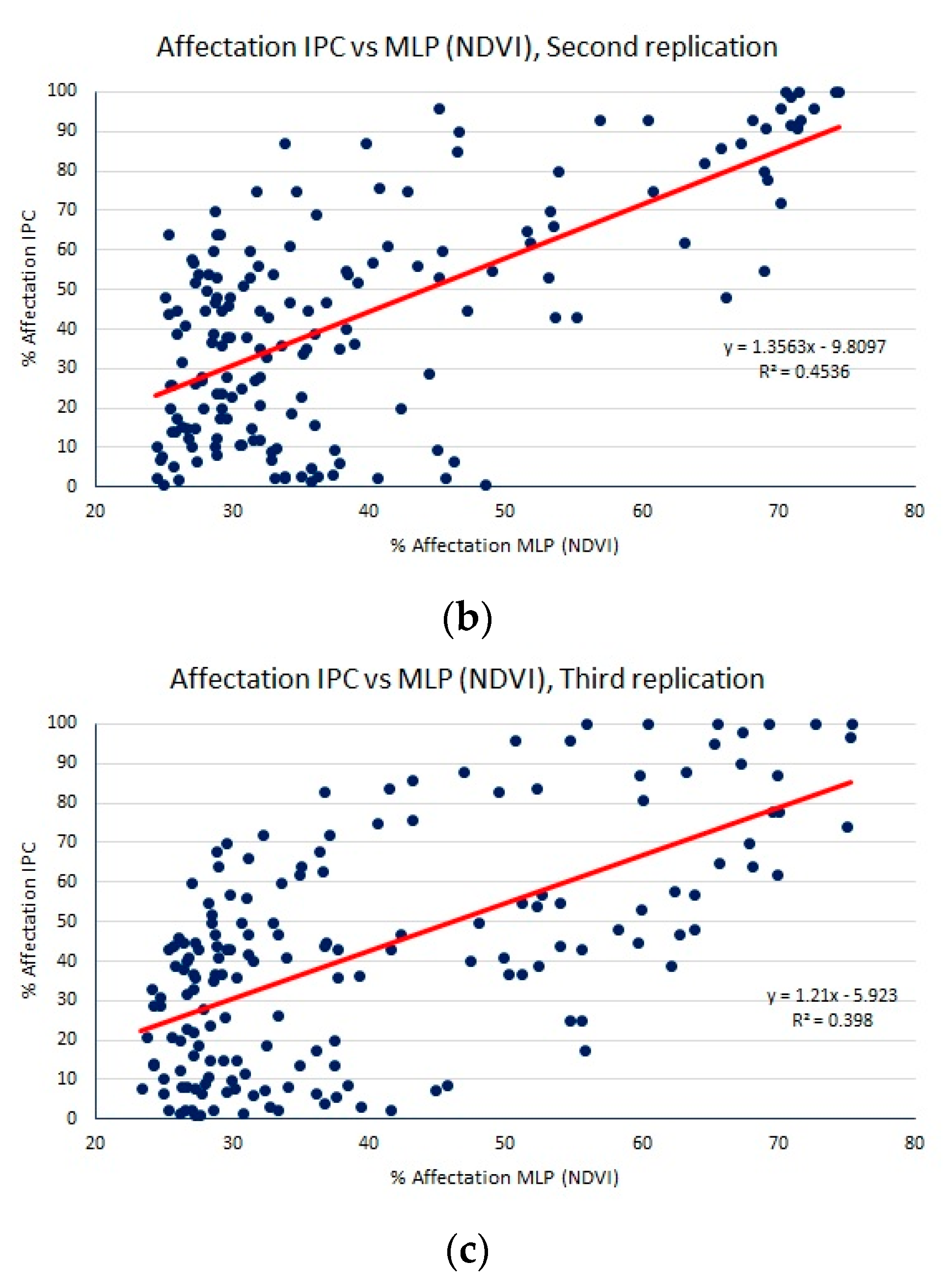
- A normalized difference vegetation index (NDVI) to obtain a dataset of samples of size 50 × 40. Since we do not have separated red and NIR bands, we must use the NIR band together with either the green or blue bands to compute the NDVI. Experimentally, we found better regression performance using NDVI = (NIR − blue)/(NIR + blue).
- The two principal components of each original multispectral plot images were extracted, and the windowing technique explained before can be used to obtain a new dataset consisting of samples of size 50 × 40 × 2. More specifically, if a plot image is of size H × W × 3, where H is the height in pixels, W the width in pixels and we have three channels, the image can be reshaped as a matrix of size P × 3 (P = H × W). Choosing the first 2 principal components, the P × 3 dataset is dimension-reduced to a P × 2 matrix, which can be reshaped as an H × W × 2 dataset, from which overlapping patches of size 50 × 40 × 2 can be extracted.
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hwang, Y.T.; Wijekoon, C.; Kalischuk, M.; Johnson, D.; Howard, R.; Prüfer, D.; Kawchuk, L. Evolution and Management of the Irish Potato Famine Pathogen Phytophthora Infestans in Canada and the United States. Am. J. Potato Res. 2014, 91, 579–593. [Google Scholar] [CrossRef]
- Vargas, A.M.; Ocampo, L.M.Q.; Céspedes, M.C.; Carreño, N.; González, A.; Rojas, A.; Zuluaga, A.P.; Myers, K.; Fry, W.E.; Jiménez, P.; et al. Characterization of Phytophthora infestans populations in Colombia: First report of the A2 mating type. Phytopathology 2009, 99, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Fry, W.E. Phytophthora infestans: New Tools (and Old Ones) Lead to New Understanding and Precision Management. Annu. Rev. Phytopathol. 2016, 54, 529–547. [Google Scholar] [CrossRef] [PubMed]
- European and Mediterranean Plant Protection Organization. Phytophthora infestans on potato. EPPO 2008, 38, 268–271. [Google Scholar] [CrossRef]
- Forbes, G.; Perez, W.; Piedra, J.A. Evaluacion de la Resistencia en Genotipos de Papa a Phytophthora infestans Bajo Condiciones de Campo: Guia Para Colaboradores Internacionales; International Potato Center: Lima, Peru, 2014. [Google Scholar]
- Henfling, J.A. El tizón tardío de la papa: Phytophthora infestans. In Boletin de Informacion Tecnica; Instituto de Censores Jurados de Cuentas de España: Madrid, Spain, 1987; p. 25. [Google Scholar]
- Ray, S.S.; Jain, N.; Arora, R.K.; Chavan, S.; Panigrahy, S. Utility of Hyperspectral Data for Potato Late Blight Disease Detection. J. Indian Soc. Remote Sens. 2011, 39, 161–169. [Google Scholar] [CrossRef]
- Franceschini, M.H.D.; Bartholomeus, H.; van Apeldoorn, D.; Suomalainen, J.; Kooistra, L. Assessing changes in potato canopy caused by late blight in organic production systems through UAV-based pushbroom imaging spectrometer. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2017, 42, 109–112. [Google Scholar] [CrossRef]
- Biswas, S.; Jagyasi, B.; Singh, B.P.; Lal, M. Severity Identification of Potato Late Blight Disease from Crop Images Captured under Uncontrolled Environment. In Proceedings of the 2014 IEEE Canada International Humanitarian Technology Conference—(IHTC), Montreal, QC, Canada, 1–4 June 2014; pp. 1–5. [Google Scholar]
- Sugiura, R.; Tsuda, S.; Tamiya, S.; Itoh, A. ScienceDirect Field phenotyping system for the assessment of potato late blight resistance using RGB imagery from an unmanned aerial vehicle. Biosyst. Eng. 2016, 148, 1–10. [Google Scholar] [CrossRef]
- Liakos, K.; Busato, P.; Moshou, D.; Pearson, S.; Bochtis, D. Machine Learning in Agriculture: A Review. Sensors 2018, 18, 2674. [Google Scholar] [CrossRef] [PubMed]
- Balducci, F.; Impedovo, D.; Pirlo, G. Machine Learning Applications on Agricultural Datasets for Smart Farm Enhancement. Machines 2018, 6, 38. [Google Scholar] [CrossRef]
- Durgabai, R.P.L.; Bhargavi, P. Pest Management using Machine Learning Algorithms: A Review. Int. J. Comput. Sci. Eng. Inf. Technol. Res. 2018, 8, 13–22. [Google Scholar]
- Corrales, D.C. Toward detecting crop diseases and pest by supervised learning. Ing. Univ. 2015, 19, 207–228. [Google Scholar] [CrossRef]
- Revathi, P.; Revathi, R.; Hemalatha, M. Comparative Study of Knowledge in Crop Diseases Using Machine Learning Techniques. Int. J. Comput. Sci. Inf. Technol. 2011, 2, 2180–2182. [Google Scholar]
- Tripathi, M.K.; Maktedar, D.D. Recent machine learning based approaches for disease detection and classification of agricultural products. In Proceedings of the 2016 International Conference on Computing Communication Control and automation (ICCUBEA), Pune, India, 12–13 August 2016; pp. 1–6. [Google Scholar]
- Alves, D.P.; Tomaz, R.S.; Laurindo, B.S.; Laurindo, R.D.F.; Cruz, C.D.; Nick, C.; Silva, D.J.H.D. Artificial neural network for prediction of the area under the disease progress curve of tomato late blight. Sci. Agric. 2017, 74, 51–59. [Google Scholar] [CrossRef]
- Moshou, D.; Bravo, C.; West, J.; Wahlen, S.; McCartney, A.; Ramon, H. Automatic detection of ‘yellow rust’ in wheat using reflectance measurements and neural networks. Comput. Electron. Agric. 2004, 44, 173–188. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, M.; Zhu, J.; Geng, S. Spectral prediction of Phytophthora infestans infection on tomatoes using artificial neural network (ANN). Int. J. Remote Sens. 2008, 29, 1693–1706. [Google Scholar] [CrossRef]
- Wasserman, P.D.; Schwartz, T. Neural networks. II. What are they and why is everybody so interested in them now? IEEE Expert 1988, 3, 10–15. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. Syst. 2012, 1097–1105. [Google Scholar] [CrossRef]
- Firdaus, P.; Arkeman, Y.; Buono, A.; Hermadi, I. Satellite image processing for precision agriculture and agroindustry using convolutional neural network and genetic algorithm. In IOP Earth and Environmental Science; IOP Publishing: Bristol, UK, 2017; p. 7. [Google Scholar]
- Sladojevic, S.; Arsenovic, M.; Anderla, A.; Culibrk, D.; Stefanovic, D. Deep Neural Networks Based Recognition of Plant Diseases by Leaf Image Classification. Comput. Intell. Neurosci. 2016, 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Smola, A.J.; Sch, B.; Schölkopf, B. A Tutorial on Support Vector Regression. Stat. Comput. 2004, 14, 199–222. [Google Scholar] [CrossRef]
- Ho, T.K. Random decision forests. In Proceedings of the IEEE Third International Conference on Document Analysis and Recognition, Montreal, QC, Canada, 14–16 August 1995; pp. 278–282. [Google Scholar]
- O’Shea, K.; Nash, R. An Introduction to Convolutional Neural Networks. 2015. Available online: https://arxiv.org/pdf/1511.08458.pdf (accessed on 7 July 2018).
- Haghighattalab, A.; Pérez, L.G.; Mondal, S.; Singh, D.; Schinstock, D.; Rutkoski, J.; Ortiz-Monasterio, I.; Singh, R.P.; Goodin, D.; Poland, J. Application of unmanned aerial systems for high throughput phenotyping of large wheat breeding nurseries. Plant. Methods 2016, 12, 35. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.M.; Milton, E.J. The use of the empirical line method to calibrate remotely sensed data to reflectance. Int. J. Remote Sens. 1999, 20, 2653–2662. [Google Scholar] [CrossRef]
- Raudys, S.; Jain, A. Small sample size effects in statistical pattern recognition: Recommendations for practitioners. IEEE Trans. Pattern Anal. Mach. Intell. 1991, 13, 252–264. [Google Scholar] [CrossRef]
- Kohavi, R. A Study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection A Study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection. In Proceedings of the 14th International Joint Conference on Artificial Intelligence, Montreal, QC, Canada, 20–25 August 1995; pp. 1137–1143. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference on Learning Representations, Banff, AB, Canada, 14–16 April 2014; pp. 1–15. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. In Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; pp. 81–87. [Google Scholar]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A Simple Way to Prevent Neural Networks from Overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Willmott, C.J.; Matsuura, K. Advantages of the mean absolute error (MAE) over the root mean square error (RMSE) in assessing average model performance. Clim. Res. 2005, 30, 79–82. [Google Scholar] [CrossRef]
- Willmott, C.J.; Matsuura, K.; Robeson, S.M. Ambiguities inherent in sums-of-squares-based error statistics. Atmos. Environ. 2009, 43, 749–752. [Google Scholar] [CrossRef]
- Fry, W.E. Phytophthora infestans: The plant (and R gene) destroyer. Mol. Plant. Pathol. 2008, 9, 385–402. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Alexandersson, E.; Sandin, M.; Resjö, S.; Lenman, M.; Hedley, P.; Levander, F.; Andreasson, E. Quantitative proteomics and transcriptomics of potato in response to Phytophthora infestans in compatible and incompatible interactions. BMC Genom. 2014, 15, 497. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Chu, B.; Zhang, C.; Liu, F.; Jiang, L.; He, Y. Hyperspectral Imaging for Presymptomatic Detection of Tobacco Disease with Successive Projections Algorithm and Machine-learning Classifiers. Sci. Rep. 2017, 7, 4125. [Google Scholar] [CrossRef] [PubMed]
- Majeed, A.; Muhammad, Z.; Ullah, Z.; Ullah, R.; Ahmad, H. Late Blight of Potato (Phytophthora infestans) I: Fungicides Application and Associated Challenges. Turk. J. Agric. Food Sci. Technol. 2017, 5, 261–266. [Google Scholar] [CrossRef]
- Van Evert, F.K.; Fountas, S.; Jakovetic, D.; Crnojevic, V.; Travlos, I.; Kempenaar, C. Big Data for weed control and crop protection. Weed Res. 2017, 57, 218–233. [Google Scholar] [CrossRef]
- Szegedy, C. Deep Neural Networks for Object Detection. In Advances in Neural Information Processing Systems; The MIT Press: Cambridge, MA, USA, 2013; pp. 2553–2561. [Google Scholar]
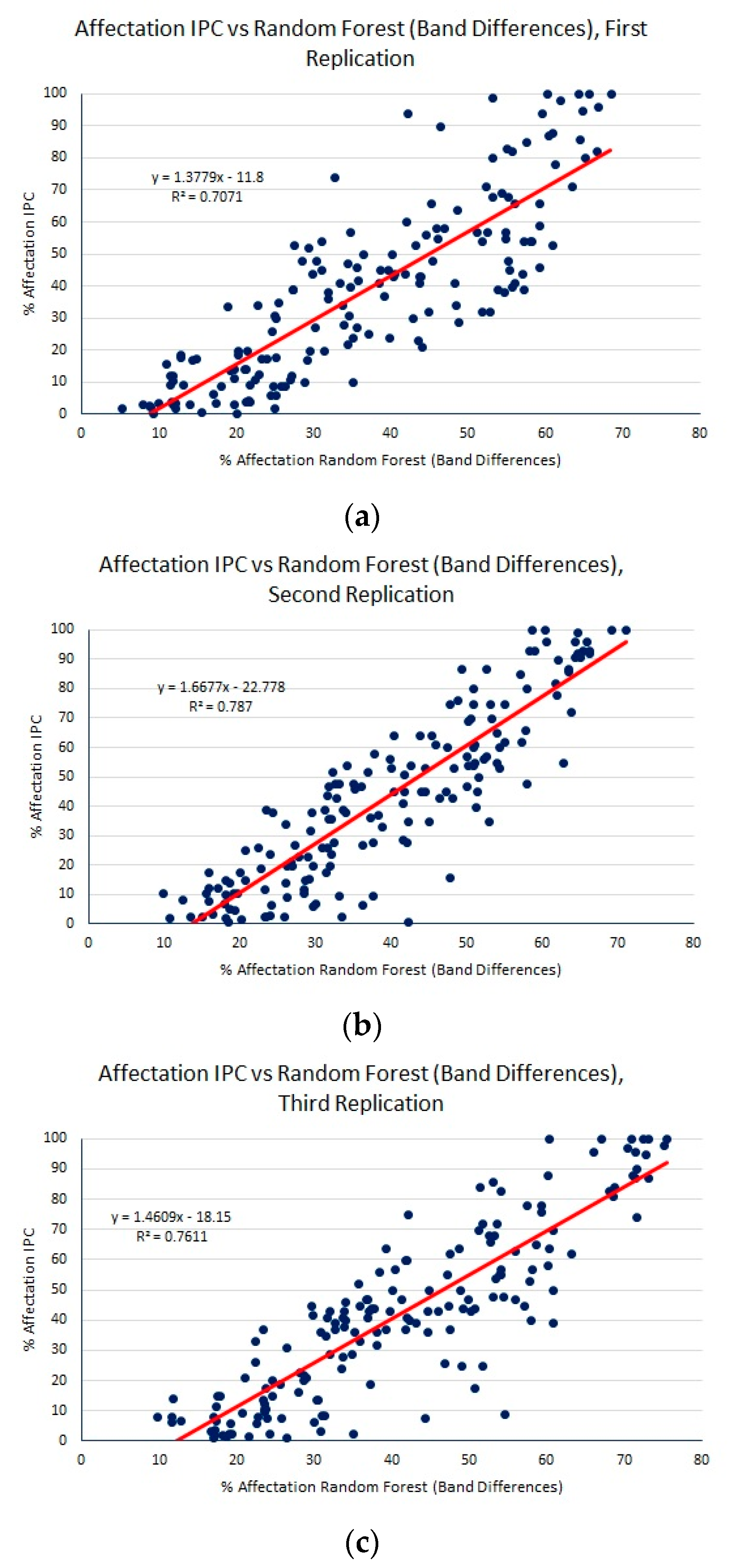
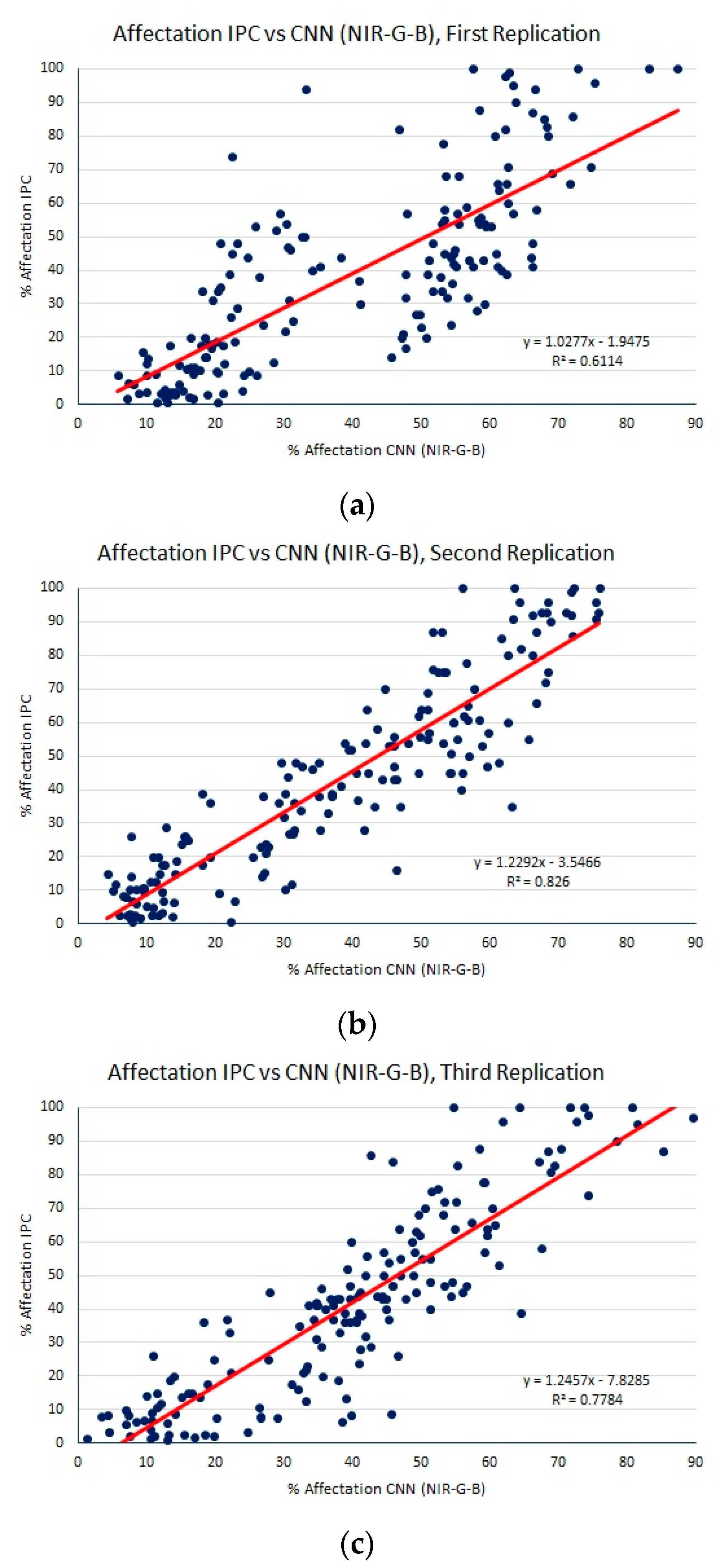
| Regression Method | MAE | RMSE | R2 |
|---|---|---|---|
| MLP (NIR-G-B) | 16.37 (1.55) | 23.25 (2.90) | 0.47 (0.17) |
| MLP (NDVI) | 18.71 (0.11) | 21.98 (0.20) | 0.44 (0.02) |
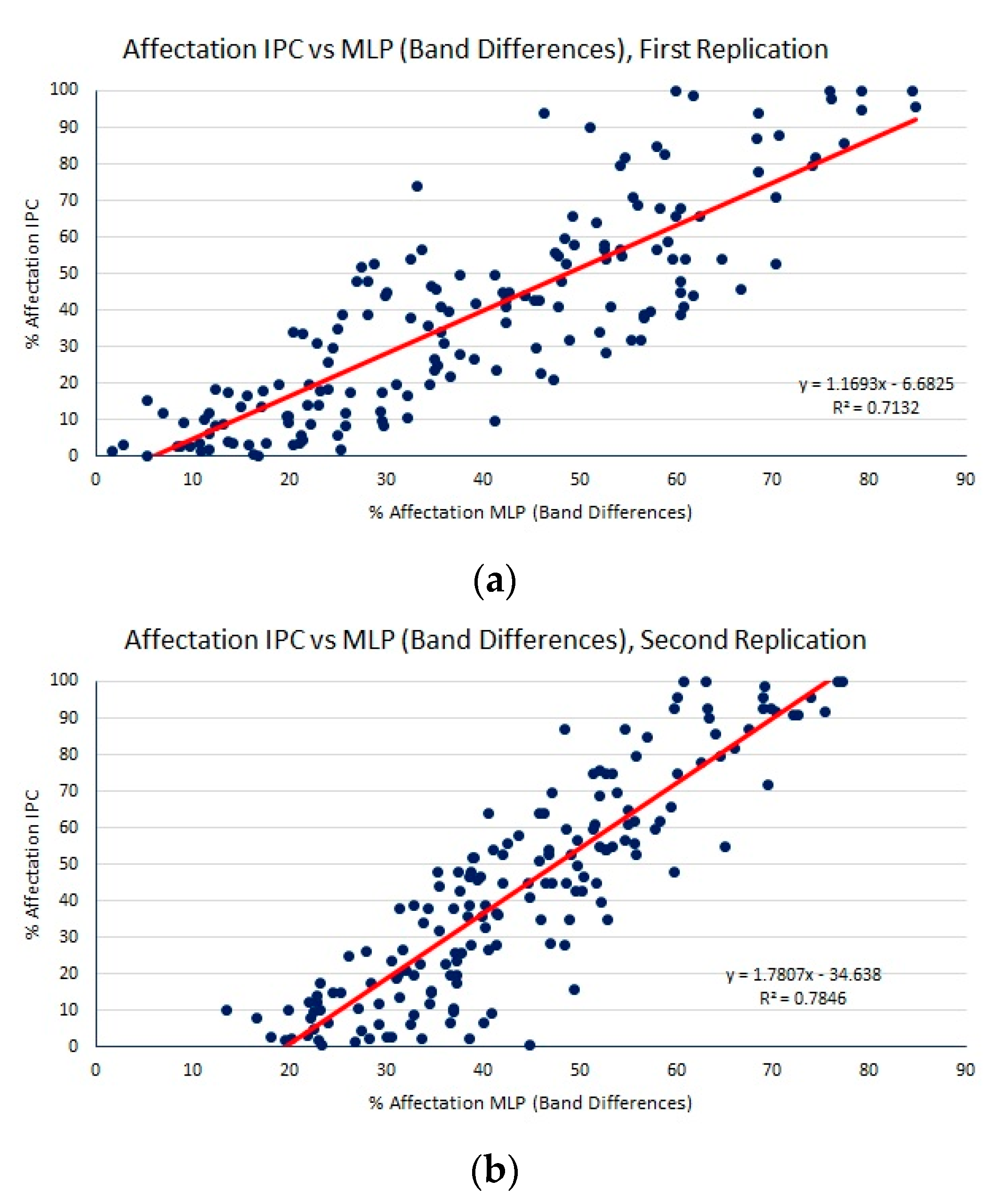
| MLP (band differences) | 13.23 (0.70) | 16.28 (0.70) | 0.75 (0.02) |
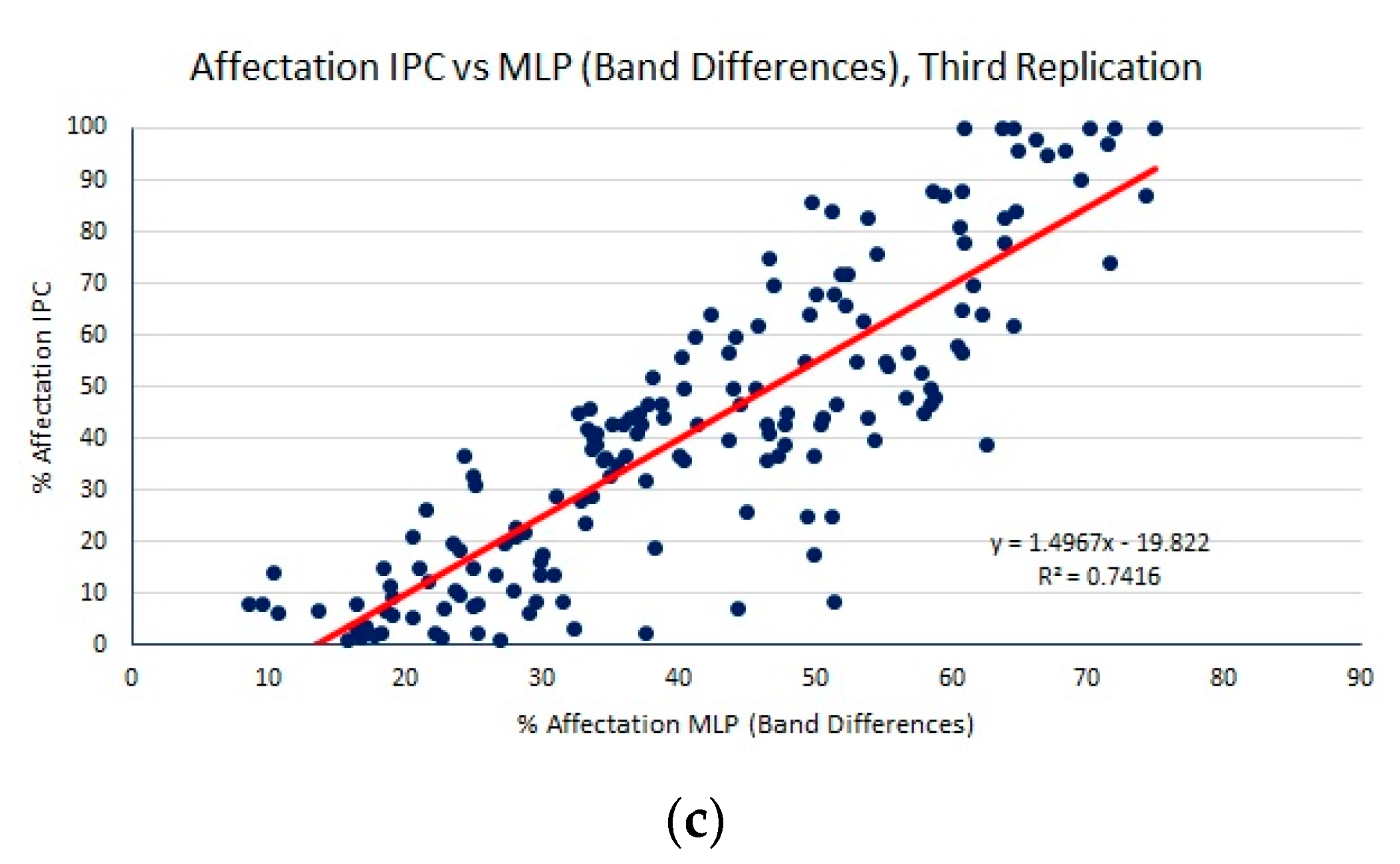
| MLP (PCA) | 16.60 (0.81) | 21.87 (1.44) | 0.48 (0.08) |
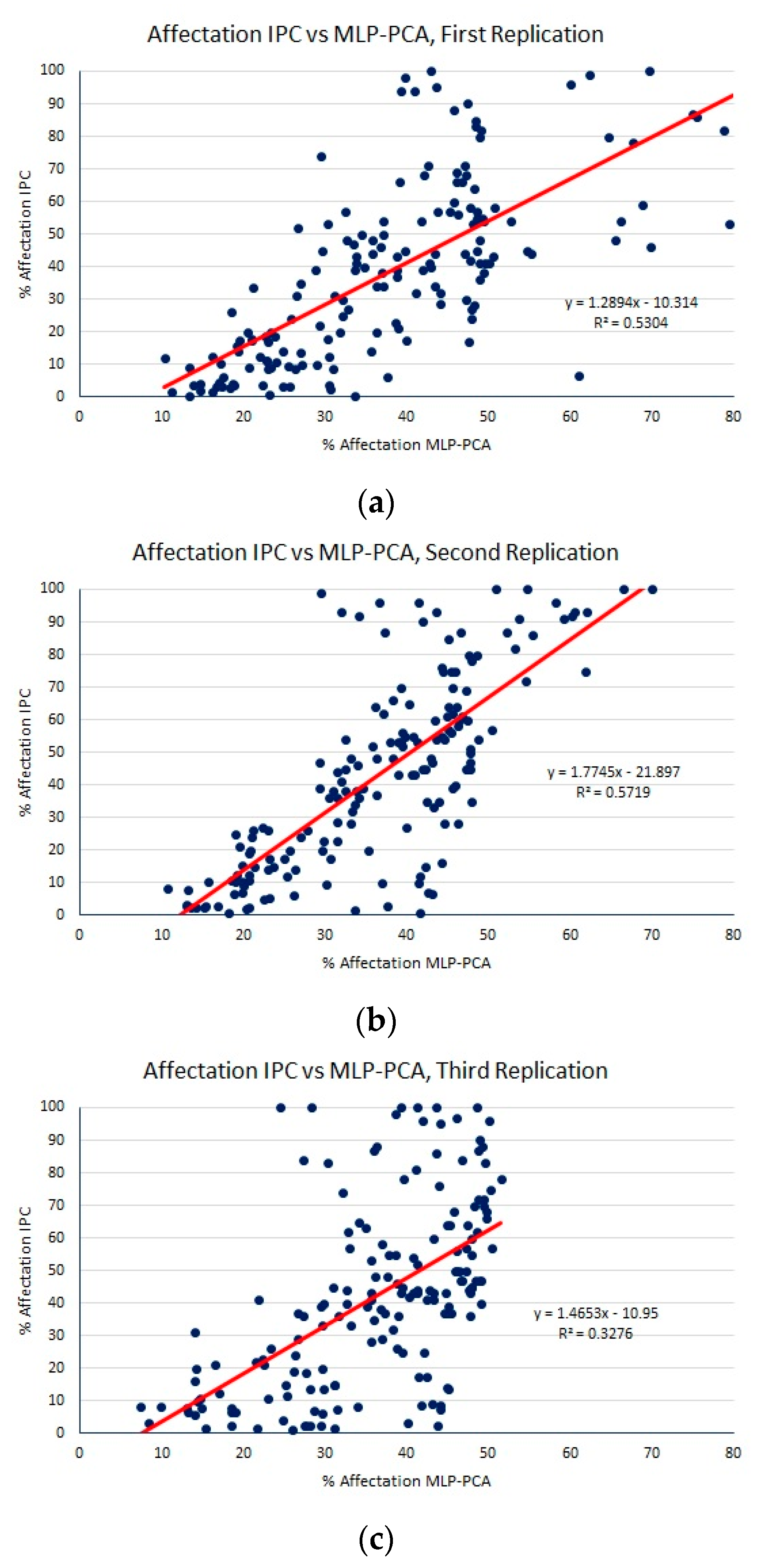
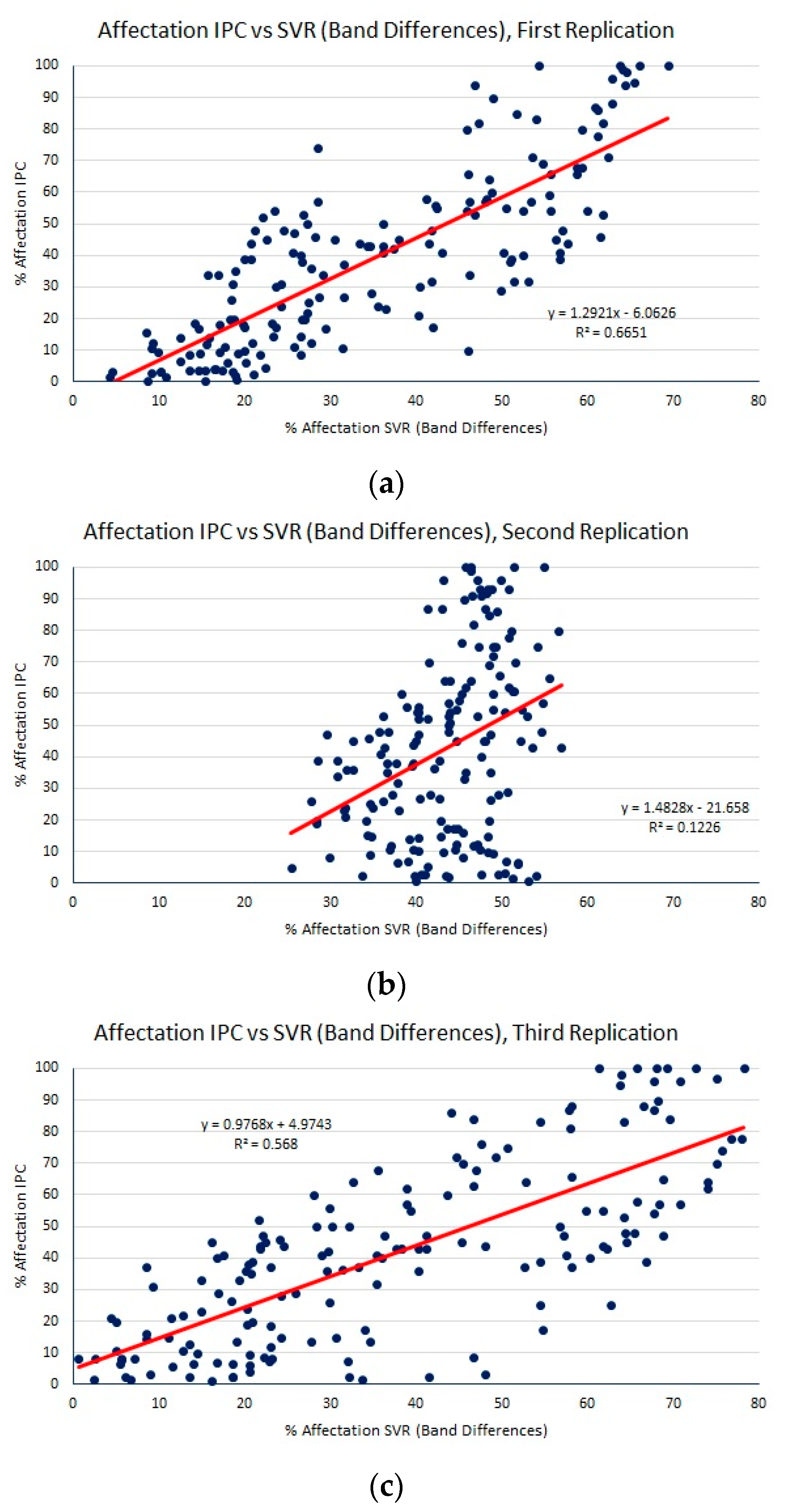
| SVR (band differences) | 17.34 (2.61) | 21.06 (3.09) | 0.45 (0.17) |
| RFs (band differences) | 12.96 (0.07) | 16.15 (0.07) | 0.75 (0.02) |
| CNNs (NIR-G-B) | 11.72 (0.92) | 15.09 (1.01) | 0.74 (0.07) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duarte-Carvajalino, J.M.; Alzate, D.F.; Ramirez, A.A.; Santa-Sepulveda, J.D.; Fajardo-Rojas, A.E.; Soto-Suárez, M. Evaluating Late Blight Severity in Potato Crops Using Unmanned Aerial Vehicles and Machine Learning Algorithms. Remote Sens. 2018, 10, 1513. https://doi.org/10.3390/rs10101513
Duarte-Carvajalino JM, Alzate DF, Ramirez AA, Santa-Sepulveda JD, Fajardo-Rojas AE, Soto-Suárez M. Evaluating Late Blight Severity in Potato Crops Using Unmanned Aerial Vehicles and Machine Learning Algorithms. Remote Sensing. 2018; 10(10):1513. https://doi.org/10.3390/rs10101513
Chicago/Turabian StyleDuarte-Carvajalino, Julio M., Diego F. Alzate, Andrés A. Ramirez, Juan D. Santa-Sepulveda, Alexandra E. Fajardo-Rojas, and Mauricio Soto-Suárez. 2018. "Evaluating Late Blight Severity in Potato Crops Using Unmanned Aerial Vehicles and Machine Learning Algorithms" Remote Sensing 10, no. 10: 1513. https://doi.org/10.3390/rs10101513
APA StyleDuarte-Carvajalino, J. M., Alzate, D. F., Ramirez, A. A., Santa-Sepulveda, J. D., Fajardo-Rojas, A. E., & Soto-Suárez, M. (2018). Evaluating Late Blight Severity in Potato Crops Using Unmanned Aerial Vehicles and Machine Learning Algorithms. Remote Sensing, 10(10), 1513. https://doi.org/10.3390/rs10101513