Candida Variety in the Oral Cavity of Mexican Subjects with Type 2 Diabetes Mellitus and TLR2 Gene Expression
Abstract
1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. Gene Expression Analysis and Genotyping
2.3. Statistical Analysis
3. Results
3.1. Characteristics of the Participants
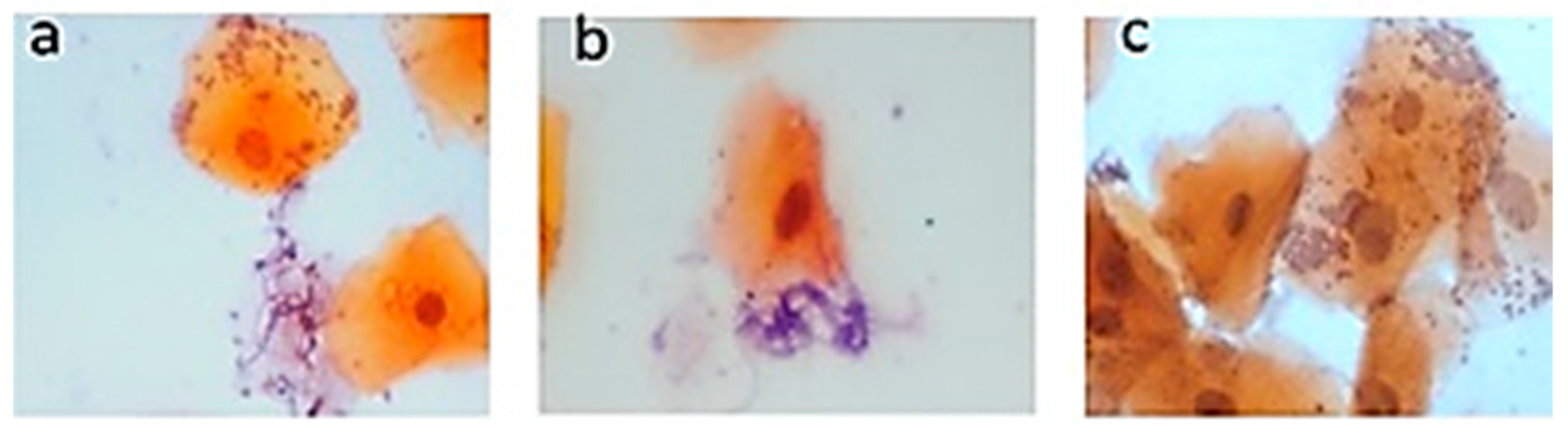
3.2. Pap Smear Results
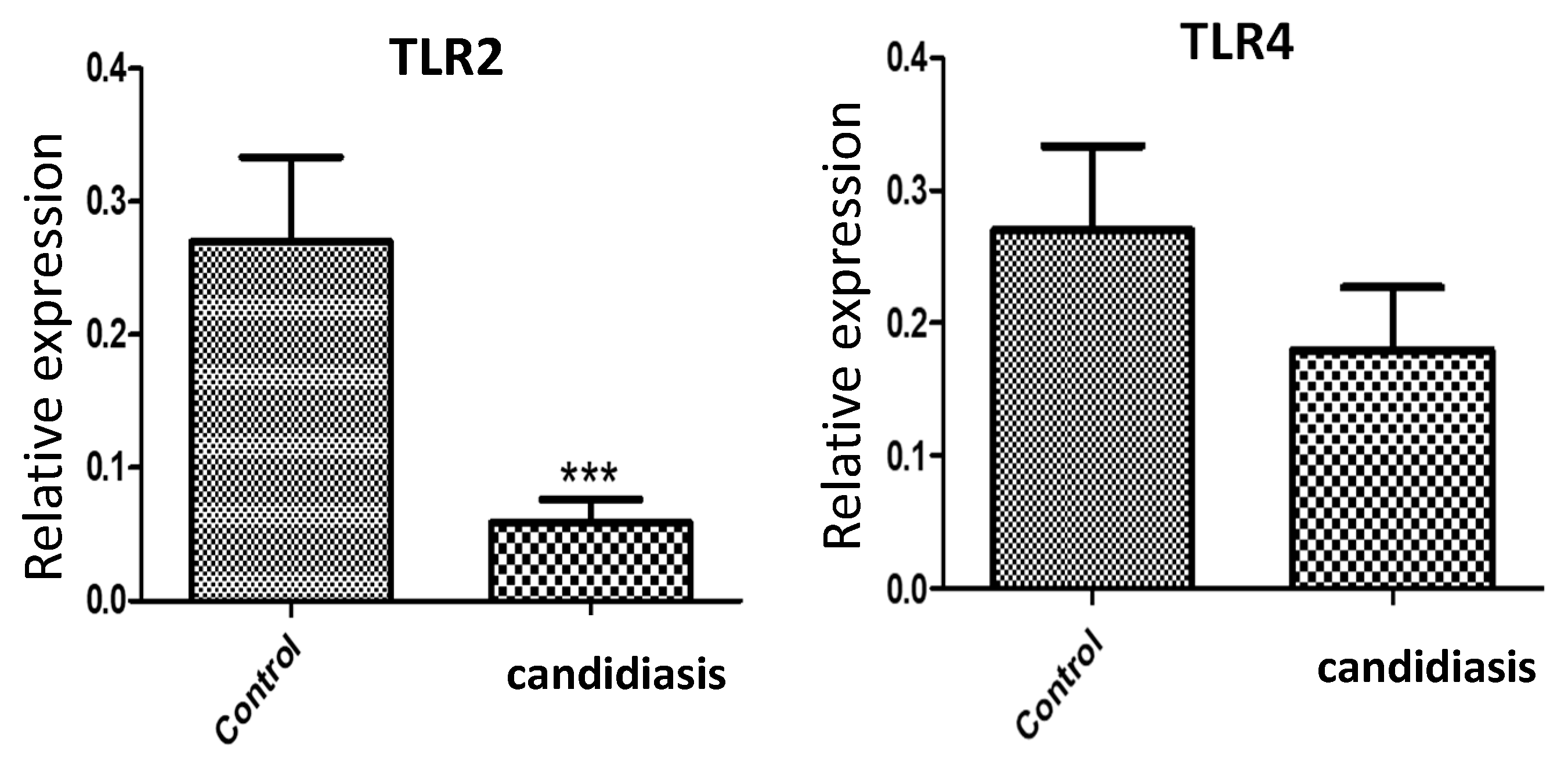
3.3. Gene Expression of TRL2 and TLR4
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arendorf, T.M.; Walker, D.M. The prevalence and intra-oral distribution of Candida albicans in man. Arch. Oral Biol. 1980, 25, 1–10. [Google Scholar] [CrossRef]
- Martins, N.; Ferreira, I.C.F.R.; Barros, L.; Silva, S.; Henriques, M. Candidiasis: Predisposing factors, prevention, diagnosis and alternative treatment. Mycopathologia 2014, 177, 223–240. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Kauffman, C.A.; Andes, D.R.; Clancy, C.J.; Marr, K.A.; Ostrosky-Zeichner, L.; Reboli, A.C.; Schuster, M.G.; Vazquez, J.A.; Walsh, T.J.; et al. Clinical practice guideline for the management of candidiasis: 2016 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 2016, 62, e1–e50. [Google Scholar] [CrossRef] [PubMed]
- Madhavan, P.; Jamal, F.; Chong, P.P. Laboratory isolation and identification of Candida species. J. Appl. Sci. 2011, 11, 2870–2877. [Google Scholar] [CrossRef][Green Version]
- de Resende, M.A.; de Sousa, L.V.N.F.; de Oliveira, R.C.B.W.; Koga-Ito, C.Y.; Lyon, J.P. Prevalence and antifungal susceptibility of yeasts obtained from the oral cavity of elderly individuals. Mycopathologia 2006, 162, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Guimarães, T.; Nucci, M.; Mendonça, J.S.; Martinez, R.; Brito, L.R.; Silva, N.; Moretti, M.L.; Salomão, R.; Colombo, A.L. Epidemiology and predictors of a poor outcome in elderly patients with candidemia. Int. J. Infect. Dis. 2012, 16, e442–e447. [Google Scholar] [CrossRef]
- Khosravi, A.R.; Yarahmadi, S.; Baiat, M.; Shokri, H.; Pourkabireh, M. Factors affecting the prevalence of yeasts in the oral cavity of patients with diabetes mellitus. J. Mycol. Médicale 2008, 18, 83–88. [Google Scholar] [CrossRef]
- Tang, H.; Liu, W.; Lin, H.; Lai, C. Epidemiology and prognostic factors of candidemia in elderly patients. Geriatr. Gerontol. Int. 2015, 15, 688–693. [Google Scholar] [CrossRef]
- Calvet, H.M.; Yoshikawa, T.T. Infections in diabetes. Infect. Dis. Clin. N. Am. 2001, 15, 407–421. [Google Scholar] [CrossRef]
- Bhuyan, L.; Hassan, S.; Dash, K.C.; Panda, A.; Behura, S.S.; Ramachandra, S. Candida species diversity in oral cavity of type 2 diabetic patients and their in vitro antifungal susceptibility. Contemp. Clin. Dent. 2018, 9 (Suppl. S1), S83–S88. [Google Scholar] [CrossRef]
- King, H.; Aubert, R.E.; Herman, W.H. Global burden of diabetes, 1995–2025: Prevalence, numerical estimates, and projections. Diabetes Care 1998, 21, 1414–1431. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, S.; Raman, R.; Paul, P.G.; Rani, P.K.; Uthra, S.; Gayathree, R.; McCarty, C.; Kumaramanickavel, G.; Sharma, T. Sankara Nethralaya—Diabetic retinopathy epidemiology and molecular genetic study (SN—DREAMS 1): Study design and research methodology. Ophthalmic Epidemiol. 2005, 12, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Davenport, J.C. The oral distribution of Candida in denture stomatitis. Br. Dent. J. 1970, 129, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Kadir, T.; Pisiriciler, R.; Akyüz, S.; Yarat, A.; Emekli, N.; Ipbüker, A. Mycological and cytological examination of oral candidal carriage in diabetic patients and non-diabetic control subjects: Thorough analysis of local aetiologic and systemic factors. J. Oral Rehabil. 2002, 29, 452–457. [Google Scholar] [CrossRef]
- Duggan, S.; Essig, F.; Hünniger, K.; Mokhtari, Z.; Bauer, L.; Lehnert, T.; Brandes, S.; Häder, A.; Jacobsen, I.D.; Martin, R.; et al. Neutrophil activation by Candida glabrata but not Candida albicans promotes fungal uptake by monocytes. Cell Microbiol. 2015, 17, 1259–1276. [Google Scholar] [CrossRef]
- Wilson, R.M.; Reeves, W.G. Neutrophil phagocytosis and killing in insulin-dependent diabetes. Clin. Exp. Immunol. 1986, 63, 478–484. [Google Scholar]
- Pinto, E.; Ribeiro, I.C.; Ferreira, N.J.; Fortes, C.E.; Fonseca, P.A.; Figueiral, M.H. Correlation between enzyme production, germ tube formation and susceptibility to fluconazole in Candida species isolated from patients with denture-related stomatitis and control individuals. J. Oral. Pathol. Med. 2008, 37, 587–592. [Google Scholar] [CrossRef]
- Motta-Silva, A.C.; Aleva, N.A.; Chavasco, J.K.; Armond, M.C.; França, J.P.; Pereira, L.J. Erythematous oral candidiasis in patients with controlled type II diabetes mellitus and complete dentures. Mycopathologia 2010, 169, 215–223. [Google Scholar] [CrossRef]
- Calderone, R.A.; Fonzi, W.A. Virulence factors of Candida albicans. Trends Microbiol. 2001, 9, 327–335. [Google Scholar] [CrossRef]
- Balan, P.; Gogineni, S.B.; Kumari, S.; Shetty, V.; Rangare, A.L.; Castelino, R.L.; Areekat K, F. Candida carriage rate and growth characteristics of saliva in diabetes mellitus patients: A case-control study. J. Dent. Res. Dent. Clin. Dent. Prospect. 2015, 9, 274. [Google Scholar] [CrossRef]
- Nowakowska, D.; Kurnatowska, A.; Stray-Pedersen, B.; Wilczyński, J. Species distribution and influence of glycemic control on fungal infections in pregnant women with diabetes. J. Infect. 2004, 48, 339–346. [Google Scholar] [CrossRef]
- Dorko, E.; Baranova, Z.; Jenča, A.; Kizek, P.; Pilipčinec, E.; Tkáčiková, L. Diabetes mellitus and candidiases. Folia Microbiol. 2005, 50, 255–261. [Google Scholar] [CrossRef]
- Feller, L.; Khammissa, R.A.G.; Chandran, R.; Altini, M.; Lemmer, J. Oral candidosis in relation to oral immunity. J. Oral Pathol. Med. 2014, 43, 563–569. [Google Scholar] [CrossRef]
- Rós Ásmundsdóttir, L.; Erlendsdóttir, H.; Haraldsson, G.; Guo, H.; Xu, J.; Gottfredsson, M. Molecular epidemiology of candidemia: Evidence of clusters of smoldering nosocomial infections. Clin. Infect. Dis. 2008, 47, e17–e24. [Google Scholar] [CrossRef]
- Moosazadeh, M.; Akbari, M.; Tabrizi, R.; Ghorbani, A.; Golkari, A.; Banakar, M.; Sekhavati, E.; Kavari, S.H.; Lankarani, K.B. Denture stomatitis and Candida albicans in Iranian population: A systematic review and meta-analysis. J. Dent. 2016, 17 (Suppl. S3), 283–292. [Google Scholar]
- Netea, M.G.; Joosten, L.A.B.; Van Der Meer, J.W.M.; Kullberg, B.J.; Van De Veerdonk, F.L. Immune defence against Candida fungal infections. Nat. Rev. Immunol. 2015, 15, 630–642. [Google Scholar] [CrossRef] [PubMed]
- Netea, M.G.; Brown, G.D.; Kullberg, B.J.; Gow, N.A.R. An integrated model of the recognition of Candida albicans by the innate immune system. Nat. Rev. Microbiol. 2008, 6, 67–78. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Gordon, S. A new receptor for β-glucans. Nature 2001, 413, 36–37. [Google Scholar] [CrossRef] [PubMed]
- Cambi, A.; Gijzen, K.; de Vries, I.J.M.; Torensma, R.; Joosten, B.; Adema, G.J.; Netea, G.; Kullberg, B.-J.; Romani, L.; Figdor, G. The C-type lectin DC-SIGN (CD209) is an antigen-uptake receptor for Candida albicans on dendritic cells. Eur. J. Immunol. 2003, 33, 532–538. [Google Scholar] [CrossRef] [PubMed]
- Netea, M.G.; Gow, N.A.R.; Munro, C.A.; Bates, S.; Collins, C.; Ferwerda, G.; Hobson, R.P.; Bertram, G.; Hughes, H.B.; Jansen, T.; et al. Immune sensing of Candida albicans requires cooperative recognition of mannans and glucans by lectin and Toll-like receptors. J. Clin. Investig. 2006, 116, 1642–1650. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, M.; Carvalho, A.; Cunha, C.; Plantinga, T.S.; Van De Veerdonk, F.; Puccetti, M.; Galosi, C.; Joosten, L.A.B.; Dupont, B.; Kullberg, B.J.; et al. Association of a variable number tandem repeat in the NLRP3 gene in women with susceptibility to RVVC. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 797–801. [Google Scholar] [CrossRef]
- Jaeger, M.; van der Lee, R.; Cheng, S.-C.; Johnson, M.D.; Kumar, V.; Ng, A.; Plantinga, T.S.; Smeekens, S.P.; Oosting, M.; Wang, X.; et al. The RIG-I-like helicase receptor MDA5 (IFIH1) is involved in the host defense against Candida infections. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 963–974. [Google Scholar] [CrossRef]
- Groslambert, M.; Py, B.F. Spotlight on the NLRP3 inflammasome pathway. J. Inflamm. Res. 2018, 2018, 359–374. [Google Scholar] [CrossRef]
- Picciani, B.L.S.; Michalski-Santos, B.; Carneiro, S.; Sampaio, A.L.; Avelleira, J.C.R.; Azulay, D.R.; Pinto, J.M.N.; Dias, E.P. Oral candidiasis in patients with psoriasis: Correlation of oral examination and cytopathological evaluation with psoriasis disease severity and treatment. J. Am. Acad. Dermatol. 2013, 68, 986–991. [Google Scholar] [CrossRef]
- Miller, D.J. Diagnosis and management of Candida and other fungal infections of the head and neck. Curr. Infect. Dis. Rep. 2002, 4, 194–200. [Google Scholar] [CrossRef]
- Hellstein, J.W.; Marek, C.L. Candidiasis: Red and white manifestations in the oral cavity. Head Neck Pathol. 2019, 13, 25–32. [Google Scholar] [CrossRef]
- Cuenca-Estrella, M.; Verweij, P.E.; Arendrup, M.C.; Arikan-Akdagli, S.; Bille, J.; Donnelly, J.P.; Jensen, H.E.; Lass-Flörl, C.; Richardson, M.D.; Akova, M.; et al. ESCMID* guideline for the diagnosis and management of Candida diseases 2012: Diagnostic procedures. Clin. Microbiol. Infect. 2012, 18, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Kauffman, C.A.; Andes, D.; Benjamin, D.K., Jr.; Calandra, T.F.; Edwards, J.E., Jr.; Filler, S.G.; Fisher, J.F.; Kullberg, B.J.; Ostrosky-Zeichner, L.; et al. Clinical practice guidelines for the management of candidiasis: 2009 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 2009, 48, 503–535. [Google Scholar] [CrossRef] [PubMed]
- González, G.M.; Elizondo, M.; Ayala, J. Trends in species distribution and susceptibility of bloodstream isolates of Candida collected in Monterrey, Mexico, to seven antifungal agents: Results of a 3-year (2004 to 2007) surveillance study. J. Clin. Microbiol. 2008, 46, 2902–2905. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, V.; Singla, R.K.; Dubey, A.K. Emerging virulence, drug resistance and future anti-fungal drugs for Candida pathogens. Curr. Top. Med. Chem. 2018, 18, 759–778. [Google Scholar] [CrossRef] [PubMed]
- Whaley, S.G.; Berkow, E.L.; Rybak, J.M.; Nishimoto, A.T.; Barker, K.S.; Rogers, P.D. Azole antifungal resistance in Candida albicans and emerging non-albicans Candida species. Front. Microbiol. 2017, 7, 2173. [Google Scholar] [CrossRef]
- Cheng, S.C.; Joosten, L.A.B.; Kullberg, B.J.; Netea, M.G. Interplay between Candida albicans and the mammalian innate host defense. Infect. Immun. 2012, 80, 1304–1313. [Google Scholar] [CrossRef]
- Richardson, J.P.; Moyes, D.L. Adaptive immune responses to Candida albicans infection. Virulence 2015, 6, 327–337. [Google Scholar] [CrossRef]
- Wuyts, J.; Van Dijck, P.; Holtappels, M. Fungal persister cells: The basis for recalcitrant infections? PLoS Pathog. 2018, 14, e1007301. [Google Scholar] [CrossRef]
- Lockhart, S.R.; Ghannoum, M.A.; Alexander, B.D. Establishment and use of epidemiological cutoff values for molds and yeasts by use of the clinical and laboratory standards institute M57 standard. J. Clin. Microbiol. 2017, 55, 1262–1268. [Google Scholar] [CrossRef]
- Pristov, K.E.; Ghannoum, M.A. Resistance of Candida to azoles and echinocandins worldwide. Clin. Microbiol. Infect. 2019, 25, 792–798. [Google Scholar] [CrossRef] [PubMed]
- Berman, J.; Krysan, D.J. Drug resistance and tolerance in fungi. Nat. Rev. Microbiol. 2020, 18, 319–331. [Google Scholar] [CrossRef]
- Liu, J.; Geng, F.; Sun, H.; Wang, X.; Zhang, H.; Yang, Q.; Zhang, J. Candida albicans induces TLR2/MyD88/NF-κB signaling and inflammation in oral lichen planus-derived keratinocytes. J. Infect. Dev. Ctries. 2018, 12, 780–786. [Google Scholar] [CrossRef] [PubMed]
- Shaoru, Z.; Jiawen, L.; Xuesong, J.; Yanqing, W. The expression of toll-like receptor 2 and 4 mRNA in local tissues of model of oropharyngeal candidiasis in mice. J. Huazhong Univ. Sci. Technol. [Med. Sci.] 2004, 24, 639–641. [Google Scholar] [CrossRef] [PubMed]
- Ye, P.; Wang, X.; Ge, S.; Chen, W.; Wang, W.; Han, X. Long-term cigarette smoking suppresses NLRP3 inflammasome activation in oral mucosal epithelium and attenuates host defense against Candida albicans in a rat model. Biomed. Pharmacother. 2019, 113, 108597. [Google Scholar] [CrossRef] [PubMed]
| Variables | Number of Patients |
|---|---|
| Gender | |
| Female | 12 (66.6%) |
| Male | 6 (33.3%) |
| Age (years), mean (SD) | 63.3 ± 14.2 |
| Smoking | 2 (11.1%) |
| Non-smoking | 16 (88.8%) |
| Hypertension | 11 (61.1%) |
| No hypertension | 7 (38.8%) |
| Type 1 stomatitis | 11 (61.1%) |
| Type 2 stomatitis | 7 (38.8%) |
| Type of candidiasis (observation of clinical pattern) | |
| Pseudomembranous | 15 (83.3%) |
| Erythematous | 3 (16.6%) |
| Glycated hemoglobin (HbA1c) mean (SD) | 9.3 ± 2.7 |
| Candida | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Patient | PAP | Albicans | Glabrata | Parapsilosis | Tropicallis | Dublinensis | Krusei | Guilermondi | Lusitanie |
| 01 | − | − | − | − | − | − | − | + | − |
| 02 | − | − | − | − | − | − | − | + | − |
| 03 | − | + | + | − | + | − | − | + | − |
| 04 | + | − | − | − | − | − | − | − | − |
| 05 | − | − | − | − | + | − | + | + | + |
| 06 | − | − | − | − | + | − | − | + | − |
| 07 | − | − | − | − | + | − | + | + | + |
| 08 | − | − | − | − | − | − | − | − | − |
| 09 | − | − | − | − | − | − | − | − | − |
| 10 | − | − | − | − | − | − | − | − | − |
| 11 | + | − | − | − | − | − | + | + | − |
| 12 | − | − | − | − | − | − | + | − | − |
| 13 | + | + | − | + | − | − | + | − | − |
| 14 | − | − | − | − | − | − | + | − | − |
| 15 | − | − | − | − | − | − | − | − | + |
| 16 | − | − | − | − | − | − | − | − | + |
| 17 | − | − | − | − | − | − | − | − | − |
| 18 | − | − | − | − | − | − | − | − | + |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pérez-Vielma, N.M.; Gómez-López, M.; Martínez-Godínez, M.d.l.Á.; Luna-Torres, A.L.; Domínguez López, A.; Miliar-García, Á. Candida Variety in the Oral Cavity of Mexican Subjects with Type 2 Diabetes Mellitus and TLR2 Gene Expression. Clin. Pract. 2024, 14, 417-425. https://doi.org/10.3390/clinpract14020031
Pérez-Vielma NM, Gómez-López M, Martínez-Godínez MdlÁ, Luna-Torres AL, Domínguez López A, Miliar-García Á. Candida Variety in the Oral Cavity of Mexican Subjects with Type 2 Diabetes Mellitus and TLR2 Gene Expression. Clinics and Practice. 2024; 14(2):417-425. https://doi.org/10.3390/clinpract14020031
Chicago/Turabian StylePérez-Vielma, Nadia Mabel, Modesto Gómez-López, María de los Ángeles Martínez-Godínez, Ana Laura Luna-Torres, Aarón Domínguez López, and Ángel Miliar-García. 2024. "Candida Variety in the Oral Cavity of Mexican Subjects with Type 2 Diabetes Mellitus and TLR2 Gene Expression" Clinics and Practice 14, no. 2: 417-425. https://doi.org/10.3390/clinpract14020031
APA StylePérez-Vielma, N. M., Gómez-López, M., Martínez-Godínez, M. d. l. Á., Luna-Torres, A. L., Domínguez López, A., & Miliar-García, Á. (2024). Candida Variety in the Oral Cavity of Mexican Subjects with Type 2 Diabetes Mellitus and TLR2 Gene Expression. Clinics and Practice, 14(2), 417-425. https://doi.org/10.3390/clinpract14020031






