Remote Sensing and Mapping of Tamarisk along the Colorado River, USA: A Comparative Use of Summer-Acquired Hyperion, Thematic Mapper and QuickBird Data
Abstract
:1. Introduction
2. Methods
2.1. Study Area
2.2. Remote Sensing
2.3. Georectification and Reflectance Calibration
2.4. Image Classification
3. Results
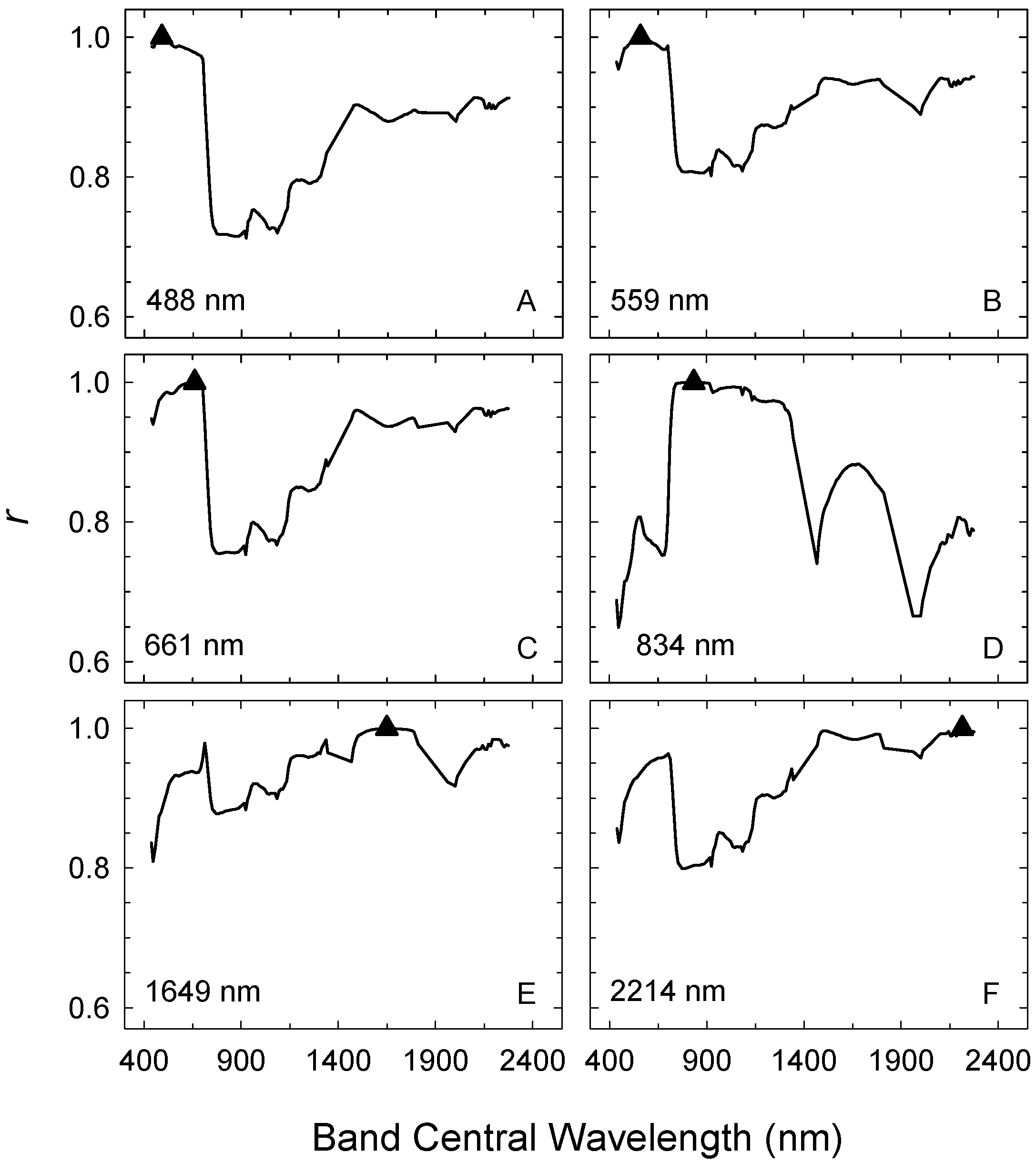
| QB or TM5 | Hyperion | |||||
|---|---|---|---|---|---|---|
| Band | Wavelength (nm) | Bandwidth (nm) | Band | Wavelength (nm) | Bandwidth (nm) | |
| 1 | 485 | 70 | 14 | 488 | 10 | |
| 2 | 560 | 80 | 21 | 559 | 10 | |
| 3 | 660 | 60 | 31 | 661 | 10 | |
| 4 | 830 | 140 | 48 | 834 | 10 | |
| TM5 | ||||||
| 5 | 1,650 | 200 | 150 | 1,649 | 10 | |
| 7 | 2,215 | 270 | 206 | 2,214 | 10 | |
| Hyperion Band | Central Wavelength (nm) | Correlated Wavelength Range (nm) |
|---|---|---|
| 14 (1) | 488 | 437–702 |
| 21 (2) | 559 | 437–712 |
| 31 (3) | 661 | 457–702 |
| 1,477–1,558 | ||
| 2,052–2,274 | ||
| 48 (4) | 834 | 722–1,326 |
| 150 (5) | 1,649 | 701–722 |
| 1,155–1,810 | ||
| 2,052–2,274 | ||
| 206 (7) | 2,214 | 610–722 |
| 1,467–2,274 |
| Band | Correlation Coefficient (r) | ||||
|---|---|---|---|---|---|
| TM5 or QB Band | |||||
| 1 | 2 | 3 | 4 | 5 | |
| TM5 | |||||
| 2 | 0.95 | ||||
| 3 | 0.94 | 0.94 | |||
| 4 | -0.14 | -0.09 | -0.14 | ||
| 5 | 0.74 | 0.79 | 0.81 | 0.13 | |
| 7 | 0.85 | 0.86 | 0.89 | -0.13 | 0.92 |
| QB | |||||
| 2 | 0.98 | ||||
| 3 | 0.96 | 0.98 | |||
| 4 | -0.05 | 0.01 | -0.05 | ||
| Hyperion Band | |||||
| 14 | 21 | 31 | 48 | 150 | |
| Hyperion | |||||
| 21 | 0.99 | ||||
| 31 | 0.98 | 0.99 | |||
| 48 | 0.72 | 0.81 | 0.76 | ||
| 150 | 0.88 | 0.93 | 0.94 | 0.88 | |
| 206 | 0.90 | 0.94 | 0.96 | 0.80 | 0.98 |
| Khat | ||||
|---|---|---|---|---|
| Algorithm | Hyperion, 2004 | TM5, 2004 | QB, 2004 | QB, 2005 |
| ML, 4-band | 0.18 | 0.18 | - | - |
| ML, 2-band | 0.23 | 0.10 | 0.69 | 0.66 |
| GNDVI | 0.36 | 0.00 | 0.73 | 0.72 |
| NDVI | 0.50 | -0.03 | 0.74 | 0.64 |
| Sensor | Algorithm | % | ||
|---|---|---|---|---|
| Accuracy | Omission Error | Commission Error | ||
| Hyperion | NDVI | 88 | 0 | 62 |
| TM5 | ML, 4-band | 80 | 40 | 83 |
| QB | NDVI | 91 | 18 | 78 |
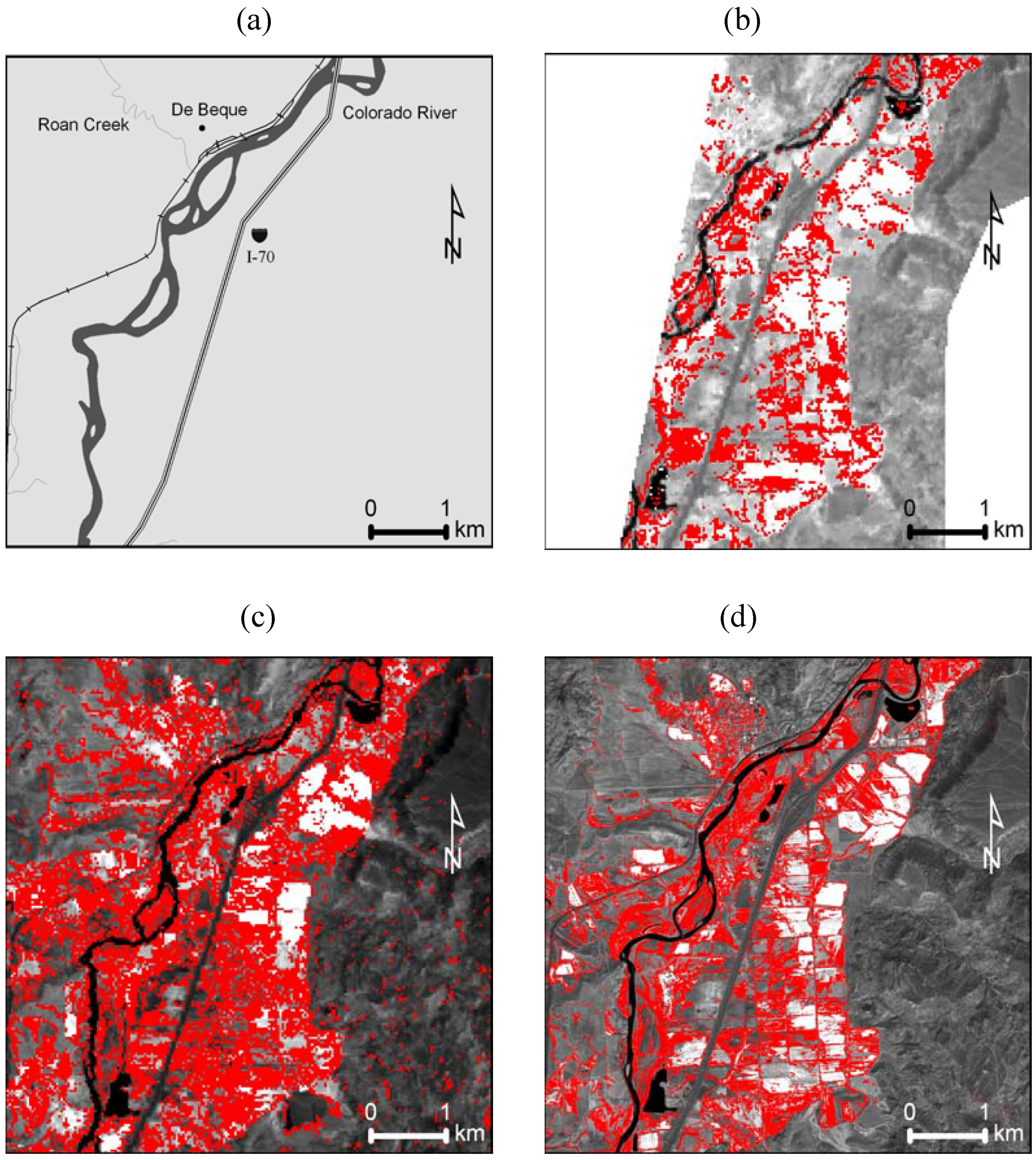
4. Discussion and Conclusions
Acknowledgements
References and Notes
- Sanders, N.J.; Gotelli, N.J.; Heller, N.E.; Gordon, D.M. Community disassembly by an invasive species. Proc. Natl. Acad. Sci.USA 2003, 100, 2474–2477. [Google Scholar] [CrossRef] [PubMed]
- Underwood, E.C.; Ustin, S.L.; DiPietro, D. Mapping nonnative plants using hyperspectral imagery. Remote Sens. Environ. 2003, 86, 150–161. [Google Scholar] [CrossRef]
- Pearce, C.M.; Smith, D.G. Saltcedar: distribution, abundance, and dispersal mechanisms, northern Montana, USA. Wetlands 2003, 23, 215–228. [Google Scholar] [CrossRef]
- Lovich, J.E. A brief review on the impacts of Tamarisk or Saltcedar on biodiversity in the New World. In Saltcedar Management and Riparian Restoration Workshop; Las Vegas, NV, USA, 1996. [Google Scholar]
- Robinson, T.W. Introduction, spread and areal extent of Saltcedar (Tamarix) in the western states. USGS Professional Paper, 1965; 491-A:1–491-A:12. [Google Scholar]
- Everitt, B.L. Ecology of saltcedar―a plea for research. Environ. Geol. 1980, 3, 77–84. [Google Scholar] [CrossRef]
- Sala, A.; Smith, S.D.; Devitt, D.A. Water use by Tamarix ramosissima and associated phreatophytes in a Mojave Dessert floodplain. Ecol. Appl. 1996, 6, 888–898. [Google Scholar] [CrossRef]
- Anderson, G.L.; Carruthers, R.I.; Shaokui, G.E.; Gong, P. Monitoring of invasive Tamarix distribution and effects of biological control with airborne hyperspectral remote sensing. Int. J. Remote Sens. 2005, 26, 2487–2489. [Google Scholar] [CrossRef]
- Everitt, J.H.; Escobar, D.E.; Alaniz, M.A.; Davis, M.R.; Richerson, J.V. Using spatial information technologies to map Chinese Tamarisk (Tamarix chinensis) infestations. Weed Sci. 1996, 44, 194–201. [Google Scholar]
- Lesica, P.; Miles, S. Ecological strategies for managing Tamarisk on the C. M. Russell National Wildlife Refuge, Montana, USA. Biol. Conserv. 2004, 119, 535–543. [Google Scholar] [CrossRef]
- Morisette, J.T.; Jarnevich, C.S.; Ullah, A.; Cai, W.; Pedelty, J.E.; Gentle, J.E.; Stohlgren, T.J.; Schnase, J.L. A tamarisk habitat suitability map for the continental United States. Front. Ecol. Environ. 2006, 4, 11–17. [Google Scholar] [CrossRef]
- Fang, H.; Xu, J. Land cover and vegetation change in the Yellow River Delta Nature Reserve analyzed with Landsat Thematic Mapper data. Geocarto Int. 2000, 15, 1–7. [Google Scholar] [CrossRef]
- Groeneveld, D.P.; Watson, R.P. Near-infrared discrimination of leafless saltcedar in wintertime Landsat TM. Int. J. Remote Sens. 2008, 29, 3577–3588. [Google Scholar] [CrossRef]
- Glenn, N.F.; Mundt, J.T.; Weber, K.T.; Prather, T.S.; Lass, L.W.; Pettingill, J. Hyperspectral data processing for repeat detection of small infestations of leafy spurge. Remote Sens. Environ. 2005, 95, 399–412. [Google Scholar] [CrossRef]
- Laes, D.; Maus, P.; Fisk, H.; Davern, T. Tamarisk remote sensing inventory and assessment. In Progress Report to the Remote Sensing Steering Committee; USDA Forest Service Engineering, Remote Sensing Applications Center: Salt Lake City, UT, USA, 2003; pp. 1–16. [Google Scholar]
- Hamada, Y.; Stow, D.A.; Coulter, L.L.; Jafolla, J.C.; Hendricks, L. W. Detecting Tamarisk species (Tamarix spp.) in riparian habitats of Southern California using high spatial resolution hyperspectral imagery. Remote Sens. Environ. 2007, 109, 237–248. [Google Scholar] [CrossRef]
- Pearlman, J.S.; Barry, P.S.; Segal, C.C.; Shepanski, J.; Beiso, D.; Carman, S.L. Hyperion, a space-based imaging spectrometer. IEEE T. Geosci. Remote 2003, 41, 1160–1173. [Google Scholar] [CrossRef]
- McGwire, K.M.; Schulz, B. Hyperspectral monitoring of invasive, non-native plant species with EO-1 Hyperion imagery. In Final Report, NASA EO-1 Science Team Grant NASA-NCC5-486; Goddard Space Flight Center: Greenbelt, MD, USA, 2002. [Google Scholar]
- Guarte, J.M.; Barrios, E.B. Estimation under purposive sampling. Commun. Stat.: Simul. C. 2006, 35, 277–284. [Google Scholar] [CrossRef]
- Mueller-Dombois, D.; Ellenberg, H. Aims and Methods of Vegetation Ecology; John Wiley & Sons: New York, NY, USA, 1974. [Google Scholar]
- Jensen, J.R. Introductory Digital Image Processing: A Remote Sensing Perspective; Prentice-Hall: Upper Saddle River, NJ, USA, 2005. [Google Scholar]
- Lillesand, T.M.; Kiefer, R.W.; Chipman, J.W. Remote Sensing and Image Interpretation; John Wiley & Sons: New York, NY, USA, 2008. [Google Scholar]
- Franklin, J.; Phinn, S.; Woodcock, C.; Rogan, J. Rationale and conceptual framework for classification approaches to assess forest resources and properties. In Remote Sensing of Forest Environments: Concepts and Case Studies; Wulder, M., Franklin, S., Eds.; Kluwer Academic Publishers: Boston, MA, USA, 2003; pp. 279–300. [Google Scholar]
- Rouse, J.; Hass, R.H.; Schell, J.A.; Deering, D.W. Monitoring vegetation systems in the Great Plains with ERTS. In Third Earth Resources Technology Satellite-1 Symposium; Greenbelt, MD, USA, NASA SP-351; 1974; pp. 3010–3317. [Google Scholar]
- Landis, J.; Koch, G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [PubMed]
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Carter, G.A.; Lucas, K.L.; Blossom, G.A.; Lassitter, C.L.; Holiday, D.M.; Mooneyhan, D.S.; Fastring, D.R.; Holcombe, T.R.; Griffith, J.A. Remote Sensing and Mapping of Tamarisk along the Colorado River, USA: A Comparative Use of Summer-Acquired Hyperion, Thematic Mapper and QuickBird Data. Remote Sens. 2009, 1, 318-329. https://doi.org/10.3390/rs1030318
Carter GA, Lucas KL, Blossom GA, Lassitter CL, Holiday DM, Mooneyhan DS, Fastring DR, Holcombe TR, Griffith JA. Remote Sensing and Mapping of Tamarisk along the Colorado River, USA: A Comparative Use of Summer-Acquired Hyperion, Thematic Mapper and QuickBird Data. Remote Sensing. 2009; 1(3):318-329. https://doi.org/10.3390/rs1030318
Chicago/Turabian StyleCarter, Gregory A., Kelly L. Lucas, Gabriel A. Blossom, Cheryl L. Lassitter, Dan M. Holiday, David S. Mooneyhan, Danielle R. Fastring, Tracy R. Holcombe, and Jerry A. Griffith. 2009. "Remote Sensing and Mapping of Tamarisk along the Colorado River, USA: A Comparative Use of Summer-Acquired Hyperion, Thematic Mapper and QuickBird Data" Remote Sensing 1, no. 3: 318-329. https://doi.org/10.3390/rs1030318
APA StyleCarter, G. A., Lucas, K. L., Blossom, G. A., Lassitter, C. L., Holiday, D. M., Mooneyhan, D. S., Fastring, D. R., Holcombe, T. R., & Griffith, J. A. (2009). Remote Sensing and Mapping of Tamarisk along the Colorado River, USA: A Comparative Use of Summer-Acquired Hyperion, Thematic Mapper and QuickBird Data. Remote Sensing, 1(3), 318-329. https://doi.org/10.3390/rs1030318





