Bioremediation of Heavy Metals from Soil and Aquatic Environment: An Overview of Principles and Criteria of Fundamental Processes
Abstract
:1. Introduction
| Heavy Metal | EPA Regulatory Limit (ppm) [9] | Toxic Effects | Ref. |
|---|---|---|---|
| Ag | 0.10 | Exposure may cause skin and other body tissues to turn gray or blue-gray, breathing problems, lung and throat irritation and stomach pain. | [10] |
| As | 0.01 | Affects essential cellular processes such asoxidative phosphorylation and ATP synthesis | [11] |
| Ba | 2.0 | Cause cardiac arrhythmias, respiratory failure, gastrointestinal dysfunction, muscle twitching and elevated blood pressure | [12] |
| Cd | 5.0 | Carcinogenic, mutagenic, endocrine disruptor, lung damage and fragile bones, affects calcium regulation in biological systems | [1,13] |
| Cr | 0.1 | Hair loss | [1] |
| Cu | 1.3 | Brain and kidney damage, elevated levels result in liver cirrhosis and chronic anemia, stomach and intestine irritation | [1,14] |
| Hg | 2.0 | Autoimmune diseases, depression, drowsiness, fatigue, hair loss, insomnia, loss of memory, restlessness, disturbance of vision, tremors, temper outbursts, brain damage, lung and kidney failure | [15,16,17] |
| Ni | 0.2 (WHO permissible limit) | Allergic skin diseases such as itching, cancer of the lungs, nose, sinuses, throat through continuous inhalation, immunotoxic, neurotoxic, genotoxic, affects fertility, hair loss | [1,18,19,20] |
| Pb | 15 | Excess exposure in children causes impaired development, reduced intelligence, short-term memory loss, disabilities in learning and coordination problems, risk of cardiovascular disease | [1,14,21] |
| Se | 50 | Dietary exposure of around 300 µg/day affects endocrine function, impairment of natural killer cells activity, hepatotoxicity and gastrointestinal disturbaces | [22] |
| Zn | 0.5 | Dizziness, fatigue etc. | [23] |
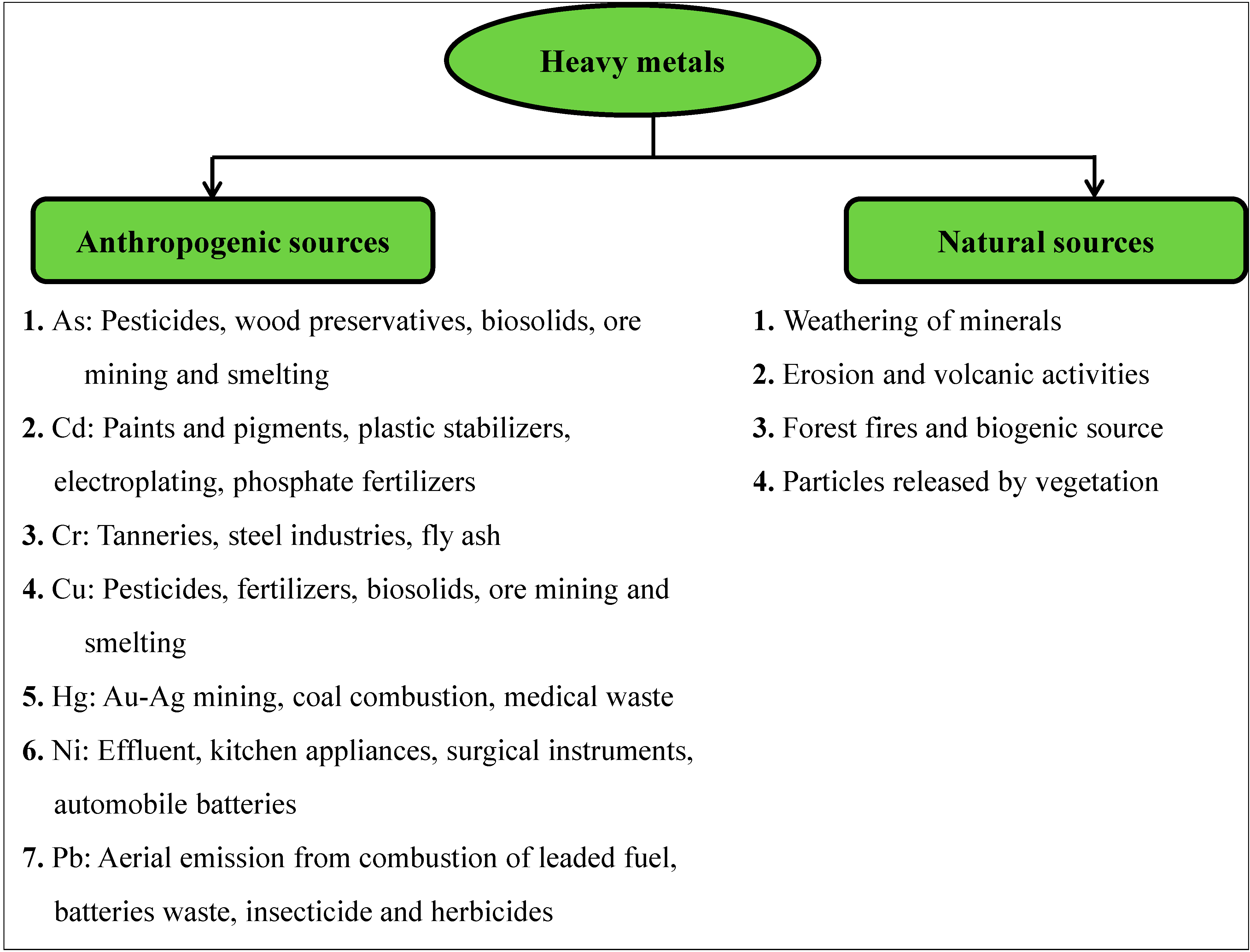
2. Sources of Heavy Metal in the Environment
3. Bioremediation: Introducing Microbe Based Clean Up System
3.1. Mechanisms of Bioremediation
3.2. Bioremediation by Adsorption
3.3. Bioremediation by Physio-Bio-Chemical Mechanism
3.4. Molecular Mechanisms Involved in Bioremediation Process
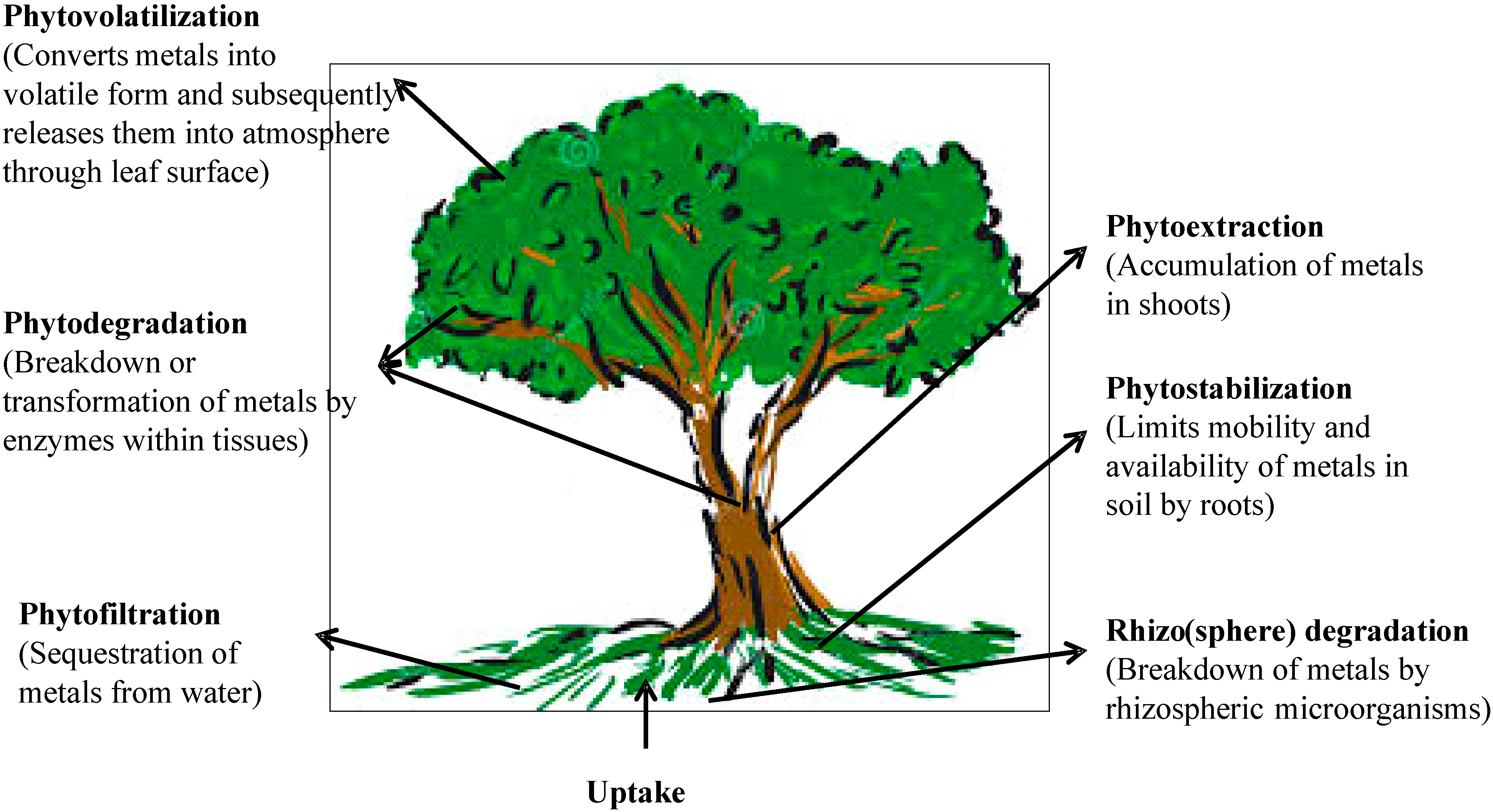
4. Phytoremediation
| Heavy Metal | Plant Species | Ref. |
|---|---|---|
| Cd, Cu, Pb, Zn | Salix spp. (Salix viminalis, Salix fragilis) | [110,111,112] |
| Cd | Castor (Ricinus communis) | [113] |
| Cd, Pb, Zn | Corn (Zea mays) | [114] |
| Cd, Cu, Pb, Zn | Populus spp. (Populus deltoides, Populus nigra, Populus trichocarpa) | [112] |
| Cd, Cu, Ni, Pb | Jatropha (Jatropha curcas L.) | [115,116] |
| Hg | Populus deltoides | [117] |
| Se | Brassica juncea, Astragalus bisulcatus | [118] |
| Zn | Populus canescens | [119] |
5. Biotechnological Intervention/Genetic Engineering in Bioremediation Processes
5.1. “Designer” Microbes Approach
| Heavy Metal | Initial Conc. (ppm) | Removal Efficiency (%) | Genetically Engineered Bacteria | Expressed Gene | Ref. |
|---|---|---|---|---|---|
| As | 0.05 | 100 | E. coli strain | Metalloregulatory protein ArsR | [127] |
| Cd2+ | - | - | E .coli strain | SpPCS | [84] |
| Cr6+ | 1.4–1000 | 100 | Methylococcus capsulatus | CrR | [128] |
| Cr | - | - | P. putida strain | Chromate reductase (ChrR) | [129] |
| Cd2+, Hg | - | - | Ralstonia eutropha CH34, Deinococcus radiodurans | merA | [82,130] |
| Hg | - | - | E. coli strain | Organomurcurial lyase | [131] |
| Hg | 7.4 | 96 | E. coli JM109 | Hg2+ transporter | [132] |
| Hg | - | - | Pseudomonas K-62 | Organomercurial lyase | [133] |
| Hg | - | - | Achromobacter sp AO22 | mer | [134] |
| Ni | 145 | 80 | P. fluorescens 4F39 | Phytochelatin synthase (PCS) | [135,136] |
5.2. Designer Plant Approach
5.3. Rhizosphere Engineering
5.4. Manipulation of Plant-Microbe Symbiosis
5.5. Application of Nano-Biotechnology
5.6. Application of Genomics
6. Future Prospects
Acknowledgments
Author Contributions
Abbreviations
| Ag | silver |
| As | arsenic |
| As(V) | arsenate |
| As(III) | arsenite |
| Au | gold |
| Ba | barium |
| Bi | bismuth |
| Cd | cadmium |
| Co | cobalt |
| Cr | chromium |
| Cu | copper |
| Fe | Iron |
| Hg | mercury |
| Mn | manganese |
| Ni | nickel |
| Pb | lead |
| Se | selenium |
| Zn | zinc |
Conflicts of Interest
References
- Salem, H.M.; Eweida, E.A.; Farag, A. Heavy metals in drinking water and their environmental impact on human health. In ICEHM 2000; Cairo University: Giza, Egypt, 2000; pp. 542–556. [Google Scholar]
- Ahluwalia, S.S.; Goyal, D. Microbial and plant derived biomass for removal of heavy metals from wastewater. Bioresour. Technol. 2007, 98, 2243–2257. [Google Scholar] [CrossRef] [PubMed]
- Hussein, H.; Farag, S.; Moawad, H. Isolation and characterization of Pseudomonas resistant to heavy metals contaminants. Arab. J. Biotechnol. 2004, 7, 13–22. [Google Scholar]
- Kapoor, A.; Viraraghvan, T. Fungal biosorption—An alternative treatment option for heavy metal bearing wastewater: A review. Bioresour. Technol. 1995, 53, 195–206. [Google Scholar]
- Doelman, P.; Jansen, E.; Michels, M.; van Til, M. Effects of heavy metals in soil on microbial diversity and activity as shown by the sensitivity-resistance index, an ecologically relevant parameter. Biol. Fertil. Soils 1994, 17, 177–1784. [Google Scholar] [CrossRef]
- Wood, J.M.; Wang, H.K. Microbiol resistance to heavy metals. Environ. Sci. Technol. 1983, 17, 582–590. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Tan, T.C. Monitoring BOD in the presence of heavy metal ions using a poly (4-vinylpyr-idine) coated microbial sensor. Biosen. Bioelectron. 1994, 9, 445–455. [Google Scholar] [CrossRef]
- Goblenz, A.; Wolf, K.; Bauda, P. The role of glutathione biosynthesis in heavy metal resistance in the fission yeast Schizosaccharomyces pombe. FEMS Microbiol. Rev. 1994, 14, 303–308. [Google Scholar] [CrossRef] [PubMed]
- US EPA. Drinking Water Contaminants; United States Environmental Protection Agency (EPA): Washington, DC, USA, 2009. [Google Scholar]
- Agency for Toxic Substances and Disease Registry (ATSDR). Toxicological Profile for Silver; U.S. Department of Health and Human Services, Public Health Service: Atlanta, GA, USA, 1990. [Google Scholar]
- Tripathi, R.D.; Srivastava, S.; Mishra, S.; Singh, N.; Tuli, R.; Gupta, D.K.; Maathuis, F.J.M. Arsenic hazards: Strategies for tolerance and remediation by plants. Trends Biotech. 2007, 25, 158–165. [Google Scholar] [CrossRef]
- Acobs, I.A.; Taddeo, J.; Kelly, K.; Valenziano, C. Poisoning as a result of barium styphnate explosion. Am. J. Ind. Med. 2002, 41, 285–288. [Google Scholar] [CrossRef] [PubMed]
- Degraeve, N. Carcinogenic, teratogenic and mutagenic effects of cadmium. Mutat. Res. 1981, 86, 115–135. [Google Scholar] [CrossRef] [PubMed]
- Wuana, R.A.; Okieimen, F.E. Heavy metals in contaminated soils: A review of sources, chemistry, risks and best available strategies for remediation. ISRN Ecol. 2011, 2011. Article 20. [Google Scholar]
- Neustadt, J.; Pieczenik, S. Toxic-metal contamination: Mercury. Integr. Med. 2007, 6, 36–37. [Google Scholar]
- Ainza, C.; Trevors, J.; Saier, M. Environmental mercury rising. Water Air Soil Poll. 2010, 205, 47–48. [Google Scholar] [CrossRef]
- Gulati, K.; Banerjee, B.; Bala Lall, S.; Ray, A. Effects of diesel exhaust, heavy metals and pesticides on various organ systems: Possible mechanisms and strategies for prevention and treatment. Indian J. Exp. Biol. 2010, 48, 710–721. [Google Scholar] [PubMed]
- Khan, M.A.; Ahmad, I.; Rahman, I. Effect of environmental pollution on heavy metals content of Withania somnifera. J. Chin. Chem. Soc. 2007, 54, 339–343. [Google Scholar]
- Das, N.; Vimala, R.; Karthika, P. Biosorption of heavy metals—An overview. Indian J. Biotechnol. 2008, 7, 159–169. [Google Scholar]
- Duda-Chodak, A.; Baszczyk, U. The impact of nickel on human health. J. Elementol. 2008, 13, 685–696. [Google Scholar]
- Padmavathiamma, P.K.; Li, L.Y. Phytoremediation technology: Hyperaccumulation metals in plants. Water Air Soil Poll. 2007, 184, 105–126. [Google Scholar] [CrossRef]
- Vinceti, M.; Wei, E.T.; Malagoli, C.; Bergomi, M.; Vivoli, G. Adverse health effects of selenium in humans. Rev. Environ. Health 2001, 16, 233–251. [Google Scholar] [CrossRef] [PubMed]
- Hess, R.; Schmid, B. Zinc supplement overdose can have toxic effects. J. Pediat. Hematol. Onc. 2002, 24, 582–584. [Google Scholar] [CrossRef]
- Gadd, G.M. Bioremedial potential of microbial mechanisms of metal mobilization and immobilization. Curr. Opin. Biotechnol. 2000, 11, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Lim, P.E.; Mak, K.Y.; Mohamed, N.; Noor, A.M. Removal and speciation of heavy metals along the treatment path of wastewater in subsurface-flow constructed wetlands. Water Sci. Technol. 2003, 48, 307–313. [Google Scholar] [PubMed]
- Malik, A. Metal bioremediation through growing cells. Environ. Int. 2004, 30, 261–278. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.C.; Lin, H.L. Remediation of soil contaminated with the heavy metal (Cd2+). J. Hazard. Mater. 2005, 122, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Elekwachi, C.O.; Andresen, J.; Hodgman, T.C. Global use of bioremediation technologies for decontamination of ecosystems. J. Bioremed. Biodeg. 2014, 5, 1–9. [Google Scholar] [CrossRef]
- Gan, S.; Lau, E.V.; Ng, H.K. Remediation of soils contaminated with polycyclic aromatic hydrocarbons (PAHs). J. Hazard. Mater. 2009, 172, 532–549. [Google Scholar] [CrossRef] [PubMed]
- Haritash, A.K.; Kaushik, C.P. Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): A review. J. Hazard. Mater. 2009, 169, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Satinder, K.B.; Verma, M.; Surampalli, R.Y.; Misra, K.; Tyagi, R.D.; Meunier, N. Bioremediation of hazardous wastes-A review. Pract. Period. Hazard. Toxic Radioact. Waste Manag. 2006, 10, 59–72. [Google Scholar] [CrossRef]
- Onwubuya, K.; Cundy, A.; Puschenreiter, M.; Kumpiene, J.; Bone, B. Developing decision support tools for the selection of “gentle” remediation approaches. Sci. Total Environ. 2009, 407, 6132–6142. [Google Scholar] [CrossRef] [PubMed]
- Environmental Agency (EA). Reporting the Evidence: Dealing with Contaminated Land in England and Wales. A Review of Progress from 2000–2007 with Part 2A of the Environmental Protection Act. Available online: https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/313964/geho0109bpha-e-e.pdf (accessed on 1 January 2015).
- US EPA. Treatment Technologies for Site Cleanup: Annual Status Report; United States Environmental Protection Agency (EPA): Washington, DC, USA, 2007. [Google Scholar]
- Hussein, H.; Krull, R.; Abou El-Ela, S.I.; Hempel, D.C. Interaction of the different heavy metal ions with immobilized bacterial culture degrading xenobiotic wastewater compounds. In Proceedings of the Second International Water Association World Water Conference, Berlin, Germany, 15–19 October 2001; pp. 15–19.
- Scott, J.A.; Karanjkar, A.M. Repeated cadmium biosorption by regenerated Enterobacter aerogenes biofilm attached to activated carbon. Biotechnol. Lett. 1992, 14, 737–740. [Google Scholar] [CrossRef]
- Ajmal, M.; Rafaqat, A.K.; Bilquees, A.S. Studies on removal and recovery of Cr (VI) from electroplating wastes. Water Res. 1996, 30, 1478–1482. [Google Scholar] [CrossRef]
- Dilek, F.B.; Gokcay, C.F.; Yetis, U. Combined effects of Ni(II) and Cr(VI) on activated sludge. Water Res. 1998, 32, 303–312. [Google Scholar] [CrossRef]
- Volesky, B.; Holan, Z.R. Biosorption of heavy metals. Biotechnol. Prog. 1995, 11, 235–250. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Huang, C.P. Application of Aspergillus oryzae and Rhizopus oryzae for Cu (II) removal. Water Res. 1996, 30, 1985–1990. [Google Scholar] [CrossRef]
- Xiao, X.; Luo, S.; Zeng, G.; Wei, W.; Wan, Y.; Chen, L.; Guo, H.; Cao, Z.; Yang, L.; Chen, J.; et al. Biosorpiton of cadmium by endophytic fungus (EF) Microsphaeropsis sp. LSE10 isolated from cadmium hyperaccumulator Solanum nigrum L. Bioresour. Technol. 2010, 101, 1668–1674. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.N.; Zhang, Y.F.; He, L.Y.; Chen, Z.J.; Wang, Q.Y.; Qian, M.; Sheng, X.F. Genetic diversity and characterization of heavy metal-resistant-endophytic bacteria from two copper-tolerant plant species on copper mine wasteland. Bioresour. Technol. 2010, 101, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Modaihsh, A.; Al-Swailem, M.; Mahjoub, M. Heavy metal contents of commercial inorganic fertilizer used in the Kingdom of Saudi Arabia. Agric. Mar. Sci. 2004, 9, 21–25. [Google Scholar]
- Chehregani, A.; Malayeri, B.E. Removal of heavy metals by native accumulator plants. Int. J. Agric. Biol. 2007, 9, 462–465. [Google Scholar]
- Fulekar, M.; Singh, A.; Bhaduri, A.M. Genetic engineering strategies for enhancing phytoremediation of heavy metals. Afr. J. Biotechnol. 2009, 8, 529–535. [Google Scholar]
- Sabiha-Javied; Mehmood, T.; Chaudhry, M.M.; Tufai, M.; Irfan, N. Heavy metal pollution from phosphate rock used for the production of fertilizer in Pakistan. Microchem. J. 2009, 91, 94–99. [Google Scholar] [CrossRef]
- Sumner, M.E. Beneficial use of effluents, wastes, and biosolids. Commun. Soil Sci. Plant Anal. 2000, 31, 1701–1715. [Google Scholar] [CrossRef]
- D’Amore, J.J.; Al-Abed, S.R.; Scheckel, K.G.; Ryan, J.A. Methods for speciation of metals in soils: A review. J. Environ. Qual. 2005, 34, 1707–1745. [Google Scholar] [CrossRef] [PubMed]
- Lombi, E.; Gerzabek, M.H. Determination of mobile heavy metal fraction in soil: Results of a pot experiment with sewage sludge. Commun. Soil Sci. Plant Anal. 1998, 29, 2545–2556. [Google Scholar] [CrossRef]
- Sposito, G.; Page, A.L. Cycling of metal ions in the soil environment. In Metal Ions in Biological Systems, Circulation of Metals in the Environment; Sigel, H., Ed.; Marcel Dekker, Inc.: New York, NY, USA, 1984; pp. 287–332. [Google Scholar]
- Ahemad, M. Remediation of metalliferous soils through the heavy metal resistant plant growth promoting bacteria: Paradigms and prospects. Arab. J. Chem. 2014. [Google Scholar] [CrossRef]
- Garbisu, C.; Alkorta, I. Phytoextraction: A cost-effective plant-based technology for the removal of metals from the environment. Bioresour. Technol. 2001, 77, 229–236. [Google Scholar] [CrossRef] [PubMed]
- Sikkema, J.; de Bont, J.A.; Poolman, B. Mechanisms of membrane toxicity of hydrocarbons. Microbiol. Rev. 1995, 59, 201–222. [Google Scholar] [PubMed]
- Roane, T.M.; Pepper, I.L. Microorganisms and metal pollution. In Environmental Microbiology; Maier, I.L., Pepper, C.B., Eds.; Gerba, Academic Press: London, UK, 2000; p. 55. [Google Scholar]
- Guiné, V.; Spadini, L.; Sarret, G.; Muris, M.; Delolme, C.; Gaudet, J.P.; Martins, J.M. Zinc sorption to three gram-negative bacteria: Combined titration, modeling and EXAFS study. Environ. Sci. Technol. 2006, 40, 1806–1813. [Google Scholar] [CrossRef] [PubMed]
- Comte, S.; Guibaud, G.; Baudu, M. Biosorption properties of extracellular polymeric substances (EPS) towards Cd, Cu and Pb for different pH values. J. Hazard. Mater. 2008, 151, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.C.; Huang, Q.Y.; Wei, X.; Liang, W.; Rong, X.M.; Chen, W.L.; Cai, P. Microcalorimetric and potentiometric titration studies on the adsorption of copper by extracellular polymeric substances (EPS), minerals and their composites. Bioresour. Technol. 2010, 101, 5774–5779. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Wei, X.; Cai, P.; Huang, Q.; Chen, H.; Liang, W.; Rong, X. Role of extracellular polymeric substances in Cu(II) adsorption on Bacillus subtilis and Pseudomonas putida. Bioresour. Technol. 2011, 102, 1137–1141. [Google Scholar] [CrossRef] [PubMed]
- Carter, P.; Cole, H.; Burton, J. Bioremediation: Successes and Shortfalls. In Proceedings of Key International Conference and Exhibition for Spill Prevention, Preparedness, Response and Restoration (Interspill), London, UK, 23 March 2006.
- Kinya, K.; Kimberly, L.D. Current use of bioremediation for TCE cleanup: Results of a survey. Remediat. J. 1996, 6, 1–14. [Google Scholar]
- Chen, C.; Wang, J.L. Characteristics of Zn2+ biosorption by Saccharomyces cerevisiae. Biomed. Environ. Sci. 2007, 20, 478–482. [Google Scholar] [PubMed]
- Talos, K.; Pager, C.; Tonk, S.; Majdik, C.; Kocsis, B.; Kilar, F.; Pernyeszi, T. Cadmium biosorption on native Saccharomyces cerevisiae cells in aqueous suspension. Acta Univ. Sapientiae Agric. Environ. 2009, 1, 20–30. [Google Scholar]
- Tigini, V.; Prigione, V.; Giansanti, P.; Mangiavillano, A.; Pannocchia, A.; Varese, G.C. Fungal biosorption, an innovative treatment for the decolourisation and detoxification of textile effluents. Water 2010, 2, 550–565. [Google Scholar] [CrossRef]
- Brierley, C.L. Bioremediation of metal-contaminated surface and groundwater. Geomicrobiol. J. 1990, 8, 201–223. [Google Scholar] [CrossRef]
- Pinedo-Rivilla, C.; Aleu, J.; Collado, I.G. Pollutants biodegradation by fungi. Curr. Org. Chem. 2009, 13, 1194–1214. [Google Scholar] [CrossRef]
- D’Annibale, A.; Leonardi, V.; Federici, E.; Baldi, F.; Zecchini, F.; Petruccioli, M. Leaching and microbial treatment of a soil contaminated by sulphide ore ashes and aromatic hydrocarbons. Appl. Microbiol. Biotechnol. 2007, 74, 1135–1144. [Google Scholar] [CrossRef] [PubMed]
- Tunali, S.; Akar, T.; Oezcan, A.S.; Kiran, I.; Oezcan, A. Equilibrium and kinetics of biosorption of lead(II) from aqueous solutions by Cephalosporium aphidicola. Sep. Purif. Technol. 2006, 47, 105–112. [Google Scholar] [CrossRef]
- Akar, T.; Tunali, S.; Cabuk, A. Study on the characterization of lead (II) biosorption by fungus Aspergillus parasiticus. Appl. Biochem. Biotech. 2007, 136, 389–406. [Google Scholar] [CrossRef]
- Kelly, D.J.A.; Budd, K.; Lefebvre, D.D. The biotransformation of mercury in pH-stat cultures of microfungi. Can. J. Bot. 2006, 84, 254–260. [Google Scholar] [CrossRef]
- Thavasi, R. Microbial biosurfactants: From an environment application point of view. J. Bioremed. Biodegrad. 2011, 2. Article 104e. [Google Scholar] [CrossRef]
- Evanko, C.R.; Dzombak, D.A. Remediation of metals-contaminated soil and groundwater. Environ. Sci. 1997, 412, 1–45. [Google Scholar]
- Silver, S. Bacterial heavy metal resistance: New surprises. Annu. Rev. Microbiol. 1996, 50, 753–789. [Google Scholar] [CrossRef] [PubMed]
- Lovley, D.R.; Phillips, E.J.P. Novel mode of microbial energy metabolism: Organic carbon oxidation to dissimilatory reduction of iron or manganese. Appl. Environ. Microbiol. 1988, 54, 1472–1480. [Google Scholar]
- Lovley, D.R.; Coates, J.D.; Blunt-Harris, E.L.; Philips, E.J.P.; Woodward, J.C. Humic substances as electron acceptors for microbial respiration. Nature 1996, 382, 445–448. [Google Scholar] [CrossRef]
- Spormann, A.M.; Widdel, F. Metabolism of alkylbenzenes, alkanes, and other hydrocarbons in anaerobic bacteria. Biodegradation 2000, 11, 85–105. [Google Scholar] [CrossRef] [PubMed]
- Lovely, D.R. Dissimilatory metal rduction: From early life to bioremediation. ASM News 2002, 68, 231–237. [Google Scholar]
- Lovley, D.R.; Philips, E.J.; Gorby, Y.A.; Landa, E.R. Microbial reduction of uranium. Nature 1991, 350, 413–416. [Google Scholar] [CrossRef]
- Gómez Jiménez-T, R.; Moliternib, E.; Rodríguezb, L.; Fernándezc, F.J.; Villaseñorc, J. Feasibility of mixed enzymatic complexes to enhanced soil bioremediation processes. Procedia Environ. Sci. 2011, 9, 54–59. [Google Scholar] [CrossRef]
- Cobbett, C.; Goldsbrough, P. Phytochelatins and metallothioneins: Role in heavy metals detoxification and homeostatis. Annu. Rev. Plant Biol. 2002, 53, 159–182. [Google Scholar] [CrossRef] [PubMed]
- Kagi, J.H.R.; Schaffer, A. Biochemistry of metallothionein. Biochemistry 1988, 27, 8509–8515. [Google Scholar] [CrossRef] [PubMed]
- Huckle, J.W.; Morby, A.P.; Turner, J.S.; Robinson, N.J. Isolation of a prokaryotic metallothionein locus and analysis of transcriptional control by trace metal ions. Mol. Microbiol. 1993, 7, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Valls, M.; Atrian, S.; de Lorenzo, V.; La, F. Engineering a mouse metallothionein on the cell surface of Ralstonia eutropha CH34 for immobilization of heavy metals in soil. Nat. Biotechnol. 2000, 18, 661–665. [Google Scholar] [CrossRef] [PubMed]
- Mejare, M.; Bulow, L. Metal binding proteins and peptides in bioremediation and phytoremediation of heavy metals. Trends Biotechnol. 2001, 19, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.H.; Singh, S.; Kim, J.Y.; Lee, W.; Mulchandani, A.; Chen, W. Bacteria metabolically engineered for enhanced phtochelatin production and cadmium accumulation. App. Environ. Microbiol. 2007, 73, 6317–6320. [Google Scholar] [CrossRef]
- Singh, S.; Kang, S.H.; Mulchandani, A.; Chen, W. Bioremediation: environmental cleanup through pathway engineering. Curr. Opin. Biotechnol. 2008, 19, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Brim, H.; Venkateshwaran, A.; Kostandarithes, H.M.; Fredrickson, J.K.; Daly, M.J. Engineering Deinococcus geothermalis for bioremediation of high temperature radioactive waste environments. App. Environ. Microbiol. 2003, 69, 4575–4582. [Google Scholar] [CrossRef]
- Rojas, L.A.; Yanez, C.; Gonzalez, M.; Lobos, S.; Smalla, K.; Seeger, M. Characterization of the metabolically modified heavy metal-resistant Cupriavidus metallidurans strain MSR33 gener-ated for mercury bioremediation. PLoS One 20 2011, 6, e17555. [Google Scholar] [CrossRef]
- Sone, Y.; Mochizuki, Y.; Koizawa, K.; Nakamura, R.; Pan-Hou, H.; Itoh, T.; Kiyono, M. Mercurial resistance determinants in Pseudomonas strain K-62 plasmid pMR68. AMB Express 2013, 3. Article 41. [Google Scholar]
- Essa, A.M.M.; Macaskie, L.E.; Brown, N.L. Mechanisms of mercury bioremediation. Biochem. Soc. Trans. 2002, 30, 672–674. [Google Scholar] [CrossRef] [PubMed]
- Brim, H.; Osborne, J.P.; Kostandarithes, H.M.; Fredrickson, J.K.; Wackett, L.P.; Daly, M.J. Deinococcus radiodurans engineered for complete toluene degradation facilities Cr(IV) reduction. Microbiology 2006, 152, 2469–2477. [Google Scholar] [CrossRef] [PubMed]
- Penny, C.; Vuilleumier, S.; Bringel, F. Microbial degradation of tetrachloromethane: Mechanisms and perspectives for bioremediation. FEMS Microbiol. Ecol. 2010, 74, 257–275. [Google Scholar] [CrossRef] [PubMed]
- Ali, H.; Khan, E.; Sajad, M.A. Phytoremediation of heavy metals-Concepts and applications. Chemosphere 2013, 91, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Robinson, B.H.; Brooks, R.R.; Howes, A.W.; Kirkman, J.H.; Gregg, P.E.H. The potential of the high biomass nickel hyperaccumulator Berkheya coddii for phytoremediation and phytomining. J. Geochem. Explor. 1997, 60, 115–126. [Google Scholar] [CrossRef]
- Martinez, M.; Bernal, P.; Almela, C.; Vélez, D.; García-Agustín, P.; Serrano, R.; Navarro-Aviñó, J. An engineered plant that accumulates higher levels of heavy metals than Thlaspi caerulescens, with yields of 100 times more biomass in mine soils. Chemosphere 2006, 64, 478–485. [Google Scholar] [CrossRef] [PubMed]
- Alkorta, I.; Hernández-Allica, J.; Becerril, J.M.; Amezaga, I.; Albizu, I.; Garbisu, C. Recent findings on the phytoremediation of soils contaminated with environmentally toxic heavy metals and metalloids such as Zinc, Cadmium, Lead, and Arsenic. Rev. Environ. Sci. Biotechnol. 2004, 3, 71–90. [Google Scholar] [CrossRef]
- Sekara, A.; Poniedzialeek, M.; Ciura, J.; Jedrszczyk, E. Cadmium and lead accumulation and distribution in the organs of nine crops: implications for phytoremediation. Pol. J. Environ. Stud. 2005, 14, 509–516. [Google Scholar]
- Mesjasz-Przybylowicz, J.; Nakonieczny, M.; Migula, P.; Augustyniak, M.; Tarnawska, M.; Reimold, W.U.; Koeberl, C.; Przybylowicz, W.; Glowacka, E. Uptake of cadmium, lead, nickel and zinc from soil and water solutions by the nickel hyperaccumulator Berkheya coddii. Acta Biol. Cracov. Bot. 2004, 46, 75–85. [Google Scholar]
- Erakhrumen, A.A. Phytoremediation: An environmentally sound technology for pollution prevention, control and remediation in developing countries. Educ. Res. Rev. 2007, 2, 151–156. [Google Scholar]
- Barceló, J.; Poschenrieder, C. Phytoremediation: principles and perspectives. Contrib. Sci. 2003, 2, 333–344. [Google Scholar]
- Vishnoi, S.R.; Srivastava, P.N. Phytoremediation-green for environmental clean. In Proceeding of Taal 2007: The 12th World Lake Conference, Jaipur, India, 29 October–2 November 2008.
- Karami, A.; Shamsuddin, Z.H. Phytoremediation of heavy metals with several efficiency enhancer methods. Afr. J. Biotechnol. 2010, 9, 3689–3698. [Google Scholar]
- Baker, A.J.M.; McGrath, S.P.; Sidoli, C.M.D.; Reeves, R.D. The possibility of in situ heavy metal decontamination of polluted soils using crops of metal accumulating plants. Resour. Conserv. Recycl. 1994, 11, 42–49. [Google Scholar]
- Prasad, M.N.V.; Freitas, H.M.D.O. Metal hyperaccumulation in plants—Biodiversity prospecting for phytoremediation technology. Electro. J. Biotechnol. 2003, 6, 285–321. [Google Scholar] [CrossRef]
- Dahmani-Muller, H.; Van-Oort, F.; Gelie, B.; Balabane, M. Strategies of heavy metal uptake by three plant species growing near a metal smelter. Environ. Pollut. 2000, 109, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.H.; Zhou, Q.X.; Wang, X. Cadmium-hyperaccumulator Solanum nigrum L. and its accumulating characteristics. Environ. Sci. 2005, 26, 167–171. [Google Scholar]
- Wei, S.; Zhou, Q.; Zhang, K.; Liang, J. Roles of rhizosphere in remediation of contaminated soils and its mechanisms. Ying Yong Sheng Tai Xue Bao 2003, 14, 143–147. [Google Scholar] [PubMed]
- Kuiper, I.; Lagendijk, E.L.; Bloemberg, G.V.; Lugtenberg, B.J.J. Rhizoremediation: A beneficial plant-microbe interaction. Mol. Plant-Microbe Interact. 2004, 17, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Glick, B.R.; Penrose, D.M.; Li, J. A model for the lowering of plant ethylene concentrations by plant growth-promoting bacteria. J. Theor. Biol. 1998, 190, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Burd, G.I.; Dixon, D.G.; Glick, B.R. A plant growth-promoting bacterium that decreases nickel toxicity in seedlings. Appl. Environ. Microbiol. 1998, 64, 3663–3668. [Google Scholar] [PubMed]
- Pulford, I.; Watson, C. Phytoremediation of heavy metal-contaminated land by trees-a review. Environ. Int. 2003, 29, 529–540. [Google Scholar]
- Volk, T.A.; Abrahamson, L.P.; Nowak, C.A.; Smart, L.B.; Tharakan, P.J.; White, E.H. The development of short-rotation willow in the northeastern United States for bioenergy and bioproducts, agroforestry and phytoremediation. Biomass Bioener. 2006, 30, 715–727. [Google Scholar] [CrossRef]
- Ruttens, A.; Boulet, J.; Weyens, N.; Smeets, K.; Adriaensen, K.; Meers, E.; van Slycken, S.; Tack, F.; Meiresonne, L.; Thewys, T.; et al. Short rotation coppice culture of willows and poplars as energy crops on metal contaminated agricultural soils. Int. J. Phytorem. 2011, 13, 194–207. [Google Scholar] [CrossRef]
- Huang, H.; Yu, N.; Wang, L.; Gupta, D.K.; He, Z.; Wang, K.; Zhu, Z.; Yan, X.; Li, T.; Yang, X.E. The phytoremediation potential of bioenergy crop Ricinus communis for DDTs and cadmium co-contaminated soil. Bioresour. Technol. 2011, 102, 11034–11038. [Google Scholar] [CrossRef] [PubMed]
- Meers, E.; van Slycken, S.; Adriaensen, K.; Ruttens, A.; Vangronsveld, J.; Du Laing, G.; Witters, N.; Thewys, T.; Tack, F.M. The use of bio-energy crops (Zea mays) for “phytoattenuation” of heavy metals on moderately contaminated soils: A field experiment. Chemosphere 2010, 78, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Abhilash, P.C.; Jamil, S.; Singh, N. Transgenic plants for enhanced biodegradation and phytoremediation of organic xenobiotics. Biotechnol. Adv. 2009, 27, 474–488. [Google Scholar] [CrossRef] [PubMed]
- Jamil, S.; Abhilash, P.C.; Singh, N.; Sharma, P.N. Jatropha curcas: A potential crop for phytoremediation of coal fly ash. J. Hazard. Mater. 2009, 172, 269–275. [Google Scholar] [CrossRef] [PubMed]
- Che, D.; Meagher, R.B.; Heaton, A.C.; Lima, A.; Rugh, C.L.; Merkle, S.A. Expression of mercuric ion reductase in Eastern cottonwood (Populus deltoides) confers mercuric ion reduction and resistance. Plant Biotechnol. J. 2003, 1, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Bitther, O.P.; Pilon-Smits, E.A.H.; Meagher, R.B.; Doty, S. Biotechnological approaches for phytoremediation. In Plant Biotechnology and Agriculture; Arie Altman, A., Hasegawa, P.M., Eds.; Academic Press: Oxford, UK, 2012; pp. 309–328. [Google Scholar]
- Bittsanszkya, A.; Kömives, T.; Gullner, G.; Gyulai, G.; Kiss, J.; Heszky, L.; Radimszky, L.; Rennenberg, H. Ability of transgenic poplars with elevated glutathione content to tolerate zinc(2+) stress. Environ. Int. 2005, 31, 251–254. [Google Scholar] [CrossRef] [PubMed]
- Sayler, G.S.; Ripp, S. Field applications of genetically engineered microorganisms for bioremediation process. Curr. Opin. Biotechnol. 2000, 11, 286–289. [Google Scholar] [CrossRef] [PubMed]
- Verma, N.; Singh, M. Biosensors for heavy metals. J. Biometals 2005, 18, 121–129. [Google Scholar] [CrossRef]
- Bruschi, M.; Goulhen, F. New bioremediation technologies to remove heavy metals and radionuclides using Fe(III)-sulfate- and sulfur reducing bacteria. In Environmental Bioremediation Technologies; Singh, S.N., Tripathi, R.D., Eds.; Springer Publication: NY, USA, 2006; pp. 35–55. [Google Scholar]
- Divya, B.; Deepak Kumar, M. Plant-Microbe interaction with enhanced bioremediation. Res. J. BioTechnol. 2011, 6, 72–79. [Google Scholar]
- Bae, W.; Mehra, R.K.; Mulchandani, A.; Chen, W. Genetic engineering of Escherichia coli for enhanced uptake and bioaccumulation of mercury. Appl. Environ. Microbiol. 2001, 67, 5335–5338. [Google Scholar] [CrossRef] [PubMed]
- Bae, W.; Wu, C.H.; Kostal, J.; Mulchandani, A.; Chen, W. Enhanced mercury biosorption by bacterial cells with surface-displayed MerR. App. Environ. Microbiol. 2003, 69, 3176–3180. [Google Scholar] [CrossRef]
- Wu, C.H.; Wood, T.K.; Mulchandani, A.; Chen, W. Engineering plant-microbe symbiosis for rhizoremediation of heavy metals. Appl. Environ. Microbiol. 2006, 72, 1129–1134. [Google Scholar] [CrossRef] [PubMed]
- Kostal, J.R.Y.; Wu, C.H.; Mulchandani, A.; Chen, W. Enhanced arsenic accumulation in engineered bacterial cells expressing ArsR. Appl. Environ. Microbiol. 2004, 70, 4582–4587. [Google Scholar] [CrossRef] [PubMed]
- Hasin, A.A.; Gurman, S.J.; Murphy, L.M.; Perry, A.; Smith, T.J.; Gardiner, P.E. Remediation of chromium (VI) by a methane-oxidizing bacterium. Environ. Sci. Technol. 2010, 44, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Ackerley, D.F.; Gonzalez, C.F.; Keyhan, M.; Blake, R.; Matin, A. Mechanism of chromate reduction by the Escherichia coli protein, NfsA, and the role of different chromate reductases in minimizing oxidative stress during chromate reduction. Environ. Microbiol. 2004, 6, 851–860. [Google Scholar] [CrossRef] [PubMed]
- Brim, H.; McFarlan, S.C.; Fredrickson, J.K.; Minton, K.W.; Zhai, M.; Wackett, L.P.; Daly, M.J. Engineering Deinococcus radiodurans for metal remediation in radioactive mixed waste environments. Nat. Biotechnol. 2000, 18, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Murtaza, I.; Dutt, A.; Ali, A. Biomolecular engineering of Escherichia coli organomercurial lyase gene and its expression. Indian J. Biotech. 2002, 1, 117–120. [Google Scholar]
- Zhao, X.W.; Zhou, M.H.; Li, Q.B.; Lu, Y.H.; He, N.; Sun, D.H.; Deng, X. Simultaneous mercury bioaccumulation and cell propagation by genetically engineered Escherichia coli. Process Biochem. 2005, 40, 1611–1616. [Google Scholar] [CrossRef]
- Kiyono, M.; Pan-Hou, H. Genetic engineering of bacteria for environmental remediation of mercury. J. Health Sci. 2006, 52, 199–204. [Google Scholar] [CrossRef]
- Ng, S.P.; Davis, B.; Polombo, E.A.; Bhave, M. A Tn5051-like mer-containing transposon identified in a heavy metal tolerant strain Achromobacter sp. AO22. BMC Res. Notes 2009, 7, 2–38. [Google Scholar]
- Lopez, A.; Lazaro, N.; Morales, S.; Margues, A.M. Nickel biosorption by free and immobilized cells of Pseudomonas fluorescens 4F39: A comparative study. Water Air Soil Pollut. 2002, 135, 157–172. [Google Scholar] [CrossRef]
- Sriprang, R.; Hayashi, M.; Ono, H.; Takagi, M.; Hirata, K.; Murooka, Y. Enhanced accumulation of Cd2+ by a Mesorhizobium sp. transformed with a gene from Arabidopsis thaliana coding for phytochelatin synthase. Appl. Environ. Microbiol. 2003, 69, 79–796. [Google Scholar] [CrossRef]
- Van Aken, B.; Tehrani, R.; Schnoor, J. Endophyte-assisted phytoremediation of explosives in poplar trees by Methylobacterium populi BJ001T. In Endophytes of Forest Trees: Biology and Applications, Forestry Sciences; Pirttilä, A.M., Frank, A.C., Eds.; Springer: Heidelberg, The Netherlands, 2011; Volume 80, pp. 217–234. [Google Scholar]
- Singh, B.K. Emerging and genomic approaches in bioremediation. In Proceedings of the 4th International Contaminated Site Remediation Conference, Adelaide, Australia, 11–15 September 2011.
- Maestri, E.; Marmiroli, M. Genetic and molecular aspects of metal tolerance and hyperaccumualtion. In Metal Toxicity in Plants: Perception, Signalling and Remediation; Gupta, D.K., Sandalio, L.M., Eds.; Springer: Berlin, Germany, 2011; pp. 41–61. [Google Scholar]
- Ruis, O.N.; Daniell, H. Genetic engineering to enhance mercury phytoremediation. Curr. Opin. Biotechnol. 2009, 20, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L. Enhanced metabolism of halogenated hydrocarbons in transgenic plants contain mammalian P450 2E1. Proc. Natl. Acad. Sci. USA 2007, 97, 6287–6291. [Google Scholar] [CrossRef]
- Gullner, G. Enhanced tolerance of transgenic poplar plants overexpressing gamma-glutamylcysteine synthetase towards chloroacetanilide herbicides. J. Exp. Bot. 2001, 52, 971–979. [Google Scholar] [CrossRef] [PubMed]
- Abhilash, P.C.; Powell, J.R.; Singh, H.B.; Singh, B.K. Plant–microbe interactions: Novel applications for exploitation in multipurpose remediation technologies. Trends Biotechnol. 2012, 30, 416–420. [Google Scholar] [CrossRef] [PubMed]
- Lovley, D.R. Cleaning up with genomics: Applying molecular biology to bioremediation. Nat. Rev. Microbiol. 2003, 1, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Prasad, M.N.; Rajkumar, M.; Freitas, H. Plant growth promoting rhizobacteria and endophytes accelerate phytoremediation of metalliferous soils. Biotechnol. Adv. 2011, 29, 248–258. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, J.; Rillig, M.C.; Thies, J.; Massielo, C.A.; Hockaday, W.C.; Crowley, D. Biochar effects on soil biota—A review. Soil Biol. Biochem. 2011, 43, 1812–1836. [Google Scholar] [CrossRef]
- Sarikaya, M.; Tamerler, C.; Jen, A.K.; Schulten, K.; Baneyx, F. Molecular biomimetics: Nanotechnology through biology. Nat. Mater. 2003, 2, 577–585. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.D.; Lennon, E.; McNeil, L.B.; Minton, K.W. Duplication insertion of drug resistance determinants in the radioresistant bacterium Deinococcus radiodurans. J. Bacteriol. 1998, 170, 2126–2135. [Google Scholar]
- Vishwanathan, B. Nanomaterials; Narosa Publishing House Pvt Ltd.: New Delhi, India, 2009. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dixit, R.; Wasiullah; Malaviya, D.; Pandiyan, K.; Singh, U.B.; Sahu, A.; Shukla, R.; Singh, B.P.; Rai, J.P.; Sharma, P.K.; et al. Bioremediation of Heavy Metals from Soil and Aquatic Environment: An Overview of Principles and Criteria of Fundamental Processes. Sustainability 2015, 7, 2189-2212. https://doi.org/10.3390/su7022189
Dixit R, Wasiullah, Malaviya D, Pandiyan K, Singh UB, Sahu A, Shukla R, Singh BP, Rai JP, Sharma PK, et al. Bioremediation of Heavy Metals from Soil and Aquatic Environment: An Overview of Principles and Criteria of Fundamental Processes. Sustainability. 2015; 7(2):2189-2212. https://doi.org/10.3390/su7022189
Chicago/Turabian StyleDixit, Ruchita, Wasiullah, Deepti Malaviya, Kuppusamy Pandiyan, Udai B. Singh, Asha Sahu, Renu Shukla, Bhanu P. Singh, Jai P. Rai, Pawan Kumar Sharma, and et al. 2015. "Bioremediation of Heavy Metals from Soil and Aquatic Environment: An Overview of Principles and Criteria of Fundamental Processes" Sustainability 7, no. 2: 2189-2212. https://doi.org/10.3390/su7022189
APA StyleDixit, R., Wasiullah, Malaviya, D., Pandiyan, K., Singh, U. B., Sahu, A., Shukla, R., Singh, B. P., Rai, J. P., Sharma, P. K., Lade, H., & Paul, D. (2015). Bioremediation of Heavy Metals from Soil and Aquatic Environment: An Overview of Principles and Criteria of Fundamental Processes. Sustainability, 7(2), 2189-2212. https://doi.org/10.3390/su7022189









