Object-Based Mangrove Species Classification Using Unmanned Aerial Vehicle Hyperspectral Images and Digital Surface Models
Abstract
:1. Introduction
2. Study Area and Data
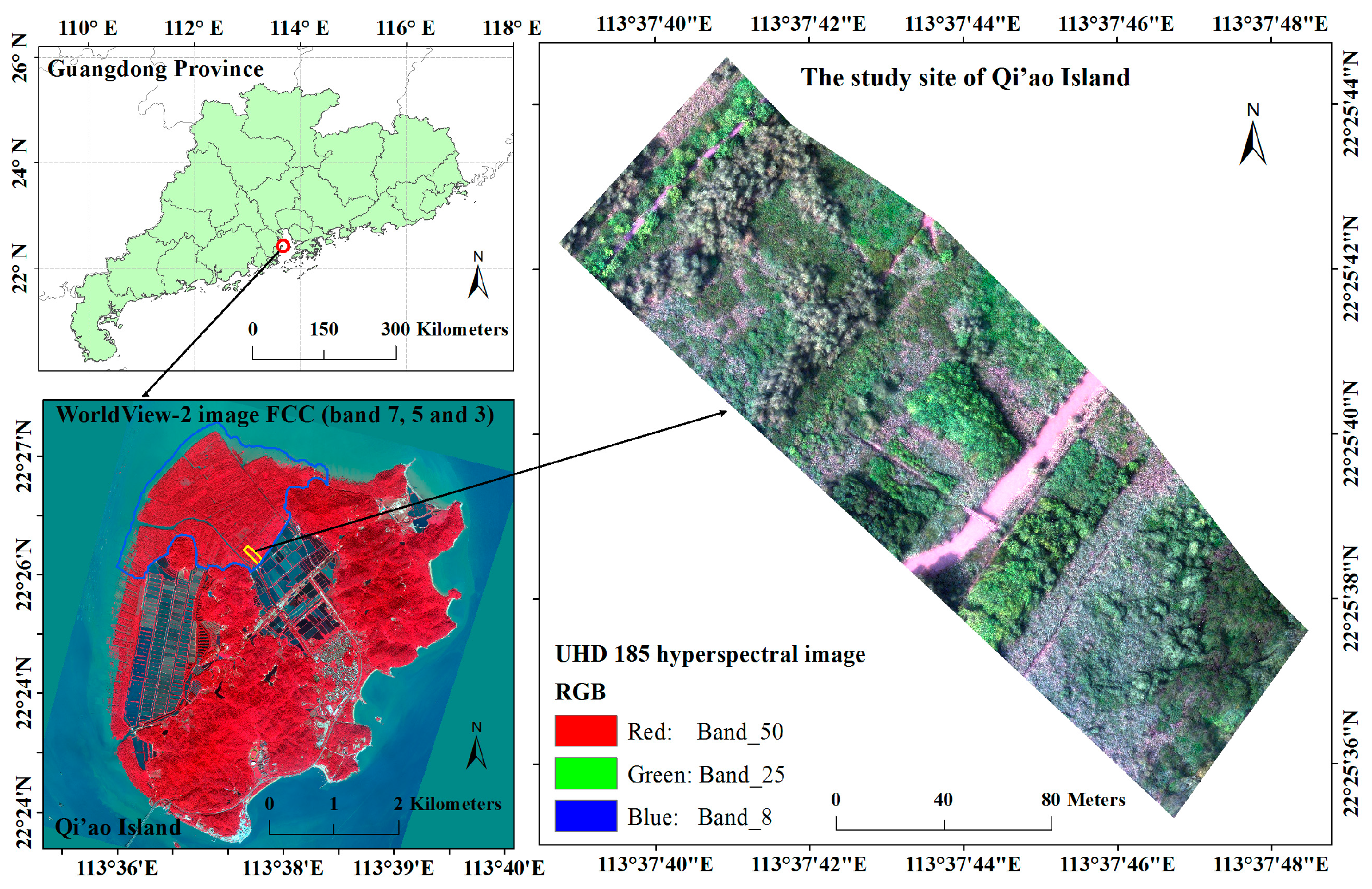
2.1. Study Area
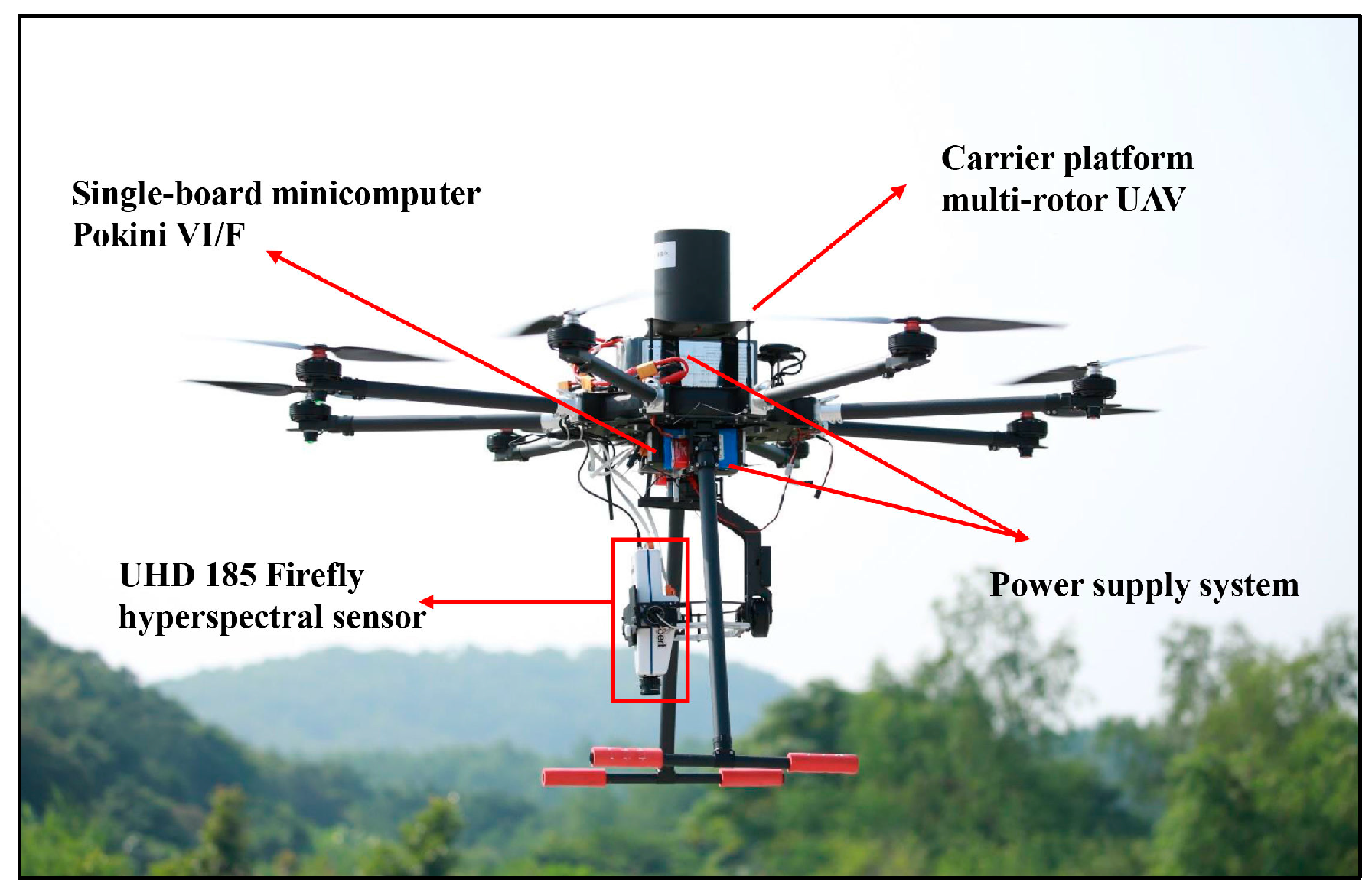
2.2. UAV Hyperspectral Data Acquisition
2.3. Image Preprocessing
2.4. Field Surveys and Sample Collection
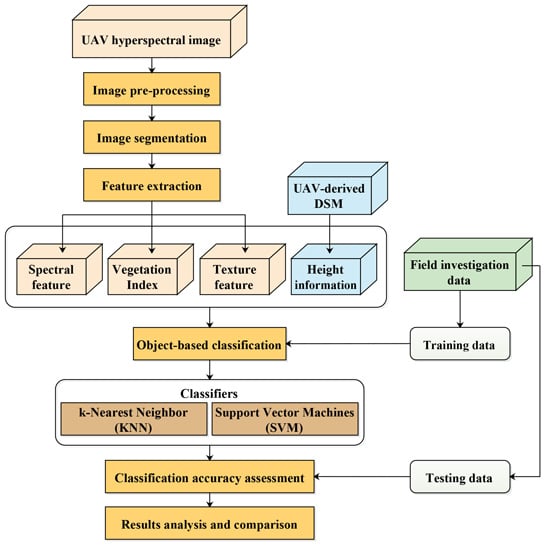
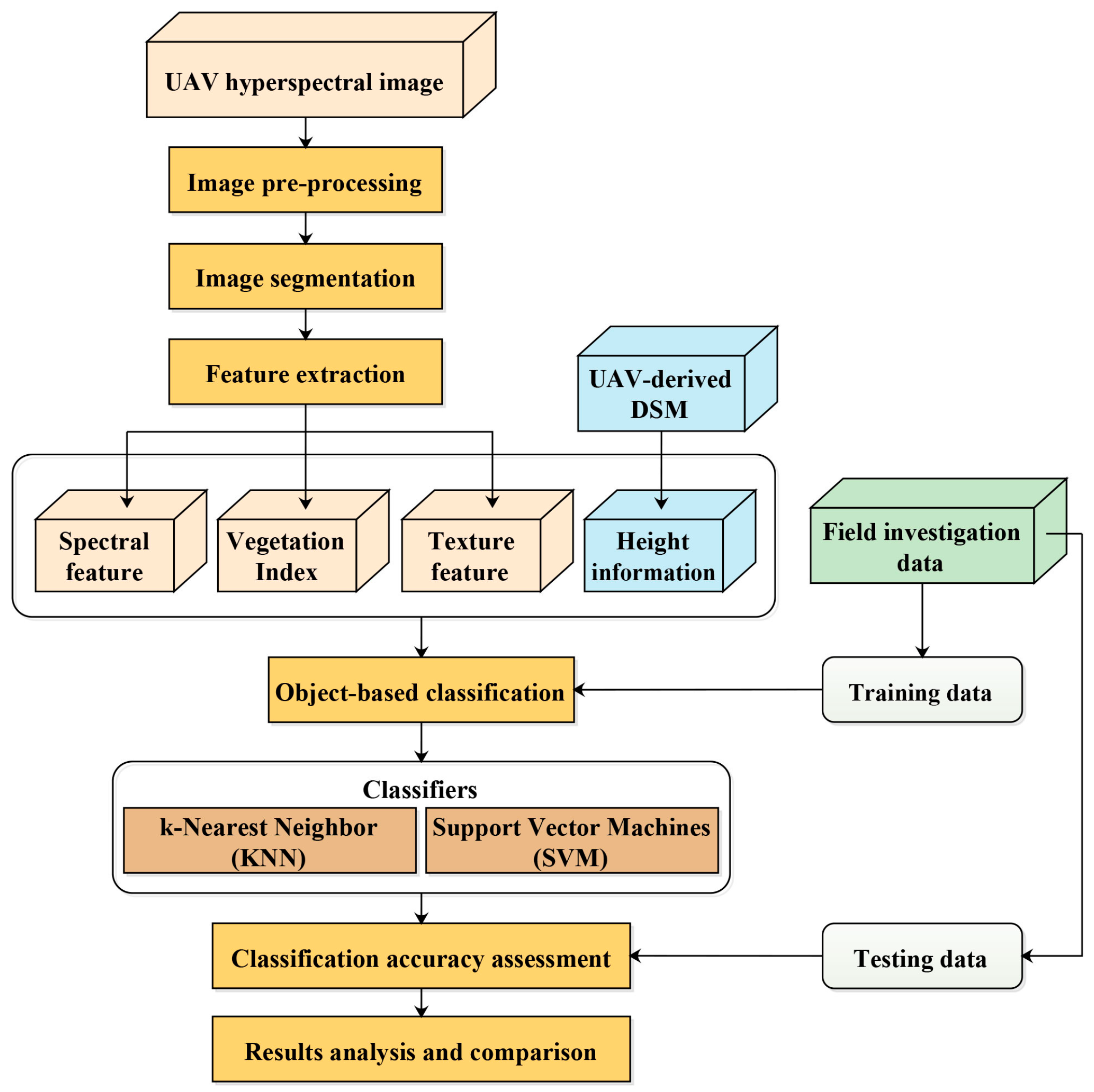
3. Methods
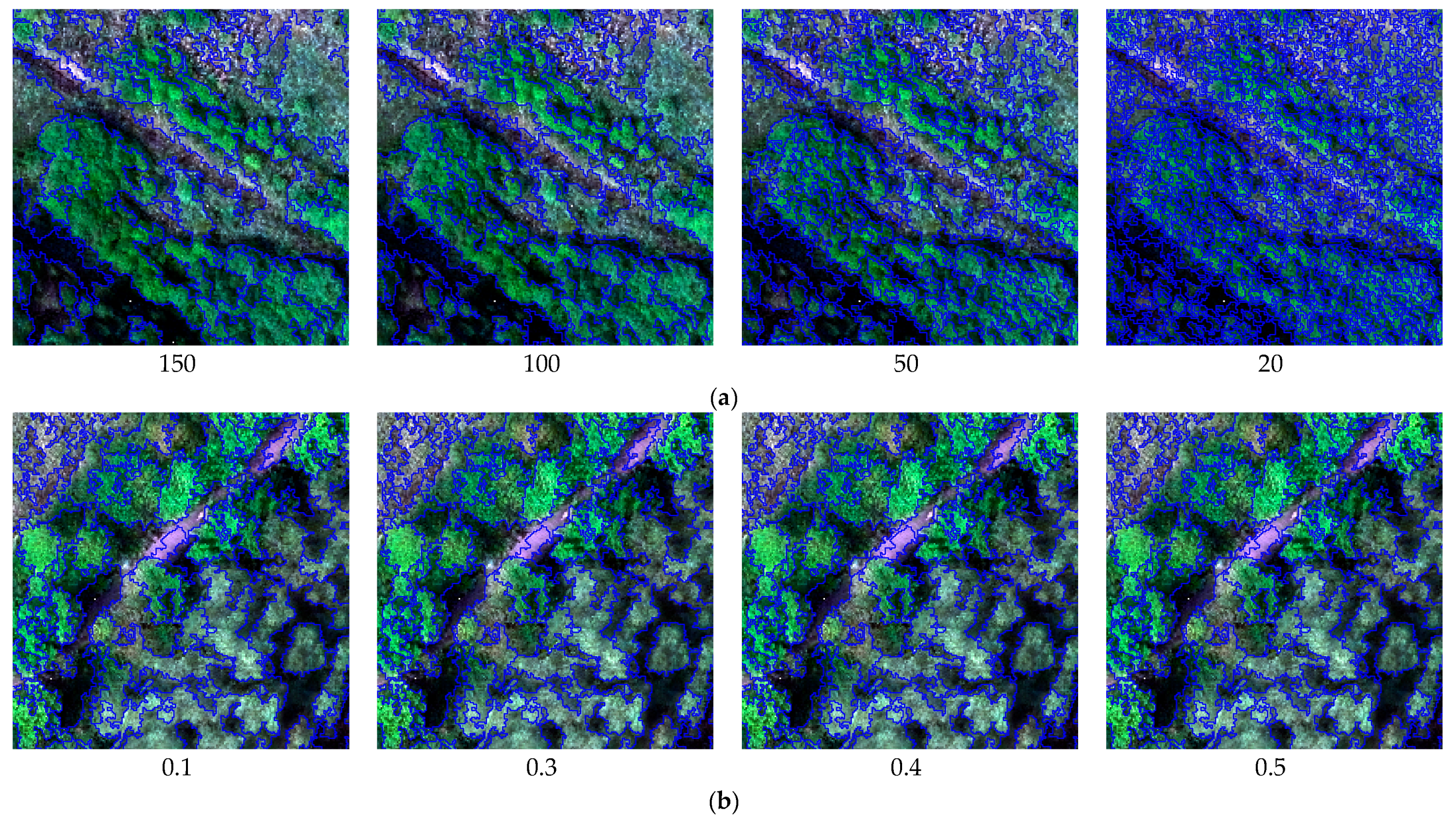
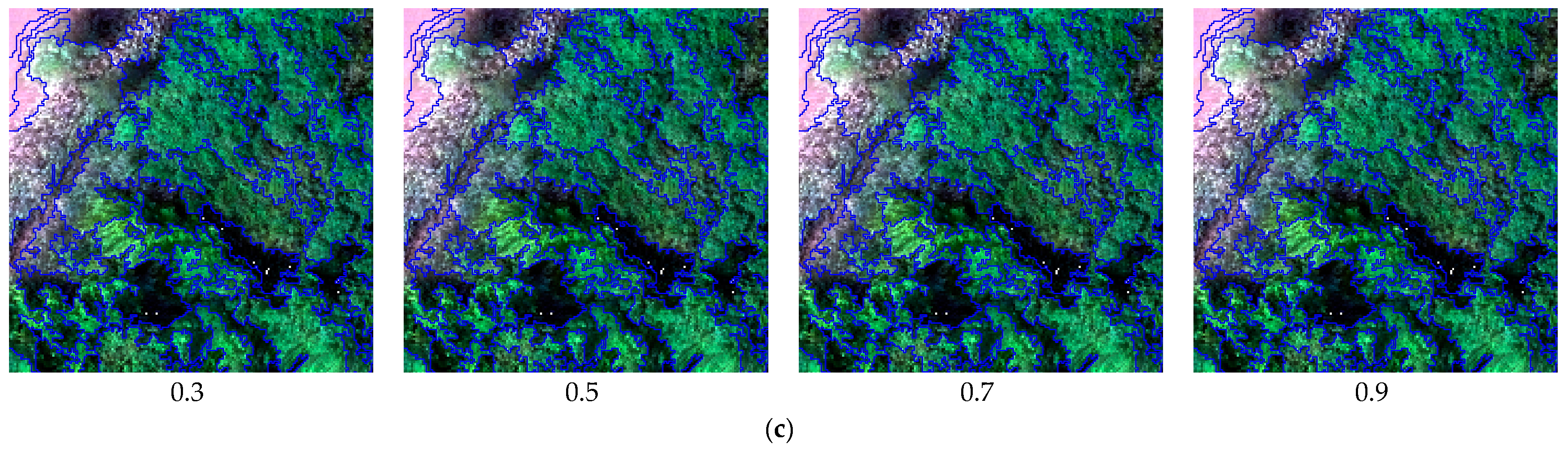
3.1. Image Segmentation
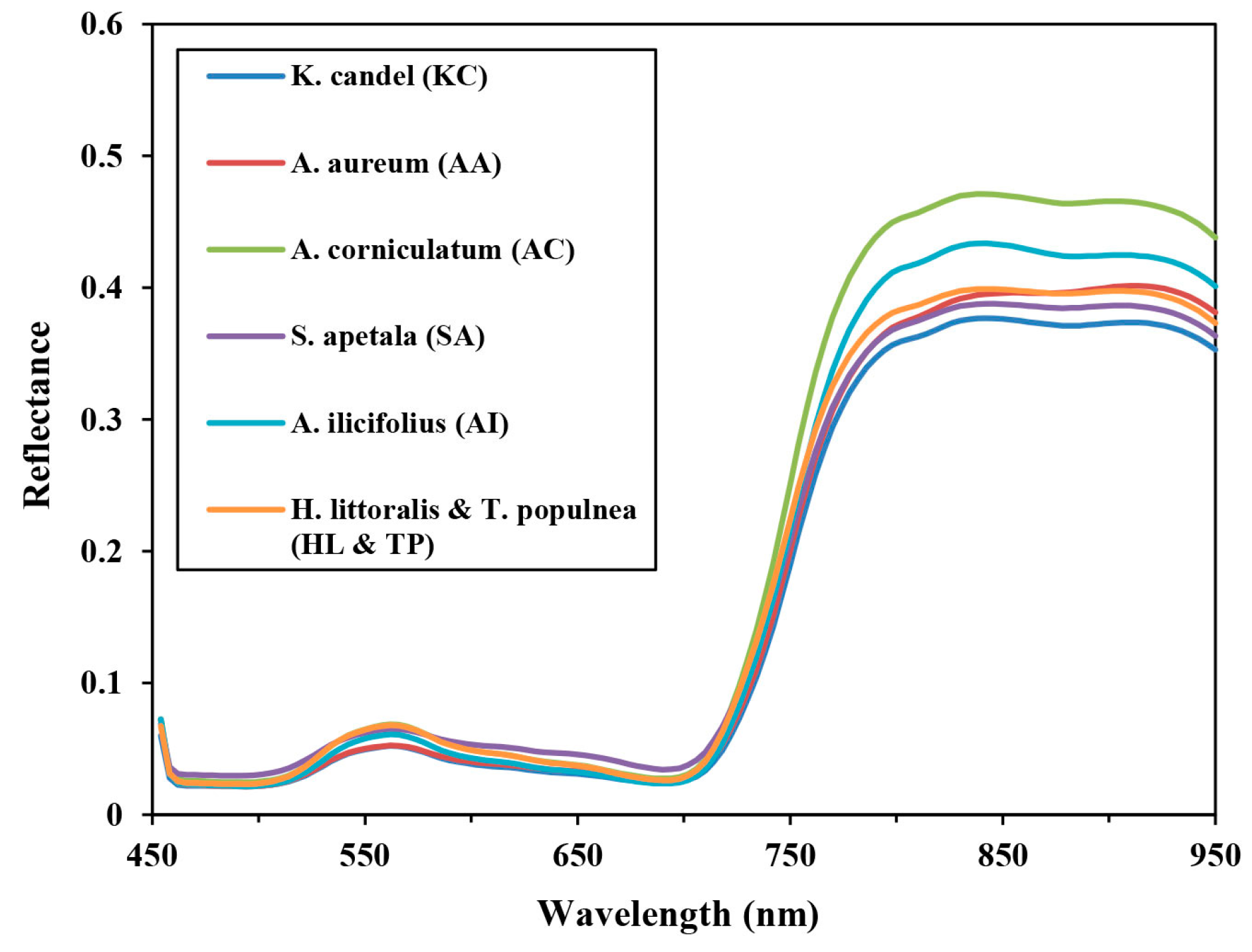
3.2. Feature Extraction and Selection
3.3. Object-Based Classification
3.3.1. KNN
3.3.2. SVM
3.4. Classification Accuracy Assessment
4. Results and Discussion
4.1. Analysis of Image Segmentation Results
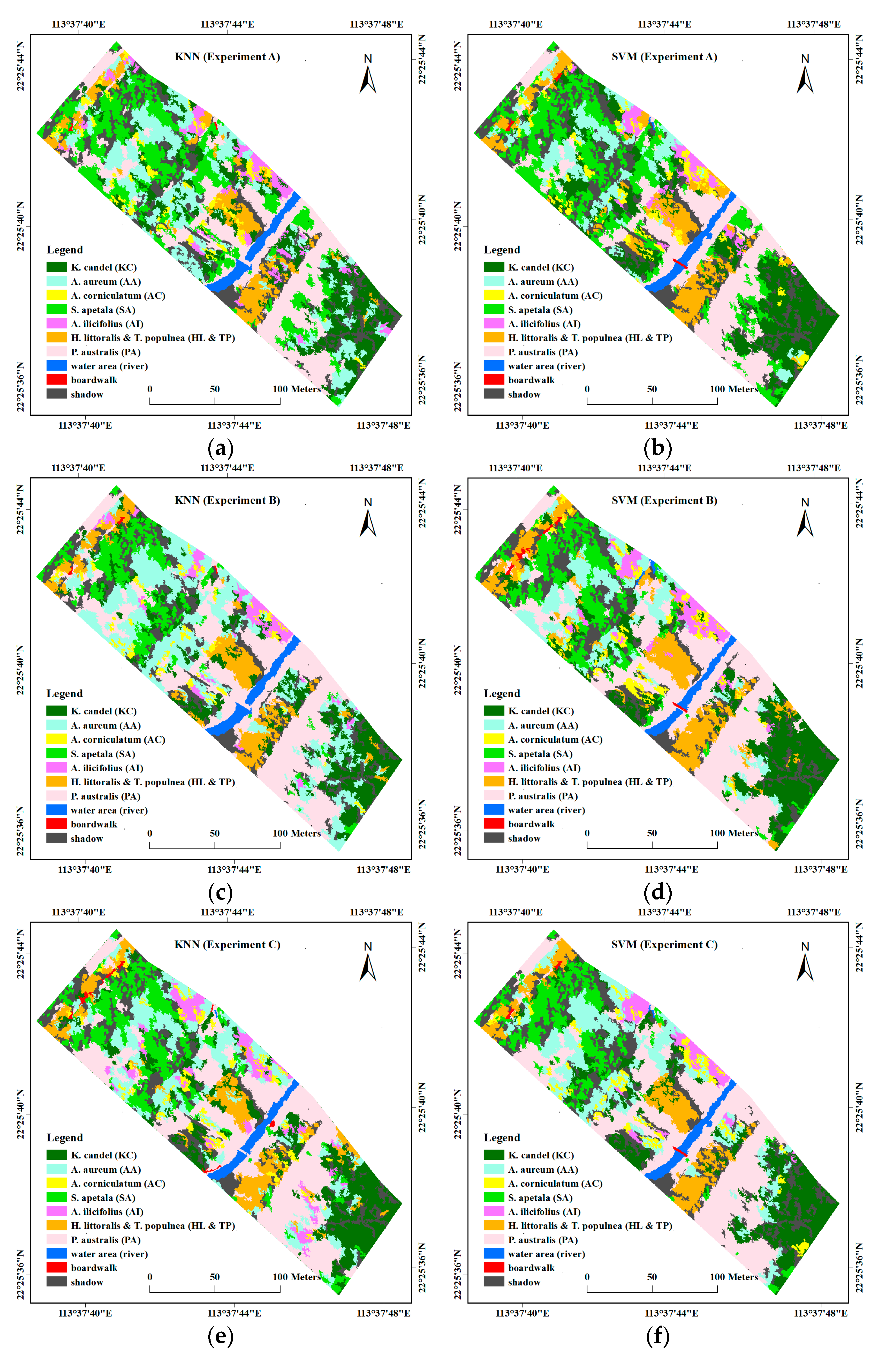
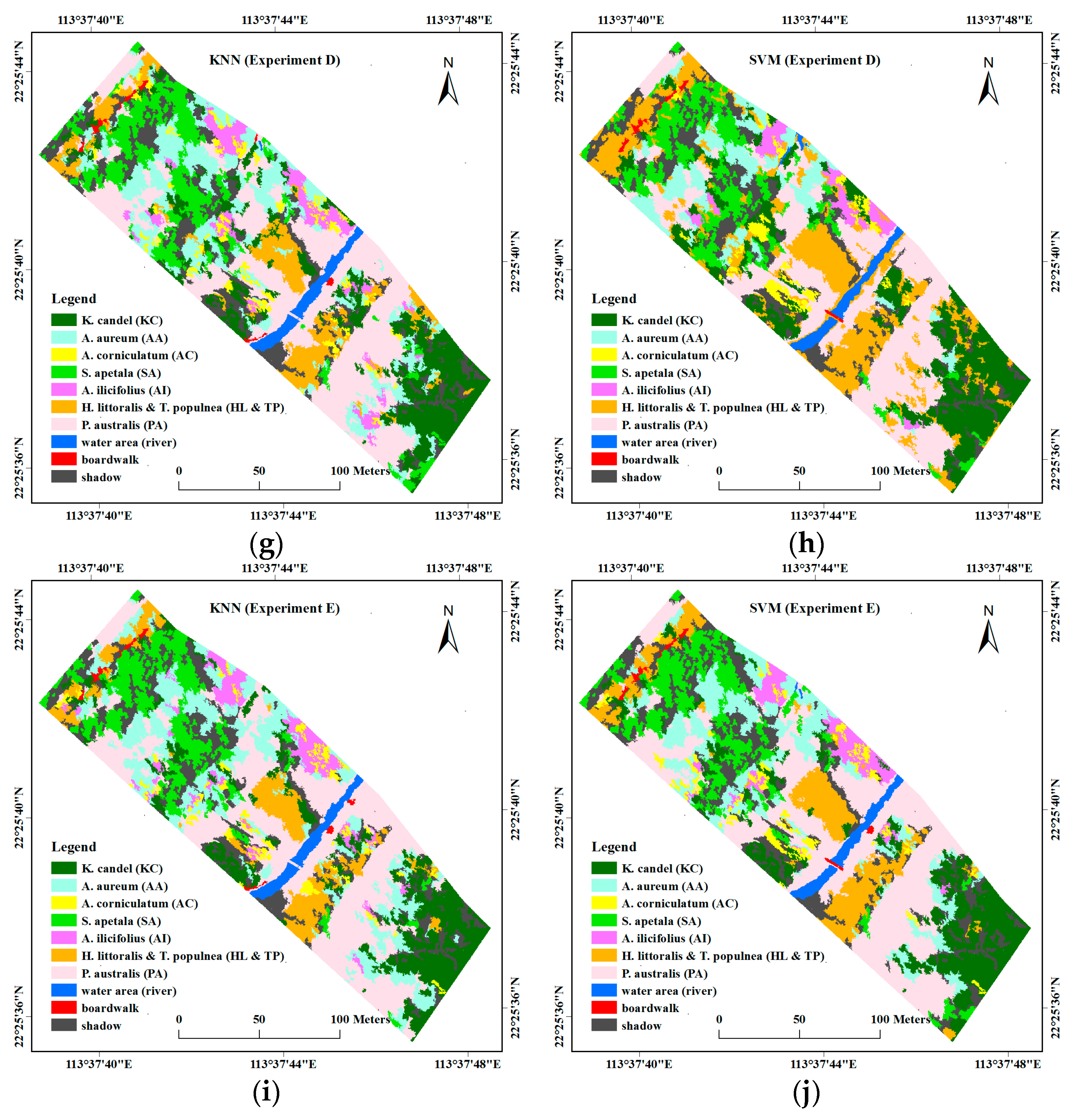
4.2. Comparison of Object-Based Classification Results
- Experiment A: spectral features, using 32 selected spectral bands (mean values) selected from the CART method, including band 1–2, band 8–10, band 14, band 17–19, band 23–24, band 26, band 28–29, band 48, band 52, band 56–57, band 62–64, band 68–70, band 72, band 75, band 79–80, band 82–83, band 91, and band 107, brightness and max.diff.
- Experiment B: stacking spectral features in Experiment A, hyperspectral VIs, and textural features.
- Experiment C: stacking spectral features in Experiment A and height information.
- Experiment D: stacking all the features together, including spectral features in Experiment A, hyperspectral VIs, textural features, and height information.
- Experiment E: 14 features selected from Experiment D using CFS, including four spectral bands, i.e., band 10, band 23, band 62 and band 91, four hyperspectral VIs, i.e., NDVI, TCARI, MCARI2, and PRI, five textural features, i.e., ASM (band 50), COR (band 8 and 25), MEAN (band 8), and StdDev (band 8), and UAV-derived DSM.
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Peng, L. A review on the mangrove research in China. J. Xiamen Univ. Nat. Sci. 2001, 40, 592–603. [Google Scholar]
- Bahuguna, A.; Nayak, S.; Roy, D. Impact of the tsunami and earthquake of 26th December 2004 on the vital coastal ecosystems of the Andaman and Nicobar Islands assessed using RESOURCESAT AWIFS data. Int. J. Appl. Earth Obs. Geoinf. 2008, 10, 229–237. [Google Scholar] [CrossRef]
- Food and Agriculture Organization (FAO). The World’s Mangroves 1980–2005; FAO: Rome, Italy, 2007. [Google Scholar]
- Li, W.; Niu, Z.; Chen, H.; Li, D.; Wu, M.; Zhao, W. Remote estimation of canopy height and aboveground biomass of maize using high-resolution stereo images from a low-cost unmanned aerial vehicle system. Ecol. Indic. 2016, 67, 637–648. [Google Scholar] [CrossRef]
- Giri, C.; Ochieng, E.; Tieszen, L.L.; Zhu, Z.; Singh, A.; Loveland, T.; Masek, J.; Duke, N. Status and distribution of mangrove forests of the world using earth observation satellite data. Glob. Ecol. Biogeogr. 2011, 20, 154–159. [Google Scholar] [CrossRef]
- Green, E.P.; Clark, C.D.; Mumby, P.J.; Edwards, A.J.; Ellis, A. Remote sensing techniques for mangrove mapping. Int. J. Remote Sens. 1998, 19, 935–956. [Google Scholar] [CrossRef]
- Held, A.; Ticehurst, C.; Lymburner, L.; Williams, N. High resolution mapping of tropical mangrove ecosystems using hyperspectral and radar remote sensing. Int. J. Remote Sens. 2003, 24, 2739–2759. [Google Scholar] [CrossRef]
- Giri, S.; Mukhopadhyay, A.; Hazra, S.; Mukherjee, S.; Roy, D.; Ghosh, S.; Ghosh, T.; Mitra, D. A study on abundance and distribution of mangrove species in Indian Sundarban using remote sensing technique. J. Coast. Conserv. 2014, 18, 359–367. [Google Scholar] [CrossRef]
- Liu, K.; Liu, L.; Liu, H.; Li, X.; Wang, S. Exploring the effects of biophysical parameters on the spatial pattern of rare cold damage to mangrove forests. Remote Sens. Environ. 2014, 150, 20–33. [Google Scholar] [CrossRef]
- Wang, L.; Silváncárdenas, J.L.; Sousa, W.P. Neural network classification of mangrove species from multi-seasonal IKONOS imagery. Photogramm. Eng. Remote Sens. 2008, 74, 921–927. [Google Scholar] [CrossRef]
- Neukermans, G.; Dahdouh-Guebas, F.; Kairo, J.G.; Koedam, N. Mangrove species and stand mapping in gazi bay (Kenya) using quickbird satellite imagery. J. Spat. Sci. 2008, 53, 75–86. [Google Scholar] [CrossRef]
- Wang, L.; Sousa, W.P.; Peng, G.; Biging, G.S. Comparison of IKONOS and quickbird images for mapping mangrove species on the caribbean coast of panama. Remote Sens. Environ. 2004, 91, 432–440. [Google Scholar] [CrossRef]
- Zhu, Y.; Liu, K.; Liu, L.; Wang, S.; Liu, H. Retrieval of mangrove aboveground biomass at the individual species level with worldview-2 images. Remote Sens. 2015, 7, 12192–12214. [Google Scholar] [CrossRef]
- Tang, H.; Liu, K.; Zhu, Y.; Wang, S.; Liu, L.; Song, S. Mangrove community classification based on worldview-2 image and SVM method. Acta Sci. Nat. Univ. Sunyatseni 2015, 54, 102–111. [Google Scholar]
- Pu, R. Mapping urban forest tree species using IKONOS imagery: Preliminary results. Environ. Monit. Assess. 2011, 172, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Tong, Q.; Zhang, B.; Zheng, L. Hyperspectral Remote Sensing: Principles, Techniques and Applications; Higher Education Press: Beijing, China, 2006. [Google Scholar]
- Guo, M.; Li, J.; Sheng, C.; Xu, J.; Wu, L. A review of wetland remote sensing. Sensors 2017, 17. [Google Scholar] [CrossRef] [PubMed]
- Hirano, A.; Madden, M.; Welch, R. Hyperspectral image data for mapping wetland vegetation. Wetlands 2003, 23, 436–448. [Google Scholar] [CrossRef]
- Koedsin, W.; Vaiphasa, C. Discrimination of tropical mangroves at the species level with EO-1 hyperion data. Remote Sens. 2013, 5, 3562–3582. [Google Scholar] [CrossRef]
- Jia, M.; Zhang, Y.; Wang, Z.; Song, K.; Ren, C. Mapping the distribution of mangrove species in the Core Zone of Mai Po Marshes Nature Reserve, Hong Kong, using hyperspectral data and high-resolution data. Int. J. Appl. Earth Obs. Geoinf. 2014, 33, 226–231. [Google Scholar] [CrossRef]
- Kamal, M.; Phinn, S. Hyperspectral data for mangrove species mapping: A comparison of pixel-based and object-based approach. Remote Sens. 2011, 3, 2222–2242. [Google Scholar] [CrossRef]
- Fauvel, M.; Tarabalka, Y.; Benediktsson, J.A.; Chanussot, J.; Tilton, J.C. Advances in spectral-spatial classification of hyperspectral images. Proc. IEEE 2013, 101, 652–675. [Google Scholar] [CrossRef]
- Matsuki, T.; Yokoya, N.; Iwasaki, A. Hyperspectral tree species classification of Japanese complex mixed forest with the aid of LiDAR data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 2177–2187. [Google Scholar] [CrossRef]
- Balas, C.; Pappas, C.; Epitropou, G. Multi/Hyper-Spectral Imaging. In Handbook of Biomedical Optics; Boas, D.A., Pitris, C., Ramanujam, N., Eds.; CRC Press: Boca Raton, FL, USA, 2011. [Google Scholar]
- Colomina, I.; Molina, P. Unmanned aerial systems for photogrammetry and remote sensing: A review. ISPRS J. Photogramm. Remote Sens. 2014, 92, 79–97. [Google Scholar] [CrossRef]
- Aasen, H.; Burkart, A.; Bolten, A.; Bareth, G. Generating 3D hyperspectral information with lightweight UAV snapshot cameras for vegetation monitoring: From camera calibration to quality assurance. ISPRS J. Photogramm. Remote Sens. 2015, 108, 245–259. [Google Scholar] [CrossRef]
- Bareth, G.; Aasen, H.; Bendig, J.; Gnyp, M.L.; Bolten, A.; Jung, A.; Michels, R.; Soukkamäki, J. Low-weight and UAV-based hyperspectral full-frame cameras for monitoring crops: Spectral comparison with portable spectroradiometer measurements. Photogramm. Fernerkund. Geoinf. 2015, 2015, 69–79. [Google Scholar] [CrossRef]
- Cui, J.; Zhang, S.; Zhang, J.; Liu, X.; Ding, R.; Liu, H. Determining surface magnetic susceptibility of loess-paleosol sections based on spectral features: Application to a UHD 185 hyperspectral image. Int. J. Appl. Earth Obs. Geoinf. 2016, 50, 159–169. [Google Scholar] [CrossRef]
- Hruska, R.; Mitchell, J.; Anderson, M.; Glenn, N.F. Radiometric and geometric analysis of hyperspectral imagery acquired from an unmanned aerial vehicle. Remote Sens. 2012, 4, 2736–2752. [Google Scholar] [CrossRef]
- Mitchell, J.J.; Glenn, N.F.; Anderson, M.O.; Hruska, R.C.; Halford, A.; Baun, C.; Nydegger, N. Unmanned aerial vehicle (UAV) hyperspectral remote sensing for dryland vegetation monitoring. In Proceedings of the Workshop on Hyperspectral Image & Signal Processing: Evolution in Remote Sensing, Shanghai, China, 4–7 June 2012; pp. 1–10. [Google Scholar]
- Zarco-Tejada, P.J.; González-Dugo, V.; Berni, J.A.J. Fluorescence, temperature and narrow-band indices acquired from a UAV platform for water stress detection using a micro-hyperspectral imager and a thermal camera. Remote Sens. Environ. 2012, 117, 322–337. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Catalina, A.; González, M.R.; Martín, P. Relationships between net photosynthesis and steady-state chlorophyll fluorescence retrieved from airborne hyperspectral imagery. Remote Sens. Environ. 2013, 136, 247–258. [Google Scholar] [CrossRef]
- Zarcotejada, P.J.; Guilléncliment, M.L.; Hernándezclemente, R.; Catalina, A.; González, M.R.; Martín, P. Estimating leaf carotenoid content in vineyards using high resolution hyperspectral imagery acquired from an unmanned aerial vehicle (UAV). Agric. For. Meteorol. 2013, 171, 281–294. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Morales, A.; Testi, L.; Villalobos, F.J. Spatio-temporal patterns of chlorophyll fluorescence and physiological and structural indices acquired from hyperspectral imagery as compared with carbon fluxes measured with eddy covariance. Remote Sens. Environ. 2013, 133, 102–115. [Google Scholar] [CrossRef]
- Calderón, R.; Navas-Cortés, J.A.; Lucena, C.; Zarco-Tejada, P.J. High-resolution airborne hyperspectral and thermal imagery for early detection of verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sens. Environ. 2013, 139, 231–245. [Google Scholar] [CrossRef]
- Yuan, H.; Yang, G.; Li, C.; Wang, Y.; Liu, J.; Yu, H.; Feng, H.; Xu, B.; Zhao, X.; Yang, X. Retrieving soybean leaf area index from unmanned aerial vehicle hyperspectral remote sensing: Analysis of RF, ANN, and SVM regression models. Remote Sens. 2017, 9. [Google Scholar] [CrossRef]
- Lu, G.; Yang, G.; Zhao, X.; Wang, Y.; Li, C.; Zhang, X. Inversion of soybean fresh biomass based on multi-payload unmanned aerial vehicles (UAVs). Soybean Sci. 2017, 36, 41–50. [Google Scholar]
- Zhao, X.; Yang, G.; Liu, J.; Zhang, X.; Xu, B.; Wang, Y.; Zhao, C.; Gai, J. Estimation of soybean breeding yield based on optimization of spatial scale of UAV hyperspectral image. Trans. Chin. Soc. Agric. Eng. 2017, 33, 110–116. [Google Scholar]
- Gao, L.; Yang, G.; Yu, H.; Xu, B.; Zhao, X.; Dong, J.; Ma, Y. Retrieving winter wheat leaf area index based on unmanned aerial vehicle hyperspectral remote sensing. Trans. Chin. Soc. Agric. Eng. 2016, 32, 113–120. [Google Scholar]
- Qin, Z.; Chang, Q.; Xie, B.; Jian, S. Rice leaf nitrogen content estimation based on hysperspectral imagery of UAV in yellow river diversion irrigation district. Trans. Chin. Soc. Agric. Eng. 2016, 32, 77–85. [Google Scholar]
- Tian, M.; Ban, S.; Chang, Q.; You, M.; Dan, L.; Li, W.; Wang, S. Use of hyperspectral images from UAV-based imaging spectroradiometer to estimate cotton leaf area index. Trans. Chin. Soc. Agric. Eng. 2016, 32, 102–108. [Google Scholar]
- Yue, J.; Yang, G.; Li, C.; Li, Z.; Wang, Y.; Feng, H.; Xu, B. Estimation of winter wheat above-ground biomass using unmanned aerial vehicle-based snapshot hyperspectral sensor and crop height improved models. Remote Sens. 2017, 9. [Google Scholar] [CrossRef]
- Sankey, T.; Donager, J.; McVay, J.; Sankey, J.B. UAV LiDAR and hyperspectral fusion for forest monitoring in the southwestern USA. Remote Sens. Environ. 2017, 195, 30–43. [Google Scholar] [CrossRef]
- Li, Q.S.; Wong, F.K.K.; Fung, T. Assessing the utility of UAV-borne hyperspectral image and photogrammetry derived 3D data for wetland species distribution quick mapping. ISPRS Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2017, XLII-2/W6, 209–215. [Google Scholar] [CrossRef]
- D’iorio, M.; Jupiter, S.D.; Cochran, S.A.; Potts, D.C. Optimizing remote sensing and GIS tools for mapping and managing the distribution of an invasive mangrove (Rhizophora mangle) on South Molokai, Hawaii. Mar. Geodesy 2007, 30, 125–144. [Google Scholar] [CrossRef]
- Yang, C.; Everitt, J.H.; Fletcher, R.S.; Jensen, R.R.; Mausel, P.W. Evaluating aisa + hyperspectral imagery for mapping black mangrove along the south texas gulf coast. Photogramm. Eng. Remote Sens. 2009, 75, 425–435. [Google Scholar] [CrossRef]
- Chakravortty, S. Analysis of end member detection and subpixel classification algorithms on hyperspectral imagery for tropical mangrove species discrimination in the Sunderbans Delta, India. J. Appl. Remote Sens. 2013, 7. [Google Scholar] [CrossRef]
- Chakravortty, S.; Shah, E.; Chowdhury, A.S. Application of spectral unmixing algorithm on hyperspectral data for mangrove species classification. In Proceedings of the International Conference on Applied Algorithms, Kolkata, India, 13–15 January 2014; Springer: New York, NY, USA, 2014; pp. 223–236. [Google Scholar]
- Chakravortty, S.; Sinha, D. Analysis of multiple scattering of radiation amongst end members in a mixed pixel of hyperspectral data for identification of mangrove species in a mixed stand. J. Indian Soc. Remote Sens. 2015, 43, 559–569. [Google Scholar] [CrossRef]
- Zhang, Z.; Kazakova, A.; Moskal, L.; Styers, D. Object-based tree species classification in urban ecosystems using LiDAR and hyperspectral data. Forests 2016, 7. [Google Scholar] [CrossRef]
- Kumar, T.; Panigrahy, S.; Kumar, P.; Parihar, J.S. Classification of floristic composition of mangrove forests using hyperspectral data: Case study of Bhitarkanika National Park, India. J. Coast. Conserv. 2013, 17, 121–132. [Google Scholar] [CrossRef]
- Wong, F.K.; Fung, T. Combining EO-1 hyperion and ENVISAT ASAR data for mangrove species classification in Mai Po Ramsar Site, Hong Kong. Int. J. Remote Sens. 2014, 35, 7828–7856. [Google Scholar] [CrossRef]
- Duro, D.C.; Franklin, S.E.; Dubé, M.G. A comparison of pixel-based and object-based image analysis with selected machine learning algorithms for the classification of agricultural landscapes using spot-5 HRG imagery. Remote Sens. Environ. 2012, 118, 259–272. [Google Scholar] [CrossRef]
- Pu, R. Tree species classification. In Remote Sensing of Natural Resources; CRC Press: Boca Raton, FL, USA, 2013; pp. 239–258. [Google Scholar]
- Li, S.S.; Tian, Q.J. Mangrove canopy species discrimination based on spectral features of geoeye-1 imagery. Spectrosc. Spectr. Anal. 2013, 33, 136–141. [Google Scholar]
- Xiao, H.Y.; Zeng, H.; Zan, Q.J.; Bai, Y.; Cheng, H.H. Decision tree model in extraction of mangrove community information using hyperspectral image data. J. Remote Sens. 2007, 11, 531–537. [Google Scholar]
- Liu, X.; Bo, Y. Object-based crop species classification based on the combination of airborne hyperspectral images and LiDAR data. Remote Sens. 2015, 7, 922–950. [Google Scholar] [CrossRef]
- Chadwick, J. Integrated LiDAR and IKONOS multispectral imagery for mapping mangrove distribution and physical properties. Int. J. Remote Sens. 2011, 32, 6765–6781. [Google Scholar] [CrossRef]
- Kamal, M.; Phinn, S.; Johansen, K. Object-based approach for multi-scale mangrove composition mapping using multi-resolution image datasets. Remote Sens. 2015, 7, 4753–4783. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Diaz-Varela, R.; Angileri, V.; Loudjani, P. Tree height quantification using very high resolution imagery acquired from an unmanned aerial vehicle (UAV) and automatic 3D photo-reconstruction methods. Eur. J. Agron. 2014, 55, 89–99. [Google Scholar] [CrossRef]
- Turner, D.; Lucieer, A.; Wallace, L. Direct georeferencing of ultrahigh-resolution UAV imagery. IEEE Trans. Geosci. Remote Sens. 2014, 52, 2738–2745. [Google Scholar] [CrossRef]
- Liao, B.W.; Wei, G.; Zhang, J.E.; Tang, G.L.; Lei, Z.S.; Yang, X.B. Studies on dynamic development of mangrove communities on Qi’ao Island, Zhuhai. J. South China Agric. Univ. 2008, 29, 59–64. [Google Scholar]
- Liu, B.E.; Liao, B.W. Mangrove reform-planting trial on Qi’ao Island. Ecol. Sci. 2013, 32, 534–539. [Google Scholar]
- Baatz, M.; Schäpe, A. Multiresolution Segmentation: An Optimization Approach for High Quality Multi-Scale Image Segmentation. Available online: http://www.ecognition.com/sites/default/files/405_baatz_fp_12.pdf (accessed on 21 November 2017).
- Cheng, J.; Bo, Y.; Zhu, Y.; Ji, X. A novel method for assessing the segmentation quality of high-spatial resolution remote-sensing images. Int. J. Remote Sens. 2014, 35, 3816–3839. [Google Scholar] [CrossRef]
- Benz, U.C.; Hofmann, P.; Willhauck, G.; Lingenfelder, I.; Heynen, M. Multi-resolution, object-oriented fuzzy analysis of remote sensing data for GIS-ready information. ISPRS J. Photogramm. Remote Sens. 2004, 58, 239–258. [Google Scholar] [CrossRef]
- Qian, Y.; Zhou, W.; Yan, J.; Li, W.; Han, L. Comparing machine learning classifiers for object-based land cover classification using very high resolution imagery. Remote Sens. 2014, 7, 153–168. [Google Scholar] [CrossRef]
- Haboudane, D.; Miller, J.R.; Tremblay, N.; Zarco-Tejada, P.J.; Dextraze, L. Integrated narrow-band vegetation indices for prediction of crop chlorophyll content for application to precision agriculture. Remote Sens. Environ. 2002, 81, 416–426. [Google Scholar] [CrossRef]
- Rondeaux, G.; Steven, M.; Baret, F. Optimization of soil-adjusted vegetation indices. Remote Sens. Environ. 1996, 55, 95–107. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Berjón, A.; López-Lozano, R.; Miller, J.R.; Martín, P.; Cachorro, V.; González, M.; De Frutos, A. Assessing vineyard condition with hyperspectral indices: Leaf and canopy reflectance simulation in a row-structured discontinuous canopy. Remote Sens. Environ. 2005, 99, 271–287. [Google Scholar] [CrossRef]
- Hernández-Clemente, R.; Navarro-Cerrillo, R.M.; Suárez, L.; Morales, F.; Zarco-Tejada, P.J. Assessing structural effects on PRI for stress detection in conifer forests. Remote Sens. Environ. 2011, 115, 2360–2375. [Google Scholar] [CrossRef]
- Rouse, J.W.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring Vegetation Systems in the Great Plains with ERTS; Paper-A20; National Aeronautics and Space Administration (NASA): Washington, DC, USA, 1974; pp. 309–317.
- Roujean, J.-L.; Breon, F.-M. Estimating par absorbed by vegetation from bidirectional reflectance measurements. Remote Sens. Environ. 1995, 51, 375–384. [Google Scholar] [CrossRef]
- Haboudane, D.; Miller, J.R.; Pattey, E.; Zarco-Tejada, P.J.; Strachan, I.B. Hyperspectral vegetation indices and novel algorithms for predicting green LAI of crop canopies: Modeling and validation in the context of precision agriculture. Remote Sens. Environ. 2004, 90, 337–352. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K. Textural features for image classification. IEEE Trans. Syst. Man Cybern. 1973, 6, 610–621. [Google Scholar] [CrossRef]
- Tan, Y.; Xia, W.; Xu, B.; Bai, L. Multi-feature classification approach for high spatial resolution hyperspectral images. J. Indian Soc. Remote Sens. 2017, 1–9. [Google Scholar] [CrossRef]
- Pu, R.; Gong, P. Hyperspectral remote sensing of vegetation bioparameters. In Advances in Environmental Remote Sensing: Sensors, Algorithm, and Applications; CRC Press: Boca Raton, FL, USA, 2011; pp. 101–142. [Google Scholar]
- Breiman, L.; Friedman, J.; Olshen, R.; Stone, C. Classification and Regresssion Trees; Wadsworth International Group: Belmont, CA, USA, 1984. [Google Scholar]
- Gomez-Chova, L.; Calpe, J.; Soria, E.; Camps-Valls, G.; Martin, J.D.; Moreno, J. CART-based feature selection of hyperspectral images for crop cover classification. In Proceedings of the International Conference on Image Processing, Barcelona, Spain, 14–17 September 2003; pp. 589–592. [Google Scholar]
- Bittencourt, H.R.; Clarke, R.T. Feature selection by using classification and regression trees (CART). Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, 35, 66–70. [Google Scholar]
- Hall, M.A. Feature selection for discrete and numeric class machine learning. In Proceedings of the Seventeenth International Conference on Machine Learning, Stanford, CA, USA, 29 June–2 July 2000; pp. 359–366. [Google Scholar]
- Wollmer, M.; Schuller, B.; Eyben, F.; Rigoll, G. Combining long short-term memory and dynamic bayesian networks for incremental emotion-sensitive artificial listening. IEEE J. Sel. Top. Signal Process. 2010, 4, 867–881. [Google Scholar] [CrossRef]
- Hall, M.A.; Holmes, G. Benchmarking attribute selection techniques for discrete class data mining. IEEE Trans. Knowl. Data Eng. 2003, 15, 1437–1447. [Google Scholar] [CrossRef]
- Piedra-Fernandez, J.A.; Canton-Garbin, M.; Wang, J.Z. Feature selection in avhrr ocean satellite images by means of filter methods. IEEE Trans. Geosci. Remote Sens. 2010, 48, 4193–4203. [Google Scholar] [CrossRef]
- Aguirre-Gutiérrez, J.; Seijmonsbergen, A.C.; Duivenvoorden, J.F. Optimizing land cover classification accuracy for change detection, a combined pixel-based and object-based approach in a mountainous area in mexico. Appl. Geogr. 2012, 34, 29–37. [Google Scholar] [CrossRef]
- Cover, T.; Hart, P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory 1967, 13, 21–27. [Google Scholar] [CrossRef]
- Hart, B.P.E. The condensed nearest neighbor rule. IEEE Trans. Inf. Theory 1968, 14, 515–516. [Google Scholar] [CrossRef]
- Yang, J.M.; Yu, P.T.; Kuo, B.C. A nonparametric feature extraction and its application to nearest neighbor classification for hyperspectral image data. IEEE Trans. Geosci. Remote Sens. 2010, 48, 1279–1293. [Google Scholar] [CrossRef]
- Féret, J.B.; Asner, G.P. Semi-supervised methods to identify individual crowns of lowland tropical canopy species using imaging spectroscopy and LiDAR. Remote Sens. 2012, 4, 2457–2476. [Google Scholar] [CrossRef]
- Mountrakis, G.; Im, J.; Ogole, C. Support vector machines in remote sensing: A review. ISPRS J. Photogramm. Remote Sens. 2011, 66, 247–259. [Google Scholar] [CrossRef]
- Chang, C.C.; Lin, C.J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2011, 2, 27. [Google Scholar] [CrossRef]
- Congalton, R.G.; Green, K. Assessing the Accuracy of Remotely Sensed Data: Principles and Practices; CRC Press: Boca Raton, FL, USA, 2008. [Google Scholar]
- Ma, L.; Cheng, L.; Li, M.; Liu, Y.; Ma, X. Training set size, scale, and features in geographic object-based image analysis of very high resolution unmanned aerial vehicle imagery. ISPRS J. Photogramm. Remote Sens. 2015, 102, 14–27. [Google Scholar] [CrossRef]
- Gomez-Chova, L.; Calpe, J.; Camps-Valls, G.; Martin, J.D.; Soria, E.; Vila, J.; Alonso-Chorda, L.; Moreno, J. Feature selection of hyperspectral data through local correlation and SFFS for crop classification. In Proceedings of the 2003 IEEE International Geoscience & Remote Sensing Symposium, Toulouse, France, 21–25 July 2003; pp. 555–557. [Google Scholar]
| Land-Cover Types | Ground Truth Samples | Total | |
|---|---|---|---|
| Training Samples | Testing Samples | ||
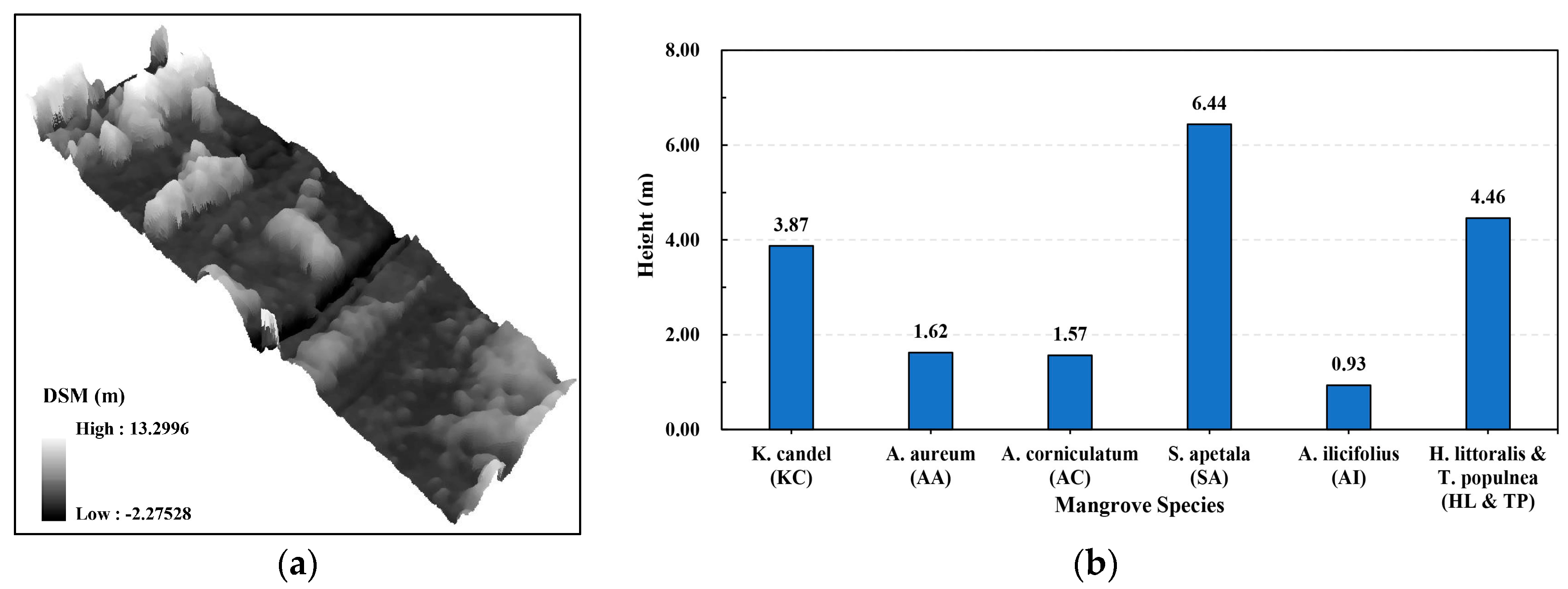
| K. candel (KC) | 68 | 58 | 126 |
| A. aureum (AA) | 46 | 31 | 77 |
| A. corniculatum (AC) | 37 | 45 | 82 |
| S. apetala (SA) | 71 | 41 | 112 |
| A. ilicifolius (AI) | 28 | 22 | 50 |
| H. littoralis & T. populnea (HL & TP) | 60 | 41 | 101 |
| water area (river) | 93 | 40 | 133 |
| P. australis (PA) | 22 | 18 | 40 |
| boardwalk | 14 | 10 | 24 |
| shadow | 54 | 29 | 83 |
| total | 493 | 335 | 828 |
| Object Features | Description |
|---|---|
| spectral bands | Mean values of 125 spectral bands for each image object, brightness, and max.diff. |
| hyperspectral vegetation indices (vis) | Eight VIs, including BGI2, NDVI, RDVI, TCARI, OSAVI, TCARI/OSAVI, MCARI2, and PRI. |
| textural features | 24 textural features, including ASM, CON, COR, ENT, HOM, MEAN, DIS, StdDev calculated using GLCM with three bands (that is band 8, band 25, and band 50). |
| height information | UAV-derived DSM (Digital Surface Model). |
| Hyperspectral Vegetation Indices (VIs) | Formulation |
|---|---|
| Blue Green Pigment Index 2 (BGI 2) | |
| Normalized Difference Vegetation Index (NDVI) | |
| Reformed Difference Vegetation Index (RDVI) | |
| Transformed Chlorophyll Absorption in Reflectance Index (TCARI) | |
| Optimized Soil-Adjusted Vegetation Index (OSAVI) | |
| TCARI/OSAVI | |
| Modified Chlorophyll Absorption Ratio Index 2 (MCARI 2) | |
| Photo-chemical Reflectance Index (PRI) |
| Textural Variables | Formulation |
|---|---|
| ASM (Angular Second Moment) | |
| CON (Contrast) | |
| COR (Correlation) | |
| ENT (Entropy) | |
| HOM (Homogeneity) | |
| Mean | |
| DIS (Dissimilarity) | |
| StdDev (Standard Deviation) |
| Spatial Resolution | Scale = 150 | Scale = 100 | Scale = 50 | Scale = 20 |
|---|---|---|---|---|
| 0.15 m | 83.58 | 88.66 | 74.63 | 60.60 |
| 0.3 m | 61.49 | 77.01 | 86.57 | 76.12 |
| 0.5 m | 35.22 | 54.93 | 82.69 | 74.93 |
| Classified Category | Experiment A | Experiment B | Experiment C | Experiment D | Experiment E | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| PA | UA | PA | UA | PA | UA | PA | UA | PA | UA | |
| K. candel (KC) | 81.03 | 69.12 | 77.59 | 70.31 | 79.31 | 76.67 | 79.31 | 73.02 | 93.55 | 67.44 |
| A. aureum (AA) | 80.65 | 49.02 | 87.10 | 65.85 | 100 | 51.67 | 90.32 | 73.68 | 59.09 | 72.22 |
| A. corniculatum (AC) | 40.00 | 75.00 | 60.00 | 90.00 | 46.67 | 84.00 | 60.00 | 93.10 | 97.50 | 72.22 |
| S. apetala (SA) | 80.49 | 76.74 | 90.24 | 86.05 | 80.49 | 94.29 | 90.24 | 88.10 | 50.00 | 100 |
| A. ilicifolius (AI) | 50.00 | 44.00 | 59.09 | 76.47 | 50.00 | 57.89 | 77.27 | 80.95 | 84.48 | 87.50 |
| H. littoralis & T. populnea (HL & TP) | 63.41 | 92.86 | 87.80 | 85.71 | 75.61 | 88.57 | 90.24 | 86.05 | 93.10 | 87.10 |
| P. australis (PA) | 92.50 | 75.51 | 92.50 | 74.00 | 97.50 | 73.58 | 95.00 | 74.51 | 57.78 | 81.25 |
| water area (river) | 77.78 | 77.78 | 77.78 | 100 | 77.78 | 77.78 | 77.78 | 100 | 77.78 | 100 |
| boardwalk | 10.00 | 100 | 50.00 | 100 | 20.00 | 100 | 50.00 | 100 | 90.24 | 92.50 |
| shadow | 93.10 | 96.43 | 89.66 | 89.66 | 93.10 | 96.43 | 89.66 | 89.66 | 85.37 | 83.33 |
| OA (%) | 71.34 | 79.70 | 76.12 | 82.09 | 81.79 | |||||
| Kappa | 0.675 | 0.770 | 0.730 | 0.797 | 0.774 | |||||
| Classified Category | Experiment A | Experiment B | Experiment C | Experiment D | Experiment E | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| PA | UA | PA | UA | PA | UA | PA | UA | PA | UA | |
| K. candel (KC) | 81.03 | 70.15 | 93.10 | 79.41 | 81.03 | 79.66 | 91.38 | 81.54 | 93.55 | 85.29 |
| A. aureum (AA) | 74.19 | 79.31 | 83.87 | 81.25 | 96.77 | 68.18 | 83.87 | 96.30 | 68.18 | 88.24 |
| A. corniculatum (AC) | 55.56 | 71.43 | 77.78 | 92.11 | 64.44 | 76.32 | 75.56 | 91.89 | 100 | 78.43 |
| S. apetala (SA) | 82.93 | 79.07 | 87.80 | 92.31 | 78.05 | 94.12 | 90.24 | 94.87 | 100 | 100 |
| A. ilicifolius (AI) | 45.45 | 66.67 | 63.64 | 93.33 | 50.00 | 61.11 | 72.73 | 88.89 | 93.10 | 90.00 |
| H. littoralis & T. populnea (HL & TP) | 85.37 | 74.47 | 90.24 | 86.05 | 92.68 | 97.44 | 92.68 | 76.00 | 100 | 90.63 |
| P. australis (PA) | 92.50 | 80.43 | 100 | 90.91 | 100 | 80.00 | 97.50 | 92.86 | 77.78 | 89.74 |
| water area (river) | 77.78 | 100 | 88.89 | 100 | 77.78 | 100 | 88.89 | 100 | 72.22 | 100 |
| boardwalk | 70.00 | 100 | 100 | 100 | 70.00 | 100 | 100 | 100 | 90.24 | 94.87 |
| shadow | 96.55 | 87.50 | 93.10 | 90.00 | 96.55 | 87.50 | 96.55 | 90.32 | 92.68 | 95.00 |
| OA (%) | 77.61 | 88.06 | 82.39 | 88.66 | 89.55 | |||||
| Kappa | 0.746 | 0.864 | 0.801 | 0.871 | 0.882 | |||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, J.; Leng, W.; Liu, K.; Liu, L.; He, Z.; Zhu, Y. Object-Based Mangrove Species Classification Using Unmanned Aerial Vehicle Hyperspectral Images and Digital Surface Models. Remote Sens. 2018, 10, 89. https://doi.org/10.3390/rs10010089
Cao J, Leng W, Liu K, Liu L, He Z, Zhu Y. Object-Based Mangrove Species Classification Using Unmanned Aerial Vehicle Hyperspectral Images and Digital Surface Models. Remote Sensing. 2018; 10(1):89. https://doi.org/10.3390/rs10010089
Chicago/Turabian StyleCao, Jingjing, Wanchun Leng, Kai Liu, Lin Liu, Zhi He, and Yuanhui Zhu. 2018. "Object-Based Mangrove Species Classification Using Unmanned Aerial Vehicle Hyperspectral Images and Digital Surface Models" Remote Sensing 10, no. 1: 89. https://doi.org/10.3390/rs10010089
APA StyleCao, J., Leng, W., Liu, K., Liu, L., He, Z., & Zhu, Y. (2018). Object-Based Mangrove Species Classification Using Unmanned Aerial Vehicle Hyperspectral Images and Digital Surface Models. Remote Sensing, 10(1), 89. https://doi.org/10.3390/rs10010089