Mapping Wild Leek through the Forest Canopy Using a UAV
Abstract
1. Introduction
2. Materials and Methods
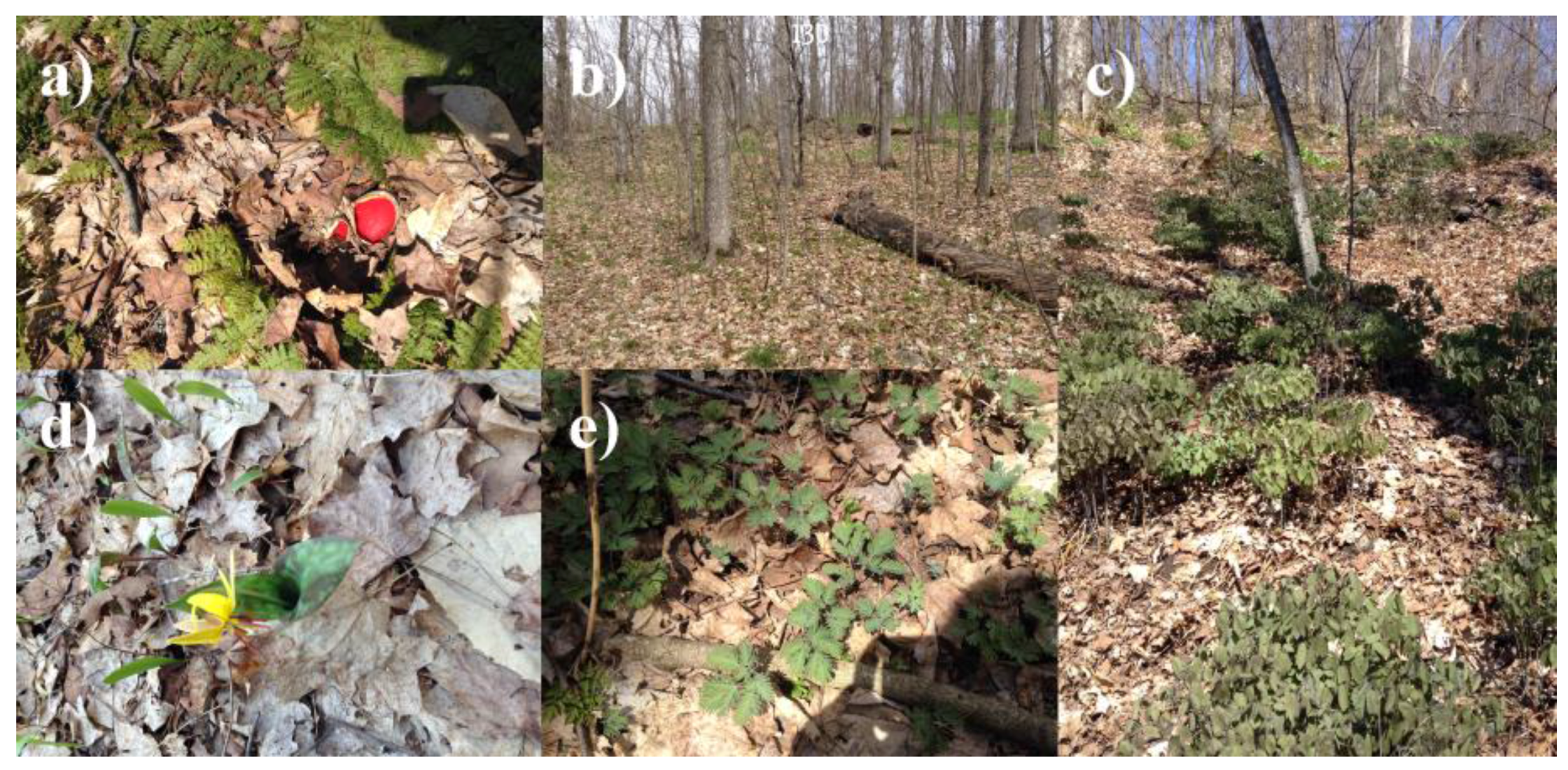
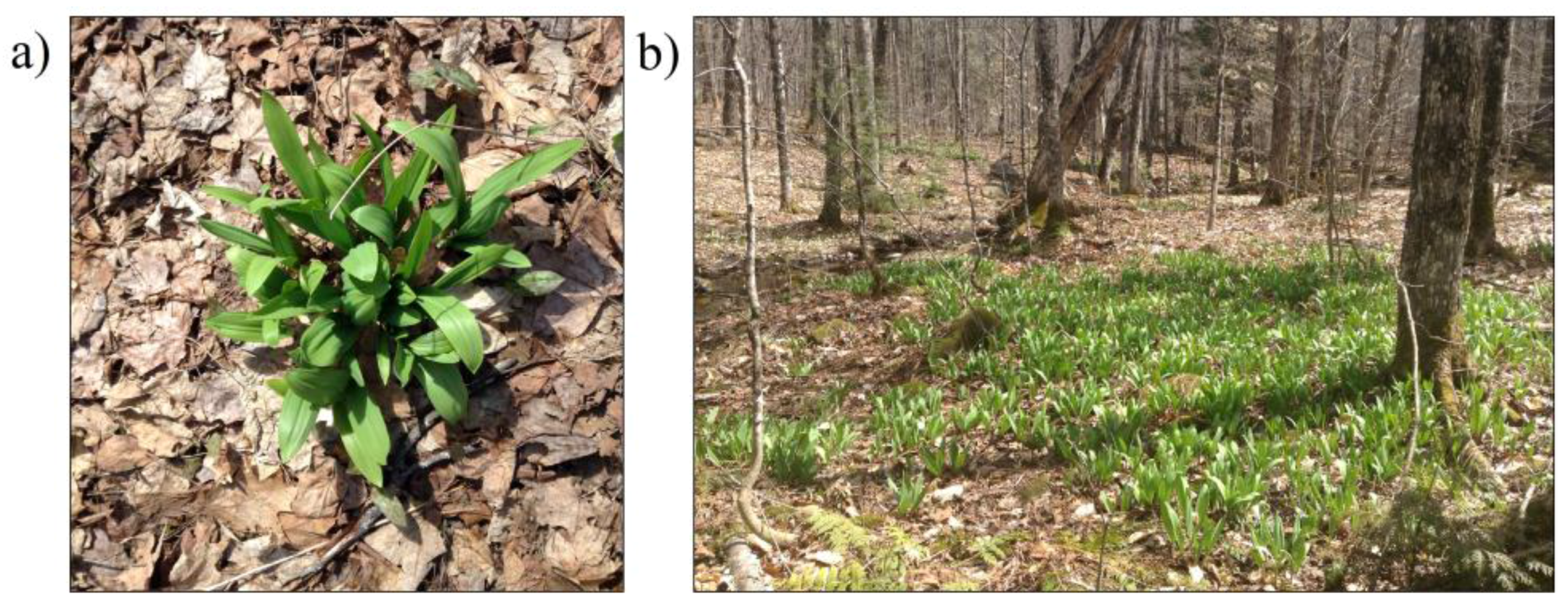
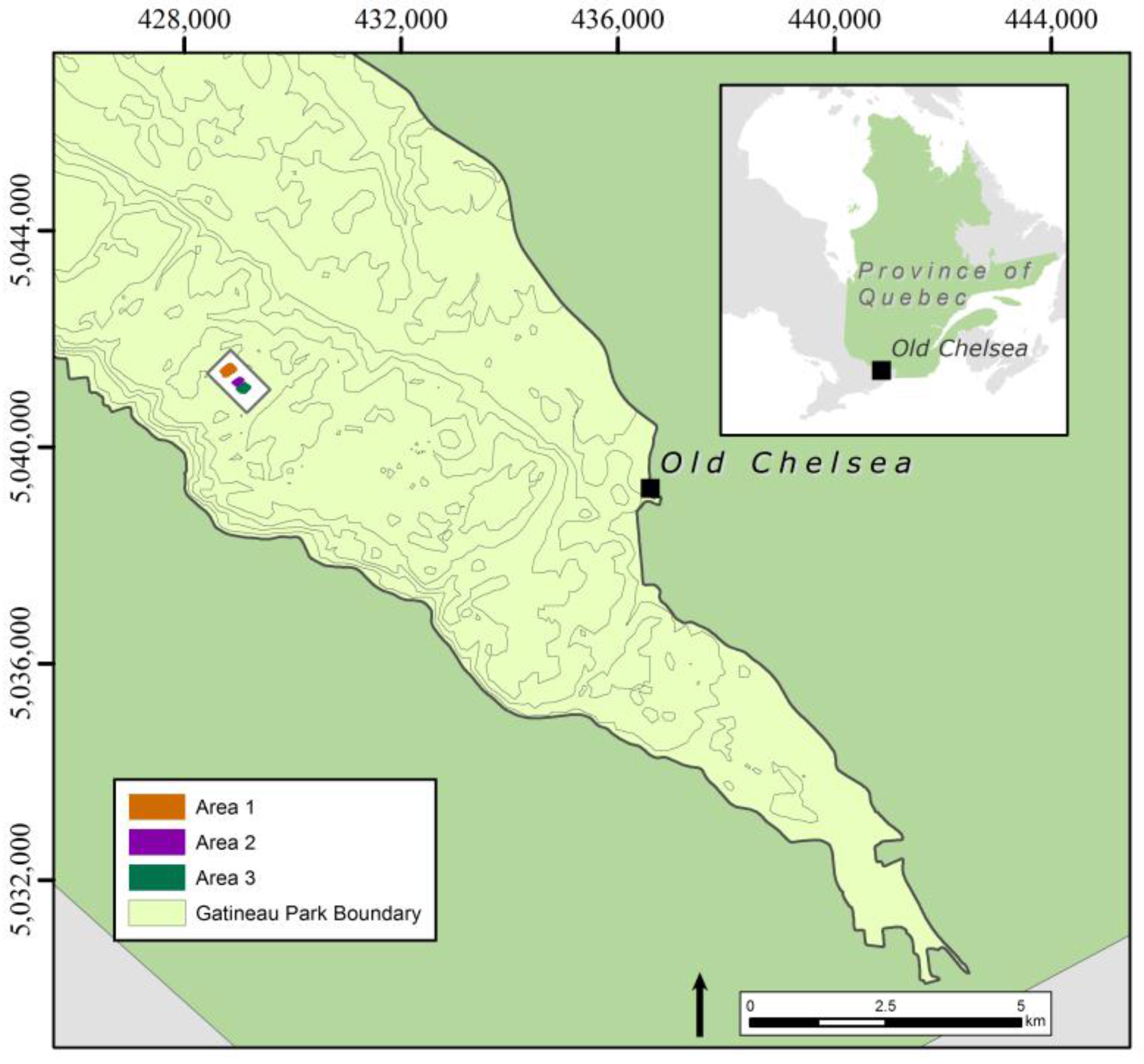
2.1. Study Area and Context
- (1)
- Area 1 (0.059 km2), contains dense and large patches of wild leek.
- (2)
- Area 2 (0.027 km2), contains smaller and more dispersed patches of wild leek.
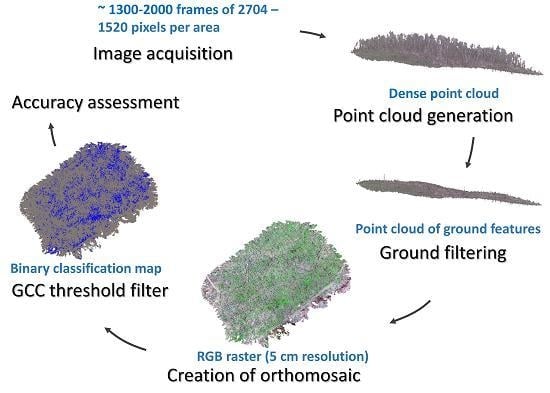
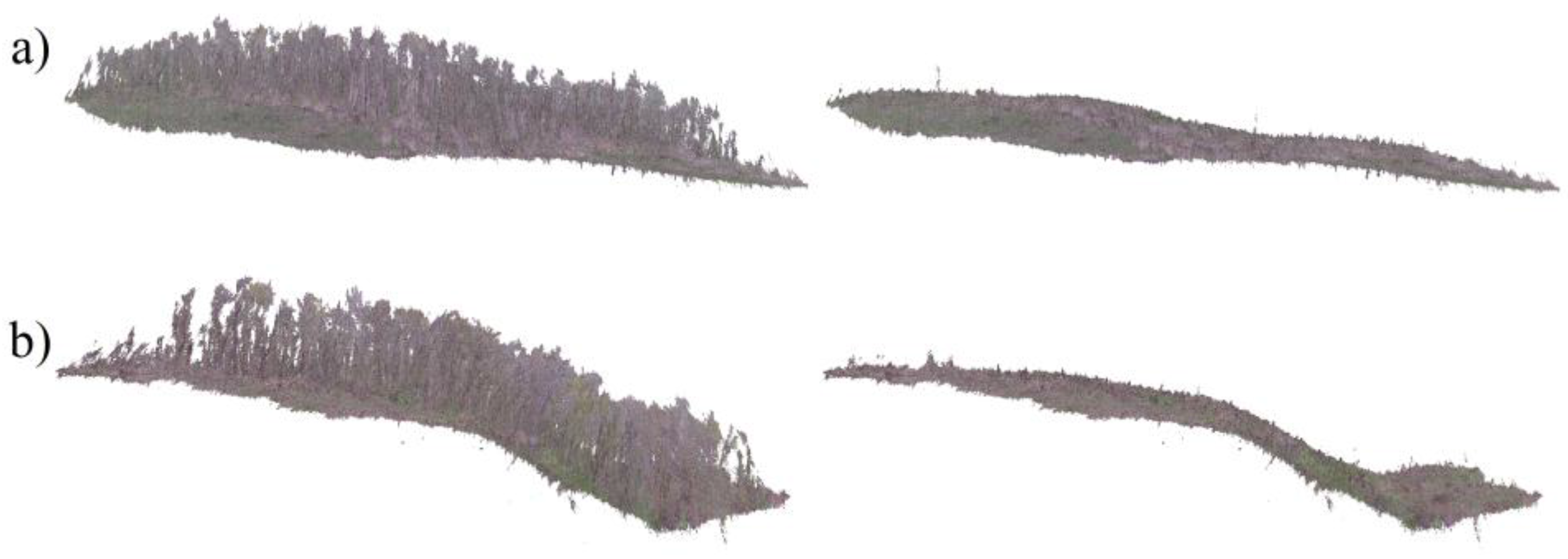
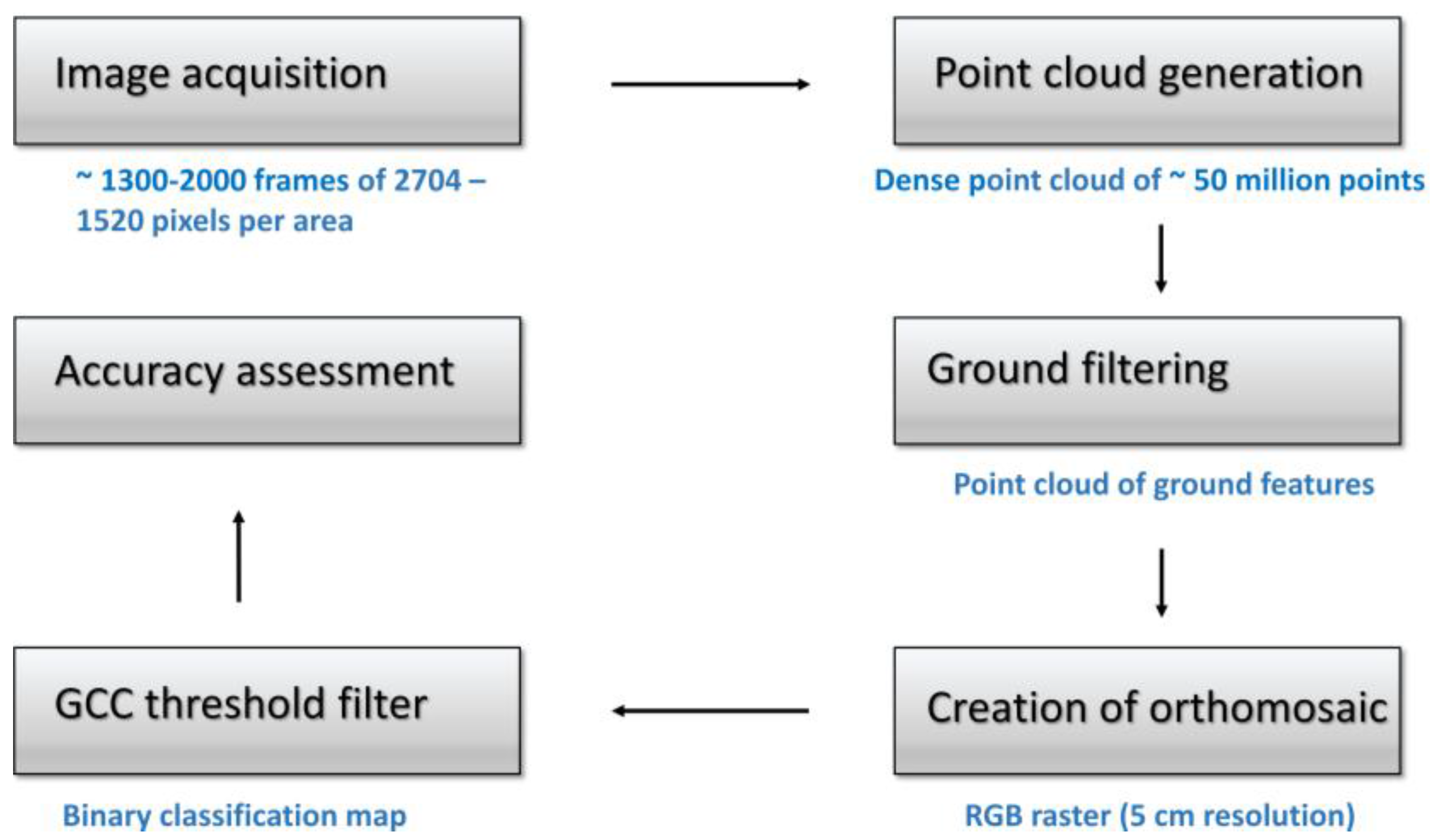
2.2. Remote Sensing Data, Acquisition and Processing
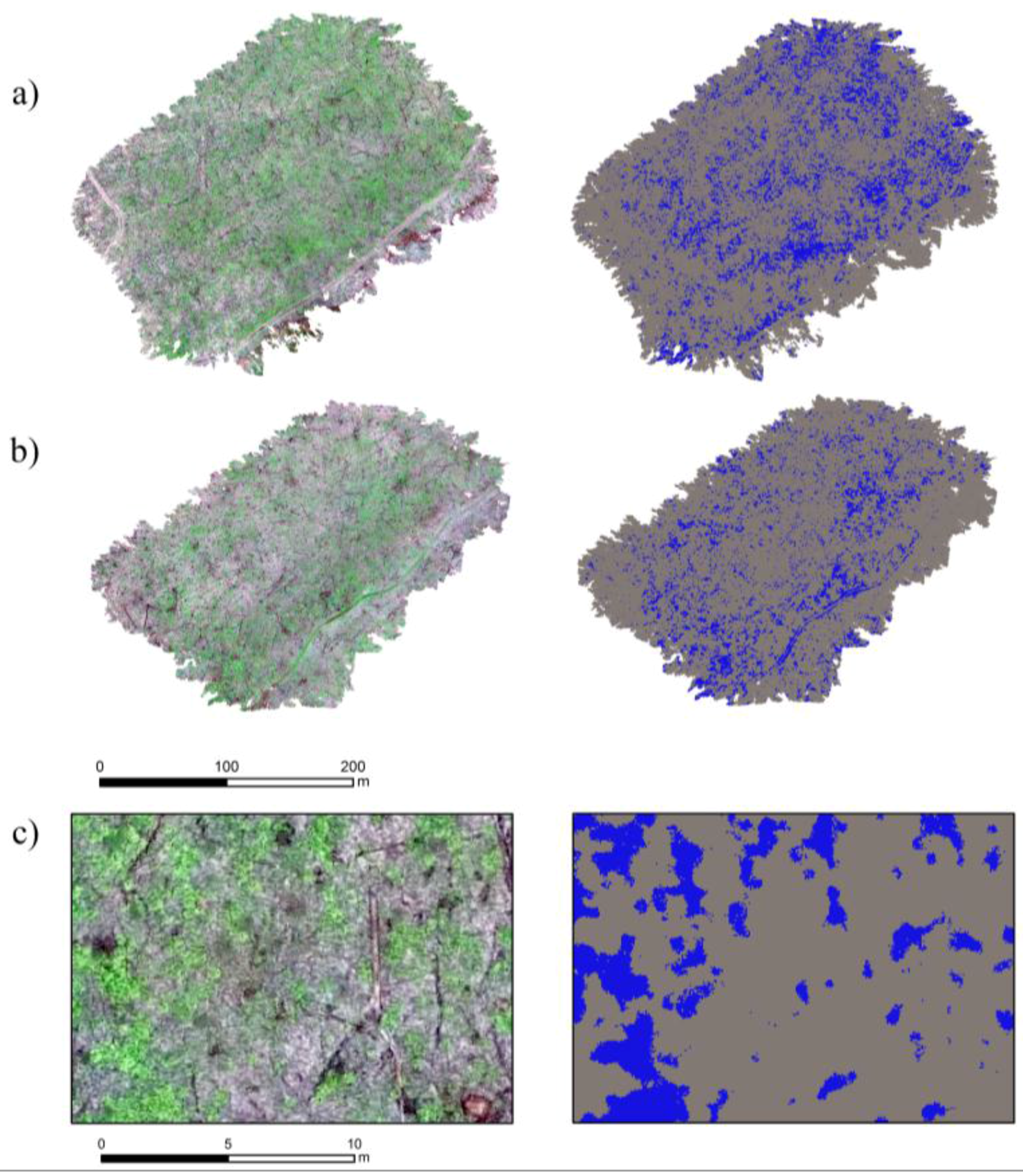
2.3. Wild Leek Detection
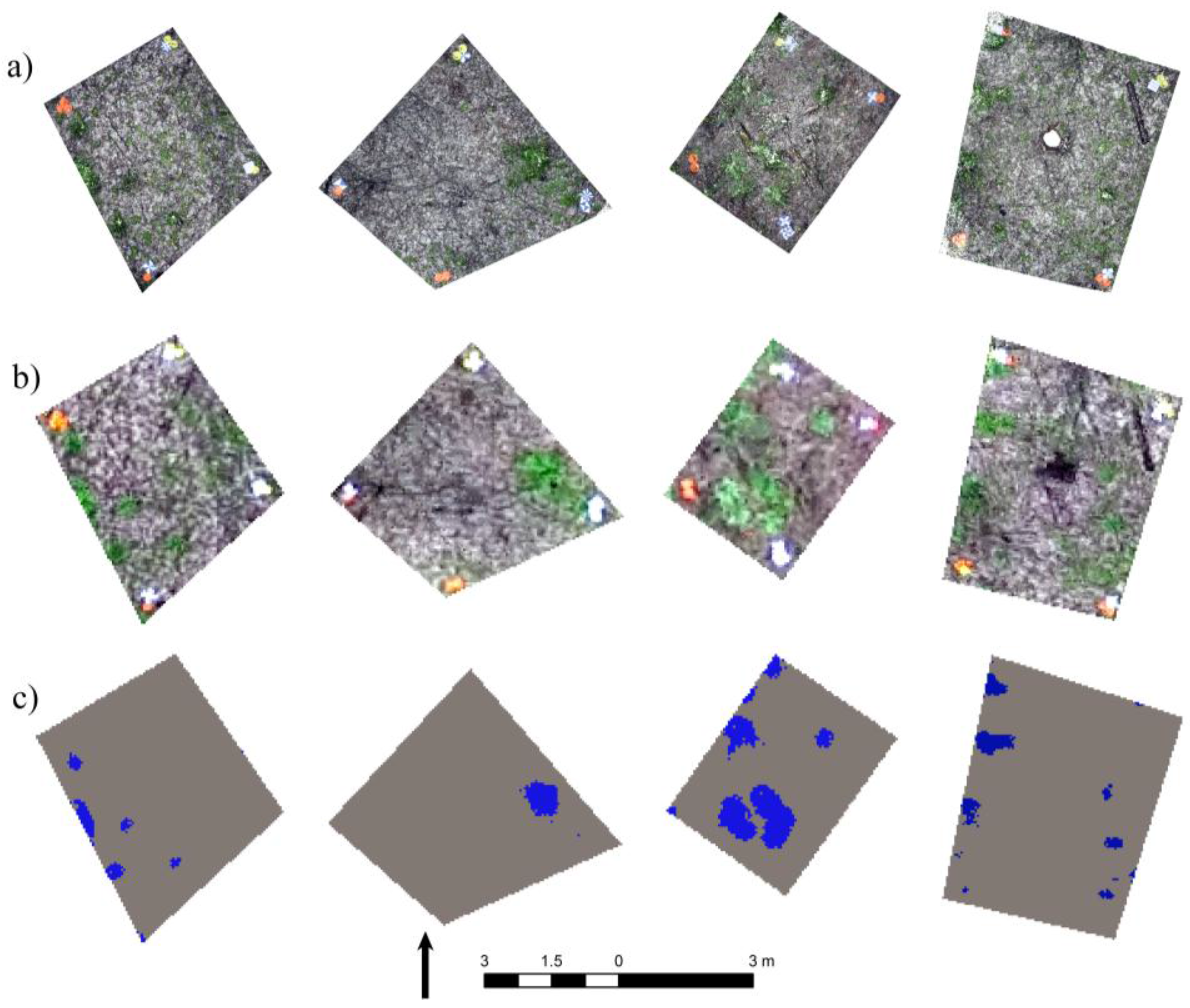
2.4. Accuracy Assessment
3. Results
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Vasseur, L.; Gagnon, D. Survival and growth of Allium tricoccum Ait. transplants in different habitats. Biol. Conserv. 1994, 68, 107–114. [Google Scholar] [CrossRef]
- BONAP North American Vascular Flora. Available online: http://www.bonap.org/ (accessed on 10 August 2017).
- eFloras Allium tricoccum. Missouri Botanical Garden, St. Louis, MO & Harvard University Herbaria, Cambridge, MA. Available online: http://www.efloras.org/ (accessed on 26 June 2017).
- MDDELCC Espèce Vulnérable au Québec. Available online: http://www.mddelcc.gouv.qc.ca/biodiversite/especes/ail/index.htm (accessed on 10 August 2017).
- Edgar, B.; Brubaker, H.; Tuminelli, K. Plugging the Leak on Wild Leeks: The Threat of Over-Harvesting Wild Leek Populations in Northern New York 2012. Available online: http://www.stlawu.edu/sites/default/files/resource/wild_leek_conservation.pdf (accessed on 10 July 2017).
- Nault, A.; Gagnon, D. Ramet demography of Allium tricoccum, a spring ephemeral, perennial forest herb. J. Ecol. 1993, 81, 101. [Google Scholar] [CrossRef]
- Nantel, P.; Gagnon, D.; Nault, A. Population Viability Analysis of American Ginseng and Wild Leek Harvested in Stochastic Environments. Conserv. Biol. 1996, 10, 608–621. [Google Scholar] [CrossRef]
- Rock, J.H.; Beckage, B.; Gross, L.J. Population recovery following differential harvesting of Allium tricoccum Ait. in the southern Appalachians. Biol. Conserv. 2004, 116, 227–234. [Google Scholar] [CrossRef]
- Regulation respecting threatened or vulnerable plant species and their habitats. Available online: http://legisquebec.gouv.qc.ca/en/ShowDoc/cr/E-12.01,%20r.%203 (accessed on 10 July 2017).
- Royo, A.A.; Carson, W.P. On the formation of dense understory layers in forests worldwide: Consequences and implications for forest dynamics, biodiversity, and succession. Can. J. For. Res. 2006, 36, 1345–1362. [Google Scholar] [CrossRef]
- Gilliam, F.S. The Ecological Significance of the Herbaceous Layer in Temperate Forest Ecosystems. BioScience 2007, 57, 845–858. [Google Scholar] [CrossRef]
- Ticktin, T. The ecological implications of harvesting non-timber forest products. J. Appl. Ecol. 2004, 41, 11–21. [Google Scholar] [CrossRef]
- Kraft, L.S.; Crow, T.R.; Buckley, D.S.; Nauertz, E.A.; Zasada, J.C. Effects of harvesting and deer browsing on attributes of understory plants in northern hardwood forests, Upper Michigan, USA. For. Ecol. Manag. 2004, 199, 219–230. [Google Scholar] [CrossRef]
- Fuller, D. Medicine from the Wild: An Overview of the U.S. Native Medicinal Plant Trade and Its Conservation Implications; World Wildlife Fund: Vaud, Switzerland, 1991; p. 28. [Google Scholar]
- Parcs Québec The Environmental Indicators Monitoring Program. Available online: https://www.sepaq.com/dotAsset/6ea4e70c-0210-4231-819f-0934a3b4f1d4.pdf (accessed on 10 August 2017).
- National Capital Commission Conservation at Gatineau Park. Available online: http://ncc-ccn.gc.ca/places-to-visit/gatineau-park/conservation-at-gatineau-park (accessed on 10 August 2017).
- National Capital Commission. Executive Summary: Report on Gatineau Park Ecosystems; NCC: Ottawa, ON, Canada, 2016.
- He, Y.; Nesbitt, N.; Tong, A.; Mui, A. Delineating the Endangered White Trillium (Trillium grandiflorum) and Improving Wild Leek (Allium tricoccum) Mapping Using Remote Sensing and GIS Technology in Gatineau Park; NCC Contribution Agreement; University of Toronto: Mississauga, ON, Canada, 2011; p. 29. [Google Scholar]
- Nault, A.; Gagnon, D. Seasonal Biomass and Nutrient Allocation patterns in Wild Leek (Allium tricoccum Ait.), a Spring Geophyte. Bull. Torrey Bot. Club 1988, 115, 45–54. [Google Scholar] [CrossRef]
- Bernatchez, A.; Lapointe, L. Cooler temperatures favour growth of wild leek (Allium tricoccum), a deciduous forest spring ephemeral. Bot.-Bot. 2012, 90, 1125–1132. [Google Scholar] [CrossRef]
- Becker, R.H.; Zmijewski, K.A.; Crail, T. Seeing the forest for the invasives: Mapping buckthorn in the Oak Openings. Biol. Invasions 2013, 15, 315–326. [Google Scholar] [CrossRef]
- Tuanmu, M.-N.; Viña, A.; Bearer, S.; Xu, W.; Ouyang, Z.; Zhang, H.; Liu, J. Mapping understory vegetation using phenological characteristics derived from remotely sensed data. Remote Sens. Environ. 2010, 114, 1833–1844. [Google Scholar] [CrossRef]
- Resasco, J.; Hale, A.N.; Henry, M.C.; Gorchov, D.L. Detecting an invasive shrub in a deciduous forest understory using late-fall Landsat sensor imagery. Int. J. Remote Sens. 2007, 28, 3739–3745. [Google Scholar] [CrossRef]
- Wilfong, B.N.; Gorchov, D.L.; Henry, M.C. Detecting an Invasive Shrub in Deciduous Forest Understories Using Remote Sensing. Weed Sci. 2009, 57, 512–520. [Google Scholar] [CrossRef]
- Johnston, S.E.; Henry, M.C.; Gorchov, D.L. Using Advanced Land Imager (ALI) and Landsat Thematic Mapper (TM) for the Detection of the Invasive Shrub Lonicera maackii in Southwestern Ohio Forests. GISci. Remote Sens. 2012, 49, 450–462. [Google Scholar] [CrossRef][Green Version]
- Hall, R.J.; Peddle, D.R.; Klita, D.L. Mapping conifer understory within boreal mixedwoods from Landsat TM satellite imagery and forest inventory information. For. Chron. 2000, 76, 887–902. [Google Scholar] [CrossRef]
- Chastain, J.; Robert, A.; Townsend, P.A. Use of Landsat ETM and Topographic Data to Characterize Evergreen Understory Communities in Appalachian Deciduous Forests. Photogramm. Eng. Remote Sens. 2007, 73, 563–575. [Google Scholar] [CrossRef]
- Pearlstine, L.; Portier, K.M.; Smith, S.E. Textural Discrimination of an Invasive Plant, Schinus terebinthifolius, from Low Altitude Aerial Digital Imagery. Photogramm. Eng. Remote Sens. 2005, 71, 289–298. [Google Scholar] [CrossRef]
- Shouse, M.; Liang, L.; Fei, S. Identification of understory invasive exotic plants with remote sensing in urban forests. Int. J. Appl. Earth Obs. Geoinf. 2013, 21, 525–534. [Google Scholar] [CrossRef]
- Jones, D.; Pike, S.; Thomas, M.; Murphy, D. Object-Based Image Analysis for Detection of Japanese Knotweed s.l. taxa (Polygonaceae) in Wales (UK). Remote Sens. 2011, 3, 319–342. [Google Scholar] [CrossRef]
- Müllerová, J.; Pergl, J.; Pyšek, P. Remote sensing as a tool for monitoring plant invasions: Testing the effects of data resolution and image classification approach on the detection of a model plant species Heracleum mantegazzianum (giant hogweed). Int. J. Appl. Earth Obs. Geoinf. 2013, 25, 55–65. [Google Scholar] [CrossRef]
- Niphadkar, M.; Nagendra, H.; Tarantino, C.; Adamo, M.; Blonda, P. Comparing Pixel and Object-Based Approaches to Map an Understorey Invasive Shrub in Tropical Mixed Forests. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Underwood, E.; Ustin, S.; DiPietro, D. Mapping nonnative plants using hyperspectral imagery. Remote Sens. Environ. 2003, 86, 150–161. [Google Scholar] [CrossRef]
- Anderson, K.; Gaston, K.J. Lightweight unmanned aerial vehicles will revolutionize spatial ecology. Front. Ecol. Environ. 2013, 11, 138–146. [Google Scholar] [CrossRef]
- Müllerová, J.; Bartaloš, T.; Brůna, J.; Dvořák, P.; Vítková, M. Unmanned aircraft in nature conservation: An example from plant invasions. Int. J. Remote Sens. 2017, 38, 2177–2198. [Google Scholar] [CrossRef]
- Lehmann, J.R.K.; Prinz, T.; Ziller, S.R.; Thiele, J.; Heringer, G.; Meira-Neto, J.A.A.; Buttschardt, T.K. Open-Source Processing and Analysis of Aerial Imagery Acquired with a Low-Cost Unmanned Aerial System to Support Invasive Plant Management. Front. Environ. Sci. 2017, 5. [Google Scholar] [CrossRef]
- Berra, E.F.; Gaulton, R.; Barr, S. Use of a digital camera onboard a UAV to monitor spring phenology at individual tree level. In Proceedings of the 2016 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Beijing, China, 10–15 July 2016; pp. 3496–3499. [Google Scholar]
- Dandois, J.P.; Ellis, E.C. High spatial resolution three-dimensional mapping of vegetation spectral dynamics using computer vision. Remote Sens. Environ. 2013, 136, 259–276. [Google Scholar] [CrossRef]
- Getzin, S.; Wiegand, K.; Schöning, I. Assessing biodiversity in forests using very high-resolution images and unmanned aerial vehicles. Methods Ecol. Evol. 2012, 3, 397–404. [Google Scholar] [CrossRef]
- Samiappan, S.; Turnage, G.; Hathcock, L.; Casagrande, L.; Stinson, P.; Moorhead, R. Using unmanned aerial vehicles for high-resolution remote sensing to map invasive Phragmites australis in coastal wetlands. Int. J. Remote Sens. 2017, 38, 2199–2217. [Google Scholar] [CrossRef]
- Hill, D.J.; Tarasoff, C.; Whitworth, G.E.; Baron, J.; Bradshaw, J.L.; Church, J.S. Utility of unmanned aerial vehicles for mapping invasive plant species: A case study on yellow flag iris (Iris pseudacorus L.). Int. J. Remote Sens. 2017, 38, 2083–2105. [Google Scholar] [CrossRef]
- Díaz-Varela, R.A.; de la Rosa, R.; León, L.; Zarco-Tejada, P.J. High-Resolution Airborne UAV Imagery to Assess Olive Tree Crown Parameters Using 3D Photo Reconstruction: Application in Breeding Trials. Remote Sens. 2015, 7, 4213–4232. [Google Scholar] [CrossRef]
- Westoby, M.J.; Brasington, J.; Glasser, N.F.; Hambrey, M.J.; Reynolds, J.M. ‘Structure-from-Motion’ photogrammetry: A low-cost, effective tool for geoscience applications. Geomorphology 2012, 179, 300–314. [Google Scholar] [CrossRef]
- Jensen, J.L.R.; Mathews, A.J. Assessment of Image-Based Point Cloud Products to Generate a Bare Earth Surface and Estimate Canopy Heights in a Woodland Ecosystem. Remote Sens. 2016, 8, 50. [Google Scholar] [CrossRef]
- Micheletti, N.; Chandler, J.H.; Lane, S.N. Structure from Motion (SFM) Photogrammetry; British Society for Geomorphology: London, UK, 2015. [Google Scholar]
- Del Degan, M. Gatineau Park Ecosystem Conservation Plan 2010; NCC: Ottawa, ON, Canada, 2010.
- Pasher, J.; King, D.J. Mapping dead wood distribution in a temperate hardwood forest using high resolution airborne imagery. For. Ecol. Manag. 2009, 258, 1536–1548. [Google Scholar] [CrossRef]
- Knudby, A.; Leduc, M.-B. Tracking the Change in Wild Leek and White Trillium Distributions in Gatineau Park; NCC Contribution Agreement; University of Ottawa: Ottawa, ON, Canada, 2017; p. 12. [Google Scholar]
- Evans, J.S.; Hudak, A.T. A Multiscale Curvature Algorithm for Classifying Discrete Return LiDAR in Forested Environments. IEEE Trans. Geosci. Remote Sens. 2007, 45, 1029–1038. [Google Scholar] [CrossRef]
- Ludwig Boltzmann Institute for Archaeological Prospection and Virtual Archaeology MCC-Lidar ALS Filtering. Available online: http://lbi-archpro.org/als-filtering/lbi-project/results/mcc-lidar/test-phase-4 (accessed on 26 June 2017).
- CloudCompare. Version 2.6; [GPL Software]. 2015. Available online: http://www.cloudcompare.org/ (accessed on 10 July 2017).
- Matiu, M.; Bothmann, L.; Steinbrecher, R.; Menzel, A. Monitoring succession after a non-cleared windthrow in a Norway spruce mountain forest using webcam, satellite vegetation indices and turbulent CO2 exchange. Agric. For. Meteorol. 2017, 244, 72–81. [Google Scholar] [CrossRef]
- Moore, C.E.; Beringer, J.; Evans, B.; Hutley, L.B.; Tapper, N.J. Tree–grass phenology information improves light use efficiency modelling of gross primary productivity for an Australian tropical savanna. Biogeosciences 2017, 14, 111–129. [Google Scholar] [CrossRef]
- Saitoh, T.M.; Nagai, S.; Saigusa, N.; Kobayashi, H.; Suzuki, R.; Nasahara, K.N.; Muraoka, H. Assessing the use of camera-based indices for characterizing canopy phenology in relation to gross primary production in a deciduous broad-leaved and an evergreen coniferous forest in Japan. Ecol. Inform. 2012, 11, 45–54. [Google Scholar] [CrossRef]
- Liu, Z.; Hu, H.; Yu, H.; Yang, X.; Yang, H.; Ruan, C.; Wang, Y.; Tang, J. Relationship between leaf physiologic traits and canopy color indices during the leaf expansion period in an oak forest. Ecosphere 2015, 6, 1–9. [Google Scholar] [CrossRef]
- Zhou, Y.; Xiao, X.; Wagle, P.; Bajgain, R.; Mahan, H.; Basara, J.B.; Dong, J.; Qin, Y.; Zhang, G.; Luo, Y.; et al. Examining the short-term impacts of diverse management practices on plant phenology and carbon fluxes of Old World bluestems pasture. Agric. For. Meteorol. 2017, 237, 60–70. [Google Scholar] [CrossRef]
- Woebbecke, D.M.; Meyer, G.E.; Von Bargen, K.; Mortensen, D.A. Color indices for weed identification under various soil, residue, and lighting conditions. Trans. ASAE USA 1995, 38, 259–269. [Google Scholar] [CrossRef]
- Reid, A.M.; Chapman, W.K.; Prescott, C.E.; Nijland, W. Using excess greenness and green chromatic coordinate colour indices from aerial images to assess lodgepole pine vigour, mortality and disease occurrence. For. Ecol. Manag. 2016, 374, 146–153. [Google Scholar] [CrossRef]
- Sonnentag, O.; Hufkens, K.; Teshera-Sterne, C.; Young, A.M.; Friedl, M.; Braswell, B.H.; Milliman, T.; O’Keefe, J.; Richardson, A.D. Digital repeat photography for phenological research in forest ecosystems. Agric. For. Meteorol. 2012, 152, 159–177. [Google Scholar] [CrossRef]
- van Rijsbergen, C.J. Information Retrieval, 2nd ed.; Butterworths: London, UK, 1979. [Google Scholar]
- Falkowski, M.J.; Hudak, A.T.; Crookston, N.L.; Gessler, P.E.; Uebler, E.H.; Smith, A.M.S. Landscape-scale parameterization of a tree-level forest growth model: A k-nearest neighbor imputation approach incorporating LiDAR data. Can. J. For. Res. 2010, 40, 184–199. [Google Scholar] [CrossRef]
- Hudak, A.T.; Crookston, N.L.; Evans, J.S.; Falkowski, M.J.; Smith, A.M.S.; Gessler, P.E.; Morgan, P. Regression modeling and mapping of coniferous forest basal area and tree density from discrete-return lidar and multispectral satellite data. Can. J. Remote Sens. 2006, 32, 126–138. [Google Scholar] [CrossRef]
- Tinkham, W.T.; Huang, H.; Smith, A.M.S.; Shrestha, R.; Falkowski, M.J.; Hudak, A.T.; Link, T.E.; Glenn, N.F.; Marks, D.G. A Comparison of Two Open Source LiDAR Surface Classification Algorithms. Remote Sens. 2011, 3, 638–649. [Google Scholar] [CrossRef]
- Podobnikar, T.; Vrečko, A. Digital Elevation Model from the Best Results of Different Filtering of a LiDAR Point Cloud. Trans. GIS 2012, 16, 603–617. [Google Scholar] [CrossRef]
- Montealegre, A.L.; Lamelas, M.T.; Riva, J. de la A Comparison of Open-Source LiDAR Filtering Algorithms in a Mediterranean Forest Environment. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 4072–4085. [Google Scholar] [CrossRef]
- Mafanya, M.; Tsele, P.; Botai, J.; Manyama, P.; Swart, B.; Monate, T. Evaluating pixel and object based image classification techniques for mapping plant invasions from UAV derived aerial imagery: Harrisia pomanensis as a case study. ISPRS J. Photogramm. Remote Sens. 2017, 129, 1–11. [Google Scholar] [CrossRef]
- Government of Canada. Flying Your Drone Safely and Legally. Available online: https://www.tc.gc.ca/eng/civilaviation/opssvs/flying-drone-safely-legally.html (accessed on 14 August 2017).
- Ouimet, C.A. (Sepaq, QC, Canada). Personal communication, 2017.





| Area | Reference Data | ||||||
|---|---|---|---|---|---|---|---|
| Classification | WL | N-WL | Total | F1 Score | TPR | FPR | |
| Area 1 | WL | 79 | 25 | 104 | 0.69 | ||
| N-WL | 45 | 251 | 296 | 0.64 | 0.09 | ||
| Total | 124 | 276 | 400 | ||||
| Area 2 | WL | 33 | 14 | 47 | 0.76 | ||
| N-WL | 7 | 346 | 353 | 0.83 | 0.04 | ||
| Total | 40 | 360 | 400 | ||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leduc, M.-B.; Knudby, A.J. Mapping Wild Leek through the Forest Canopy Using a UAV. Remote Sens. 2018, 10, 70. https://doi.org/10.3390/rs10010070
Leduc M-B, Knudby AJ. Mapping Wild Leek through the Forest Canopy Using a UAV. Remote Sensing. 2018; 10(1):70. https://doi.org/10.3390/rs10010070
Chicago/Turabian StyleLeduc, Marie-Bé, and Anders J. Knudby. 2018. "Mapping Wild Leek through the Forest Canopy Using a UAV" Remote Sensing 10, no. 1: 70. https://doi.org/10.3390/rs10010070
APA StyleLeduc, M.-B., & Knudby, A. J. (2018). Mapping Wild Leek through the Forest Canopy Using a UAV. Remote Sensing, 10(1), 70. https://doi.org/10.3390/rs10010070