Parameter Estimation of the Farquhar—von Caemmerer—Berry Biochemical Model from Photosynthetic Carbon Dioxide Response Curves
Abstract
1. Introduction
2. Materials and Methods
2.1. The FvCB Model and Characteristics
2.2. Data Generation
2.2.1. Constraints for the Parameters
2.2.2. Criteria for Deriving Parameters
2.2.3. Generation of Datasets
- (i)
- Datasets with varied data points. The accuracy of this dataset was eight decimal places; and the numbers of data points were 4, 5, 6, 7, 8, 9, and 12. The varied data point dataset was used to evaluate the impact of the number of data points on parameterization. It should be noted that these datasets are included in the high accuracy dataset.
- (ii)
- Datasets with varied accuracy. These datasets were with either eight or 12 data points, and accuracies were from one to eight decimal places. The varied accuracy dataset was used to identify the impact of accuracy on parameterization.
2.3. Fitting Methods
2.3.1. Gu et al.’s Method
- (1)
- The enumeration of all possible data point distributions of three states of a given dataset. The three limited states must follow a certain pattern along the Ci axis in an order dictated by the FvCB model. The minimum numbers of data points (3 or 0, 2 or 0, 3 or 0) and the number of data points higher than the number of parameters to be derived are required for resolvable parameters. Under these conditions, the resolvable parameters are defined as Gu et al.’s resolvable parameters to differentiate them from resolvable parameters as a general case. Thus, the minimum numbers of data points (3, 2, 3) and a minimum number of nine observed data points are required for all eight parameters to be resolvable. We refer to these as Gu et al.’s requirements for all eight resolvable parameters. If only one or two states are Gu et al.’s resolvable states, the dataset is Gu et al.’s partially resolvable dataset and the resolvable parameters are Gu et al.’s partially resolvable parameters. If a state does not meet the minimum data point requirement of Gu et al., Gu et al.’s method forces the dataset to meet the requirements by moving data points from one state to another. If the number of observed data points is zero in the Ap state, α = 0 and Tp = (asymptote of An + Rd)/three.
- (2)
- Fitting the FvCB model to each limited state distribution separately. In this step, the transition points are never calculated and the carboxylation rates in different states are never compared. The An is calculated with the submodel of the limited state to which the data point is assigned.
- (3)
- Detection and correction of inadmissible fits. Gu et al., [2] defined “inadmissible fits” as cases where the limitation states of the points in the A/Ci curve have not consistently or correctly identified the derived parameters. This step is only used for a dataset that contains multiple limited states. If the calculated limited state distribution is the same as the assigned limited state distribution, then the fit is admissible; otherwise, the fit is inadmissible. If the fit is inadmissible, the fit will be corrected via a penalization strategy.
- (4)
- Section of best fit. The best fit for an observed set of data is the method that gives the smallest value for the minimized objective function. If the values of the minimized objective function are equal when comparing across different limited state distributions, the one with fewer parameters is selected.
2.3.2. Sharkey et al.’s Method
2.3.3. Parameter Calculations
3. Results
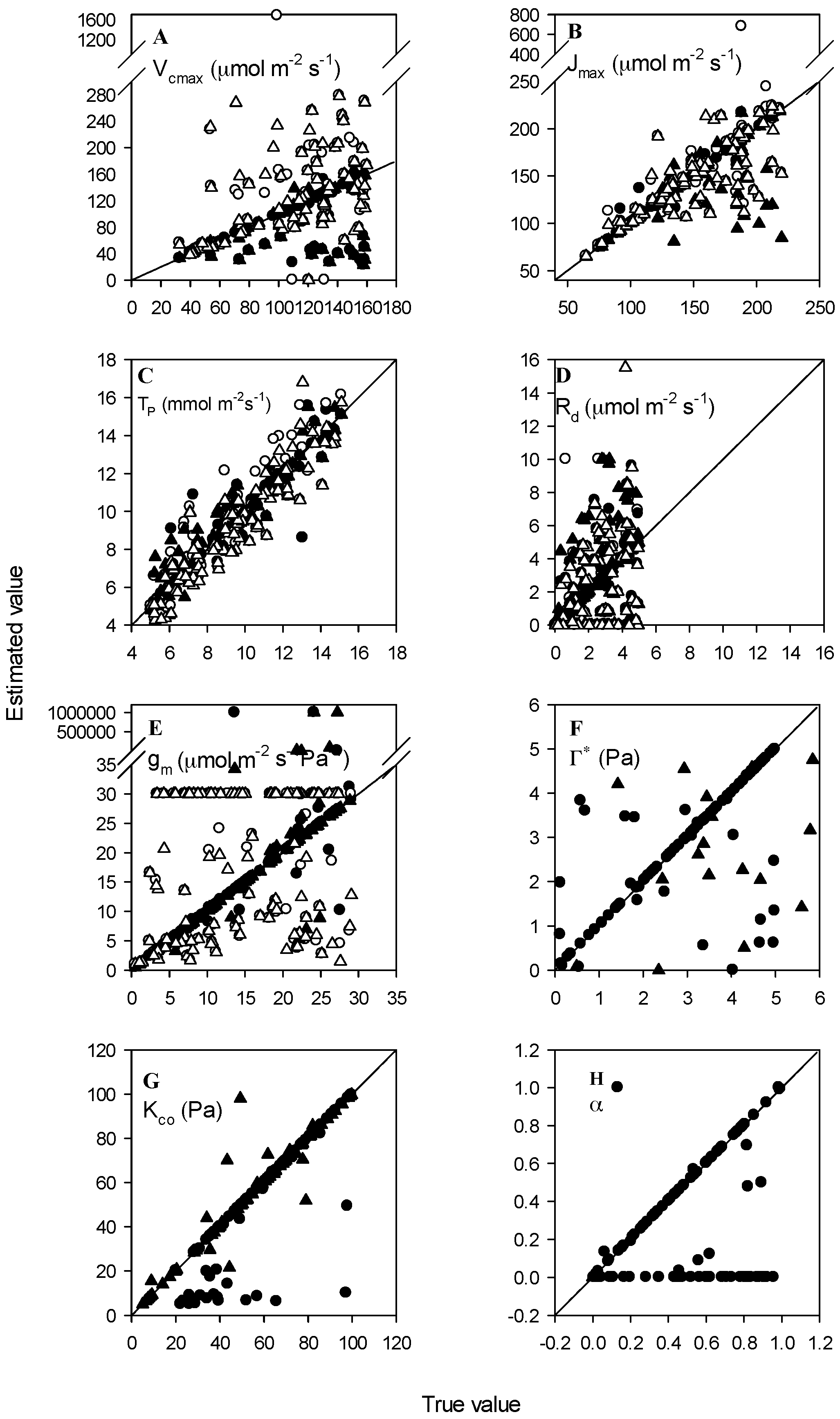
3.1. High Accuracy Dataset
3.2. Normal Accuracy Datasets
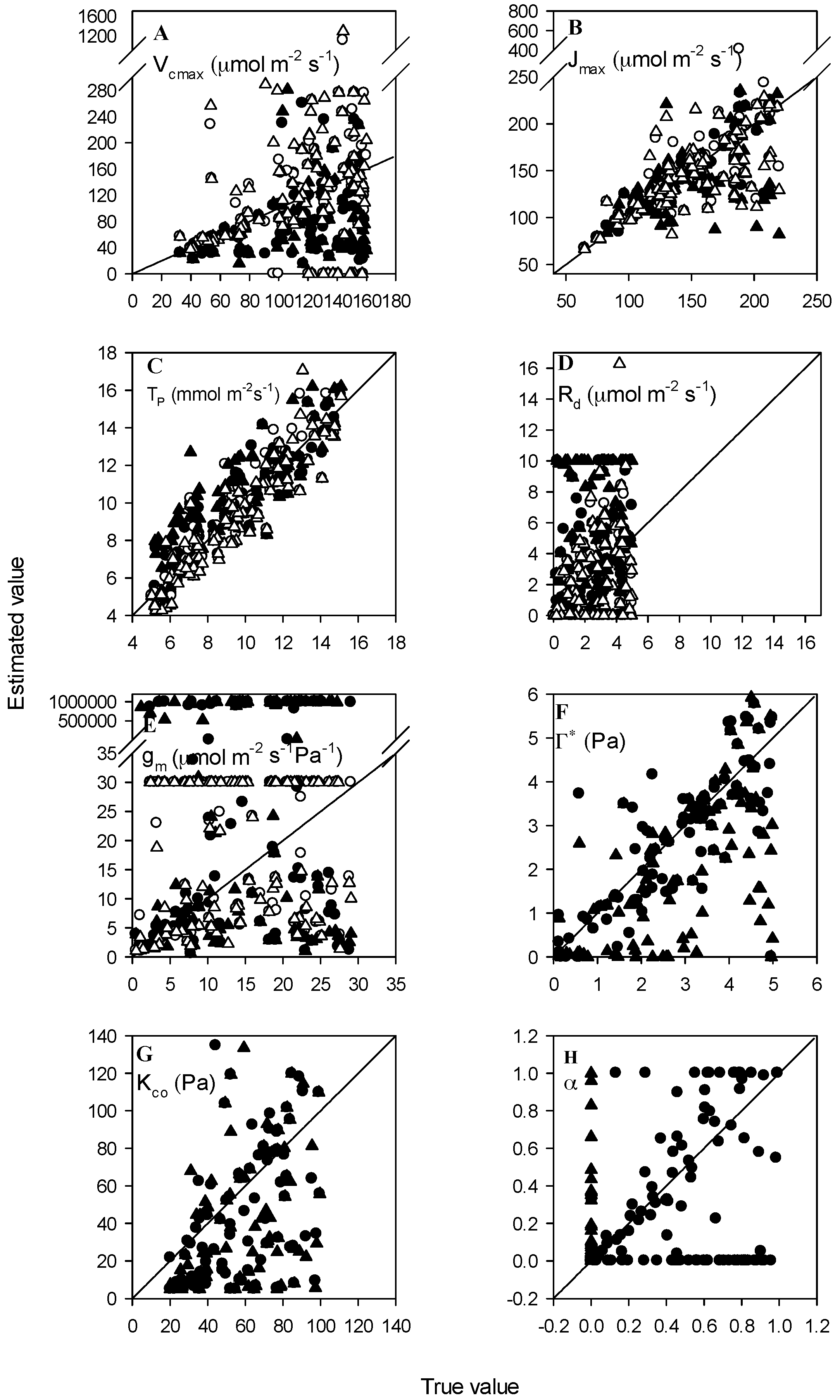
3.3. Datasets with Measurement Errors
3.4. Datasets with Varied Data Points
3.5. Datasets with Varied Accuracy
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Nomenclature
| Ac | Net assimilation rate assuming Rubisco-limited state, μmol m−2 s−1 |
| Acmi | Measured net assimilation rates in the Ac state point i, μmol m−2 s−1 |
| Ajj | Calculated net assimilation rates in the Aj state point j, μmol m−2 s−1 |
| An | Net assimilation rate, μmol m−2 s−1 |
| Anj | Net assimilation rate at Aj state, μmol m−2 s−1 |
| Ap | Net assimilation rate assuming triose phosphate utilization (TPU) limited state, μmol m−2 s−1 |
| Apk | Calculated net assimilation rates in the Ap state point k, μmol m−2 s−1 |
| Cc | Chloroplastic CO2 partial pressure, Pa |
| Ci | Intercellular CO2 partial pressure, Pa |
| gm | Internal (mesophyll) conductance to CO2 transport, μmol m−2 s−1 Pa−1 |
| Jmax | Maximum rate of electron transport, μmol m−2 s−1 |
| Ko | Michaelis–Menten constant for Rubisco for O2, Pa |
| Rd | Day respiration, μmol m−2 s−1 |
| Tp | Rate of triose phosphate export from the chloroplast, μmol m−2 s−1 |
| Wc | Maximal Rubisco carboxylation rate, μmol m−2 s−1 |
| Wp | TPU-limited carboxylation rate, μmol m−2 s−1 |
| Aci | Calculated net assimilation rates in the Ac state point i, μmol m−2 s−1 |
| Aj | Net assimilation rate assuming RuBP regeneration limited state, μmol m−2 s−1 |
| Ajmj | Measured net assimilation rates in the Aj state point j, μmol m−2 s−1 |
| Anc | Net assimilation rate at Ac state, μmol m−2 s−1 |
| Anm | Measured net assimilation rate, μmol m−2 s−1 |
| Γ* | Chloroplastic CO2 photocompensation point, Pa |
| Apmk | Measured net assimilation rates in the Ap state point k, μmol m−2 s−1 |
| Ccc_cj | Chloroplastic CO2 partial pressure at transition point between Ac and Aj state, Pa |
| Cs | Intercellular CO2 partial pressure, Pa |
| J | Potential electron transport rate at the measurement light level, μmol m−2 s−1 |
| Kc | Michaelis-Menten constant for Rubisco for CO2, Pa |
| Kco | Effective Michaelis–Menten coefficient for CO2, Kco = Kc(1 + O/Ko), Pa |
| O | Oxygen partial pressure, Pa |
| Vcmax | Maximal Rubisco carboxylation rate, μmol m−2 s−1 |
| Wj | RuBP regeneration-limited carboxylation rate, μmol m−2 s−1 |
| α | Non-returned fraction of the glycolate carbon recycled in the photorespiratory cycle (dimensionless) |
Appendix A. Errors Superimposed to an Ideal Dataset
References
- Farquhar, G.D.; Von Caemmerer, S.; Berry, J.A. A biochemical model of photosynthetic CO2 assimilation in leaves of C3 species. Planta 1980, 149, 78–90. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Pallardy, S.G.; Tu, K.; Law, B.E.; Wullschleger, S.D. Reliable estimation of biochemical parameters from C3 leaf photosynthesis-intercellular carbon dioxide response curves. Plant Cell Environ. 2010, 33, 1852–1874. [Google Scholar] [CrossRef] [PubMed]
- Long, S.P. Modification of the response of photosynthetic productivity to rising temperature by atmospheric CO2 concentrations: Has its importance been underestimated? Plant Cell Environ. 1991, 14, 729–739. [Google Scholar] [CrossRef]
- Amthor, J. Scaling CO2-photosynthesis relationships from the leaf to the canopy. Photosynth. Res. 1994, 39, 321–350. [Google Scholar] [CrossRef] [PubMed]
- Field, C.B.; Avissar, R. Bidirectional interactions between the biosphere and the atmosphere—Introduction. Glob. Chang. Biol. 1998, 4, 459–460. [Google Scholar] [CrossRef]
- Jones, J.W.; Hoogenboom, G.; Porter, C.H.; Boote, K.J.; Batchelor, W.D.; Hunt, L.A.; Wilkens, P.W.; Singh, U.; Gijsman, A.J.; Ritchie, J.T. The DSSAT Cropping System Model. Eur. J. Agron. 2003, 18, 235–265. [Google Scholar] [CrossRef]
- De Pury, D.G.G.; Farquhar, D.G. Simple scaling of photosynthesis from leaves to canopy without the errors of bigleaf models. Plant Cell Environ. 1997, 20, 537–557. [Google Scholar] [CrossRef]
- Sellers, P.J.; Bounoua, L.; Collatz, G.J.; Randall, D.A.; Dazlich, D.A.; Los, S.O; Berry, J.A.; Fung, I.; Tucker, C.J.; Field, C.B.; et al. Comparison of radiative and physiological effects of doubled atmospheric CO2 on climate. Science 1996, 271, 1402–1406. [Google Scholar] [CrossRef]
- Sellers, P.J.; Dickinson, R.E.; Randall, D.A.; Betts, A.K.; Hall, F.G.; Berry, J.A; Collatz, G.J.; Denning, A.S.; Mooney, H.A.; Nobre, C.A.; et al. Modeling the exchanges of energy, water, and carbon between continents and the atmosphere. Science 1997, 275, 502–509. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.P.; Leuning, R. A two-leaf model for canopy conductance, photosynthesis and portioning of available energy I: Model description and comparison with a multi-layered model. Agric. For. Meteorol. 1998, 91, 89–111. [Google Scholar] [CrossRef]
- Wittig, V.E.; Bernacchi, C.J.; Zhu, X.-G.; Calfapietra, C.; Ceulemans, R.; Deangelis, P.; Gielen, B.; Miglietta, F.; Morgan, P.B.; Long, S.P. Gross primary production is stimulated for three Populus species grown under free-air CO2 enrichment from planting through canopy closure. Glob. Chang. Biol. 2005, 11, 644–656. [Google Scholar] [CrossRef]
- Ethier, G.J.; Livingston, N.J. On the need to incorporate sensitivity to CO2 transfer conductance into the Farquhar-von Caemmerer-Berry leaf photosynthesis model. Plant Cell Environ. 2004, 27, 137–153. [Google Scholar] [CrossRef]
- Bunce, J.A. Acclimation of photosynthesis to temperature in eight cool and warm climate herbaceous C3 species: Temperature dependence of parameters of a biochemical photosynthesis model. Photosynth. Res. 2000, 63, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Harley, P.C.; Loreto, F.; Di Marco, G.; Sharkey, T.D. Theoretical consideration when estimating the mesophyll conductance to CO2 flux by analysis of the response of Photosynthesis to CO2. Plant Physiol. 1992, 98, 1429–1436. [Google Scholar] [CrossRef] [PubMed]
- Harley, P.C.; Thomas, R.B.; Reynolds, J.F.; Strain, B.R. Modelling photosynthesis of cotton grown in elevated CO2. Plant Cell Environ. 1992, 15, 271–282. [Google Scholar] [CrossRef]
- Hikosaka, K.; Ishikawa, K.; Borjigidai, A.; Muller, O.; Onoda, Y. Temperature acclimation of photosynthesis: Mechanisms involved in the changes in temperature dependence of photosynthetic rate. J. Exp. Bot. 2006, 57, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Possell, M.; Hewitt, C.N. Gas exchange and photosynthetic performance of the tropical tree Acacia nigrescens when grown in different CO2 concentrations. Planta 2009, 229, 837–846. [Google Scholar] [CrossRef] [PubMed]
- Von Caemmerer, S.; Berry, J.; Farquhar, G.D. Biochemical model of C3 photosynthesis. In Photosynthesis in Silico: Understanding Complexity from Molecules to Ecosystems; Laisk, A., NedbalGovindjee, L., Eds.; Springer Science + Business Media, B.V.: Dordrecht, The Netherlands, 2009; pp. 209–230. [Google Scholar]
- Bernacchi, C.J.; Bagley, J.E.; Serbin, S.P.; Ruiz-Vera, U.M.; Rosenthal, D.M.; Vanloocke, A. Modelling C3 photosynthesis from the chloroplast to the ecosystem. Plant Cell Environ. 2013, 36, 1641–1657. [Google Scholar] [CrossRef] [PubMed]
- Farquhar, G.D.; von Caemmerer, S. Modelling of photosynthetic responses to environmental conditions. In Physiological Plant Ecology II. Encyclopedia of Plant Physiology; Lange, O.L., Nobel, P.S., Osmond, C.B., Ziegler, H., Eds.; New Series; Springer: Heidelberg, Germany, 1982; Volume 12B, pp. 550–587. [Google Scholar]
- Sharkey, T.D.; Berry, J.A.; Raschke, K. Starch and sucrose synthesis in Phaseolus vulgaris as affected by light, CO2, and abscisic acid. Plant Physiol. 1985, 77, 617–620. [Google Scholar] [CrossRef] [PubMed]
- Von Caemmerer, S. Biochemical Models of Leaf Photosynthesis; Techniques in Plant Sciences No. 2; CSIRO Publishing: Collingwood, Australia, 2000. [Google Scholar]
- Von Caemmerer, S.; Farquhar, G.D. Some relationships between the biochemistry of photosynthesis and the gas exchange of leaves. Planta 1981, 153, 376–387. [Google Scholar] [CrossRef] [PubMed]
- Dubois, J.J.B.; Fiscus, E.L.; Booker, F.L.; Flowers, M.D.; Reid, C.D. Optimizing the statistical estimation of the parameters of the Farquhar-von Caemmerer-Berry model of photosynthesis. New Phytol. 2007, 176, 402–414. [Google Scholar] [CrossRef] [PubMed]
- Manter, D.K.; Kerrigan, J. A/Ci curve analysis across a range of woody plant species: Influence of regression analysis parameters andmesophyll conductance. J. Exp. Bot. 2004, 55, 2581–2588. [Google Scholar] [CrossRef] [PubMed]
- Miao, Z.W.; Xu, M.; Lathrop, R.G.; Wang, Y. Comparison of the A-Cc curve fitting methods in determining maximum ribulose 1.5-bisphosphate carboxylase/oxygenase carboxylation rate, potential light saturated electron transport rate and leaf dark respiration. Plant Cell Environ. 2009, 32, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Sharkey, T.D.; Bernacchi, C.J.; Farquhar, G.D.; Singsaas, E.L. Fitting photosynthetic carbon dioxide response curves for C-3 leaves. Plant Cell Environ. 2007, 30, 1035–1040. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.H.; Zhu, G.F.; Miao, Z.W.; Feng, Q.; Chang, Z. Estimation of parameters of a biochemically based model of photosynthesis using a genetic algorithm. Plant Cell Environ. 2009, 32, 1710–1723. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Struik, P.C. Theoretical reconsiderations when estimating the mesophyll conductance to CO2 diffusion in leaves of C3 plants by analysis of combined gas exchange and chlorophyll fluorescence measurements. Plant Cell Environ. 2009, 32, 1513–1524. [Google Scholar] [CrossRef] [PubMed]
- Using the Li-6400/Li-6400XT Portable Photosynthesis System, version 6; Li-Cor, Inc.: Lincoln, NE, USA, 2008.
- Flexas, J.; Ribas-Carbó, M.; Díaz-Espejo, A.; Galmés, J.; Medrano, H. Mesophyll conductance to CO2: Current knowledge and future prospects. Plant Cell Environ. 2008, 31, 602–621. [Google Scholar] [CrossRef] [PubMed]
- Hanson, P.J.; Amthor, J.S.; Wullschleger, S.D.; Wilson, K.B.; Grant, R.F.; Hartley, A.; Hui, D.; Hunt, E.R., Jr.; Johnson, D.W.; Kimballet, J.S.; et al. Oak forest carbon and water simulations: Model intercomparisons and evaluations against independent data. Ecol. Monogr. 2004, 74, 443–489. [Google Scholar] [CrossRef]
- Harley, P.C.; Baldocchi, D.D. Scaling carbon dioxide and water vapour exchange from leaf to canopy in a deciduous forest. I. leaf model parameterization. Plant Cell Environ. 1995, 18, 1146–1156. [Google Scholar] [CrossRef]
- Wang, Q.; Fleisher, D.; Reddy, V.R.; Timlin, D.; Chun, J.A. Quantifying the Measurement Errors in a Potable Open Gas Exchange System and Their Effects on the Parameterization of Farquhar et al. model for C3 Leaves. Photosynthetica 2012, 50, 22–238. [Google Scholar] [CrossRef]

| α | Dataset | Method | Number of Parameters | gm | Γ* | Rd | Vcmax | Kco | Jmax | Tp | α |
|---|---|---|---|---|---|---|---|---|---|---|---|
| α > 0 | HDS | Gu et al. | Resolvable a | 100 | 90 | 90 | 71 | 80 | 72 | 61 | 61 |
| Correctly estimated | 72 | 62 | 62 | 52 | 62 | 49 | 42 | 42 | |||
| Total estimated | 100 | 100 | 100 | 91 | 91 | 74 | 100 | 100 | |||
| Error within ±10% | 89 | 80 | 75 | 71 | 70 | 69 | 89 | 55 | |||
| Sharkey et al. | Resolvable a | 100 | NA | 100 | 88 | NA | 79 | 72 | NA | ||
| Correctly estimated b | 0 | NA | 0 | 0 | NA | 0 | 0 | NA | |||
| Total estimated d | 100 | NA | 100 | 100 | NA | 100 | 100 | NA | |||
| Error within ±10% | 41 | NA | 6 | 22 | NA | 65 | 7 | NA | |||
| NDS | Gu et al. | Correctly estimated e | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Total estimated | 100 | 100 | 100 | 98 | 98 | 76 | 100 | 100 | |||
| Error within ±10% | 26 | 48 | 16 | 55 | 40 | 65 | 70 | 26 | |||
| Sharkey et al. | Error within ±10% | 6 | NA | 6 | 24 | NA | 70 | 57 | NA | ||
| DSE | Gu et al. | Total estimated | 100 | 100 | 100 | 98 | 98 | 86 | 100 | 100 | |
| Error within ±10% | 7 | 36 | 7 | 30 | 10 | 61 | 64 | 23 | |||
| Sharkey et al. | Error within ±10% | 8 | NA | 5 | 24 | NA | 70 | 60 | NA | ||
| α = 0 | HDS | Gu et al. | Resolvable a | 98 | 62 | 62 | 44 | 72 | 62 | 45 | 45 |
| Correctly estimated | 72 | 46 | 46 | 37 | 63 | 46 | 30 | 30 | |||
| Total estimated | 100 | 100 | 100 | 86 | 86 | 92 | 100 | 100 | |||
| Error within ±10% | 82 | 61 | 57 | 64 | 65 | 68 | 74 | 100 | |||
| Sharkey et al. | Resolvable a | 100 | NA | 100 | 88 | NA | 95 | 79 | NA | ||
| Error within ±10% | 63 | NA | 5 | 23 | NA | 64 | 8 | NA | |||
| NDS | Gu et al. | Total estimated | 100 | 100 | 100 | 93 | 93 | 90 | 100 | 100 | |
| Error within ±10% | 20 | 42 | 12 | 43 | 27 | 64 | 67 | 99 | |||
| Sharkey et al. | Error within ±10% | 8 | NA | 8 | 22 | NA | 59 | 51 | NA | ||
| DSE | Gu et al. | Total estimated | 100 | 100 | 100 | 98 | 98 | 93 | 100 | 100 | |
| Error within ±10% | 5 | 23 | 8 | 22 | 7 | 50 | 51 | 78 | |||
| Sharkey et al. | Error within ±10% | 4 | NA | 7 | 26 | NA | 61 | 63 | NA |
| Number of Data Points | Number of Datasets | Methods | Number of Values | gm | Γ* | Rd | Vcmax | Kco | Jmax | Tp | α |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 4 | 15 | Gu et al. | Resolvable | 3 | 1 | 1 | 0 | 1 | 0 | 0 | 0 |
| Correctly estimated | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | |||
| Total estimated | 15 | 15 | 15 | 2 | 2 | 12 | 15 | 15 | |||
| Sharkey et al. | Resolvable | 7 | NA | 7 | 5 | NA | 5 | 2 | NA | ||
| Correctly estimated a | 0 | 0 | 0 | NA | 0 | 0 | NA | NA | |||
| Total estimated | 15 | NA | 15 | 11 | NA | 13 | 10 | NA | |||
| 5 | 21 | Gu et al. | Resolvable | 9 | 3 | 3 | 0 | 3 | 3 | 0 | 0 |
| Correctly estimated | 3 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | |||
| Total estimated | 21 | 21 | 21 | 10 | 10 | 19 | 21 | 21 | |||
| Sharkey et al. | Resolvable | 13 | NA | 13 | 10 | NA | 10 | 5 | NA | ||
| Total estimated | 21 | NA | 21 | 21 | 21 | NA | 19 | NA | |||
| 6 | 28 | Gu et al. | Resolvable | 20 | 10 | 10 | 3 | 8 | 10 | 3 | 3 |
| Correctly estimated | 4 | 3 | 3 | 2 | 3 | 3 | 0 | 0 | |||
| Total estimated | 28 | 28 | 28 | 16 | 16 | 25 | 28 | 28 | |||
| Sharkey et al. | Resolvable | 21 | NA | 21 | 17 | NA | 17 | 14 | NA | ||
| Total estimated | 28 | NA | 28 | 27 | NA | 27 | 27 | NA | |||
| 7 | 36 | Gu et al. | Resolvable | 32 | 20 | 20 | 9 | 15 | 19 | 8 | 8 |
| Correctly estimated | 7 | 5 | 5 | 3 | 4 | 5 | 1 | 1 | |||
| Total estimated | 36 | 36 | 36 | 23 | 23 | 31 | 36 | 36 | |||
| Sharkey et al. | Resolvable | 28 | NA | 28 | 23 | NA | 23 | 20 | NA | ||
| Total estimated | 28 | NA | 28 | 28 | NA | 28 | 27 | NA | |||
| 8 | 45 | Gu et al. | Resolvable | 45 | 33 | 33 | 20 | 26 | 26 | 20 | 20 |
| Correctly estimated | 7 | 6 | 6 | 4 | 4 | 5 | 3 | 3 | |||
| Total estimated | 45 | 45 | 45 | 31 | 31 | 40 | 45 | 45 | |||
| Sharkey et al. | Resolvable | 37 | NA | 37 | 31 | NA | 31 | 28 | NA | ||
| Total estimated | 45 | NA | 44 | 45 | NA | 45 | 45 | NA | |||
| 9 | 55 | Gu et al. | Resolvable | 55 | 43 | 43 | 28 | 34 | 39 | 28 | 28 |
| Correctly estimated | 15 | 13 | 13 | 10 | 11 | 11 | 8 | 8 | |||
| Total estimated | 55 | 55 | 55 | 37 | 37 | 49 | 55 | 55 | |||
| Sharkey et al. | Resolvable | 47 | NA | 47 | 40 | NA | 40 | 37 | NA | ||
| Total estimated | 55 | NA | 55 | 53 | NA | 55 | 55 | NA | |||
| 12 | 73 | Gu et al. | Resolvable | 73 | 71 | 71 | 56 | 57 | 64 | 56 | 56 |
| Correctly estimated | 30 | 30 | 30 | 25 | 25 | 26 | 24 | 24 | |||
| Total estimated | 73 | 73 | 73 | 61 | 61 | 67 | 73 | 73 | |||
| Sharkey et al. | Resolvable | 72 | NA | 72 | 64 | NA | 65 | 65 | NA | ||
| Total estimated | 73 | NA | 73 | 73 | NA | 73 | 73 | NA |
| Ci (μmol mol−1) | Ajmj (μmol m−2 s−1) | AjjI (μmol m−2 s−1) | AjjII (μmol m−2 s−1) | Parameter/SSE | True Parameter | Estimated Parameter I | Estimated Parameter II |
|---|---|---|---|---|---|---|---|
| 373.56422385 | 20.52707868 | 20.53126975 | 20.52533184 | Jmax (μmol m−2 s−1) | 120.494 | 126.278 | 117.536 |
| 559.64521334 | 22.90931588 | 22.90327060 | 22.91250945 | Rd (μmol m−2 s−1) | 1.674 | 3.014 | 1.039 |
| 672.56218564 | 23.76820169 | 23.76423592 | 23.7701254 | gm (μmol m−2 s−1Pa−1) | 9.564 | 30.000 | 7.170 |
| 909.96541253 | 24.92146362 | 24.92728744 | 24.91809557 | SSE (μmol m−2 s−1)2 | - | 0.000 | 0.000 |
| An | Ci | Parameter/SSE | True Value | I a | II b | III c |
|---|---|---|---|---|---|---|
| 2.17 | 38.9 | Vcmax (μmol m−2 s−1) | 99.4 | 108.9 | 108.9 | 112.0 |
| 4.47 | 58.1 | Jmax (μmol m−2 s−1) | 136.2 | 143.1 | 146.1 | 145.5 |
| 9.60 | 99.4 | Tp (μmol m−2 s−1) | - | 10.1 | 10.1 | |
| 15.3 | 150 | Rd (μmol m−2 s−1) | 1.1 | 0.0 | 0.0 | 0.0 |
| 20.6 | 201 | gm (μmol m−2 s−1 Pa−1) | 3.7 | 30.0 | 30.0 | 30.0 |
| 25.3 | 284 | Kco (Pa) | 42.9 | - | - | - |
| 27.0 | 371 | Γ* (Pa) | 1.9 | - | - | - |
| 28.2 | 415 | SSE | - | 9.939 | 7.315 | 7.266 |
| 29.4 | 552 | - | - | - | - | - |
| 30.1 | 673 | - | - | - | - | - |
| 30.1 | 730 | - | - | - | - | - |
| 30.6 | 908 | - | - | - | - | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Q.; Chun, J.A.; Fleisher, D.; Reddy, V.; Timlin, D.; Resop, J. Parameter Estimation of the Farquhar—von Caemmerer—Berry Biochemical Model from Photosynthetic Carbon Dioxide Response Curves. Sustainability 2017, 9, 1288. https://doi.org/10.3390/su9071288
Wang Q, Chun JA, Fleisher D, Reddy V, Timlin D, Resop J. Parameter Estimation of the Farquhar—von Caemmerer—Berry Biochemical Model from Photosynthetic Carbon Dioxide Response Curves. Sustainability. 2017; 9(7):1288. https://doi.org/10.3390/su9071288
Chicago/Turabian StyleWang, Qingguo, Jong Ahn Chun, David Fleisher, Vangimalla Reddy, Dennis Timlin, and Jonathan Resop. 2017. "Parameter Estimation of the Farquhar—von Caemmerer—Berry Biochemical Model from Photosynthetic Carbon Dioxide Response Curves" Sustainability 9, no. 7: 1288. https://doi.org/10.3390/su9071288
APA StyleWang, Q., Chun, J. A., Fleisher, D., Reddy, V., Timlin, D., & Resop, J. (2017). Parameter Estimation of the Farquhar—von Caemmerer—Berry Biochemical Model from Photosynthetic Carbon Dioxide Response Curves. Sustainability, 9(7), 1288. https://doi.org/10.3390/su9071288







