PCR Species-Specific Primers for Molecular Gut Content Analysis to Determine the Contribution of Generalist Predators to the Biological Control of the Vector of Xylella fastidiosa
Abstract
1. Introduction
2. Materials and Methods
3. Results and Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- EFSA (European Food Safety Authority). Workshop on Xylella fastidiosa: Knowledge Gaps and Research Priorities for the EU. EFSA Supporting Publication 2016: EN-1039. 2016. Available online: https://efsa.onlinelibrary.wiley.com/doi/epdf/10.2903/sp.efsa.2016.EN-1039 (accessed on 3 June 2018).
- IOC (International Olive Council). 2016. Available online: http://www.internationaloliveoil.org/ (accessed on 3 June 2018).
- Saponari, M.; Boscia, D.; Nigro, D.; Martelli, G.P. Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (Southern Italy). J. Plant Pathol. 2013, 95, 668. [Google Scholar] [CrossRef]
- EPPO (European and Mediterranean Plant Protection Organization). Available online: http://www.eppo.int/QUARANTINE/special_topics/Xylella_fastidiosa/Xylella_fastidiosa.htm (accessed on 3 June 2018).
- MAPAMA (Ministerio de Agricultura y Pesca, Alimentación y Medio Ambiente). Available online: http://www.mapama.gob.es/es/agricultura/temas/sanidad-vegetal/xylella-fastidiosa/ (accessed on 3 June 2018).
- Boletín Oficial de la Comunidad de Madrid. No. 86. Available online: https://www.bocm.es/boletin/CM_Orden_BOCM/2018/04/11/BOCM-20180411-16.PDF (accessed on 3 June 2018).
- Boletín Oficial de la Junta de Andalucía. No. 81. Available online: http://www.juntadeandalucia.es/eboja/2018/81/BOJA18-081-00005-7337-01_00134737.pdf (accessed on 3 June 2018).
- Plant Health and Biosecurity, European Commission. Available online: https://ec.europa.eu/food/plant/plant_health_biosecurity/legislation/emergency_measures/xylella-fastidiosa_en (accessed on 3 June 2018).
- Saponari, M.; Loconsole, G.; Cornara, D.; Yokomi, R.K.; De Stradis, A.; Boscia, D.; Bosco, D.; Martelli, G.P.; Krugner, R.; Porcelli, F. Infectivity and Transmission of Xylella fastidiosa by Philaenus spumarius (Hemiptera: Aphrophoridae) in Apulia, Italy. J. Econ. Entomol. 2014, 107, 1316–1319. [Google Scholar] [CrossRef] [PubMed]
- Luvisi, A.; Nicolì, F.; De Bellis, L. Sustainable Management of Plant Quarantine Pests: The Case of Olive Quick Decline Syndrome. Sustainability 2017, 9, 659. [Google Scholar] [CrossRef]
- Bucci, E.M. Xylella fastidiosa, a new plant pathogen that threatens global farming: Ecology, molecular biology, search for remedies. Biochem. Biophys. Res. Commun. 2018, 502, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Soubeyrand, S.; Jerphanion, P.; Martin, O.; Saussac, M.; Manceau, C.; Hendrikx, P.; Lannou, C. Inferring pathogen dynamics from temporal count data: The emergence of Xylella fastidiosa in France is probably not recent. New Phytol. 2018, 219, 824–836. [Google Scholar] [CrossRef] [PubMed]
- MAPAMA (Ministerio de Agricultura y Pesca, Alimentación y Medio Ambiente). Available online: http://www.mapama.gob.es/es/agricultura/temas/sanidad-vegetal/xylellafastidiosa_contingencia_febrero2018_tcm30-445867.pdf (accessed on 23 June 2018).
- Seabra, S.G.; Pina-Martins, F.; Marabuto, E.; Yurtsever, S.; Halkka, O.; Quartau, J.A.; Paulo, O.S. Molecular phylogeny and DNA barcoding in the meadow-spittlebug Philaenus spumarius (Hemiptera, Cercopidae) and its related species. Mol. Phylogenet. Evol. 2010, 56, 462–467. [Google Scholar] [CrossRef] [PubMed]
- Cornara, D.; Saponari, M.; Zeilinger, A.R.; de Stradis, A.; Boscia, D.; Loconsole, G.; Bosco, D.; Martelli, G.P.; Almeida, R.P.P.; Porcelli, F. Spittlebugs as vectors of Xylella fastidiosa in olive orchards in Italy. J. Pest Sci. 2017, 90, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Albertini, A.; Pizoloto, R.; Petacchi, R. Carabid patterns in olive orchards and woody semi-natural habitats: First implications for conservation biological control against Bactrocera oleae. BioControl 2017, 62, 71. [Google Scholar] [CrossRef]
- Rosenheim, J.A.; Limburg, D.D.; Colfer, R.G. Impact of generalist predators on a biological control agent, Chrysoperla carnea: Direct observations. Ecol. Appl. 1999, 9, 409–417. [Google Scholar] [CrossRef]
- Symondson, W.O.C. Molecular identification of prey in predator diets. Mol. Ecol. 2002, 11, 627–641. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.M.J.; Ingels, B.; Wheatley, A.; Rhule, E.L.; de Clercq, P.; van Leeuwen, P.; Thomas, A. Intraguild predation by Harmonia axyridis (Coleoptera: Coccinellidae) on native insects in Europe: Molecular detection from field samples. Entomol. Sci. 2015, 18, 130–133. [Google Scholar] [CrossRef]
- MacDonald, A.J.; Young, M.J.; Lintermans, M.; Sarre, S.D. Primers for detection of Macquarie perch from environmental and trace DNA samples. Conserv. Genet. Resour. 2014, 6, 551–553. [Google Scholar] [CrossRef]
- Pentinsaari, M.; Salmela, H.; Mutanen, M.; Roslin, T. Molecular evolution of a widely-adopted taxonomic marker (COI) across the animal tree of life. Sci. Rep. 2016, 6, 35275. [Google Scholar] [CrossRef] [PubMed]
- BOLD Database. Available online: http://v3.boldsystems.org/index.php/databases (accessed on 3 June 2018).
- GenBank Database. Available online: https://www.ncbi.nlm.nih.gov/genbank/ (accessed on 3 June 2018).
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/ NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- EMBL-EBI (European Bioinformatics Institute) Tools: Multiple Sequence Alignment. Available online: https://www.ebi.ac.uk/Tools/msa/ (accessed on 3 June 2018).
- Lantero, E.; Matallanas, B.; Ochando, M.D.; Pascual, S.; Callejas, C. Specific and sensitive primers for the detection of predated olive fruit flies, Bactrocera oleae (Diptera: Tephritidae). Span. J. Agric. Res. 2017, 15, e1002. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Harper, G.L.; King, R.A.; Dodd, C.S.; Harwood, J.D.; Glen, D.M.; Bruford, M.W.; Symondson, W.O.C. Rapid screening of invertebrate predators for multiple prey DNA targets. Mol. Ecol. 2005, 14, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Harwood, J.D.; Desneux, N.; Yoo, H.J.; Rowley, D.I.; Greenstone, M.H.; Obrycki, J.J.; O’Neil, R.J. Tracking the role of alternative prey in soybean aphid predation by Orius insidiosus: A molecular approach. Mol. Ecol. 2007, 16, 4390–4400. [Google Scholar] [CrossRef] [PubMed]
- King, R.A.; Moreno-Ripoll, R.; Agusti, N.; Shayler, S.P.; Bell, J.R.; Bohan, D.A.; Symondson, W.O.C. Multiplex reactions for the molecular detection of predation on pest and non pest invertebrates in agroecosystems. Mol. Ecol. Resour. 2010, 11, 370–373. [Google Scholar] [CrossRef] [PubMed]
- Monzó, C.; Sabater-Muñóz, B.; Urbaneja, A.; Casta-era, P. Tracking medfly predation by the wolf spider, Pardosa cribata Simon, in citrus orchards using PCR-based gut content analysis. Bull. Entomol. Res. 2010, 100, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Ripoll, R.; Gabarra, R.; Symondson, W.O.C.; King, R.A.; Agustí, N. Trophic relationships between predators, whiteflies and their parasitoids in tomato greenhouses: A molecular approach. Bull. Entomol. Res. 2012, 102, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Hajibabaei, M.; Janzen, D.H.; Burns, J.M.; Hallwachs, W.; Hebert, P.D.N. DNA barcodes distinguish species of tropical Lepidoptera. PNAS 2005, 103, 968–971. [Google Scholar] [CrossRef] [PubMed]
- León, J.H.; Fournier, V.; Hagler, J.R.; Daane, K.M. Development of molecular diagnostic markers for sharpshooters Homalodisca coagulate and Homalodisca liturata for use in predator gut content examinations. Entomol. Exp. Appl. 2006, 119, 109–119. [Google Scholar] [CrossRef]
- Rowley, C.; Cherrill, A.J.; Leather, S.R.; McCormack, A.W.; Skarp, J.E.; Pope, T.W. PCR-based gut content analysis to identify arthropod predators of Haplodiplosis marginata. Biol. Control 2017, 115, 112–118. [Google Scholar] [CrossRef]
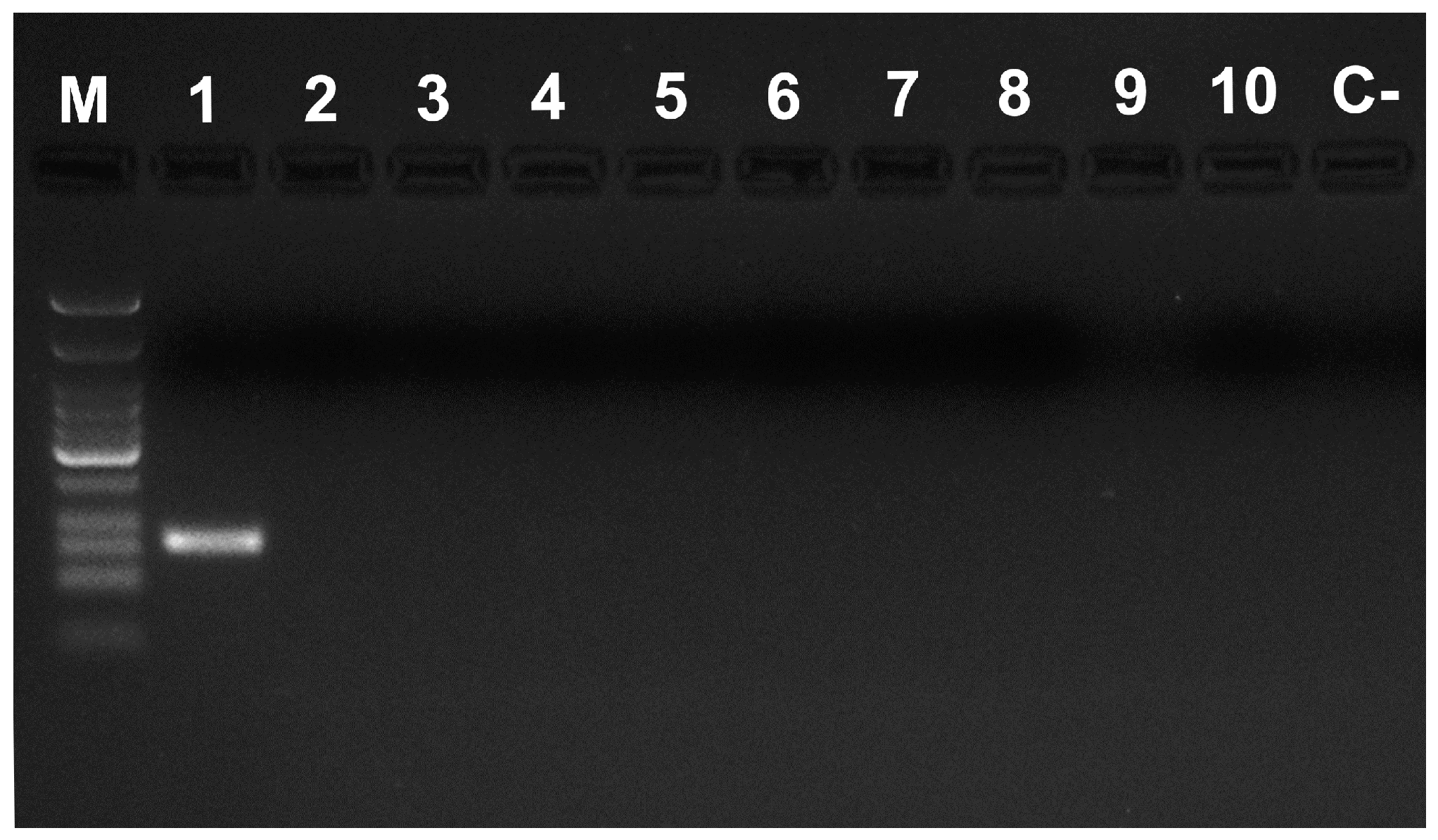
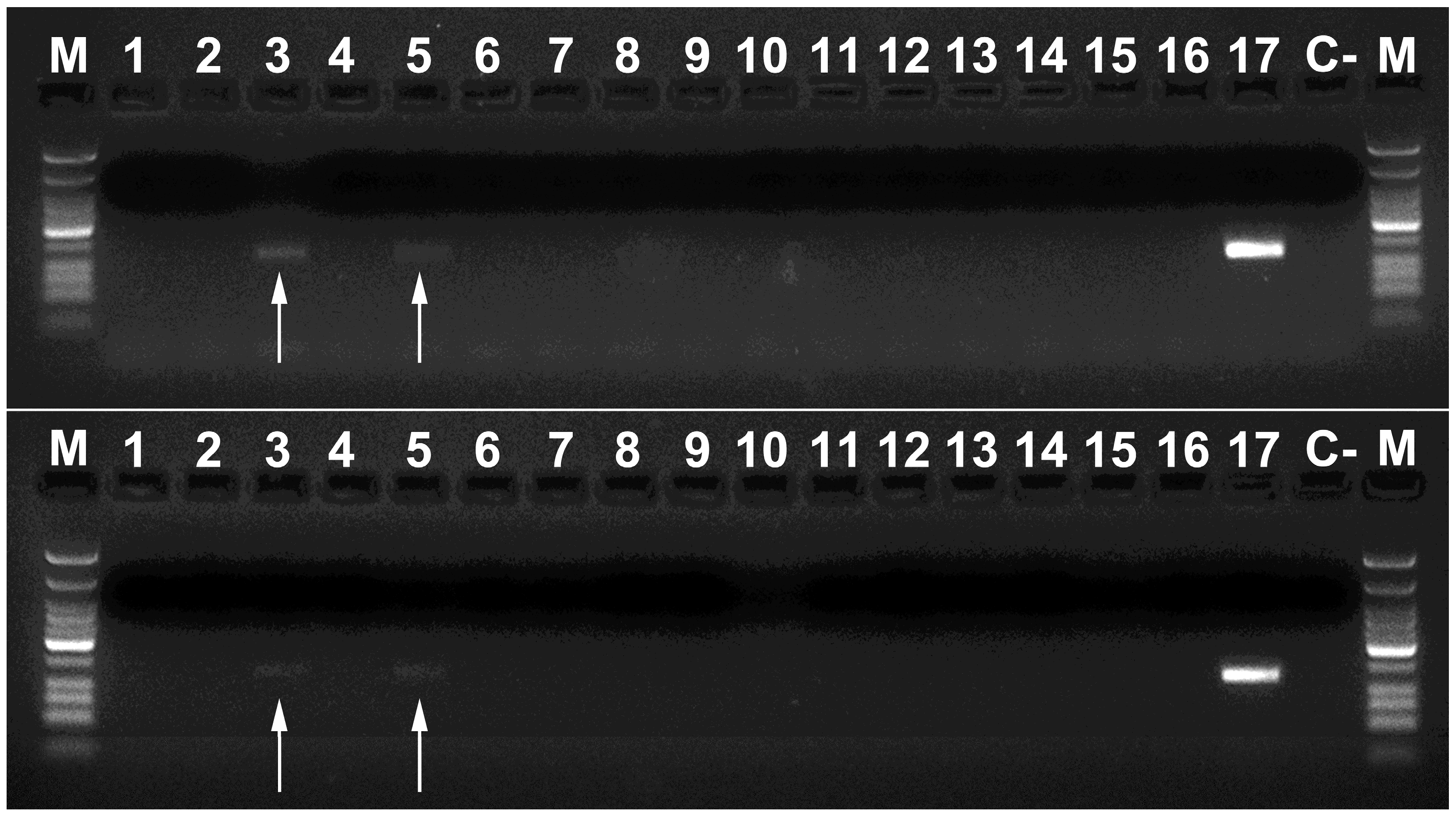
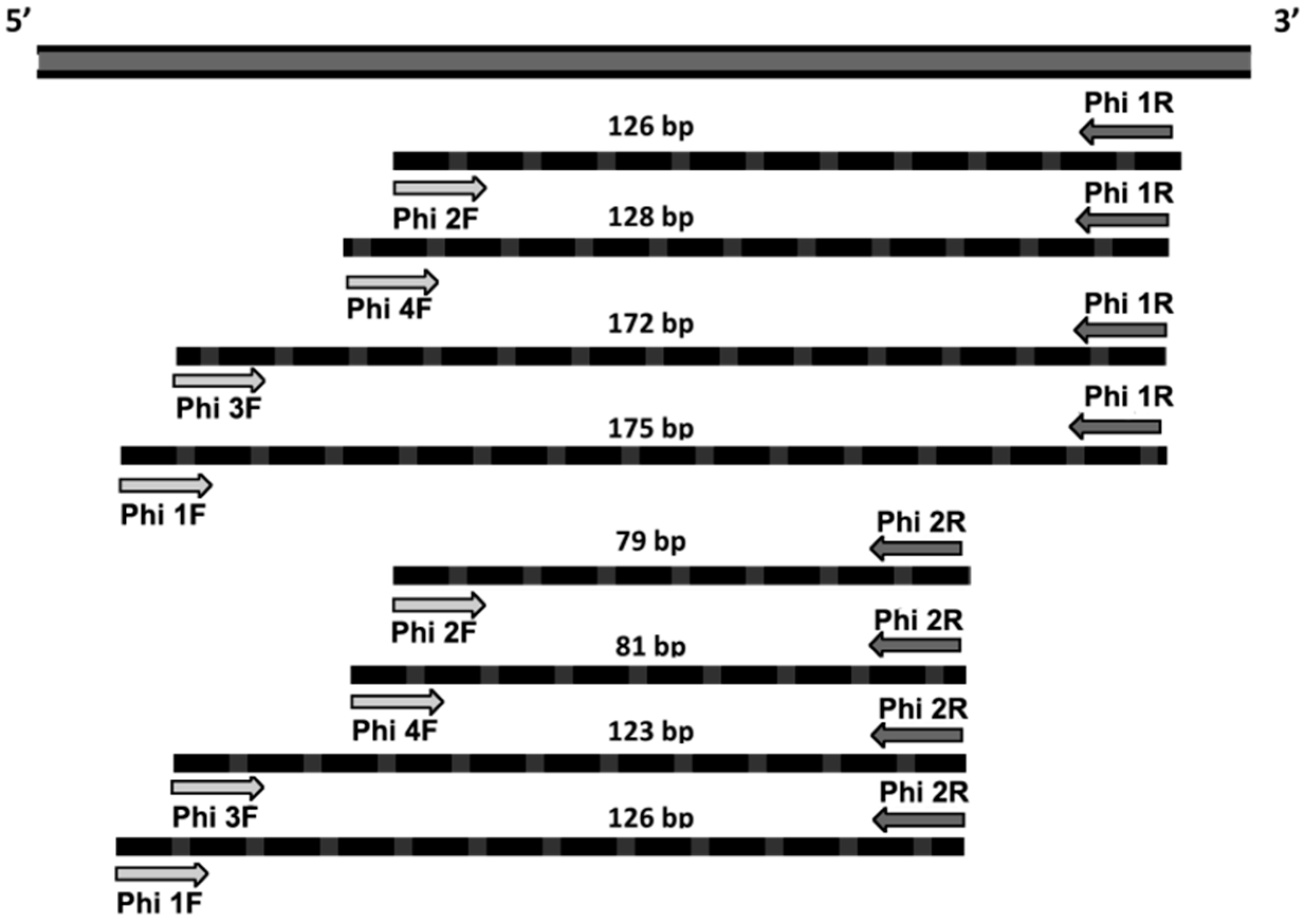


| Primer Name | Primer Sequence 5′–3′ |
|---|---|
| Phi 1F | GCTCCTGACATAGCATTCCCA |
| Phi 2F | GCTTCCTCCTTCATTAACGCTT |
| Phi 3F | CCTGACATAGCATTCCCACGA |
| Phi 4F | TGCTTCCTCCTTCATTAACGCTT |
| Phi 1R | TAGCTAAATCAACACATGCACCAG |
| Phi 2R | GGAGGATAAACTGTTCATCCC |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lantero, E.; Matallanas, B.; Pascual, S.; Callejas, C. PCR Species-Specific Primers for Molecular Gut Content Analysis to Determine the Contribution of Generalist Predators to the Biological Control of the Vector of Xylella fastidiosa. Sustainability 2018, 10, 2207. https://doi.org/10.3390/su10072207
Lantero E, Matallanas B, Pascual S, Callejas C. PCR Species-Specific Primers for Molecular Gut Content Analysis to Determine the Contribution of Generalist Predators to the Biological Control of the Vector of Xylella fastidiosa. Sustainability. 2018; 10(7):2207. https://doi.org/10.3390/su10072207
Chicago/Turabian StyleLantero, Esther, Beatriz Matallanas, Susana Pascual, and Carmen Callejas. 2018. "PCR Species-Specific Primers for Molecular Gut Content Analysis to Determine the Contribution of Generalist Predators to the Biological Control of the Vector of Xylella fastidiosa" Sustainability 10, no. 7: 2207. https://doi.org/10.3390/su10072207
APA StyleLantero, E., Matallanas, B., Pascual, S., & Callejas, C. (2018). PCR Species-Specific Primers for Molecular Gut Content Analysis to Determine the Contribution of Generalist Predators to the Biological Control of the Vector of Xylella fastidiosa. Sustainability, 10(7), 2207. https://doi.org/10.3390/su10072207






