Optimization of Cellulase Production by a Novel Endophytic Fungus Penicillium oxalicum R4 Isolated from Taxus cuspidata
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Microorganisms
2.2. Screening for Cellulase-Producing Fungi
2.3. Identification of the Selected Fungal Strain
2.4. Culture Conditions for Cellulase Production through Single-Factor Test
2.5. Response Surface Methodology
2.6. Artificial Neural Network and Genetic Algorithm
2.7. Determination of Cellulases Activity
2.8. Statistical Analysis
3. Results
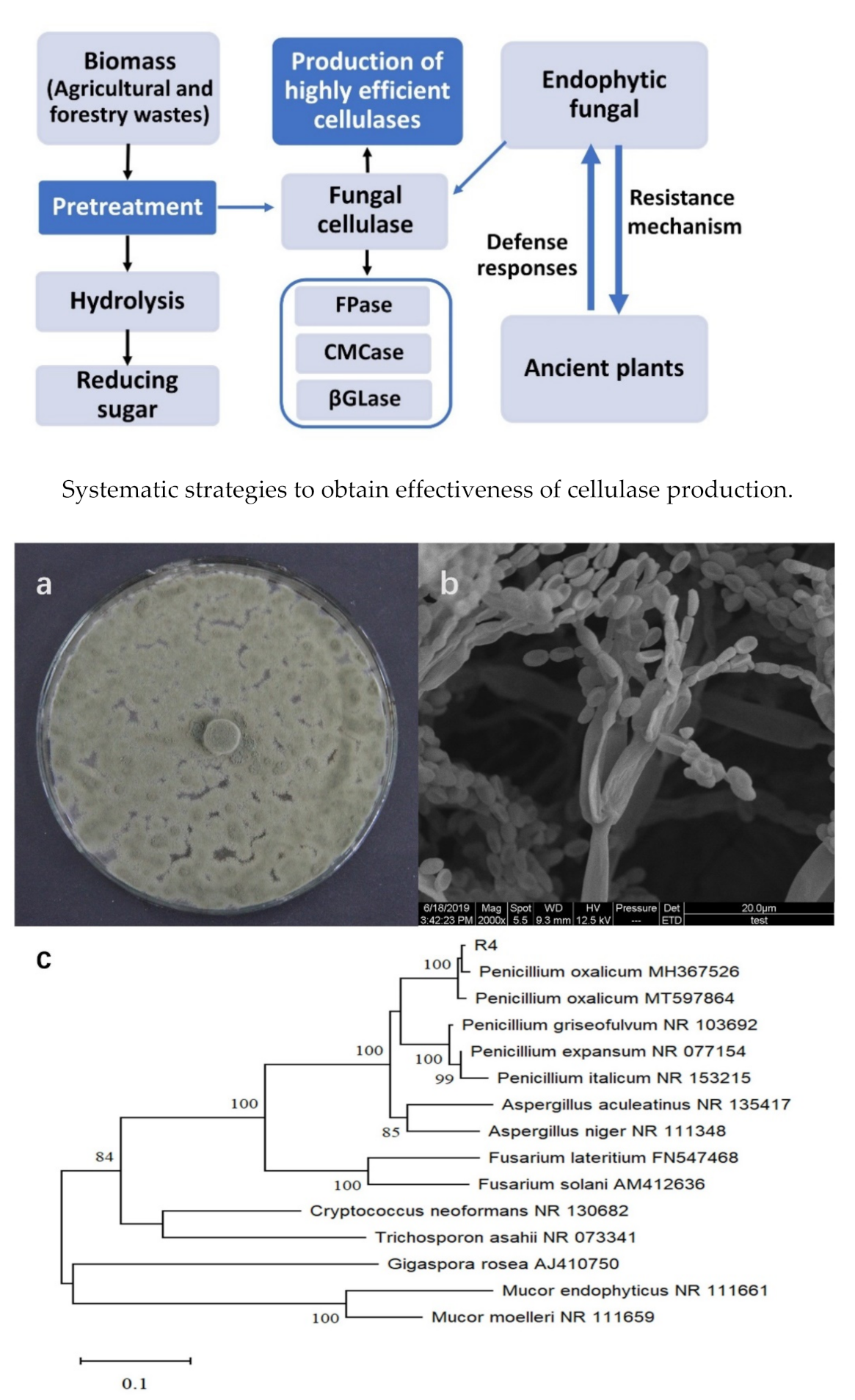
3.1. Screening and Identification of Fungal Isolate
3.2. Identification of the Most Potent Fungal Isolate
3.3. Optimization of Submerged Fermentation Conditions for Maximum Cellulases Production by P. oxalicum R4
3.3.1. Effect of Nitrogen Source
3.3.2. Optimization of Initial pH Value
3.3.3. Effect of Incubation Temperature
3.3.4. Effect of Incubation Period
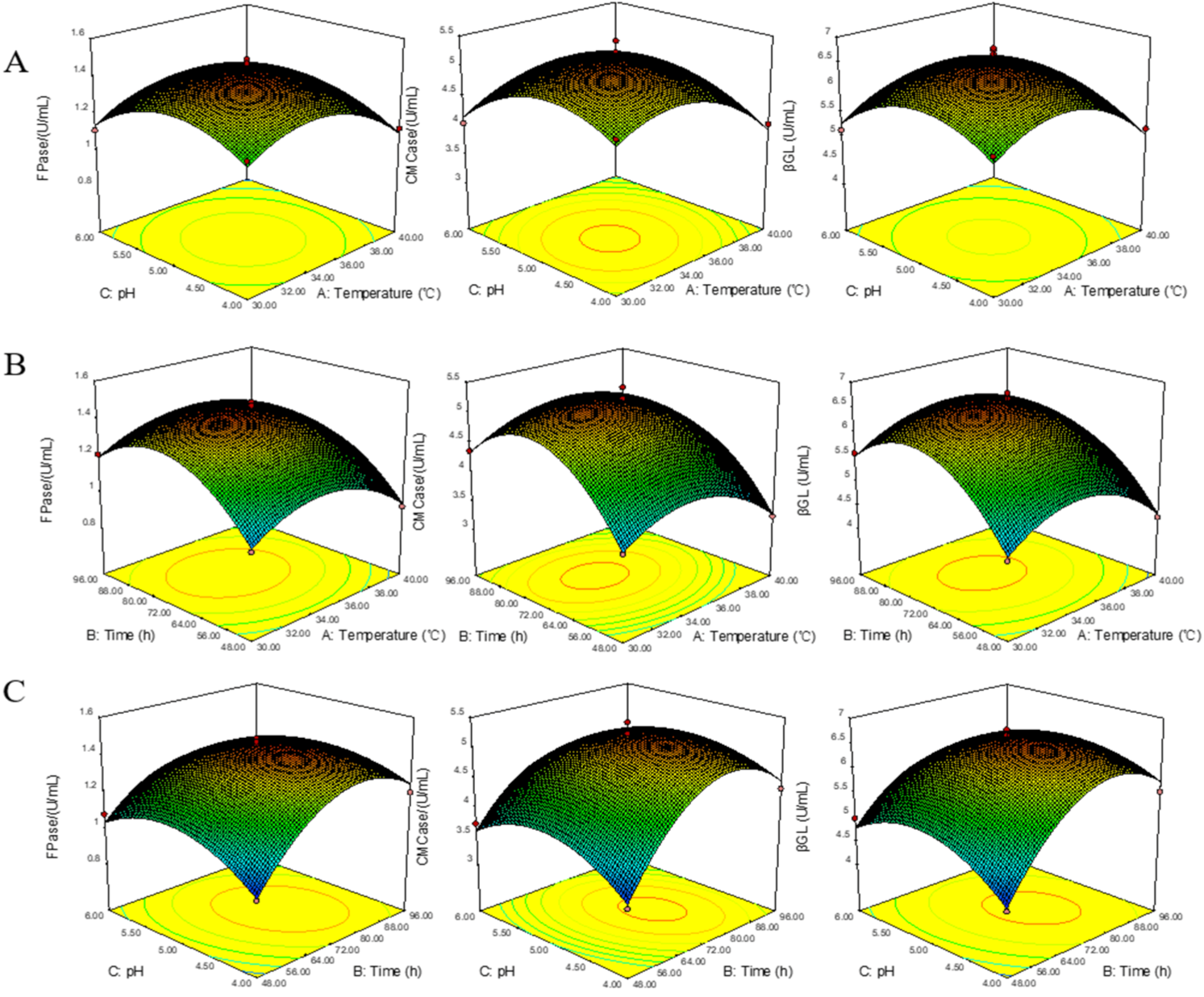
3.4. Optimization and Experimental Design of BBD Test
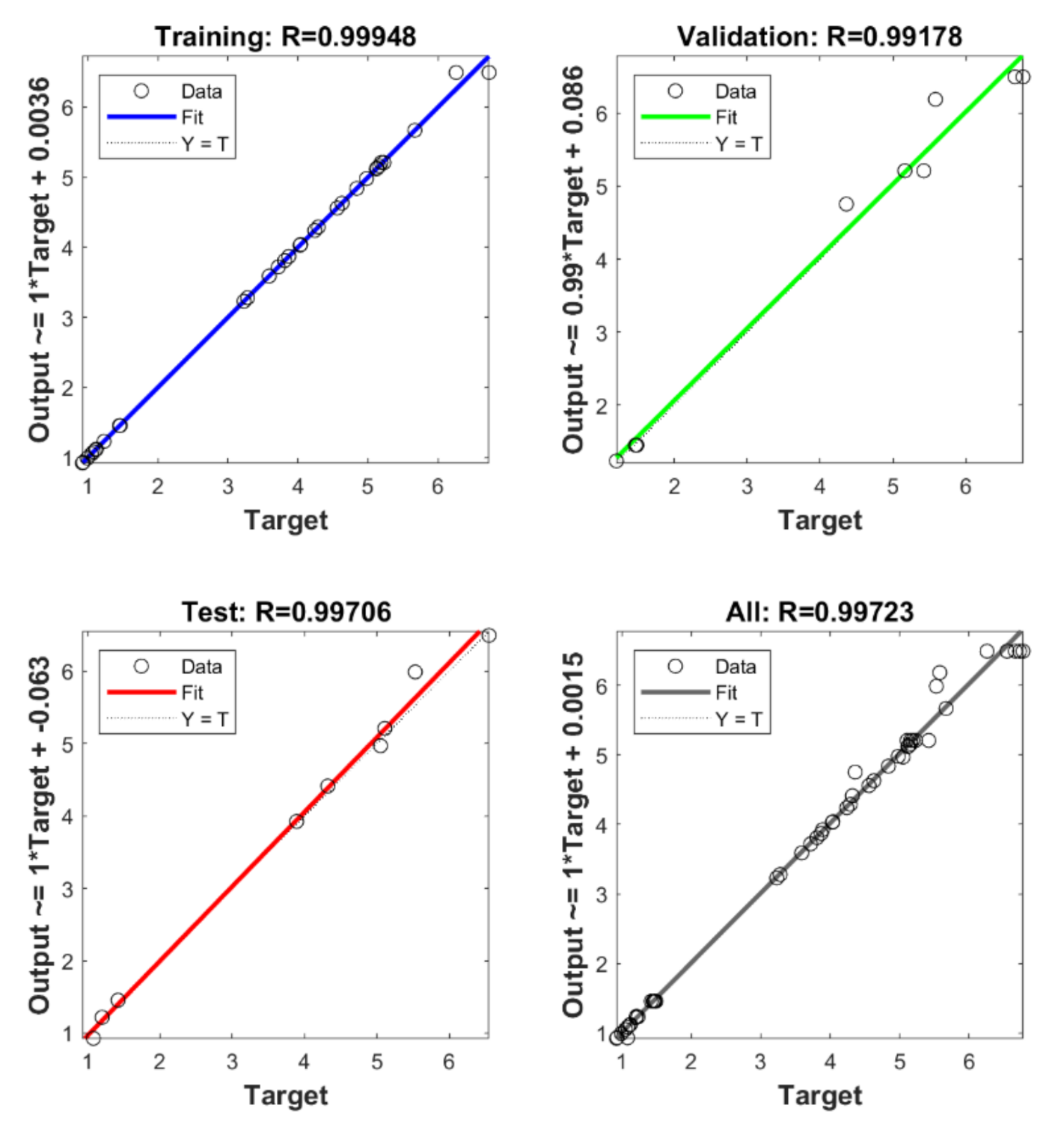
3.5. Artificial Neural Network and Genetic Algorithm
3.6. Verification and Comparison of Models
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| BBD | Box–Behnken design |
| ANN–GA | Artificial neural network–genetic algorithm |
| FPase | Filter Paperase |
| CMCase | Carboxymethyl Cellulase |
| βGLase | β-glucosidase |
| RSM | Response surface methodology |
References
- Yousef, S.; Kuliesiene, N.; Sakalauskaite, S.; Nenartavicius, T.; Daugelavicius, R. Sustainable green strategy for recovery of glucose from end-of-life euro banknotes. Waste Manag. 2021, 123, 23–32. [Google Scholar] [CrossRef]
- Yousef, S.; Eimontas, J.; Zakarauskas, K.; Striugas, N.; Mohamed, A. A new strategy for using lint-microfibers generated from clothes dryer as a sustainable source of renewable energy. Sci. Total Environ. 2021, 762. [Google Scholar] [CrossRef]
- Darwesh, O.M.; El-Maraghy, S.H.; Abdel-Rahman, H.M.; Zaghloul, R.A. Improvement of paper wastes conversion to bioethanol using novel cellulose degrading fungal isolate. Fuel 2020, 262, 116518. [Google Scholar] [CrossRef]
- Moreira, J.R.; Romeiro, V.; Fuss, S.; Kraxner, F.; Pacca, S.A. BECCS potential in Brazil: Achieving negative emissions in ethanol and electricity production based on sugar cane bagasse and other residues. Appl. Energy 2016, 179, 55–63. [Google Scholar] [CrossRef]
- Mansson, A.; Sanches-Pereira, A.; Hermann, S. Biofuels for road transport: Analysing evolving supply chains in Sweden from an energy security perspective. Appl. Energy 2014, 123, 349–357. [Google Scholar] [CrossRef]
- Sindhu, R.; Gnansounou, E.; Rebello, S.; Binod, P.; Varjani, S.; Thakur, I.S.; Nair, R.B.; Pandey, A. Conversion of food and kitchen waste to value-added products. J. Environ. Manag. 2019, 241, 619–630. [Google Scholar] [CrossRef]
- de Andrade, R.R.; Maugeri Filho, F.; Maciel Filho, R.; da Costa, A.C. Kinetics of ethanol production from sugarcane bagasse enzymatic hydrolysate concentrated with molasses under cell recycle. Bioresour. Technol. 2013, 130, 351–359. [Google Scholar] [CrossRef]
- Houfani, A.A.; Anders, N.; Spiess, A.C.; Baldrian, P.; Benallaoua, S. Insights from enzymatic degradation of cellulose and hemicellulose to fermentable sugars—A review. Biomass Bioenergy 2020, 134, 105481. [Google Scholar] [CrossRef]
- Sivarathnakumar, S.; Jayamuthunagai, J.; Baskar, G.; Praveenkumar, R.; Selvakumari, I.A.E.; Bharathiraja, B. Bioethanol production from woody stem Prosopis juliflora using thermo tolerant yeast Kluyveromyces marxianus and its kinetics studies. Bioresour. Technol. 2019, 293, 122060. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhao, Z.; Zhang, Y. Using straw as a bio-ethanol source to promote anaerobic digestion of waste activated sludge. Bioresour. Technol. 2019, 286. [Google Scholar] [CrossRef] [PubMed]
- Visser, E.M.; Falkoski, D.L.; de Almeida, M.N.; Maitan-Alfenas, G.P.; Guimarães, V.M. Production and application of an enzyme blend from Chrysoporthe cubensis and Penicillium pinophilum with potential for hydrolysis of sugarcane bagasse. Bioresour. Technol. 2013, 144, 587–594. [Google Scholar] [CrossRef]
- Saini, R.; Saini, J.K.; Adsul, M.; Patel, A.K.; Mathur, A.; Tuli, D.; Singhania, R.R. Enhanced cellulase production by Penicillium oxalicum for bio-ethanol application. Bioresour. Technol. 2015, 188, 240–246. [Google Scholar] [CrossRef]
- Troiano, D.; Orsat, V.; Dumont, M.J. Status of filamentous fungi in integrated biorefineries. Renew. Sust. Energ. Rev. 2020, 117, 109472. [Google Scholar] [CrossRef]
- Sopalun, K.; Iamtham, S. Isolation and screening of extracellular enzymatic activity of endophytic fungi isolated from Thai orchids. S. Afr. J. Bot. 2020, 134, 273–279. [Google Scholar] [CrossRef]
- Amobonye, A.; Bhagwat, P.; Singh, S.; Pillai, S. Enhanced xylanase and endoglucanase production from Beauveria bassiana SAN01, an entomopathogenic fungal endophyte. Fungal Biol. 2021, 125, 39–48. [Google Scholar] [CrossRef]
- Jing, L.; Zhao, S.; Xue, J.-L.; Zhang, Z.; Yang, Q.; Xian, L.; Feng, J.-X. Isolation and characterization of a novel Penicillium oxalicum strain Z1-3 with enhanced cellobiohydrolase production using cellulase-hydrolyzed sugarcane bagasse as carbon source. Ind. Crops. Prod. 2015, 77, 666–675. [Google Scholar] [CrossRef]
- Zhang, Z.; Liu, J.L.; Lan, J.Y.; Duan, C.J.; Ma, Q.S.; Feng, J.X. Predominance of Trichoderma and Penicillium in cellulolytic aerobic filamentous fungi from subtropical and tropical forests in China, and their use in finding highly efficient beta-glucosidase. Biotechnol. Biofuels 2014, 7, 14. [Google Scholar] [CrossRef]
- Gusakov, A.V. Alternatives to Trichoderma reesei in biofuel production. Trends Biotechnol. 2011, 29, 419–425. [Google Scholar] [CrossRef]
- Marques, N.P.; de Cassia Pereira, J.; Gomes, E.; da Silva, R.; Araújo, A.R.; Ferreira, H.; Rodrigues, A.; Dussán, K.J.; Bocchini, D.A. Cellulases and xylanases production by endophytic fungi by solid state fermentation using lignocellulosic substrates and enzymatic saccharification of pretreated sugarcane bagasse. Ind. Crops. Prod. 2018, 122, 66–75. [Google Scholar] [CrossRef]
- Kaushal, R.; Sharma, N.; Dogra, V. Molecular characterization of glycosyl hydrolases of Trichoderma harzianum WF5—A potential strain isolated from decaying wood and their application in bioconversion of poplar wood to ethanol under separate hydrolysis and fermentation. Biomass Bioenergy 2016, 85, 243–251. [Google Scholar] [CrossRef]
- Amobonye, A.; Bhagwat, P.; Pandey, A.; Singh, S.; Pillai, S. Biotechnological potential of Beauveria bassiana as a source of novel biocatalysts and metabolites. Crit. Rev. Biotechnol. 2020, 40, 1019–1034. [Google Scholar] [CrossRef]
- Cui, Q.; Liu, J.-Z.; Yu, L.; Gao, M.-Z.; Wang, L.-T.; Wang, W.; Zhao, X.-H.; Fu, Y.-J.; Jiang, J.-C. Experimental and simulative studies on the implications of natural and green surfactant for extracting flavonoids. J. Clean. Prod. 2020, 274. [Google Scholar] [CrossRef]
- Hankin, L.; Freed, S.Z. Bacteria in enucleated prostate. J. Urol. 1975, 6, 703–705. [Google Scholar] [CrossRef]
- Miller, G.A.I.L. Use of Dinitrosalicylic Acid Reagent for Detection of Reducing Sugars. Anal. Chem. 1959, 31. [Google Scholar] [CrossRef]
- Leite, R.S.R.; Alves-Prado, H.F.; Cabral, H.; Pagnocca, F.C.; Gomes, E.; Da-Silva, R. Production and characteristics comparison of crude β-glucosidases produced by microorganisms Thermoascus aurantiacus e Aureobasidium pullulans in agricultural wastes. Enzyme Microb. Technol. 2008, 43, 391–395. [Google Scholar] [CrossRef]
- Yao, L.; Anne Naeth, M.; Jobson, A. Soil microbial response to waste potassium silicate drilling fluid. J. Environ. Sci. 2015, 29, 189–198. [Google Scholar] [CrossRef]
- Jiang, W.; Chang, S.; Qu, Y.; Zhang, Z.; Xu, J. Changes on structural properties of biomass pretreated by combined deacetylation with liquid hot water and its effect on enzymatic hydrolysis. Bioresour. Technol. 2016, 220, 448–456. [Google Scholar] [CrossRef]
- Darwesh, O.M.; Sultan, Y.Y.; Seif, M.M.; Marrez, D.A. Bio-evaluation of crustacean and fungal nano-chitosan for applying as food ingredient. Toxicol. Rep. 2018, 5, 348–356. [Google Scholar] [CrossRef]
- Nishimura, M.; Ohkawara, T.; Tetsuka, K.; Kawasaki, Y.; Nakagawa, R.; Satoh, H.; Sato, Y.; Nishihira, J. Effects of yogurt containing Lactobacillus plantarum HOKKAIDO on immune function and stress markers. J. Tradit. Complement. Med. 2016, 6, 275–280. [Google Scholar] [CrossRef]
- Zhao, X.H.; Wang, W.; Tong, B.; Zhang, S.P.; Wei, D.Z. A newly isolated Penicillium oxalicum 16 cellulase with high efficient synergism and high tolerance of monosaccharide. Appl. Biochem. Biotechnol. 2016, 178, 173–183. [Google Scholar] [CrossRef]
- Huang, Q.X.; Wang, K.X.; Li, H.X.; Yi, S.; Zhao, X.H. Enhancing cellulosic ethanol production through coevolution of multiple enzymatic characteristics of beta-glucosidase from Penicillium oxalicum 16. Appl. Microbiol. Biotechnol. 2020, 104, 8299–8308. [Google Scholar] [CrossRef]
- Yao, G.S.; Wu, R.M.; Kan, Q.B.; Gao, L.W.; Liu, M.; Yang, P.; Du, J.; Li, Z.H.; Qu, Y.B. Production of a high-efficiency cellulase complex via beta-glucosidase engineering in Penicillium oxalicum. Biotechnol. Biofuels 2016, 9. [Google Scholar] [CrossRef]
- Sarkar, N.; Ghosh, S.K.; Bannerjee, S.; Aikat, K. Bioethanol production from agricultural wastes: An overview. Renew. Energy 2012, 37, 19–27. [Google Scholar] [CrossRef]
- Singhania, R.R.; Patel, A.K.; Saini, R.; Pandey, A. Industrial Enzymes: β-Glucosidases. In Current Developments in Biotechnology and Bioengineering; Pandey, A., Negi, S., Soccol, C.R., Eds.; Elsevier: Amsterdam, The Netherlands, 2017; pp. 103–125. [Google Scholar]
- Goukanapalle, P.K.R.; Kanderi, D.K.; Rajoji, G.; Shanthi Kumari, B.S.; Bontha, R.R. Optimization of cellulase production by a novel endophytic fungus Pestalotiopsis microspora TKBRR isolated from Thalakona forest. Cellulose 2020, 27, 6299–6316. [Google Scholar] [CrossRef]
- Pradhan, D.; Abdullah, S.; Pradhan, R.C. Chironji (Buchanania lanzan) fruit juice extraction using cellulase enzyme: Modelling and optimization of process by artificial neural network and response surface methodology. J. Food Sci. Tech. 2020. [Google Scholar] [CrossRef]
- Abdullah, S.; Pradhan, R.C.; Pradhan, D.; Mishra, S. Modeling and optimization of pectinase-assisted low-temperature extraction of cashew apple juice using artificial neural network coupled with genetic algorithm. Food Chem. 2021, 339, 127862. [Google Scholar] [CrossRef]
- Singh, R.; Kumar, R.; Bishnoi, K.; Bishnoi, N.R. Optimization of synergistic parameters for thermostable cellulase activity of Aspergillus heteromorphus using response surface methodology. Biochem. Eng. J. 2009, 48, 28–35. [Google Scholar] [CrossRef]
- Bhuyan, P.M.; Sandilya, S.P.; Nath, P.K.; Gandotra, S.; Subramanian, S.; Kardong, D.; Gogoi, D.K. Optimization and characterization of extracellular cellulase produced by Bacillus pumilus MGB05 isolated from midgut of muga silkworm (Antheraea assamensis Helfer). J. Asia Pacif. Entomol. 2018, 21, 1171–1181. [Google Scholar] [CrossRef]
- El-Deeb, B.; Fayez, K.; Gherbawy, Y. Isolation and characterization of endophytic bacteria from Plectranthus tenuiflorus medicinal plant in Saudi Arabia desert and their antimicrobial activities. J. Plant Interact. 2013, 8, 56–64. [Google Scholar] [CrossRef]
- Giauque, H.; Connor, E.W.; Hawkes, C.V. Endophyte traits relevant to stress tolerance, resource use and habitat of origin predict effects on host plants. New Phytol. 2019, 221, 2239–2249. [Google Scholar] [CrossRef] [PubMed]
- Li-Ming, H.U.; Yu, X.; Jie, L.I.; Qing-Cheng, Z. Study on the culture conditions of Aspergillus to produce cellulases. J. Henan Univ. Tech. 2007, 1, 70–74. [Google Scholar] [CrossRef]
- Han, F.; Sun, C.; Song, G.; Song, X.; Yuan, S. Conditions of cellulases from Trichoderma pseudokoningii UV III by liquid state fermentation. Ind. Microbiol. 2002, 32, 28–30. [Google Scholar]
- Jin, J.; Zhu, Y.; Jang, J.; Wang, S.; Xing, J.; Chiang, P.-C.; Fan, S.; Long, S. Enhancement of the polynomial functions response surface model for real-time analyzing ozone sensitivity. Front. Environ. Sci. Eng. 2021, 15. [Google Scholar] [CrossRef]
| Standad | Temperature | Time | pH | Actual Value (U/mL) | Predicted Value (U/mL) | ANN–GA (U/mL) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Order | (X1, °C) | (X2, h) | (X3) | Y1 | Y2 | Y3 | Y1 | Y2 | Y3 | Y1 | Y2 | Y3 |
| 1 | −1(30) | −1(48) | 0(5) | 0.99 | 3.59 | 4.56 | 1.01 | 3.61 | 4.64 | 0.99 | 3.59 | 4.56 |
| 2 | 1(40) | −1(48) | 0(5) | 0.92 | 3.23 | 4.24 | 0.94 | 3.26 | 4.31 | 0.92 | 3.23 | 4.24 |
| 3 | −1(30) | 1(96) | 0(5) | 1.21 | 4.36 | 5.58 | 1.19 | 4.33 | 5.51 | 1.21 | 4.36 | 5.58 |
| 4 | 1(40) | 1(96) | 0(5) | 1.11 | 3.87 | 5.12 | 1.09 | 3.85 | 5.04 | 1.11 | 3.87 | 5.12 |
| 5 | −1(30) | 0(72) | −1(4) | 1.23 | 4.63 | 5.67 | 1.2 | 4.53 | 5.53 | 1.23 | 4.63 | 5.67 |
| 6 | 1(40) | 0(72) | −1(4) | 1.12 | 4.03 | 5.16 | 1.09 | 3.92 | 5.03 | 1.12 | 4.03 | 5.16 |
| 7 | −1(30) | 0(72) | 1(6) | 1.11 | 4.04 | 5.13 | 1.14 | 4.15 | 5.26 | 1.11 | 4.04 | 5.13 |
| 8 | 1(40) | 0(72) | 1(6) | 1.05 | 3.81 | 4.84 | 1.08 | 3.91 | 4.98 | 1.05 | 3.81 | 4.84 |
| 9 | 0(35) | −1(48) | −1(4) | 0.93 | 3.28 | 4.29 | 0.94 | 3.36 | 4.35 | 0.93 | 3.28 | 4.29 |
| 10 | 0(35) | 1(96) | −1(4) | 1.2 | 4.32 | 5.53 | 1.24 | 4.45 | 5.74 | 1.20 | 4.32 | 5.53 |
| 11 | 0(35) | −1(48) | 1(6) | 1.08 | 3.72 | 4.98 | 1.03 | 3.59 | 4.77 | 1.08 | 3.72 | 4.98 |
| 12 | 0(35) | 1(96) | 1(6) | 1.08 | 3.89 | 5.05 | 1.07 | 3.81 | 4.99 | 1.08 | 3.89 | 5.05 |
| 13 | 0(35) | 0(72) | 0(5) | 1.46 | 5.19 | 6.73 | 1.46 | 5.22 | 6.6 | 1.47 | 5.27 | 6.57 |
| 14 | 0(35) | 0(72) | 0(5) | 1.49 | 5.42 | 6.67 | 1.46 | 5.22 | 6.6 | 1.47 | 5.27 | 6.57 |
| 15 | 0(35) | 0(72) | 0(5) | 1.45 | 5.23 | 6.26 | 1.46 | 5.22 | 6.6 | 1.47 | 5.27 | 6.57 |
| 16 | 0(35) | 0(72) | 0(5) | 1.42 | 5.11 | 6.55 | 1.46 | 5.22 | 6.6 | 1.47 | 5.27 | 6.57 |
| 17 | 0(35) | 0(72) | 0(5) | 1.47 | 5.16 | 6.78 | 1.46 | 5.22 | 6.6 | 1.47 | 5.27 | 6.57 |
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value Prob > F | ||
|---|---|---|---|---|---|---|---|
| Fpase activity | Model | 0.59 | 9 | 0.066 | 41.34 | <0.0001 | significant |
| X1 | 0.014 | 1 | 0.014 | 9.05 | 0.0197 | ||
| X2 | 0.058 | 1 | 0.058 | 36.19 | 0.0005 | ||
| X3 | 3.200 × 10−3 | 1 | 3.200 × 10−3 | 2.00 | 0.1998 | ||
| X1X2 | 2.250 × 10−4 | 1 | 2.250 × 10−4 | 0.14 | 0.7185 | ||
| X1X3 | 6.250 × 10−4 | 1 | 6.250 × 10−4 | 0.39 | 0.5514 | ||
| X2X3 | 0.018 | 1 | 0.018 | 11.41 | 0.0118 | ||
| X12 | 0.13 | 1 | 0.13 | 78.67 | <0.0001 | ||
| X22 | 0.22 | 1 | 0.22 | 136.74 | <0.0001 | ||
| X32 | 0.10 | 1 | 0.10 | 65.60 | <0.0001 | ||
| Residual | 0.011 | 7 | 1.578 | ||||
| Lack of Fit | 8.500 | 3 | 2.833 | 4.23 | 0.0987 | not significant | |
| Pure Error | 2.680 | 4 | 6.700 | ||||
| Cor Total | 0.61 | 16 | |||||
| CMCase activity | Source | Sum of Squares | df | Mean Square | F-value | p-value Prob > F | |
| Model | 8.03 | 9 | 0.89 | 42.85 | <0.0001 | significant | |
| X1 | 0.35 | 1 | 0.35 | 16.94 | 0.0045 | ||
| X2 | 0.86 | 1 | 0.86 | 41.20 | 0.0004 | ||
| X3 | 0.080 | 1 | 0.080 | 3.84 | 0.0908 | ||
| X1X2 | 4.225 × 10−3 | 1 | 4.225 × 10−3 | 0.20 | 0.6660 | ||
| X1X3 | 0.034 | 1 | 0.034 | 1.64 | 0.2407 | ||
| X2X3 | 0.19 | 1 | 0.19 | 9.09 | 0.0195 | ||
| X12 | 1.35 | 1 | 1.35 | 65.06 | <0.0001 | ||
| X22 | 3.35 | 1 | 3.35 | 160.96 | <0.0001 | ||
| X32 | 1.17 | 1 | 1.17 | 56.20 | 0.0001 | ||
| Residual | 0.15 | 7 | 0.021 | ||||
| Lack of Fit | 0.089 | 3 | 0.030 | 2.10 | 0.2435 | not significant | |
| Pure Error | 0.057 | 4 | 0.014 | ||||
| Cor Total | 8.18 | 16 | |||||
| βGLase activity | Source | Sum of Squares | df | Mean Square | F-value | p-value Prob > F | |
| Model | 11.11 | 9 | 1.23 | 24.07 | 0.0001 | significant | |
| X1 | 0.31 | 1 | 0.31 | 6.09 | 0.0430 | ||
| X2 | 1.29 | 1 | 1.29 | 25.12 | 0.0015 | ||
| X3 | 0.053 | 1 | 0.053 | 1.03 | 0.3440 | ||
| X1X2 | 4.900 × 10−3 | 1 | 4.900 × 10−3 | 0.096 | 0.7662 | ||
| X1X3 | 0.012 | 1 | 0.012 | 0.24 | 0.6420 | ||
| X2X3 | 0.34 | 1 | 0.34 | 6.67 | 0.0363 | ||
| X12 | 2.32 | 1 | 2.32 | 45.30 | 0.0003 | ||
| X22 | 4.05 | 1 | 4.05 | 78.90 | <0.0001 | ||
| X32 | 1.81 | 1 | 1.81 | 35.25 | 0.0006 | ||
| Residual | 0.36 | 7 | 0.051 | ||||
| Lack of Fit | 0.19 | 3 | 0.062 | 1.44 | 0.3550 | not significant | |
| Pure Error | 0.17 | 4 | 0.043 | ||||
| Cor Total | 11.47 | 16 |
| Method | Temperature (°C) | Time (h) | pH | Predictive Value | Actual Value | ||||
|---|---|---|---|---|---|---|---|---|---|
| Fpase | CMCase | βGLase | Fpase | CMCase | βGLase | ||||
| RAW | 0.87 | 3.31 | 2.83 | ||||||
| RSM | 34.22 | 77.09 | 4.87 | 1.47 | 5.28 | 6.66 | 1.45 | 5.24 | 6.34 |
| ANN–GA | 34.83 | 78.11 | 4.98 | 1.45 | 5.27 | 6.35 | 1.45 | 5.27 | 6.35 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, H.; Dou, M.; Wang, X.; Guo, N.; Kou, P.; Jiao, J.; Fu, Y. Optimization of Cellulase Production by a Novel Endophytic Fungus Penicillium oxalicum R4 Isolated from Taxus cuspidata. Sustainability 2021, 13, 6006. https://doi.org/10.3390/su13116006
Li H, Dou M, Wang X, Guo N, Kou P, Jiao J, Fu Y. Optimization of Cellulase Production by a Novel Endophytic Fungus Penicillium oxalicum R4 Isolated from Taxus cuspidata. Sustainability. 2021; 13(11):6006. https://doi.org/10.3390/su13116006
Chicago/Turabian StyleLi, Hongkun, Meijia Dou, Xinyu Wang, Na Guo, Ping Kou, Jiao Jiao, and Yujie Fu. 2021. "Optimization of Cellulase Production by a Novel Endophytic Fungus Penicillium oxalicum R4 Isolated from Taxus cuspidata" Sustainability 13, no. 11: 6006. https://doi.org/10.3390/su13116006
APA StyleLi, H., Dou, M., Wang, X., Guo, N., Kou, P., Jiao, J., & Fu, Y. (2021). Optimization of Cellulase Production by a Novel Endophytic Fungus Penicillium oxalicum R4 Isolated from Taxus cuspidata. Sustainability, 13(11), 6006. https://doi.org/10.3390/su13116006







