Stability of the Anaerobic Digestion Process during Switch from Parallel to Serial Operation—A Microbiome Study
Abstract
:1. Introduction
2. Materials and Methods
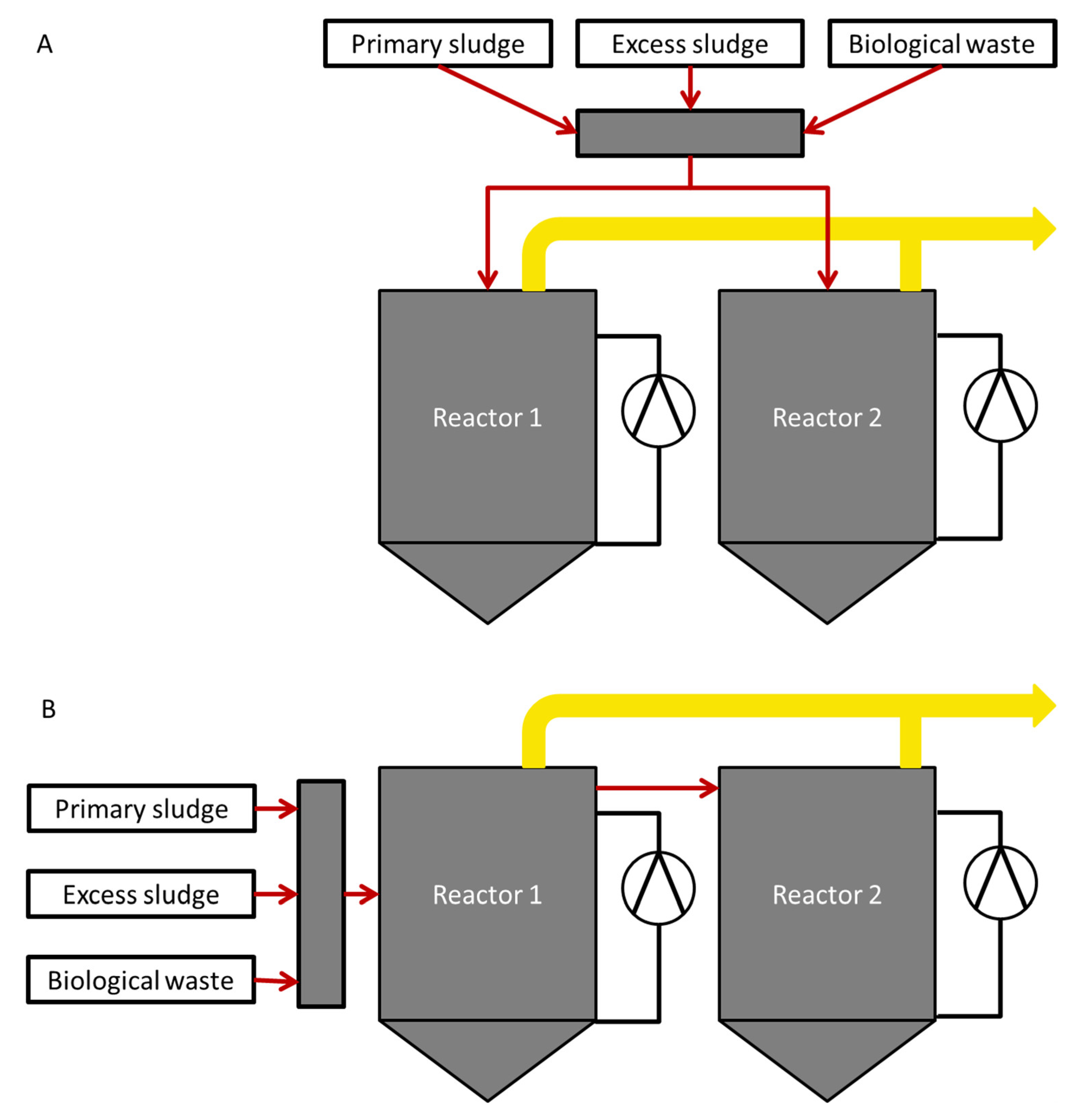
2.1. Reactor Setup and Operation
2.2. Sampling and Physicochemical Analysis
2.3. DNA Extraction
2.4. Community Profiling
2.5. Statistical Analyses
3. Results and Discussion
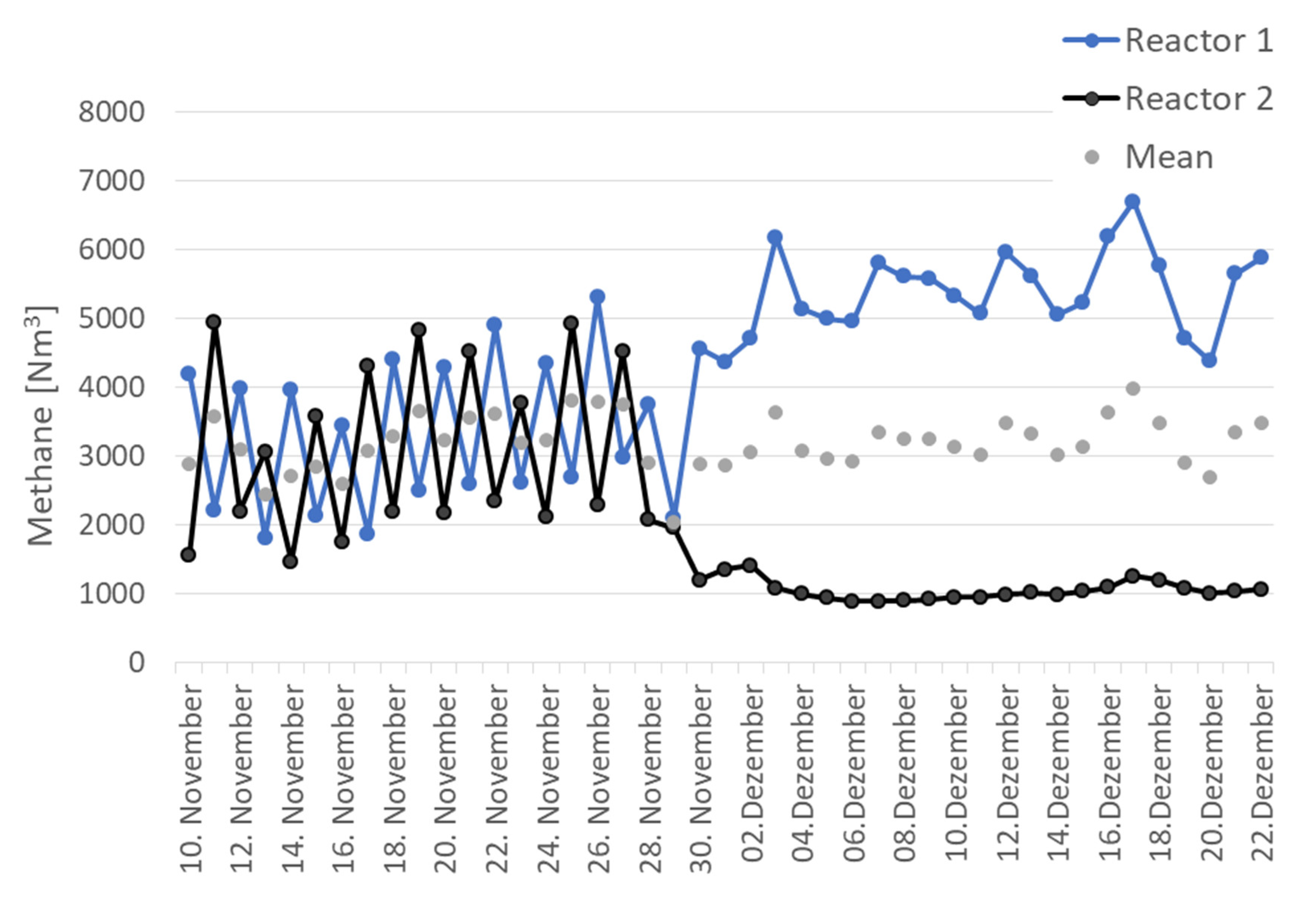
3.1. Process Characteristics in Parallel and Serial Operation
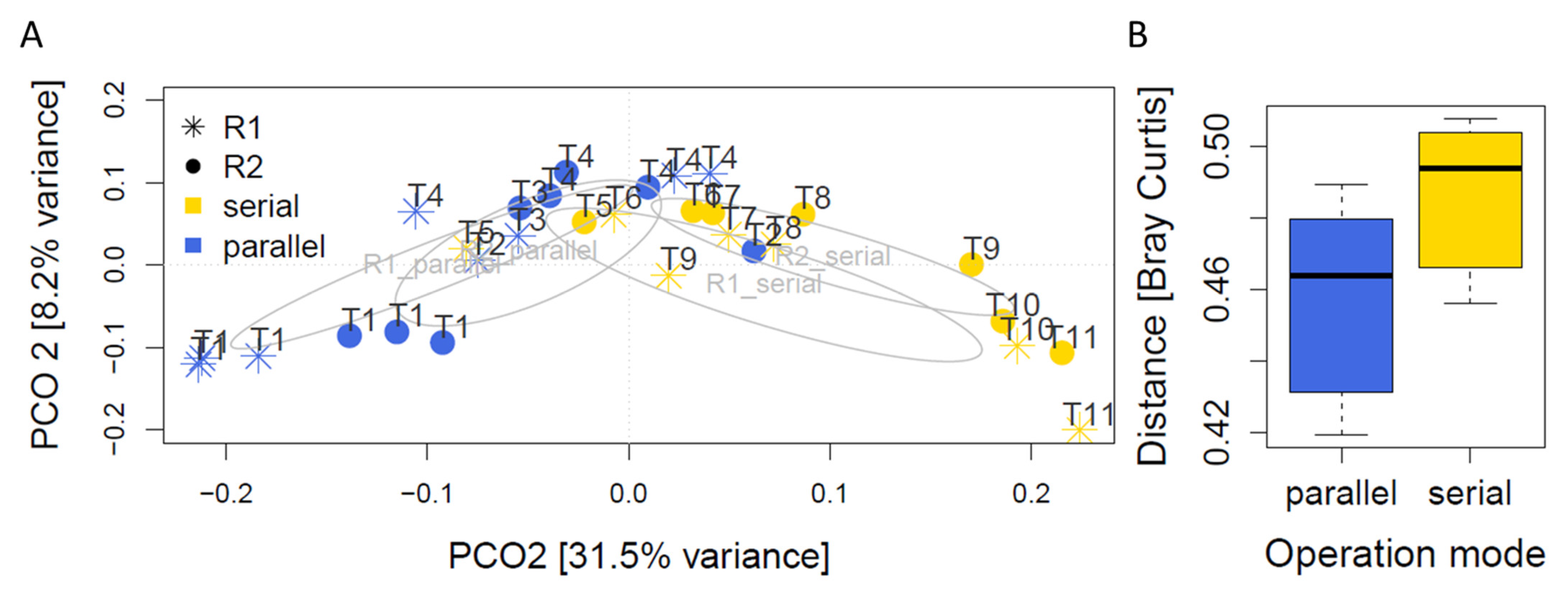
3.2. Microbial Community Profiles in Parallel and Serial Operation
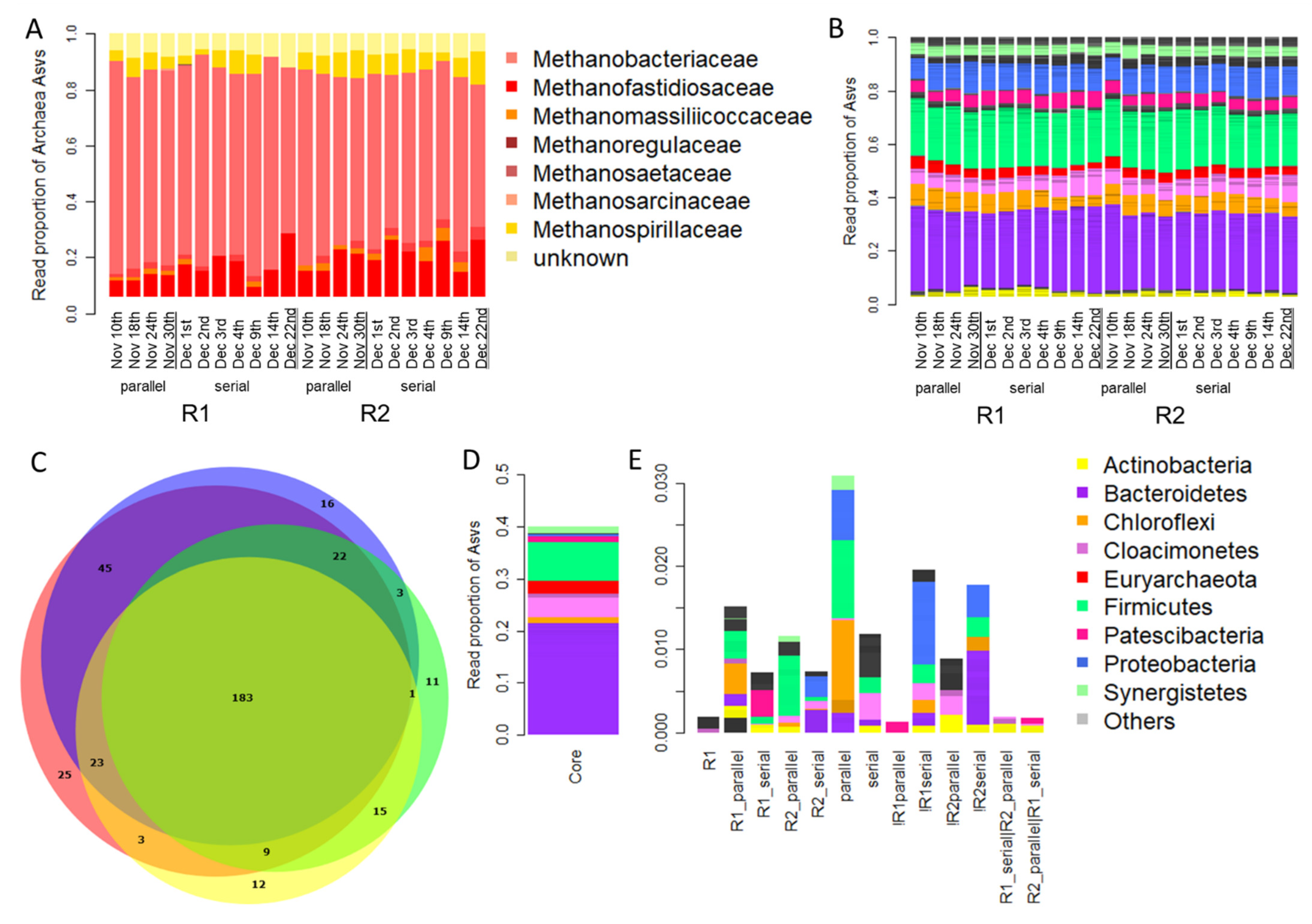
3.3. Taxonomic Composition of the Microbial Community Present in Parallel and Serial Operation
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fountoulakis, M.S.; Petousi, I.; Manios, T. Co-digestion of sewage sludge with glycerol to boost biogas production. Waste Manag. 2010, 30, 1849–1853. [Google Scholar] [CrossRef]
- Murto, M.; Björnsson, L.; Mattiasson, B. Impact of food industrial waste on anaerobic co-digestion of sewage sludge and pig manure. J. Environ. Manag. 2004, 70, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Appels, L.; Baeyens, J.; Degrève, J.; Dewil, R. Principles and potential of the anaerobic digestion of waste-activated sludge. Prog. Energy Combust. Sci. 2008, 34, 755–781. [Google Scholar] [CrossRef]
- Deublein, D.; Steinhauser, A. Biogas from Waste and Renewable Resources: An Introduction; John Wiley & Sons: Hoboken, NJ, USA, 2011; ISBN 3527643710. [Google Scholar]
- Weiland, P. Biogas production: Current state and perspectives. Appl. Microbiol. Biotechnol. 2010, 85, 849–860. [Google Scholar] [CrossRef] [PubMed]
- Insam, H.; Franke-Whittle, I.; Goberna, M. Microbes in aerobic and anaerobic waste treatment. In Microbes at Work; Springer: Berlin/Heidelberg, Germany, 2010; pp. 1–34. [Google Scholar]
- de Crescenzo, C.; Marzocchella, A.; Karatza, D.; Molino, A.; Ceron-Chafla, P.; Lindeboom, R.E.F.; van Lier, J.B.; Chianese, S.; Musmarra, D. Modelling of autogenerative high-pressure anaerobic digestion in a batch reactor for the production of pressurised biogas. Biotechnol. Biofuels Bioprod. 2022, 15, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Su, X.; Zhao, W.; Xia, D.; Fu, H.; Wang, G. Culture medium optimization for producing biomethane by coal anaerobic digestion. J. Biotechnol. 2022, 348, 26–35. [Google Scholar] [CrossRef]
- Kaparaju, P.; Ellegaard, L.; Angelidaki, I. Optimisation of biogas production from manure through serial digestion: Lab-scale and pilot-scale studies. Bioresour. Technol. 2009, 100, 701–709. [Google Scholar] [CrossRef]
- Boe, K.; Angelidaki, I. Serial CSTR digester configuration for improving biogas production from manure. Water Res. 2009, 43, 166–172. [Google Scholar] [CrossRef]
- Li, Y.; Liu, C.; Wachemo, A.C.; Yuan, H.; Zou, D.; Liu, Y.; Li, X. Serial completely stirred tank reactors for improving biogas production and substance degradation during anaerobic digestion of corn stover. Bioresour. Technol. 2017, 235, 380–388. [Google Scholar] [CrossRef]
- Bensmann, A.; Hanke-Rauschenbach, R.; Sundmacher, K. Reactor configurations for biogas plants—A model based analysis. Chem. Eng. Sci. 2013, 104, 413–426. [Google Scholar] [CrossRef]
- Angelidaki, I.; Boe, K.; Ellegaard, L. Effect of operating conditions and reactor configuration on efficiency of full-scale biogas plants. Water Sci. Technol. 2005, 52, 189–194. [Google Scholar] [CrossRef]
- Boe, K.; Karakashev, D.; Trably, E.; Angelidaki, I. Effect of post-digestion temperature on serial CSTR biogas reactor performance. Water Res. 2009, 43, 669–676. [Google Scholar] [CrossRef]
- Wagner, A.O.; Markt, R.; Puempel, T.; Illmer, P.; Insam, H.; Ebner, C. Sample preparation, preservation, and storage for volatile fatty acid quantification in biogas plants. Eng. Life Sci. 2017, 17, 132–139. [Google Scholar] [CrossRef]
- Podmirseg, S.M.; Waldhuber, S.; Knapp, B.A.; Insam, H.; Goberna, M. Robustness of the autochthonous microbial soil community after amendment of cattle manure or its digestate. Biol. Fertil. Soils 2019, 55, 565–576. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.R-project.org (accessed on 7 January 2022).
- Oksanen, J.; Blanchet, F.J.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D. Community Ecology Package, R Package, version 2.4-2. 2017. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 7 January 2022).
- Hulsen, T. BioVenn–an R and Python package for the comparison and visualization of biological lists using area-proportional Venn diagrams. Data Sci. 2021, 4, 51–66. [Google Scholar] [CrossRef]
- Schöftner, R.; Valentin, K.; Schmiedinger, B.; Trogisch, S.; Haberbauer, M.; Katzlinger, K.; Schnitzhofer, W.; Weran, N. Best Biogas Practise. Rep. Energy Environ. Res. 2007, 45, 2006. [Google Scholar]
- Drosg, B. Process Monitoring in Biogas Plants; IEA Bioenergy: Paris, France, 2013. [Google Scholar]
- Blume, F.; Bergmann, I.; Nettmann, E.; Schelle, H.; Rehde, G.; Mundt, K.; Klocke, M. Methanogenic population dynamics during semi-continuous biogas fermentation and acidification by overloading. J. Appl. Microbiol. 2010, 109, 441–450. [Google Scholar] [CrossRef]
- Zuo, Z.; Wu, S.; Zhang, W.; Dong, R. Effects of organic loading rate and effluent recirculation on the performance of two-stage anaerobic digestion of vegetable waste. Bioresour. Technol. 2013, 146, 556–561. [Google Scholar] [CrossRef]
- Babaee, A.; Shayegan, J. Effect of organic loading rates (OLR) on production of methane from anaerobic digestion of vegetables waste. In World Renewable Energy Congress-Sweden; Linköping University Electronic Press: Linköping, Sweden, 2011. [Google Scholar]
- Mauky, E.; Weinrich, S.; Jacobi, H.-F.; Nägele, H.-J.; Liebetrau, J.; Nelles, M. Demand-driven biogas production by flexible feeding in full-scale–Process stability and flexibility potentials. Anaerobe 2017, 46, 86–95. [Google Scholar] [CrossRef]
- Nghiem, L.D.; Nguyen, T.T.; Manassa, P.; Fitzgerald, S.K.; Dawson, M.; Vierboom, S. Co-digestion of sewage sludge and crude glycerol for on-demand biogas production. Int. Biodeterior. Biodegrad. 2014, 95, 160–166. [Google Scholar] [CrossRef]
- Athanasoulia, E.; Melidis, P.; Aivasidis, A. Optimization of biogas production from waste activated sludge through serial digestion. Renew. Energy 2012, 47, 147–151. [Google Scholar] [CrossRef]
- Adekunle, K.F.; Okolie, J.A. A review of biochemical process of anaerobic digestion. Adv. Biosci. Biotechnol. 2015, 6, 205. [Google Scholar] [CrossRef]
- Yang, Y.; Yu, K.; Xia, Y.; Lau, F.T.K.; Tang, D.T.W.; Fung, W.C.; Fang, H.H.P.; Zhang, T. Metagenomic analysis of sludge from full-scale anaerobic digesters operated in municipal wastewater treatment plants. Appl. Microbiol. Biotechnol. 2014, 98, 5709–5718. [Google Scholar] [CrossRef]
- Guo, J.; Peng, Y.; Ni, B.-J.; Han, X.; Fan, L.; Yuan, Z. Dissecting microbial community structure and methane-producing pathways of a full-scale anaerobic reactor digesting activated sludge from wastewater treatment by metagenomic sequencing. Microb. Cell Factories 2015, 14, 33. [Google Scholar] [CrossRef]
- Li, A.; Chu, Y.; Wang, X.; Ren, L.; Yu, J.; Liu, X.; Yan, J.; Zhang, L.; Wu, S.; Li, S. A pyrosequencing-based metagenomic study of methane-producing microbial community in solid-state biogas reactor. Biotechnol. Biofuels 2013, 6, 3. [Google Scholar] [CrossRef]
- Treu, L.; Kougias, P.G.; de Diego-Díaz, B.; Campanaro, S.; Bassani, I.; Fernández-Rodríguez, J.; Angelidaki, I. Two-year microbial adaptation during hydrogen-mediated biogas upgrading process in a serial reactor configuration. Bioresour. Technol. 2018, 264, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Calusinska, M.; Goux, X.; Fossépré, M.; Muller, E.E.L.; Wilmes, P.; Delfosse, P. A year of monitoring 20 mesophilic full-scale bioreactors reveals the existence of stable but different core microbiomes in bio-waste and wastewater anaerobic digestion systems. Biotechnol. Biofuels 2018, 11, 196. [Google Scholar] [CrossRef]
- Traversi, D.; Villa, S.; Lorenzi, E.; Degan, R.; Gilli, G. Application of a real-time qPCR method to measure the methanogen concentration during anaerobic digestion as an indicator of biogas production capacity. J. Environ. Manag. 2012, 111, 173–177. [Google Scholar] [CrossRef]
- Garcia-Peña, E.I.; Parameswaran, P.; Kang, D.W.; Canul-Chan, M.; Krajmalnik-Brown, R. Anaerobic digestion and co-digestion processes of vegetable and fruit residues: Process and microbial ecology. Bioresour. Technol. 2011, 102, 9447–9455. [Google Scholar] [CrossRef]
- Jaenicke, S.; Ander, C.; Bekel, T.; Bisdorf, R.; Dröge, M.; Gartemann, K.-H.; Jünemann, S.; Kaiser, O.; Krause, L.; Tille, F. Comparative and joint analysis of two metagenomic datasets from a biogas fermenter obtained by 454-pyrosequencing. PLoS ONE 2011, 6, e14519. [Google Scholar] [CrossRef] [PubMed]
- Mudhoo, A. Biogas Production: Pretreatment Methods in Anaerobic Digestion; John Wiley & Sons: Hoboken, NJ, USA, 2012; ISBN 1118404076. [Google Scholar]
- Kuever, J. The Family Syntrophaceae. In The Prokaryotes: Deltaproteobacteria and Epsilonproteobacteria; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer Berlin Heidelberg: Berlin/Heidelberg, Germany, 2014; pp. 281–288. ISBN 978-3-642-39044-9. [Google Scholar]
- Kuever, J. The Family Syntrophorhabdaceae. In The Prokaryotes: Deltaproteobacteria and Epsilonproteobacteria; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer Berlin Heidelberg: Berlin/Heidelberg, Germany, 2014; pp. 301–303. ISBN 978-3-642-39044-9. [Google Scholar]
- Yu, Y.; Lee, C.; Hwang, S. Analysis of community structures in anaerobic processes using a quantitative real-time PCR method. Water Sci. Technol. 2005, 52, 85–91. [Google Scholar] [CrossRef] [PubMed]
- McMahon, K.D.; Zheng, D.; Stams, A.J.M.; Mackie, R.I.; Raskin, L. Microbial population dynamics during start-up and overload conditions of anaerobic digesters treating municipal solid waste and sewage sludge. Biotechnol. Bioeng. 2004, 87, 823–834. [Google Scholar] [CrossRef] [PubMed]
- Walter, A.; Probst, M.; Franke-Whittle, I.H.; Ebner, C.; Podmirseg, S.M.; Etemadi-Shalamzari, M.; Hupfauf, S.; Insam, H. Microbiota in anaerobic digestion of sewage sludge with and without co-substrates. Water Environ. J. 2019, 33, 214–222. [Google Scholar] [CrossRef]
- Bedoya, K.; Hoyos, O.; Zurek, E.; Cabarcas, F.; Alzate, J.F. Annual microbial community dynamics in a full-scale anaerobic sludge digester from a wastewater treatment plant in Colombia. Sci. Total Environ. 2020, 726, 138479. [Google Scholar] [CrossRef]
- Nobu, M.K.; Narihiro, T.; Kuroda, K.; Mei, R.; Liu, W.-T. Chasing the elusive Euryarchaeota class WSA2: Genomes reveal a uniquely fastidious methyl-reducing methanogen. ISME J. 2016, 10, 2478–2487. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Walter, A.; Hanser, M.; Ebner, C.; Insam, H.; Markt, R.; Hupfauf, S.; Probst, M. Stability of the Anaerobic Digestion Process during Switch from Parallel to Serial Operation—A Microbiome Study. Sustainability 2022, 14, 7161. https://doi.org/10.3390/su14127161
Walter A, Hanser M, Ebner C, Insam H, Markt R, Hupfauf S, Probst M. Stability of the Anaerobic Digestion Process during Switch from Parallel to Serial Operation—A Microbiome Study. Sustainability. 2022; 14(12):7161. https://doi.org/10.3390/su14127161
Chicago/Turabian StyleWalter, Andreas, Maria Hanser, Christian Ebner, Heribert Insam, Rudolf Markt, Sebastian Hupfauf, and Maraike Probst. 2022. "Stability of the Anaerobic Digestion Process during Switch from Parallel to Serial Operation—A Microbiome Study" Sustainability 14, no. 12: 7161. https://doi.org/10.3390/su14127161
APA StyleWalter, A., Hanser, M., Ebner, C., Insam, H., Markt, R., Hupfauf, S., & Probst, M. (2022). Stability of the Anaerobic Digestion Process during Switch from Parallel to Serial Operation—A Microbiome Study. Sustainability, 14(12), 7161. https://doi.org/10.3390/su14127161







