Abstract
The European Union (EU) is facing the recent arrival of the bacterium Xylella fastidiosa. Its fast spread has caused great alarm because of the economic impact it implies for the agroeconomy of European countries. Among its insect vectors, Philaenus spumarius has been demonstrated to transmit the bacterium from infected to uninfected trees in the EU, where different measures have been established to control it. One of the proposals to manage this vector is the augmentation of natural enemy populations. However, the identification of candidate predator species is essential if such a management system is to be introduced. The present paper describes a set of species-specific primers designed to detect the presence of P. spumarius DNA in soil arthropod fauna generalist predators’ gut which can reveal candidate species for the pest’s biological control. Such primers have been proven to be a useful and reliable taxonomic tool for P. spumarius identification at any life stage, i.e., nymphs. This rapid and accurate identification is essential for control strategies designed to avoid the spread of the pest and consequently the considerable economic losses it causes in crops.
1. Introduction
Olive quick decline syndrome (OQDS) is a new severe disease with which the bacterium Xylella fastidiosa subsp pauca Wells et al. (1987) is strongly associated. In early stages, the disease can be asymptomatic but usually causes leaf scorching, extensive dieback, and finally, the death of the entire plant affected [1]. As the European Union (EU) produces 73.2% and 34.4% of the world’s olive oil and table olives, respectively [2], the recent arrival of X. fastidiosa to Europe has triggered an alarm due to the huge impact it could have for the economies of those countries that produce and export table olives and olive oil. The first detection of the bacterium X. fastidiosa subsp pauca in the EU territory was in olive trees in the Italian region of Apulia in 2013, with nearly 10,000 ha affected by the strain ST53 [3]. In July 2015, the bacterium X. fastidiosa subsp multiplex was reported for the first time in France [4]. Spain, the main producer and exporter of olive products worldwide, reported the first X. fastidiosa subsp fastidiosa outbreak in Prunus avium in the Balearic Islands in October 2016 and confirmed the presence of X. fastidiosa subsp multiplex in the continental territory in June 2017 [5]. By 2018, the presence of this subspecies of X. fastidiosa has been confirmed in olive groves in Madrid and Andalucía, the main olive oil and table olives producer region in Spain [6] (pp. 72–76), [7] (pp. 146–150).
Until now, at least these three different subspecies of X. fastidiosa have been detected affecting different plant species in the EU. Transmission of this wide host spectrum bacterium, with more than 300 different hosts, occurs through vector insects that inhabit the EU territory. Strict rules have been introduced on the basis of a full pest risk assessment by the European Food Safety Authority [1]. The European Union adopted the executive decision 2015/789/UE to avoid the propagation of X. fastidiosa. These measures have severe impact on the local agroeconomy and the environment and include the removal and destruction of all host plants of X. fastidiosa and harsh phytosanitary treatments against the vector [8].
In the EU, the meadow spittlebug Philaenus spumarius (Linnaeus, 1758) has been reported to be a vector of X. fastidiosa, transmitting the bacteria from infected to uninfected plants, including olive trees [9] (see also [10,11,12]), and P. italosignus and Neophilaenus campestris also possibly act as vectors [13]. P. spumarius is a ubiquitous and highly polyphagous species [14] that can feed on a large variety of plant species, including olive trees [15]. Control programs for this vector integrate different measures such as soil and vegetation management, application of insecticides, and augmentation of natural enemy populations [1].
In this context, biological control mediated by predators may offer a solution to manage P. spumarius populations, being the most environmentally friendly control strategy against X. fastidiosa. The soil arthropod fauna in olive groves is composed of different taxa, and some of them, such as spiders, carabids, or staphylinids, are generalist predators [16]. However, selecting the most appropriate predator is often a difficult task, given that the trophic web is complex and that human presence in the ecosystem alters the arthropods natural behaviour [17]. Also, postmortem visual examination of the gut contents of candidate predators is usually unsuccessful given their feeding habits and the size of preys’ remains [18]. To overcome this issue, PCR-based techniques allow for the reliable identification of prey species DNA in their predators’ gut and require a species-specific primers design. Gut content analysis, however, implies the detection of minute quantities of the prey’s DNA, which is expected to be degraded by the digestive process [19,20]. Thus, amplifying small fragments of multicopy genes can improve detection rates. It is for this reason that the mitochondrial genome is a suitable target for primers design because of the presence of several copies per organelle and also several organelles per cell. Likewise, the interspecific variability of the mitochondrial cytochrome oxidase I (COI) gene, considered as the universal barcode for animal species identification [21], makes it suitable for distinguishing at species level.
In addition, P. spumarius traditionally has been identified based on morphological characteristics. To distinguish between the meadow spittlebug and other closely related species, such as those from genus Neophilaenus, adults’ characteristics lend themselves for identification purposes. However, the immature stages are hard to identify unambiguously because both of those genera are very similar. Therefore, discrimination of these species using their morphological features is difficult at the early stages and morphological characteristics alone are not reliable for proper identification, especially if both species coexist in the same area, as in this case.
The aims of this work are, firstly, to develop several species-specific primer pairs of P. spumarius DNA for molecular gut content analysis to identify future candidate predators of this species in the field for biological control. Secondly, to use these primers as useful and reliable taxonomic tools for P. spumarius identification at any life stage. A rapid an accurate identification of effective predators is vital for control strategies designed to avoid the spread of this pest.
2. Materials and Methods
The 5′ region of the mitochondrial COI gene was checked for in silico primer designing. Over 200 sequences from the Philaenus genus (P. spumarius and closely related species), Aphrophoridae members, and other potential predators arthropods present in the olive agroecosystem were retrieved from BOLD [22] and GenBank databases [23] (Table S1). The set of sequences was grouped using BioEdit v.7.0.9.0 [24] and aligned employing the K-Alignment tool available on EMBL-EBI [25]. Six species-specific primers were developed (Table 1) and their combination amplified eight different fragments whose sizes encompassed 79–175 bp (Figure 1).

Table 1.
PCR primers designed to amplify P. spumarius COI gene fragment.
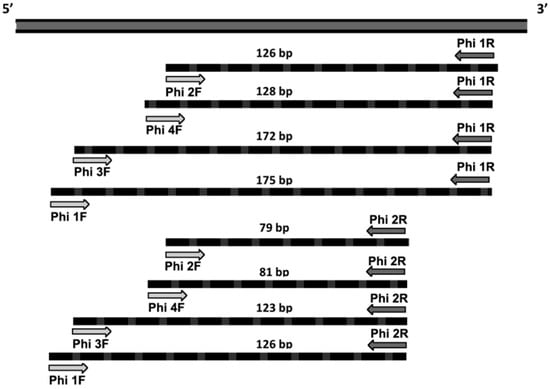
Figure 1.
Diagram showing primer combinations and COI amplicon sizes in base pairs. Dark and light grey arrows indicate the designed primers (names in bold).
Specificity assays were conducted using arthropod specimens of: Philaenus spumarius, Neophilaenus campestris (Fallen, 1805), Neophilaenus lineatus (Linnaeus, 1758), Cercopis intermedia (Latreille 1829), Alopecosa cuneata (Clerck, 1758), Synema globosum (Fabricius, 1775), Tapinoma nigerrimum (Nylander, 1856), Forficula auricularia (Linnaeus, 1758), Harpalus griseus (Panzer, 1796), Orthomus barbarus (Dejean, 1828), and Bactrocera oleae (Rossi, 1790). Eight Aphrophoridae nymphs were also used for specificity and identification assay. Cercopids and aphrophorids were collected and provided from Valencia by Dra. Cristina Navarro and Dra. Altea Calabuig (Elytra Agroscience Services). We collected the remaining specimens from a previous study [26].
All the genomic DNAs were extracted using the Speedtools Tissue DNA Extraction Kit (Biotools, B&M Labs, Madrid, Spain) and their integrity checked in 0.8% agarose gels stained with ethidium bromide (0.5 mg/mL). The quality of all DNA extracted was examined on a first set of PCR reactions using the universal COI primers LCO1490 and HCO2198 [27] under the conditions reported in Lantero et al. [26]. Amplified products with these primes from Aphrophoridae nymphs were used as templates for sequencing reactions. These reactions were carried out with the “BIG Dye Terminator Cycle Sequencing Ready Reaction Kit” (Applied Biosystems, Inc., Foster City, CA, USA) on a 3730 DNA Analyzer (Applied Biosystems, Inc., Foster City, CA, USA), using the primers employed for the amplification step, at the Genomic Unit of The Complutense University of Madrid.
Then, the specificity of the designed primers for P. spumarius was assessed in PCR reactions using 10 ng of DNA from the arthropod species stated above. The final volume was 12.5 µL with 6.25 µL Taq PCR Master Mix (Qiagen, Hilden, Germany), 0.8 mM of each primer, and 1.5 µM of MgCl2. The PCR program included an initial denaturation of 5 min at 94 °C, followed by 30 cycles of 94 °C for 45 s, 62 °C for 1 min, and 72 °C for 1 min.
The sensitivity assays were carried out in PCR reactions performed as in the specificity assay. These involved control samples from P. spumarius and the potential predator A. cuneata (5 ng/µL), plus samples with the dilution of spittlebugs’ DNA in spiders’ DNA in ratios of 1:100 (0.05 ng/µL to 5 ng/µL), 1:1000 (5 pg/µL to 5 ng/µL) 1:2000 (2.5 pg/µL to 5 ng/µL), and 1:4000 (1.25 pg/µL to 5 ng/µL). All PCR products from specificity and sensitivity assays were visualized in 3.5% agarose gels stained with ethidium bromide (0.5 mg/mL).
Once the specificity and sensitivity of the primer pairs were tested, a first approach in the field was performed. Sixty-three spiders were collected by hand as potential predators of P. spumarius in an olive-growing area in the southeast of Madrid, located near the locality (Villarejo de Salvanés, Spain) of Xylellas’ outbreak in Madrid [6] at the beginning of May, expecting high levels of vectors’ infestation. The DNA from the spiders’ gut was isolated and amplified as described in [26]. PCR reactions were conducted at least twice using the two primer pairs selected in the previous tests.
3. Results and Discussion
The designed primer pairs amplified only DNA from P. spumarius, revealing their high specificity (Figure 2). Amplicon sizes arose from 79 bp (Phi 1F–Phi 1R) to 175 bp (Phi 2F–Phi 2R) as expected (Figure 1). They are within the range for postdigestion amplification products reported in previous surveys with other species (78–242 bp [28], 160–281 bp [29], 101–274 bp [30], 130–330 bp [31], and 150–345 bp [32]). The amplicon size is crucial for postmortem gut content analyses to be successful. Given the preys’ DNA degradation during the digestive processes, amplifying small fragments from multicopy genes improves the detection rates of these minute quantities of preys’ DNA in the predators’ gut [19,20,26].
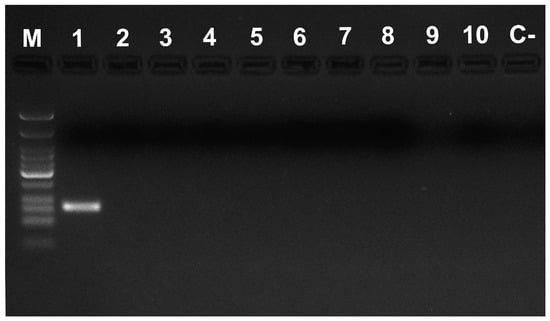

Figure 2.
Specificity assays with the primer pair Phi 3F–Phi 2R. M: Low Molecular Weight DNA Ladder (New England Biolabs); Lanes 1–12 PCR reactions using as template 10 ng of DNA from: 1, P. spumarius; 2, Neophilaenus campestris; 3, Neophilaenus lineatus; 4, Cercopis intermedia; 5, T. nigerrimum; 6, F. auricularia; 7, H. griseus; 8, O. barbarus; 9, B. oleae; 10, S. globosum; C-PCR negative control.
The proper identification of the meadow spittlebug at any life stage is particularly important when prompt identification is needed (e.g., in rapid action for pest control management in quarantine areas). Thus, the species-specific primers were also employed on DNA from nymphs sampled in the field that could not be assigned unambiguously to P. spumarius based on their morphological characteristics. Outcomes showed that three nymphs from Valencia and one from Madrid could be accurately assigned to P. spumarius after PCR reactions with our designed primers (Figure 3). Sequences amplified with the universal COI primers PCRs confirmed that result since these four DNA sequences were identical to the P. spumarius sequence deposited at GenBank [23] (accession number KR042125). DNAs obtained from nymphs that were not amplified with the species-specific primers for P. spumarius were identified as Neophilaenus lineatus after sequencing with the COI universal primers and querying genetic databases [22,23]. Only a previous work about phylogeny and DNA barcoding of P. spumarius and its related species is found in the literature [14]. The authors used primer pairs designed for amplifying DNA from butterflies and moths [33], and they observed that the amplicon of 399 bp had enough interspecific variability to necessarily distinguish among the Philaenus species after sequencing all of them.
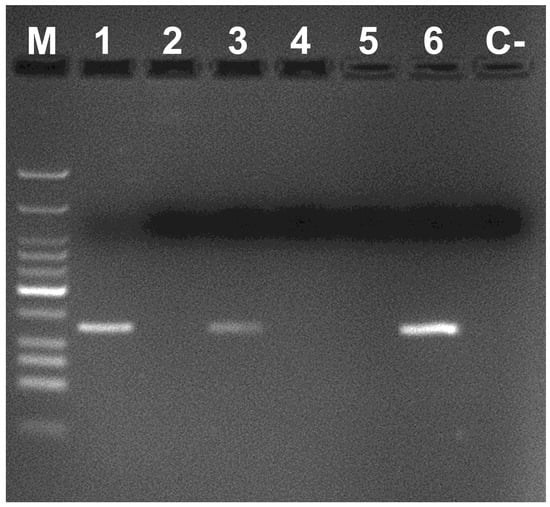
Figure 3.
Identification of the meadow spittlebug nymphs with the primer combination Phi 1F–Phi 1R. M: Low Molecular Weight DNA Ladder (New England Biolabs). Lanes 1–5 DNA of nymphs from Valencia region, lane 6 P. spumarius (positive control) and C-PCR negative control.
Although all primer combinations were highly specific, primer pairs Phi 1F–Phi 1R and Phi 2F–Phi 2R produced the highest PCR efficiency and they were selected to perform the sensitivity tests to figure out the limits of the meadow spittlebug DNA detection. Both primer pairs were very sensitive given that P. spumarius DNA was amplified in all the heterospecific DNA mixes, even when potential predator DNA was in great excess (Ratio 1:4000) (Figure 4). Outcomes returned a detection limit of 2.5 pg/µL of P. spumarius DNA, surpassing that obtained for the same gene on sharpshooters (6 pg/µL [34]), equaling the one obtained for Haplodiplosis marginata [35] and slightly higher than that for the olive pest, Bactrocera oleae (1 pg/µL [26]).
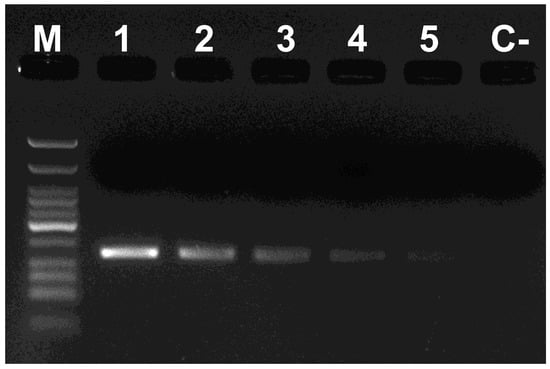
Figure 4.
Sensitivity assays with the primer pair Phi 1F–Phi 1R. M: Low Molecular Weight DNA Ladder (New England Biolabs); Lanes 1–8 PCR reactions using as template DNA from: 1, P. spumarius (5 ng/μL); 2, dilution 1:100 of DNA of P. spumarius with DNA of A. cuneata; 3, dilution 1:1000; 4, dilution 1:2000; 5, dilution 1:4000; C-PCR negative control.
These two primer combinations were thus very appropriate for molecular detection of P. spumarius DNA under laboratory conditions. However, it was mandatory to test their efficacy in DNA analysis of the gut content of spiders collected in olive orchards, the most abundant potential predators during spring according our previous surveys (data unpublished). In this first approach, the DNA of P. spumarius was detected in 4 out of the 63 spiders tested, representing the 6.34% of the sample (Figure 5). For this outcome, it is important to note that (a) no meadow spittlebugs were deployed in the field, unlike other surveys [31], meaning that predation occurred at natural ratios, and (b) most spiders were collected at an olive grove where, despite the high density of foam in the vegetation—a sign of meadow spittlebug activity in the field—the number of visualized emerged adults (real spiders’ prey) was low. The relevant predatory activity of spiders has already been demonstrated with other spider species and preys [30], stressing the utility of this group to mitigate the spread of some pests.
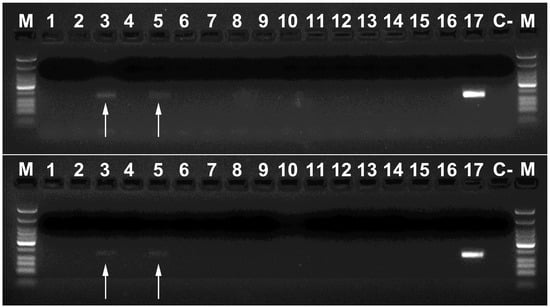

Figure 5.
Detection of P. spumarius DNA with Phi 2R–Phi 2F (two independent replications are shown, positive reactions indicated with arrows). M: Low Molecular Weight DNA Ladder (New England Biolabs). Lanes 1–16 PCR reactions using as template DNAs from guts of collected spiders in the field. Lane 17 P. spumarius (positive control) DNA and C-PCR negative control.
In short, the primers presented here constitute a reliable, sensitive, and robust tool for the specific detection of P. spumarius DNA. This PCR based method proved to be a useful taxonomic tool for species identification at immature stages. Since the current status relies almost exclusively on adult characteristics, PCR analysis is particularly useful in cases requiring a prompt identification. The designed primers also have shown high specificity and sensitivity in critical conditions of a very limiting template, stressing the advantage of using DNA markers to track predation against insect pests or disease vectors present on crops. Spiders have been identified as predators of Philaenus and, thus, as potential natural enemies to control its populations in the olive groves as an alternative to insecticides. Research is ongoing to find more potential predators of P. spumarius, which additionally will improve the knowledge of food webs at this agroecosystem.
Supplementary Materials
The following are available online at http://www.mdpi.com/2071-1050/10/7/2207/s1, Table S1: Philaenus and other arthropod sequences used in the alignment procedure for the development of species-specific primers for P. spumarius.
Author Contributions
Conceptualization, E.L., B.M. and C.C.; Methodology, E.L. and C.C.; Writing-Original Draft Preparation, E.L. and B.M.; Writing-Review & Editing, B.M., S.P. and C.C.; Funding Acquisition, S.P.
Funding
This research was funded by Ministerio de Economía y Competitividad of Spain (part of project RTA2013-00039-C03-03, Biological control of Bactrocera oleae: Effect of landscape structure and importance of predation) and European Regional Development Fund.
Acknowledgments
The authors thank to Cristina Navarro, Altea Calabuig (Elytra Agroscience Services), Ismael Sanchez-Ramos, Marta Ortega, Alejandro Rescia, Manuel González-Núñez, África Sanchiz, Santos Domínguez and Ignacio Juarez for their sampling support. We would like, also, to thanks to two anonymous referees for their useful comments on a first version of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- EFSA (European Food Safety Authority). Workshop on Xylella fastidiosa: Knowledge Gaps and Research Priorities for the EU. EFSA Supporting Publication 2016: EN-1039. 2016. Available online: https://efsa.onlinelibrary.wiley.com/doi/epdf/10.2903/sp.efsa.2016.EN-1039 (accessed on 3 June 2018).
- IOC (International Olive Council). 2016. Available online: http://www.internationaloliveoil.org/ (accessed on 3 June 2018).
- Saponari, M.; Boscia, D.; Nigro, D.; Martelli, G.P. Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (Southern Italy). J. Plant Pathol. 2013, 95, 668. [Google Scholar] [CrossRef]
- EPPO (European and Mediterranean Plant Protection Organization). Available online: http://www.eppo.int/QUARANTINE/special_topics/Xylella_fastidiosa/Xylella_fastidiosa.htm (accessed on 3 June 2018).
- MAPAMA (Ministerio de Agricultura y Pesca, Alimentación y Medio Ambiente). Available online: http://www.mapama.gob.es/es/agricultura/temas/sanidad-vegetal/xylella-fastidiosa/ (accessed on 3 June 2018).
- Boletín Oficial de la Comunidad de Madrid. No. 86. Available online: https://www.bocm.es/boletin/CM_Orden_BOCM/2018/04/11/BOCM-20180411-16.PDF (accessed on 3 June 2018).
- Boletín Oficial de la Junta de Andalucía. No. 81. Available online: http://www.juntadeandalucia.es/eboja/2018/81/BOJA18-081-00005-7337-01_00134737.pdf (accessed on 3 June 2018).
- Plant Health and Biosecurity, European Commission. Available online: https://ec.europa.eu/food/plant/plant_health_biosecurity/legislation/emergency_measures/xylella-fastidiosa_en (accessed on 3 June 2018).
- Saponari, M.; Loconsole, G.; Cornara, D.; Yokomi, R.K.; De Stradis, A.; Boscia, D.; Bosco, D.; Martelli, G.P.; Krugner, R.; Porcelli, F. Infectivity and Transmission of Xylella fastidiosa by Philaenus spumarius (Hemiptera: Aphrophoridae) in Apulia, Italy. J. Econ. Entomol. 2014, 107, 1316–1319. [Google Scholar] [CrossRef] [PubMed]
- Luvisi, A.; Nicolì, F.; De Bellis, L. Sustainable Management of Plant Quarantine Pests: The Case of Olive Quick Decline Syndrome. Sustainability 2017, 9, 659. [Google Scholar] [CrossRef]
- Bucci, E.M. Xylella fastidiosa, a new plant pathogen that threatens global farming: Ecology, molecular biology, search for remedies. Biochem. Biophys. Res. Commun. 2018, 502, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Soubeyrand, S.; Jerphanion, P.; Martin, O.; Saussac, M.; Manceau, C.; Hendrikx, P.; Lannou, C. Inferring pathogen dynamics from temporal count data: The emergence of Xylella fastidiosa in France is probably not recent. New Phytol. 2018, 219, 824–836. [Google Scholar] [CrossRef] [PubMed]
- MAPAMA (Ministerio de Agricultura y Pesca, Alimentación y Medio Ambiente). Available online: http://www.mapama.gob.es/es/agricultura/temas/sanidad-vegetal/xylellafastidiosa_contingencia_febrero2018_tcm30-445867.pdf (accessed on 23 June 2018).
- Seabra, S.G.; Pina-Martins, F.; Marabuto, E.; Yurtsever, S.; Halkka, O.; Quartau, J.A.; Paulo, O.S. Molecular phylogeny and DNA barcoding in the meadow-spittlebug Philaenus spumarius (Hemiptera, Cercopidae) and its related species. Mol. Phylogenet. Evol. 2010, 56, 462–467. [Google Scholar] [CrossRef] [PubMed]
- Cornara, D.; Saponari, M.; Zeilinger, A.R.; de Stradis, A.; Boscia, D.; Loconsole, G.; Bosco, D.; Martelli, G.P.; Almeida, R.P.P.; Porcelli, F. Spittlebugs as vectors of Xylella fastidiosa in olive orchards in Italy. J. Pest Sci. 2017, 90, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Albertini, A.; Pizoloto, R.; Petacchi, R. Carabid patterns in olive orchards and woody semi-natural habitats: First implications for conservation biological control against Bactrocera oleae. BioControl 2017, 62, 71. [Google Scholar] [CrossRef]
- Rosenheim, J.A.; Limburg, D.D.; Colfer, R.G. Impact of generalist predators on a biological control agent, Chrysoperla carnea: Direct observations. Ecol. Appl. 1999, 9, 409–417. [Google Scholar] [CrossRef]
- Symondson, W.O.C. Molecular identification of prey in predator diets. Mol. Ecol. 2002, 11, 627–641. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.M.J.; Ingels, B.; Wheatley, A.; Rhule, E.L.; de Clercq, P.; van Leeuwen, P.; Thomas, A. Intraguild predation by Harmonia axyridis (Coleoptera: Coccinellidae) on native insects in Europe: Molecular detection from field samples. Entomol. Sci. 2015, 18, 130–133. [Google Scholar] [CrossRef]
- MacDonald, A.J.; Young, M.J.; Lintermans, M.; Sarre, S.D. Primers for detection of Macquarie perch from environmental and trace DNA samples. Conserv. Genet. Resour. 2014, 6, 551–553. [Google Scholar] [CrossRef]
- Pentinsaari, M.; Salmela, H.; Mutanen, M.; Roslin, T. Molecular evolution of a widely-adopted taxonomic marker (COI) across the animal tree of life. Sci. Rep. 2016, 6, 35275. [Google Scholar] [CrossRef] [PubMed]
- BOLD Database. Available online: http://v3.boldsystems.org/index.php/databases (accessed on 3 June 2018).
- GenBank Database. Available online: https://www.ncbi.nlm.nih.gov/genbank/ (accessed on 3 June 2018).
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/ NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- EMBL-EBI (European Bioinformatics Institute) Tools: Multiple Sequence Alignment. Available online: https://www.ebi.ac.uk/Tools/msa/ (accessed on 3 June 2018).
- Lantero, E.; Matallanas, B.; Ochando, M.D.; Pascual, S.; Callejas, C. Specific and sensitive primers for the detection of predated olive fruit flies, Bactrocera oleae (Diptera: Tephritidae). Span. J. Agric. Res. 2017, 15, e1002. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Harper, G.L.; King, R.A.; Dodd, C.S.; Harwood, J.D.; Glen, D.M.; Bruford, M.W.; Symondson, W.O.C. Rapid screening of invertebrate predators for multiple prey DNA targets. Mol. Ecol. 2005, 14, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Harwood, J.D.; Desneux, N.; Yoo, H.J.; Rowley, D.I.; Greenstone, M.H.; Obrycki, J.J.; O’Neil, R.J. Tracking the role of alternative prey in soybean aphid predation by Orius insidiosus: A molecular approach. Mol. Ecol. 2007, 16, 4390–4400. [Google Scholar] [CrossRef] [PubMed]
- King, R.A.; Moreno-Ripoll, R.; Agusti, N.; Shayler, S.P.; Bell, J.R.; Bohan, D.A.; Symondson, W.O.C. Multiplex reactions for the molecular detection of predation on pest and non pest invertebrates in agroecosystems. Mol. Ecol. Resour. 2010, 11, 370–373. [Google Scholar] [CrossRef] [PubMed]
- Monzó, C.; Sabater-Muñóz, B.; Urbaneja, A.; Casta-era, P. Tracking medfly predation by the wolf spider, Pardosa cribata Simon, in citrus orchards using PCR-based gut content analysis. Bull. Entomol. Res. 2010, 100, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Ripoll, R.; Gabarra, R.; Symondson, W.O.C.; King, R.A.; Agustí, N. Trophic relationships between predators, whiteflies and their parasitoids in tomato greenhouses: A molecular approach. Bull. Entomol. Res. 2012, 102, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Hajibabaei, M.; Janzen, D.H.; Burns, J.M.; Hallwachs, W.; Hebert, P.D.N. DNA barcodes distinguish species of tropical Lepidoptera. PNAS 2005, 103, 968–971. [Google Scholar] [CrossRef] [PubMed]
- León, J.H.; Fournier, V.; Hagler, J.R.; Daane, K.M. Development of molecular diagnostic markers for sharpshooters Homalodisca coagulate and Homalodisca liturata for use in predator gut content examinations. Entomol. Exp. Appl. 2006, 119, 109–119. [Google Scholar] [CrossRef]
- Rowley, C.; Cherrill, A.J.; Leather, S.R.; McCormack, A.W.; Skarp, J.E.; Pope, T.W. PCR-based gut content analysis to identify arthropod predators of Haplodiplosis marginata. Biol. Control 2017, 115, 112–118. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).