Preventing Respiratory Viral Diseases with Antimicrobial Peptide Master Regulators in the Lung Airway Habitat
Abstract
1. Introduction
2. Major Antimicrobial Peptides Expressed in Lung Airways
2.1. Defensins
2.2. Cathelicidin
2.3. Lactoferrin
2.4. Secretory Leucoprotease Inhibitor (SLPI)
2.5. Lysozyme
2.6. Lactoperoxidase
2.7. CCL20
3. Regulation of AMPs Expression in the Respiratory Tract and Innate Immunity
3.1. Toll-like and Cytokine Receptors Mediate Regulation
3.2. Vitamin-D-Dependent Regulation
3.3. Signaling Pathways Involved in the Regulation of Defensins and Cathelicidins
4. Nutrients Involved in the Regulation of Defensins and Cathelicidins
4.1. Amino Acids
4.2. Fatty Acids and Analogs
4.3. Carbohydrates and Conjugates
4.4. Plant Extracts
4.5. Other Factors and Mechanisms
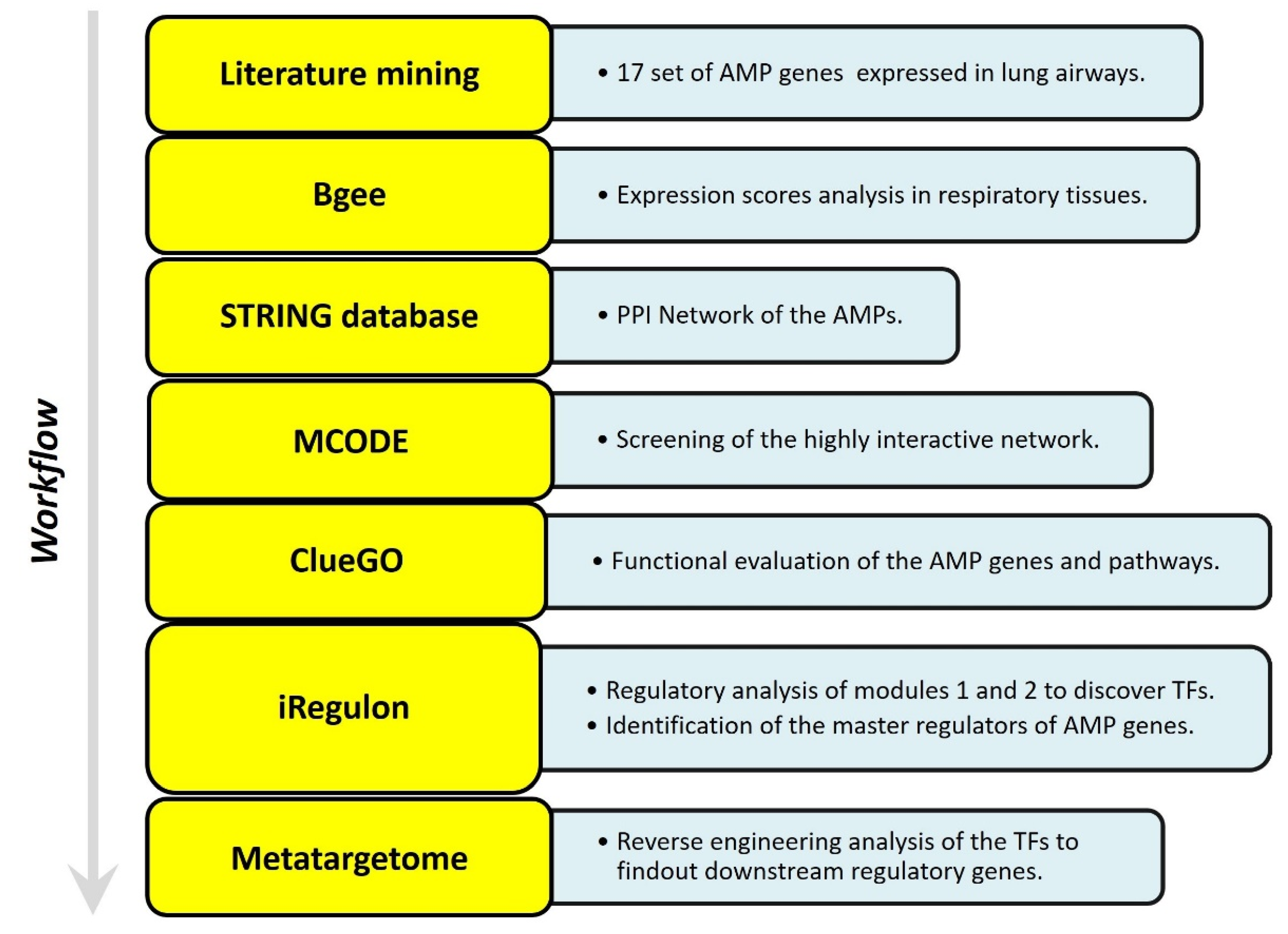
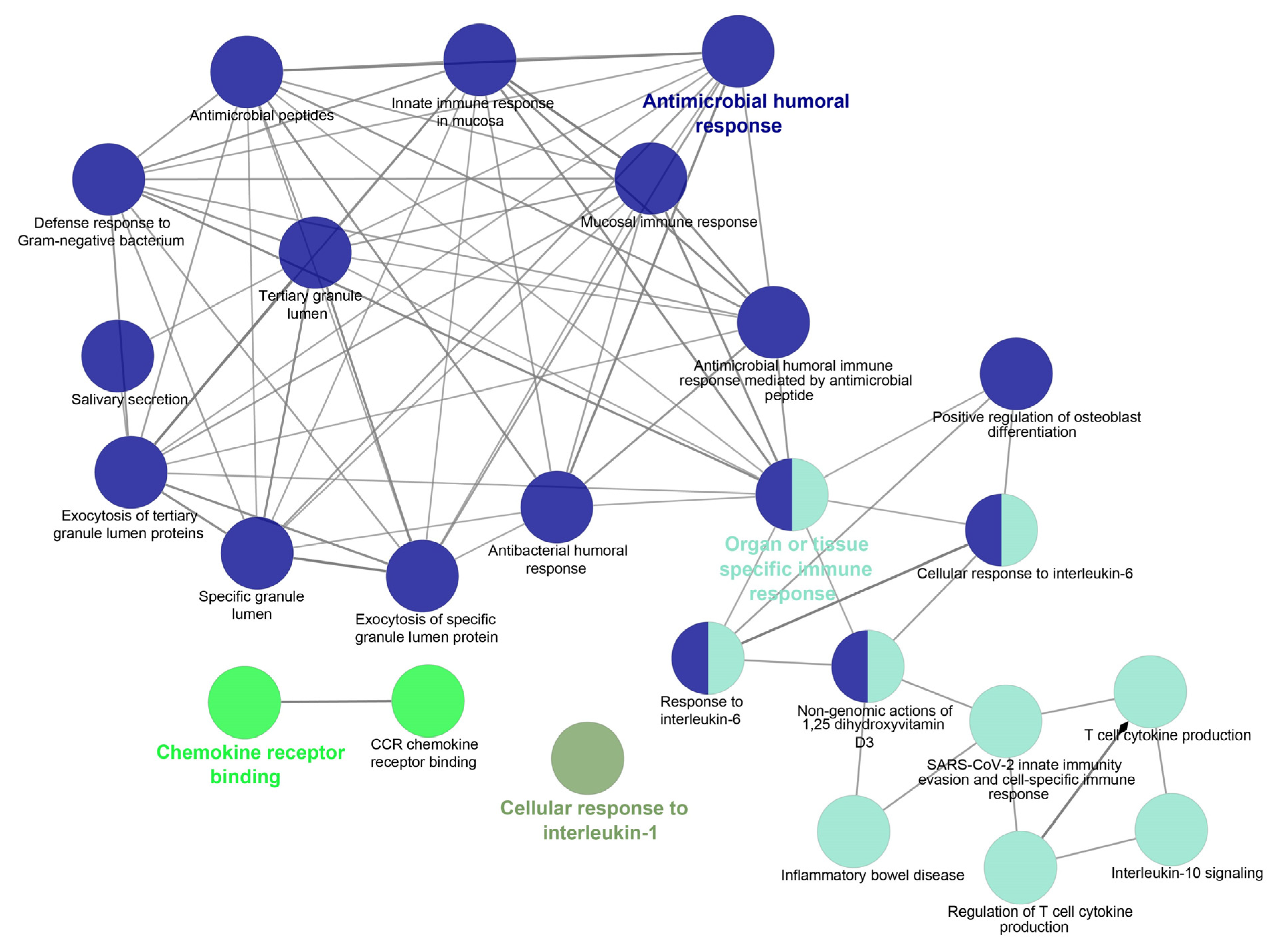
5. Studying the Expression of HDPs and Protein–Protein Interaction (PPI) Networks Construction
5.1. Identifying the Transcription Factors and Validation
5.2. In Silico Analysis and Expression Profiling of Lung Airways Antimicrobial Peptides
6. Conclusions
7. Limitations of the Study
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nicolas, P. Multifunctional host defense peptides: Intracellular-targeting antimicrobial peptides. FEBS J. 2009, 276, 6483–6496. [Google Scholar] [CrossRef] [PubMed]
- Nijnik, A.; Hancock, R. Host defence peptides: Antimicrobial and immunomodulatory activity and potential applications for tackling antibiotic-resistant infections. Emerg. Heal. Threat. J. 2009, 2, 7078. [Google Scholar] [CrossRef]
- Drayton, M.; Deisinger, J.P.; Ludwig, K.C.; Raheem, N.; Müller, A.; Schneider, T.; Straus, S.K. Host Defense Peptides: Dual Antimicrobial and Immunomodulatory Action. Int. J. Mol. Sci. 2021, 22, 11172. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.J.; Gallo, R.L. Antimicrobial peptides. Curr. Biol. 2016, 6, 1543–1575. [Google Scholar] [CrossRef]
- Robinson, K.; Deng, Z.; Hou, Y.; Zhang, G. Regulation of the Intestinal Barrier Function by Host Defense Peptides. Front. Veter- Sci. 2015, 2, 57. [Google Scholar] [CrossRef]
- Baindara, P.; Chakraborty, R.; Holliday, Z.; Mandal, S.; Schrum, A. Oral probiotics in coronavirus disease 2019: Connecting the gut–lung axis to viral pathogenesis, inflammation, secondary infection and clinical trials. New Microbes New Infect. 2021, 40, 100837. [Google Scholar] [CrossRef]
- Manna, S.; Chowdhury, T.; Chakraborty, R.; Mandal, S.M. Probiotics-Derived Peptides and Their Immunomodulatory Molecules Can Play a Preventive Role Against Viral Diseases Including COVID-19. Probiotics Antimicrob. Proteins 2020, 13, 611–623. [Google Scholar] [CrossRef]
- Haney, E.F.; Mansour, S.C.; Hancock, R.E.W. Antimicrobial peptides: An introduction. In Methods in Molecular Biology; Humana Press: New York, NY, USA, 2017; Volume 1548, pp. 3–22. ISBN 978-1-60761-593-4. [Google Scholar]
- Hilchie, A.L.; Wuerth, K.; Hancock, R.E.W. Immune modulation by multifaceted cationic host defense (antimicrobial) peptides. Nat. Chem. Biol. 2013, 9, 761–768. [Google Scholar] [CrossRef]
- Manna, S.; Baindara, P.; Mandal, S.M. Molecular pathogenesis of secondary bacterial infection associated to viral infections including SARS-CoV-2. J. Infect. Public Health 2020, 13, 1397–1404. [Google Scholar] [CrossRef]
- Rogan, M.P.; Geraghty, P.; Greene, C.M.; O’Neill, S.J.; Taggart, C.C.; McElvaney, N.G. Antimicrobial proteins and polypeptides in pulmonary innate defence. Respir. Res. 2006, 7, 29. [Google Scholar] [CrossRef]
- Mookherjee, N.; Anderson, M.A.; Haagsman, H.P.; Davidson, D.J. Antimicrobial host defence peptides: Functions and clinical potential. Nat. Rev. Drug Discov. 2020, 19, 311–332. [Google Scholar] [CrossRef]
- Beisswenger, C.; Bals, R. Antimicrobial Peptides in Lung Inflammation. Chem. Immunol. Allergy 2005, 86, 55–71. [Google Scholar] [CrossRef] [PubMed]
- Lai, Y.P.; Gallo, R.L. AMPed up immunity: How antimicrobial peptides have multiple roles in immune defense. Trends Immunol. 2009, 30, 131–141. [Google Scholar] [CrossRef]
- Manna, P.B.A.S.M.M.S.; Chowdhury, T.; Baindara, P.; Mandal, S.M. Fusion Protein Targeted Antiviral Peptides: Fragment-Based Drug Design (FBDD) Guided Rational Design of Dipeptides Against SARS-CoV-2. Curr. Protein Pept. Sci. 2020, 21, 938–947. [Google Scholar] [CrossRef]
- Baindara, P.; Roy, D.; Mandal, S.M.; Schrum, A.G. Conservation and Enhanced Binding of SARS-CoV-2 Omicron Spike Protein to Coreceptor Neuropilin-1 Predicted by Docking Analysis. Infect. Dis. Rep. 2022, 14, 29. [Google Scholar] [CrossRef] [PubMed]
- McCray, P.B., Jr.; Bentley, L. Human airway epithelia express a beta-defensin. Am. J. Respir Cell Mol. Biol. 1997, 16, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Diamond, G.; Legarda, D.; Ryan, L.K. The innate immune response of the respiratory epithelium. Immunol. Rev. 2000, 173, 27–38. [Google Scholar] [CrossRef] [PubMed]
- Schutte, B.C.; McCray, P.B. β-Defensins in Lung Host Defense. Annu. Rev. Physiol. 2002, 64, 709–748. [Google Scholar] [CrossRef]
- Bals, R.; Wang, X.; Zasloff, M.; Wilson, J.M. The peptide antibiotic ll-37/hcap-18 is expressed in epithelia of the human lung where it has broad antimicrobial activity at the airway surface. Proc. Natl. Acad. Sci. USA 1998, 95, 9541–9546. [Google Scholar] [CrossRef]
- Zanetti, M. Cathelicidins, multifunctional peptides of the innate immunity. J. Leukoc. Biol. 2004, 75, 39–48. [Google Scholar] [CrossRef]
- Crouch, E.; Parghi, D.; Kuan, S.F.; Persson, A. Surfactant protein D: Subcellular localization in nonciliated bronchiolar epithelial cells. Am. J. Physiol. Cell. Mol. Physiol. 1992, 263, L60–L66. [Google Scholar] [CrossRef]
- Kuroki, Y.; Takahashi, M.; Nishitani, C. Pulmonary collectins in innate immunity of the lung. Cell. Microbiol. 2007, 9, 1871–1879. [Google Scholar] [CrossRef] [PubMed]
- Dubin, R.F.; Robinson, S.K.; Widdicombe, J.H. Secretion of lactoferrin and lysozyme by cultures of human airway epithelium. Am. J. Physiol. Cell. Mol. Physiol. 2004, 286, L750–L755. [Google Scholar] [CrossRef] [PubMed]
- Ganz, T. Antimicrobial polypeptides. J. Leukoc. Biol. 2003, 75, 34–38. [Google Scholar] [CrossRef] [PubMed]
- Sallenave, J.M. Antimicrobial activity of antiproteinases. Biochem. Soc. Trans. 2002, 30, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Moraes, T.J.; Chow, C.W.; Downey, G.P. Proteases and lung injury. Crit. Care Med. 2003, 31, S189–S194. [Google Scholar] [CrossRef]
- Lindbom, J.; Ljungman, A.G.; Lindahl, M.; Tagesson, C. Increased Gene Expression of Novel Cytosolic and Secretory Phospholipase A2Types in Human Airway Epithelial Cells Induced by Tumor Necrosis Factor-α and IFN-γ. J. Interf. Cytokine Res. 2002, 22, 947–955. [Google Scholar] [CrossRef]
- Gimenez, A.P.; Wu, Y.-Z.; Paya, M.; Delclaux, C.; Touqui, L.; Goossens, P.L. High Bactericidal Efficiency of Type IIA Phospholipase A2 against Bacillus anthracis and Inhibition of Its Secretion by the Lethal Toxin. J. Immunol. 2004, 173, 521–530. [Google Scholar] [CrossRef]
- Christensen, T.G.; Blanchard, G.C.; Nolley, G.; Hayes, J.A. Ultrastructural localization of endogenous peroxidase in the lower respiratory tract of the guinea pig. Cell Tissue Res. 1981, 214, 407–415. [Google Scholar] [CrossRef]
- Starner, T.D.; Barker, C.K.; Jia, H.P.; Kang, Y.; McCray, P.B. CCL20 Is an Inducible Product of Human Airway Epithelia with Innate Immune Properties. Am. J. Respir. Cell Mol. Biol. 2003, 29, 627–633. [Google Scholar] [CrossRef]
- Sørensen, O.E.; Thapa, D.R.; Rosenthal, A.; Liu, L.; Roberts, A.A.; Ganz, T. Differential Regulation of β-Defensin Expression in Human Skin by Microbial Stimuli. J. Immunol. 2005, 174, 4870–4879. [Google Scholar] [CrossRef]
- Shestakova, T.; Zhuravel, E.; Bolgova, L.; Alekseenko, O.; Soldatkina, M.; Pogrebnoy, P. Expression of human beta-defensins-1, 2 and 4 mRNA in human lung tumor tissue: A pilot study. Exp. Oncol. 2008, 30, 153–156. [Google Scholar]
- Otte, J.-M.; Neumann, H.M.; Brand, S.; Schrader, H.; Schmidt, W.E.; Schmitz, F. Expression of beta-defensin 4 is increased in human gastritis. Eur. J. Clin. Investig. 2009, 39, 126–138. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Biragyn, A.; Hoover, D.M.; Lubkowski, J.; Oppenheim, J.J. Multiple Roles of Antimicrobial Defensins, Cathelicidins, and Eosinophil-Derived Neurotoxin in Host Defense. Annu. Rev. Immunol. 2004, 22, 181–215. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.L.; Hancock, R.E. Cationic host defense (antimicrobial) peptides. Curr. Opin. Immunol. 2006, 18, 24–30. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Chino, T.; Horst, O.V.; Hacker, B.M.; Clark, E.A.; Dale, B.A.; Chung, W.O. Differential and coordinated expression of defensins and cytokines by gingival epithelial cells and dendritic cells in response to oral bacteria. BMC Immunol. 2010, 11, 37. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Wang, A.; Marin, M.; Honnen, W.; Ramasamy, S.; Porter, E.; Subbian, S.; Pinter, A.; Melikyan, G.; Lu, W.; et al. Human Defensins Inhibit SARS-CoV-2 Infection by Blocking Viral Entry. Viruses 2021, 13, 1246. [Google Scholar] [CrossRef]
- Wang, C.; Wang, S.; Li, D.; Wei, D.-Q.; Zhao, J.; Wang, J. Human Intestinal Defensin 5 Inhibits SARS-CoV-2 Invasion by Cloaking ACE2. Gastroenterology 2020, 159, 1145–1147.e4. [Google Scholar] [CrossRef]
- Kudryashova, E.; Zani, A.; Vilmen, G.; Sharma, A.; Lu, W.; Yount, J.S.; Kudryashov, D.S. Inhibition of SARS-CoV-2 Infection by Human Defensin HNP1 and Retrocyclin RC-101. J. Mol. Biol. 2022, 434, 167225. [Google Scholar] [CrossRef]
- Holly, M.K.; Diaz, K.; Smith, J.G. Defensins in Viral Infection and Pathogenesis. Annu. Rev. Virol. 2017, 4, 369–391. [Google Scholar] [CrossRef]
- Matsumura, T.; Sugiyama, N.; Murayama, A.; Yamada, N.; Shiina, M.; Asabe, S.; Wakita, T.; Imawari, M.; Kato, T. Antimicrobial peptide LL-37 attenuates infection of hepatitis C virus. Hepatol. Res. 2015, 46, 924–932. [Google Scholar] [CrossRef]
- Agerberth, B.; Gunne, H.; Odeberg, J.; Kogner, P.; Boman, H.G.; Gudmundsson, G.H. FALL-39, a putative human peptide antibiotic, is cysteine-free and expressed in bone marrow and testis. Proc. Natl. Acad. Sci. USA 1995, 92, 195–199. [Google Scholar] [CrossRef] [PubMed]
- Harcourt, J.L.; McDonald, M.; Svoboda, P.; Pohl, J.; Tatti, K.; Haynes, L.M. Human cathelicidin, LL-37, inhibits respiratory syncytial virus infection in polarized airway epithelial cells. BMC Res. Notes 2016, 9, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Agier, J.; Efenberger, M.; Brzezińska-Błaszczyk, E. Review paper Cathelicidin impact on inflammatory cells. Central Eur. J. Immunol. 2015, 2, 225–235. [Google Scholar] [CrossRef]
- Bandurska, K.; Berdowska, A.; Barczyńska-Felusiak, R.; Krupa, P. Unique features of human cathelicidin LL-37. Biofactors 2015, 41, 289–300. [Google Scholar] [CrossRef]
- Sousa, F.H.; Casanova, V.; Findlay, F.; Stevens, C.; Svoboda, P.; Pohl, J.; Proudfoot, L.; Barlow, P.G. Cathelicidins display conserved direct antiviral activity towards rhinovirus. Peptides 2017, 95, 76–83. [Google Scholar] [CrossRef]
- Currie, S.M.; Findlay, E.G.; McFarlane, A.J.; Fitch, P.M.; Böttcher, B.; Colegrave, N.; Paras, A.; Jozwik, A.; Chiu, C.; Schwarze, J.; et al. Cathelicidins Have Direct Antiviral Activity against Respiratory Syncytial Virus In Vitro and Protective Function In Vivo in Mice and Humans. J. Immunol. 2016, 196, 2699–2710. [Google Scholar] [CrossRef] [PubMed]
- Parker, D.; Prince, A. Innate Immunity in the Respiratory Epithelium. Am. J. Respir. Cell Mol. Biol. 2011, 45, 189–201. [Google Scholar] [CrossRef]
- Telang, S. Lactoferrin: A Critical Player in Neonatal Host Defense. Nutrients 2018, 10, 1228. [Google Scholar] [CrossRef]
- Travis, S.M.; Conway, B.-A.D.; Zabner, J.; Smith, J.J.; Anderson, N.N.; Singh, P.K.; Greenberg, E.P.; Welsh, M.J. Activity of Abundant Antimicrobials of the Human Airway. Am. J. Respir. Cell Mol. Biol. 1999, 20, 872–879. [Google Scholar] [CrossRef]
- Singh, P.K.; Tack, B.F.; McCray, P.; Welsh, M. Synergistic and additive killing by antimicrobial factors found in human airway surface liquid. Am. J. Physiol. Cell. Mol. Physiol. 2000, 279, L799–L805. [Google Scholar] [CrossRef]
- Berlutti, F.; Pantanella, F.; Natalizi, T.; Frioni, A.; Paesano, R.; Polimeni, A.; Valenti, P. Antiviral Properties of Lactoferrin—A Natural Immunity Molecule. Molecules 2011, 16, 6992–7018. [Google Scholar] [CrossRef] [PubMed]
- Hirashima, N.; Orito, E.; Ohba, K.; Kondo, H.; Sakamoto, T.; Matsunaga, S.; Kato, A.; Nukaya, H.; Sakakibara, K.; Ohno, T.; et al. A randomized controlled trial of consensus interferon with or without lactoferrin for chronic hepatitis C patients with genotype 1b and high viral load. Hepatol. Res. 2004, 29, 9–12. [Google Scholar] [CrossRef] [PubMed]
- Shin, K.; Wakabayashi, H.; Yamauchi, K.; Teraguchi, S.; Tamura, Y.; Kurokawa, M.; Shiraki, K. Effects of orally administered bovine lactoferrin and lactoperoxidase on influenza virus infection in mice. J. Med Microbiol. 2005, 54, 717–723. [Google Scholar] [CrossRef] [PubMed]
- Elass, E.; Masson, M.; Mazurier, J.; Legrand, D. Lactoferrin Inhibits the Lipopolysaccharide-Induced Expression and Proteoglycan-Binding Ability of Interleukin-8 in Human Endothelial Cells. Infect. Immun. 2002, 70, 1860–1866. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, H.; Oda, H.; Yamauchi, K.; Abe, F. Lactoferrin for prevention of common viral infections. J. Infect. Chemother. 2014, 20, 666–671. [Google Scholar] [CrossRef]
- Kell, D.B.; Heyden, E.L.; Pretorius, E. The Biology of Lactoferrin, an Iron-Binding Protein That Can Help Defend Against Viruses and Bacteria. Front. Immunol. 2020, 11, 1221. [Google Scholar] [CrossRef]
- Berthon, B.S.; Williams, L.M.; Williams, E.J.; Wood, L.G. Effect of Lactoferrin Supplementation on Inflammation, Immune Function, and Prevention of Respiratory Tract Infections in Humans: A Systematic Review and Meta-analysis. Adv. Nutr. Int. Rev. J. 2022, 13, 1799–1819. [Google Scholar] [CrossRef]
- Ali, A.S.; Hasan, S.S.; Know, C.S.; Merchant, H.A. Lactoferrin reduces the risk of respiratory tract infections: A meta-analysis of randomized controlled trials. Clin. Nutr. ESPEN 2021, 45, 26–32. [Google Scholar] [CrossRef]
- Kouchi, I.; Yasuoka, S.; Ueda, Y.; Ogura, T. Analysis of secretory leukocyte protease inhibitor (SLPI) in bronchial secretions from patients with hypersecretory respiratory diseases. Tokushima J. Exp. Med. 1993, 40, 95–107. [Google Scholar]
- Majchrzak-Gorecka, M.; Majewski, P.; Grygier, B.; Murzyn, K.; Cichy, J. Secretory leukocyte protease inhibitor (SLPI), a multifunctional protein in the host defense response. Cytokine Growth Factor Rev. 2016, 28, 79–93. [Google Scholar] [CrossRef] [PubMed]
- Weldon, S.; McNally, P.; McElvaney, N.G.; Elborn, J.S.; McAuley, D.F.; Wartelle, J.; Belaaouaj, A.; Levine, R.L.; Taggart, C.C. Decreased Levels of Secretory Leucoprotease Inhibitor in the Pseudomonas-Infected Cystic Fibrosis Lung Are Due to Neutrophil Elastase Degradation. J. Immunol. 2009, 183, 8148–8156. [Google Scholar] [CrossRef]
- Hiemstra, P.S.; Maassen, R.J.; Stolk, J.; Heinzel-Wieland, R.; Steffens, G.J.; Dijkman, J.H. Antibacterial activity of antileukoprotease. Infect. Immun. 1996, 64, 4520–4524. [Google Scholar] [CrossRef]
- Lentsch, A.B.; Jordan, J.A.; Czermak, B.J.; Diehl, K.M.; Younkin, E.M.; Sarma, V.; Ward, P.A. Inhibition of NF-κB Activation and Augmentation of IκBβ by Secretory Leukocyte Protease Inhibitor during Lung Inflammation. Am. J. Pathol. 1999, 154, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Lentsch, A.B.; Yoshidome, H.; Warner, R.L.; Ward, P.A.; Edwards, M.J. Secretory leukocyte protease inhibitor in mice regulates local and remote organ inflammatory injury induced by hepatic ischemia/reperfusion. Gastroenterology 1999, 117, 953–961. [Google Scholar] [CrossRef] [PubMed]
- Greene, C.; Taggart, C.; Lowe, G.; Gallagher, P.; McElvaney, N.; O’Neill, S. Local Impairment of Anti–Neutrophil Elastase Capacity in Community-Acquired Pneumonia. J. Infect. Dis. 2003, 188, 769–776. [Google Scholar] [CrossRef]
- Meyer, M.; Kesic, M.J.; Clarke, J.; Ho, E.; Simmen, R.C.; Diaz-Sanchez, D.; Noah, T.L.; Jaspers, I. Sulforaphane induces SLPI secretion in the nasal mucosa. Respir. Med. 2013, 107, 472–475. [Google Scholar] [CrossRef]
- Ganz, T. Lysozyme. In Encyclopedia of Respiratory Medicine, Four-Volume Set; Academic Press: Cambridge, MA, USA, 2006; ISBN 9780123708793. [Google Scholar]
- Akinbi, H.T.; Epaud, R.; Bhatt, H.; Weaver, T.E. Bacterial Killing Is Enhanced by Expression of Lysozyme in the Lungs of Transgenic Mice. J. Immunol. 2000, 165, 5760–5766. [Google Scholar] [CrossRef]
- Dajani, R.; Zhang, Y.; Taft, P.J.; Travis, S.M.; Starner, T.D.; Olsen, A.; Zabner, J.; Welsh, M.J.; Engelhardt, J.F. Lysozyme Secretion by Submucosal Glands Protects the Airway from Bacterial Infection. Am. J. Respir. Cell Mol. Biol. 2005, 32, 548–552. [Google Scholar] [CrossRef]
- Song, Y.; Zhang, H.; Zhu, Y.; Zhao, X.; Lei, Y.; Zhou, W.; Yu, J.; Dong, X.; Wang, X.; Du, M.; et al. Lysozyme Protects Against Severe Acute Respiratory Syndrome Coronavirus 2 Infection and Inflammation in Human Corneal Epithelial Cells. Investig. Opthalmology Vis. Sci. 2022, 63, 16. [Google Scholar] [CrossRef]
- Brunaugh, A.D.; Seo, H.; Warnken, Z.; Ding, L.; Seo, S.H.; Smyth, H.D.C. Development and evaluation of inhalable composite niclosamide-lysozyme particles: A broad-spectrum, patient-adaptable treatment for coronavirus infections and sequalae. PLoS ONE 2021, 16, e0246803. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Li, Y.; Wang, L.; Guo, J.; Liu, W.; Meng, G.; Zhang, L.; Li, M.; Cong, L.; Sun, M. Recent Insights Into the Prognostic and Therapeutic Applications of Lysozymes. Front. Pharmacol. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Wijkstrom-Frei, C.; El-Chemaly, S.; Ali-Rachedi, R.; Gerson, C.; Cobas, M.A.; Forteza, R.; Salathe, M.; Conner, G.E. Lactoperoxidase and Human Airway Host Defense. Am. J. Respir. Cell Mol. Biol. 2003, 29, 206–212. [Google Scholar] [CrossRef] [PubMed]
- Fischer, A.J.; Lennemann, N.J.; Krishnamurthy, S.; Pócza, P.; Durairaj, L.; Launspach, J.L.; Rhein, B.A.; Wohlford-Lenane, C.; Lorentzen, D.; Bánfi, B.; et al. Enhancement of Respiratory Mucosal Antiviral Defenses by the Oxidation of Iodide. Am. J. Respir. Cell Mol. Biol. 2011, 45, 874–881. [Google Scholar] [CrossRef] [PubMed]
- Patel, U.; Gingerich, A.; Widman, L.; Sarr, D.; Tripp, R.A.; Rada, B. Susceptibility of influenza viruses to hypothiocyanite and hypoiodite produced by lactoperoxidase in a cell-free system. PLoS ONE 2018, 13, e0199167. [Google Scholar] [CrossRef]
- Homey, B.; Dieu-Nosjean, M.-C.; Wiesenborn, A.; Massacrier, C.; Pin, J.-J.; Oldham, E.; Catron, D.; Buchanan, M.E.; Müller, A.; deWaal Malefyt, R.; et al. Up-Regulation of Macrophage Inflammatory Protein-3α/CCL20 and CC Chemokine Receptor 6 in Psoriasis. J. Immunol. 2000, 164, 6621–6632. [Google Scholar] [CrossRef]
- Harant, H.; Eldershaw, S.A.; Lindley, I.J. Human macrophage inflammatory protein-3α/CCL20/LARC/Exodus/SCYA20 is transcriptionally upregulated by tumor necrosis factor-α via a non-standard NF-κB site. FEBS Lett. 2001, 509, 439–445. [Google Scholar] [CrossRef]
- Ghosh, M.; Shen, Z.; Schaefer, T.M.; Fahey, J.V.; Gupta, P.; Wira, C.R. ORIGINAL ARTICLE: CCL20/MIP3α is a Novel Anti-HIV-1 Molecule of the Human Female Reproductive Tract. Am. J. Reprod. Immunol. 2009, 62, 60–71. [Google Scholar] [CrossRef]
- Khalil, B.A.; Elemam, N.M.; Maghazachi, A.A. Chemokines and chemokine receptors during COVID-19 infection. Comput. Struct. Biotechnol. J. 2021, 19, 976–988. [Google Scholar] [CrossRef]
- Selsted, M.E.; Ouellette, A.J. Mammalian defensins in the antimicrobial immune response. Nat. Immunol. 2005, 6, 551–557. [Google Scholar] [CrossRef]
- Frye, M.; Bargon, J.; Dauletbaev, N.; Weber, A.; Wagner, T.O.F.; Gropp, R. Expression of human alpha-defensin 5 (HD5) mRNA in nasal and bronchial epithelial cells. J. Clin. Pathol. 2000, 53, 770–773. [Google Scholar] [CrossRef] [PubMed]
- Kao, C.Y.; Chen, Y.; Zhao, Y.H.; Wu, R. ORFeome-Based Search of Airway Epithelial Cell-Specific Novel Human β-Defensin Genes. Am. J. Respir. Cell Mol. Biol. 2003, 29, 71–80. [Google Scholar] [CrossRef] [PubMed]
- Yanagi, S.; Ashitani, J.-I.; Ishimoto, H.; Date, Y.; Mukae, H.; Chino, N.; Nakazato, M. Isolation of human β-defensin-4 in lung tissue and its increase in lower respiratory tract infection. Respir. Res. 2005, 6, 130. [Google Scholar] [CrossRef] [PubMed]
- Harder, J.; Bartels, J.; Christophers, E.; Schröder, J.-M. Isolation and Characterization of Human μ-Defensin-3, a Novel Human Inducible Peptide Antibiotic. J. Biol. Chem. 2001, 276, 5707–5713. [Google Scholar] [CrossRef] [PubMed]
- Littmann, M.; Albiger, B.; Frentzen, A.; Normark, S.; Henriques-Normark, B.; Plant, L. Streptococcus pneumoniae evades human dendritic cell surveillance by pneumolysin expression. EMBO Mol. Med. 2009, 1, 211–222. [Google Scholar] [CrossRef]
- Benincasa, M.; Mattiuzzo, M.; Herasimenka, Y.; Cescutti, P.; Rizzo, R.; Gennaro, R. Activity of antimicrobial peptides in the presence of polysaccharides produced by pulmonary pathogens. J. Pept. Sci. 2009, 15, 595–600. [Google Scholar] [CrossRef] [PubMed]
- Scharf, S.; Vardarova, K.; Lang, F.; Schmeck, B.; Opitz, B.; Flieger, A.; Heuner, K.; Hippenstiel, S.; Suttorp, N.; N’Guessan, P.D. Legionella pneumophila induces human beta Defensin-3 in pulmonary cells. Respir. Res. 2010, 11, 93. [Google Scholar] [CrossRef]
- Liu, L.; Wang, L.; Jia, H.P.; Zhao, C.; Heng, H.H.; Schutte, B.C.; McCray, P.; Ganz, T. Structure and mapping of the human β-defensin HBD-2 gene and its expression at sites of inflammation. Gene 1998, 222, 237–244. [Google Scholar] [CrossRef]
- Koczulla, A.R.; Bals, R. Antimicrobial peptides: Current status and therapeutic potential. Drugs 2003, 63, 389–406. [Google Scholar] [CrossRef]
- Jia, H.P.; Schutte, B.C.; Schudy, A.; Linzmeier, R.; Guthmiller, J.M.; Johnson, G.K.; Tack, B.F.; Mitros, J.P.; Rosenthal, A.; Ganz, T.; et al. Discovery of new human β-defensins using a genomics-based approach. Gene 2001, 263, 211–218. [Google Scholar] [CrossRef]
- Hancock, R.E.; Diamond, G. The role of cationic antimicrobial peptides in innate host defences. Trends Microbiol. 2000, 8, 402–410. [Google Scholar] [CrossRef]
- Harder, J.; Meyer-Hoffert, U.; Teran, L.M.; Schwichtenberg, L.; Bartels, J.; Maune, S.; Schröder, J.-M. Mucoid Pseudomonas aeruginosa, TNF- α, and IL-1 β, but Not IL-6, Induce Human β -Defensin-2 in Respiratory Epithelia. Am. J. Respir. Cell Mol. Biol. 2000, 22, 714–721. [Google Scholar] [CrossRef]
- Scott, M.G.; Vreugdenhil, A.C.E.; Buurman, W.A.; Hancock, R.E.W.; Gold, M.R. Cutting Edge: Cationic Antimicrobial Peptides Block the Binding of Lipopolysaccharide (LPS) to LPS Binding Protein. J. Immunol. 2000, 164, 549–553. [Google Scholar] [CrossRef] [PubMed]
- Scott, M.G.; Hancock, R.E.W. Cationic antimicrobial peptides and their multifunctional role in the immune system. Crit. Rev. Immunol. 2000, 20, 24. [Google Scholar] [CrossRef]
- Harder, R.; Meyer-Hoffert, U.; Wehkamp, K. Differential Gene Induction of Human β-Defensins (hBD-1, -2, -3, and -4) in Keratinocytes Is Inhibited by Retinoic Acid. J. Investig. Dermatol. 2004, 123, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Van Es, J.H.; Jay, P.; Gregorieff, A.; Van Gijn, M.E.; Jonkheer, S.; Hatzis, P.; Thiele, A.; van den Born, M.; Begthel, H.; Brabletz, T.; et al. Wnt signalling induces maturation of Paneth cells in intestinal crypts. Nat. Cell Biol. 2005, 7, 381–386. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, O.E.; Thapa, D.R.; Roupé, K.M.; Valore, E.V.; Sjöbring, U.; Roberts, A.A.; Schmidtchen, A.; Ganz, T. Injury-induced innate immune response in human skin mediated by transactivation of the epidermal growth factor receptor. J. Clin. Investig. 2006, 116, 1878–1885. [Google Scholar] [CrossRef]
- Rivas-Santiago, B.; Hernandez-Pando, R.; Carranza, C.; Juarez, E.; Contreras, J.L.; Aguilar-Leon, D.; Torres, M.; Sada, E. Expression of Cathelicidin LL-37 during Mycobacterium tuberculosis Infection in Human Alveolar Macrophages, Monocytes, Neutrophils, and Epithelial Cells. Infect. Immun. 2008, 76, 935–941. [Google Scholar] [CrossRef]
- Di Nardo, A.; Vitiello, A.; Gallo, R.L. Cutting Edge: Mast Cell Antimicrobial Activity Is Mediated by Expression of Cathelicidin Antimicrobial Peptide. J. Immunol. 2003, 170, 2274–2278. [Google Scholar] [CrossRef]
- Wang, T.-T.; Nestel, F.P.; Bourdeau, V.; Nagai, Y.; Wang, Q.; Liao, J.; Tavera-Mendoza, L.; Lin, R.; Hanrahan, J.W.; Mader, S.; et al. Cutting Edge: 1,25-Dihydroxyvitamin D3 Is a Direct Inducer of Antimicrobial Peptide Gene Expression. J. Immunol. 2004, 173, 2909–2912. [Google Scholar] [CrossRef]
- Liu, P.T.; Stenger, S.; Li, H.; Wenzel, L.; Tan, B.H.; Krutzik, S.R.; Ochoa, M.T.; Schauber, J.; Wu, K.; Meinken, C.; et al. Toll-Like Receptor Triggering of a Vitamin D-Mediated Human Antimicrobial Response. Science 2006, 311, 1770–1773. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Mallampalli, R.K. The Role of Surfactant in Lung Disease and Host Defense against Pulmonary Infections. Ann. Am. Thorac. Soc. 2015, 12, 765–774. [Google Scholar] [CrossRef] [PubMed]
- Wright, J.R. Immunoregulatory functions of surfactant proteins. Nat. Rev. Immunol. 2005, 5, 58–68. [Google Scholar] [CrossRef] [PubMed]
- King, R.J.; Ruch, J.; Gikas, E.G.; Platzker, A.C.; Creasy, R.K. Appearance of paoproteins of pulmonary surfactant in human amniotic fluid. J. Appl. Physiol. 1975, 39, 735–741. [Google Scholar] [CrossRef]
- Tomita, T.; Nagase, T.; Ohga, E.; Yamaguchi, Y.; Yoshizumi, M.; Ouchi, Y. Molecular mechanisms underlying human beta-defensin-2 gene expression in a human airway cell line (LC2/ad). Respirology 2002, 7, 305–310. [Google Scholar] [CrossRef]
- Hertz, C.J.; Wu, Q.; Porter, E.M.; Zhang, Y.J.; Weismüller, K.-H.; Godowski, P.J.; Ganz, T.; Randell, S.H.; Modlin, R.L. Activation of Toll-Like Receptor 2 on Human Tracheobronchial Epithelial Cells Induces the Antimicrobial Peptide Human β Defensin-2. J. Immunol. 2003, 171, 6820–6826. [Google Scholar] [CrossRef]
- MacRedmond, R.; Greene, C.; Taggart, C.C.; McElvaney, N.; O’Neill, S. Respiratory epithelial cells require Toll-like receptor 4 for induction of Human β-defensin 2 by Lipopolysaccharide. Respir. Res. 2005, 6, 116. [Google Scholar] [CrossRef]
- Froy, O. Regulation of mammalian defensin expression by Toll-like receptor-dependent and independent signalling pathways. Cell. Microbiol. 2005, 7, 1387–1397. [Google Scholar] [CrossRef]
- Hess, C.; Herr, C.; Beisswenger, C.; Zakharkina, T.; Schmid, R.M.; Bals, R. Myeloid RelA regulates pulmonary host defense networks. Eur. Respir. J. 2009, 35, 343–352. [Google Scholar] [CrossRef]
- Kao, C.-Y.; Chen, Y.; Thai, P.; Wachi, S.; Huang, F.; Kim, C.; Harper, R.W.; Wu, R. IL-17 Markedly Up-Regulates β-Defensin-2 Expression in Human Airway Epithelium via JAK and NF-κB Signaling Pathways. J. Immunol. 2004, 173, 3482–3491. [Google Scholar] [CrossRef]
- Albanesi, C.; Fairchild, H.R.; Madonna, S.; Scarponi, C.; De Pità, O.; Leung, D.Y.M.; Howell, M.D. IL-4 and IL-13 Negatively Regulate TNF-α- and IFN-γ-Induced β-Defensin Expression through STAT-6, Suppressor of Cytokine Signaling (SOCS)-1, and SOCS-3. J. Immunol. 2007, 179, 984–992. [Google Scholar] [CrossRef]
- Scharf, S.; Hippenstiel, S.; Flieger, A.; Suttorp, N.; N’Guessan, P.D. Induction of human β-defensin-2 in pulmonary epithelial cells by Legionella pneumophila: Involvement of TLR2 and TLR5, p38 MAPK, JNK, NF-κB, and AP-1. Am. J. Physiol. Cell. Mol. Physiol. 2010, 298, L687–L695. [Google Scholar] [CrossRef]
- van der Does, A.M.; Kenne, E.; Koppelaar, E.; Agerberth, B.; Lindbom, L. Vitamin D3 and phenylbutyrate promote development of a human dendritic cell subset displaying enhanced antimicrobial properties. J. Leukoc. Biol. 2014, 95, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Yim, S.; Dhawan, P.; Ragunath, C.; Christakos, S.; Diamond, G. Induction of cathelicidin in normal and CF bronchial epithelial cells by 1,25-dihydroxyvitamin D3. J. Cyst. Fibros. 2007, 6, 403–410. [Google Scholar] [CrossRef]
- Schauber, J.; Dorschner, R.A.; Coda, A.B.; Büchau, A.S.; Liu, P.T.; Kiken, D.; Helfrich, Y.R.; Kang, S.; Elalieh, H.Z.; Steinmeyer, A.; et al. Injury enhances TLR2 function and antimicrobial peptide expression through a vitamin D–dependent mechanism. J. Clin. Investig. 2007, 117, 803–811. [Google Scholar] [CrossRef]
- Schauber, J.; Oda, Y.; Büchau, A.S.; Yun, Q.-C.; Steinmeyer, A.; Zügel, U.; Bikle, D.D.; Gallo, R.L. Histone Acetylation in Keratinocytes Enables Control of the Expression of Cathelicidin and CD14 by 1,25-Dihydroxyvitamin D3. J. Investig. Dermatol. 2008, 128, 816–824. [Google Scholar] [CrossRef] [PubMed]
- Dhawan, P.; Wei, R.; Sun, C.; Gombart, A.F.; Koeffler, H.P.; Diamond, G.; Christakos, S. C/EBPα and the Vitamin D Receptor Cooperate in the Regulation of Cathelicidin in Lung Epithelial Cells. J. Cell. Physiol. 2014, 230, 464–472. [Google Scholar] [CrossRef] [PubMed]
- Gombart, A.F.; Borregaard, N.; Koeffler, H.P. Human cathelicidin antimicrobial peptide (CAMP) gene is a direct target of the vitamin D receptor and is strongly up-regulated in myeloid cells by 1,25-dihydroxyvitamin D3. FASEB J. 2005, 19, 1067–1077. [Google Scholar] [CrossRef]
- Wang, T.-T.; Dabbas, B.; Laperriere, D.; Bitton, A.J.; Soualhine, H.; Tavera-Mendoza, L.E.; Dionne, S.; Servant, M.J.; Bitton, A.; Seidman, E.G.; et al. Direct and Indirect Induction by 1,25-Dihydroxyvitamin D3 of the NOD2/CARD15-Defensin β2 Innate Immune Pathway Defective in Crohn Disease. J. Biol. Chem. 2010, 285, 2227–2231. [Google Scholar] [CrossRef]
- Hertting, O.; Holm, Å.; Lüthje, P.; Brauner, H.; Dyrdak, R.; Jonasson, A.F.; Wiklund, P.; Chromek, M.; Brauner, A. Vitamin D Induction of the Human Antimicrobial Peptide Cathelicidin in the Urinary Bladder. PLoS ONE 2010, 5, e15580. [Google Scholar] [CrossRef]
- García-Quiroz, J.; García-Becerra, R.; Santos-Martínez, N.; Avila, E.; Larrea, F.; Díaz, L. Calcitriol stimulates gene expression of cathelicidin antimicrobial peptide in breast cancer cells with different phenotype. J. Biomed. Sci. 2016, 23, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Castañeda-Delgado, J.E.; Araujo, Z.; Gonzalez-Curiel, I.; Serrano, C.J.; Santiago, C.R.; Enciso-Moreno, J.A.; Rivas-Santiago, B.; Information, R. Vitamin D and L-Isoleucine Promote Antimicrobial Peptide hBD-2 Production in Peripheral Blood Mononuclear Cells from Elderly Individuals. Int. J. Vitam. Nutr. Res. 2016, 86, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Paria, K.; Paul, D.; Chowdhury, T.; Pyne, S.; Chakraborty, R.; Mandal, S.M. Synergy of melanin and vitamin-D may play a fundamental role in preventing SARS-CoV-2 infections and halt COVID-19 by inactivating furin protease. Transl. Med. Commun. 2020, 5, 1–14. [Google Scholar] [CrossRef]
- Wancket, L.M.; Frazier, W.J.; Liu, Y. Mitogen-activated protein kinase phosphatase (MKP)-1 in immunology, physiology, and disease. Life Sci. 2012, 90, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S.M.; Manna, S.; Mondal, S.; Ghosh, A.K.; Chakraborty, R. Transcriptional regulation of human defense peptides: A new direction in infection control. Biol. Chem. 2018, 399, 1277–1284. [Google Scholar] [CrossRef] [PubMed]
- Robinson, K.; Ma, X.; Liu, Y.; Qiao, S.; Hou, Y.; Zhang, G. Dietary modulation of endogenous host defense peptide synthesis as an alternative approach to in-feed antibiotics. Anim. Nutr. 2018, 4, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Kida, Y.; Shimizu, T.; Kuwano, K. Sodium butyrate up-regulates cathelicidin gene expression via activator protein-1 and histone acetylation at the promoter region in a human lung epithelial cell line, EBC-1. Mol. Immunol. 2006, 43, 1972–1981. [Google Scholar] [CrossRef]
- Schauber, J.; Svanholm, C.; Termén, S.; Iffland, K.; Menzel, T.; Scheppach, W.; Melcher, R.; Agerberth, B.; Lührs, H.; Gudmundsson, G.H. Expression of the cathelicidin LL-37 is modulated by short chain fatty acids in colonocytes: Relevance of signalling pathways. Gut 2003, 52, 735–741. [Google Scholar] [CrossRef]
- Schwab, M.; Reynders, V.; Shastri, Y.; Loitsch, S.; Stein, J.; Schröder, O. Role of nuclear hormone receptors in butyrate-mediated up-regulation of the antimicrobial peptide cathelicidin in epithelial colorectal cells. Mol. Immunol. 2007, 44, 2107–2114. [Google Scholar] [CrossRef]
- Steinmann, J.; Halldórsson, S.; Agerberth, B.; Gudmundsson, G.H. Phenylbutyrate Induces Antimicrobial Peptide Expression. Antimicrob. Agents Chemother. 2009, 53, 5127–5133. [Google Scholar] [CrossRef]
- Peric, M.; Koglin, S.; Dombrowski, Y.; Groß, K.; Bradac, E.; Ruzicka, T.; Schauber, J. VDR and MEK-ERK dependent induction of the antimicrobial peptide cathelicidin in keratinocytes by lithocholic acid. Mol. Immunol. 2009, 46, 3183–3187. [Google Scholar] [CrossRef] [PubMed]
- Baker, R.G.; Hayden, M.S.; Ghosh, S. NF-κB, inflammation, and metabolic disease. Cell Metab. 2011. [Google Scholar] [CrossRef] [PubMed]
- Fehlbaum, P.; Rao, M.; Zasloff, M.; Anderson, G.M. An essential amino acid induces epithelial β-defensin expression. Proc. Natl. Acad. Sci. USA 2000, 97, 12723–12728. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.-J.; Zheng, X.-W.; Feng, T.; Huang, J.; Chen, J.; Wu, Y.-Y.; Zhou, L.-M.; Tu, W.-W.; Li, H. Andrographolide exerted its antimicrobial effects by upregulation of human β-defensin-2 induced through p38 MAPK and NF-κB pathway in human lung epithelial cells. Can. J. Physiol. Pharmacol. 2012, 90, 647–653. [Google Scholar] [CrossRef]
- Gan, Y.; Cui, X.; Ma, T.; Liu, Y.; Li, A.; Huang, M. Paeoniflorin Upregulates β-Defensin-2 Expression in Human Bronchial Epithelial Cell Through the p38 MAPK, ERK, and NF-κB Signaling Pathways. Inflammation 2014, 37, 1468–1475. [Google Scholar] [CrossRef]
- Xiong, W.-B.; Shao, Z.-J.; Xiong, Y.; Chen, J.; Sun, Y.; Zhu, L.; Zhou, L.-M. Dehydroandrographolide enhances innate immunity of intestinal tract through up-regulation the expression of hBD-2. DARU J. Pharm. Sci. 2015, 23, 1–7. [Google Scholar] [CrossRef]
- Park, K.; Elias, P.M.; Hupe, M.; Borkowski, A.W.; Gallo, R.L.; Shin, K.-O.; Lee, Y.-M.; Holleran, W.M.; Uchida, Y. Resveratrol Stimulates Sphingosine-1-Phosphate Signaling of Cathelicidin Production. J. Investig. Dermatol. 2013, 133, 1942–1949. [Google Scholar] [CrossRef]
- Park, K.; Kim, Y.-I.; Shin, K.-O.; Seo, H.S.; Kim, J.Y.; Mann, T.; Oda, Y.; Lee, Y.-M.; Holleran, W.M.; Elias, P.M.; et al. The dietary ingredient, genistein, stimulates cathelicidin antimicrobial peptide expression through a novel S1P-dependent mechanism. J. Nutr. Biochem. 2014, 25, 734–740. [Google Scholar] [CrossRef]
- Xiong, H.; Guo, B.; Gan, Z.; Song, D.; Lu, Z.; Yi, H.; Wu, Y.; Wang, Y.; Du, H. Butyrate upregulates endogenous host defense peptides to enhance disease resistance in piglets via histone deacetylase inhibition. Sci. Rep. 2016, 6, 27070. [Google Scholar] [CrossRef]
- Wang, J.; Huang, N.; Xiong, J.; Wei, H.; Jiang, S.; Peng, J. Caprylic acid and nonanoic acid upregulate endogenous host defense peptides to enhance intestinal epithelial immunological barrier function via histone deacetylase inhibition. Int. Immunopharmacol. 2018, 65, 303–311. [Google Scholar] [CrossRef]
- Andou, A.; Takehana, K. Regulatory Roles of Amino Acids in Immune Response. Curr. Rheumatol. Rev. 2009, 5, 252–258. [Google Scholar] [CrossRef]
- Sun, J.; Furio, L.; Mecheri, R.; van der Does, A.M.; Lundeberg, E.; Saveanu, L.; Chen, Y.; van Endert, P.; Agerberth, B.; Diana, J. Pancreatic β-Cells Limit Autoimmune Diabetes via an Immunoregulatory Antimicrobial Peptide Expressed under the Influence of the Gut Microbiota. Immunity 2015, 43, 304–317. [Google Scholar] [CrossRef] [PubMed]
- Sherman, H.; Chapnik, N.; Froy, O. Albumin and amino acids upregulate the expression of human beta-defensin 1. Mol. Immunol. 2006, 43, 1617–1623. [Google Scholar] [CrossRef]
- Mao, X.; Qi, S.; Yu, B.; Huang, Z.; Chen, H.; Mao, Q.; Han, G.; Chen, D. Dietary l-arginine supplementation enhances porcine β-defensins gene expression in some tissues of weaned pigs. Livest. Sci. 2012, 148, 103–108. [Google Scholar] [CrossRef]
- Melano, I.; Kuo, L.-L.; Lo, Y.-C.; Sung, P.-W.; Tien, N.; Su, W.-C. Effects of Basic Amino Acids and Their Derivatives on SARS-CoV-2 and Influenza-A Virus Infection. Viruses 2021, 13, 1301. [Google Scholar] [CrossRef] [PubMed]
- Sunkara, L.T.; Jiang, W.; Zhang, G. Modulation of Antimicrobial Host Defense Peptide Gene Expression by Free Fatty Acids. PLoS ONE 2012, 7, e49558. [Google Scholar] [CrossRef]
- Alva-Murillo, N.; Ochoa-Zarzosa, A.; López-Meza, J.E. Short chain fatty acids (propionic and hexanoic) decrease Staphylococcus aureus internalization into bovine mammary epithelial cells and modulate antimicrobial peptide expression. Veter- Microbiol. 2012, 155, 324–331. [Google Scholar] [CrossRef]
- Roy, C.; Mandal, S.M.; Mondal, S.K.; Mukherjee, S.; Mapder, T.; Ghosh, W.; Chakraborty, R. Trends of mutation accumulation across global SARS-CoV-2 genomes: Implications for the evolution of the novel coronavirus. Genomics 2020, 112, 5331–5342. [Google Scholar] [CrossRef]
- Baindara, P.; Ghosh, A.K.; Mandal, S.M. Coevolution of Resistance Against Antimicrobial Peptides. Microb. Drug Resist. 2020, 26, 880–899. [Google Scholar] [CrossRef]
- Bentley-Hewitt, K.L.; Blatchford, P.A.; Parkar, S.G.; Ansell, J.; Pernthaner, A. Digested and Fermented Green Kiwifruit Increases Human β-Defensin 1 and 2 Production In vitro. Plant Foods Hum. Nutr. 2012, 67, 208–214. [Google Scholar] [CrossRef]
- Jiang, W.; Sunkara, L.T.; Zeng, X.; Deng, Z.; Myers, S.M.; Zhang, G. Differential regulation of human cathelicidin LL-37 by free fatty acids and their analogs. Peptides 2013, 50, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Nylén, F.; Miraglia, E.; Cederlund, A.; Ottosson, H.; Strömberg, R.; Gudmundsson, G.H.; Agerberth, B. Boosting innate immunity: Development and validation of a cell-based screening assay to identify LL-37 inducers. J. Endotoxin Res. 2013, 20, 364–376. [Google Scholar] [CrossRef] [PubMed]
- Cederlund, A.; Kai-Larsen, Y.; Printz, G.; Yoshio, H.; Alvelius, G.; Lagercrantz, H.; Strömberg, R.; Jörnvall, H.; Gudmundsson, G.H.; Agerberth, B. Lactose in Human Breast Milk an Inducer of Innate Immunity with Implications for a Role in Intestinal Homeostasis. PLoS ONE 2013, 8, e53876. [Google Scholar] [CrossRef] [PubMed]
- Malik, A.N.; Al-Kafaji, G. Glucose regulation of β-defensin-1 mRNA in human renal cells. Biochem. Biophys. Res. Commun. 2007, 353, 318–323. [Google Scholar] [CrossRef]
- Díaz, L.A.C.; Miramontes, M.G.F.; Hurtado, P.C.; Allen, K.; Ávila, M.G.; de Oca, E.P.M. Ascorbic Acid, Ultraviolet C Rays, and Glucose but not Hyperthermia Are Elicitors of Humanβ-Defensin 1 mRNA in Normal Keratinocytes. BioMed Res. Int. 2015, 2015, 714580. [Google Scholar] [CrossRef] [PubMed]
- Barnea, M.; Madar, Z.; Froy, O. Glucose and insulin are needed for optimal defensin expression in human cell lines. Biochem. Biophys. Res. Commun. 2008, 367, 452–456. [Google Scholar] [CrossRef]
- Lan, C.-C.E.; Wu, C.-S.; Huang, S.-M.; Kuo, H.-Y.; Wu, I.-H.; Wen, C.-H.; Chai, C.-Y.; Fang, A.-H.; Chen, G.-S. High-Glucose Environment Inhibits p38MAPK Signaling and Reduces Human β-3 Expression in Keratinocytes. Mol. Med. 2011, 17, 771–779. [Google Scholar] [CrossRef]
- Donnarumma, G.; Buommino, E.; Baroni, A.; Auricchio, L.; De Filippis, A.; Cozza, V.; Msika, P.; Piccardi, N.; Tufano, M.A. Effects of AV119, a natural sugar from avocado, on Malassezia furfur invasiveness and on the expression of HBD-2 and cytokines in human keratinocytes. Exp. Dermatol. 2007, 16, 912–919. [Google Scholar] [CrossRef]
- Paoletti, I.; Buommino, E.; Fusco, A.; Baudouin, C.; Msika, P.; Tufano, M.A.; Baroni, A.; Donnarumma, G. Patented natural avocado sugar modulates the HBD-2 and HBD-3 expression in human keratinocytes through Toll-like receptor-2 and ERK/MAPK activation. Arch. Dermatol. Res. 2012, 304, 619–625. [Google Scholar] [CrossRef]
- Žugčić, T.; Abdelkebir, R.; Alcantara, C.; Collado, M.C.; Garcia-Perez, J.V.; Meléndez-Martínez, A.J.; Režek Jambrak, A.; Lorenzo, J.M.; Barba, F.J. From extraction of valuable compounds to health promoting benefits of olive leaves through bioaccessibility, bioavailability and impact on gut microbiota. Trends Food Sci. Technol. 2019, 83, 63–77. [Google Scholar] [CrossRef]
- Van Hai, N. The use of medicinal plants as immunostimulants in aquaculture: A review. Aquaculture 2015, 446, 88–96. [Google Scholar] [CrossRef]
- Cabrera, C.; Artacho, R.; Giménez, R. Beneficial Effects of Green Tea—A Review. J. Am. Coll. Nutr. 2006, 25, 79–99. [Google Scholar] [CrossRef] [PubMed]
- Cooper, R. Green tea and theanine: Health benefits. Int. J. Food Sci. Nutr. 2011, 63, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Bedran, T.B.L.; Feghali, K.; Zhao, L.; Spolidorio, D.M.P.; Grenier, D. Green tea extract and its major constituent, epigallocatechin-3-gallate, induce epithelial beta-defensin secretion and prevent beta-defensin degradation by Porphyromonas gingivalis. J. Periodontal Res. 2013, 49, 615–623. [Google Scholar] [CrossRef]
- Bedran, T.B.L.; Morin, M.-P.; Spolidorio, D.P.; Grenier, D. Black Tea Extract and Its Theaflavin Derivatives Inhibit the Growth of Periodontopathogens and Modulate Interleukin-8 and β-Defensin Secretion in Oral Epithelial Cells. PLoS ONE 2015, 10, e0143158. [Google Scholar] [CrossRef]
- Promsong, A.; Chung, W.O.; Satthakarn, S.; Nittayananta, W. Ellagic acid modulates the expression of oral innate immune mediators: Potential role in mucosal protection. J. Oral Pathol. Med. 2014, 44, 214–221. [Google Scholar] [CrossRef]
- Lin, C.-W.; Tsai, F.-J.; Tsai, C.-H.; Lai, C.-C.; Wan, L.; Ho, T.-Y.; Hsieh, C.-C.; Chao, P.-D.L. Anti-SARS coronavirus 3C-like protease effects of Isatis indigotica root and plant-derived phenolic compounds. Antivir. Res. 2005, 68, 36–42. [Google Scholar] [CrossRef]
- Mao, X.; Qi, S.; Yu, B.; He, J.; Yu, J.; Chen, D. Zn2+ and l-isoleucine induce the expressions of porcine β-defensins in IPEC-J2 cells. Mol. Biol. Rep. 2012, 40, 1547–1552. [Google Scholar] [CrossRef]
- Pernet, I.; Reymermier, C.; Guezennec, A.; Branka, J.-E.; Guesnet, J.; Perrier, E.; Dezutter-Dambuyant, C.; Schmitt, D.; Viac, J. Calcium triggers beta-defensin (hBD-2 and hBD-3) and chemokine macrophage inflammatory protein-3alpha (MIP-3alpha/CCL20) expression in monolayers of activated human keratinocytes. Exp. Dermatol. 2003, 12, 755–760. [Google Scholar] [CrossRef]
- Chen, K.; Chen, H.; Faas, M.M.; de Haan, B.J.; Li, J.; Xiao, P.; Zhang, H.; Diana, J.; de Vos, P.; Sun, J. Specific inulin-type fructan fibers protect against autoimmune diabetes by modulating gut immunity, barrier function, and microbiota homeostasis. Mol. Nutr. Food Res. 2017, 61, 1601006. [Google Scholar] [CrossRef]
- He, Y.; Wu, C.; Li, J.; Li, H.; Sun, Z.; Zhang, H.; De Vos, P.; Pan, L.-L.; Sun, J. Inulin-Type Fructans Modulates Pancreatic–Gut Innate Immune Responses and Gut Barrier Integrity during Experimental Acute Pancreatitis in a Chain Length-Dependent Manner. Front. Immunol. 2017, 8, 1209. [Google Scholar] [CrossRef]
- Termén, S.; Tollin, M.; Rodriguez, E.; Sveinsdóttir, S.H.; Jóhannesson, B.; Cederlund, A.; Sjövall, J.; Agerberth, B.; Gudmundsson, G.H. PU.1 and bacterial metabolites regulate the human gene CAMP encoding antimicrobial peptide LL-37 in colon epithelial cells. Mol. Immunol. 2008, 45, 3947–3955. [Google Scholar] [CrossRef] [PubMed]
- Ottosson, H.; Nylén, F.; Sarker, P.; Miraglia, E.; Bergman, P.; Gudmundsson, G.H.; Raqib, R.; Agerberth, B.; Strömberg, R. Potent Inducers of Endogenous Antimicrobial Peptides for Host Directed Therapy of Infections. Sci. Rep. 2016, 6, 36692. [Google Scholar] [CrossRef] [PubMed]
- Holstein, J.; Fehrenbacher, B.; Brück, J.; Müller-Hermelink, E.; Schäfer, I.; Carevic, M.; Schittek, B.; Schaller, M.; Ghoreschi, K.; Eberle, F.C. Anthralin modulates the expression pattern of cytokeratins and antimicrobial peptides by psoriatic keratinocytes. J. Dermatol. Sci. 2017, 87, 236–245. [Google Scholar] [CrossRef]
- Lyu, W.; Deng, Z.; Sunkara, L.T.; Becker, S.; Robinson, K.; Matts, R.; Zhang, G. High Throughput Screening for Natural Host Defense Peptide-Inducing Compounds as Novel Alternatives to Antibiotics. Front. Cell. Infect. Microbiol. 2018, 8, 191. [Google Scholar] [CrossRef]
- Zhao, Y.; Chen, F.; Wu, W.; Sun, M.; Bilotta, A.J.; Yao, S.; Xiao, Y.; Huang, X.; Eaves-Pyles, T.D.; Golovko, G.; et al. GPR43 mediates microbiota metabolite SCFA regulation of antimicrobial peptide expression in intestinal epithelial cells via activation of mTOR and STAT3. Mucosal Immunol. 2018, 11, 752–762. [Google Scholar] [CrossRef]
- Guo, C.; Rosoha, E.; Lowry, M.B.; Borregaard, N.; Gombart, A.F. Curcumin induces human cathelicidin antimicrobial peptide gene expression through a vitamin D receptor-independent pathway. J. Nutr. Biochem. 2012, 24, 754–759. [Google Scholar] [CrossRef]
- Kauko, O.; Laajala, T.D.; Jumppanen, M.; Hintsanen, P.; Suni, V.; Haapaniemi, P.; Corthals, G.; Aittokallio, T.; Westermarck, J.; Imanishi, S. Label-free quantitative phosphoproteomics with novel pairwise abundance normalization reveals synergistic RAS and CIP2A signaling. Sci. Rep. 2015, 5, 13099. [Google Scholar] [CrossRef]
- Bader, G.D.; Hogue, C.W.V. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4, 2–27. [Google Scholar] [CrossRef] [PubMed]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.-H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef]
- Janky, R.; Verfaillie, A.; Imrichova, H.; Van de Sande, B.; Standaert, L.; Christiaens, V.; Hulselmans, G.; Herten, K.; Sanchez, M.N.; Potier, D.; et al. iRegulon: From a Gene List to a Gene Regulatory Network Using Large Motif and Track Collections. PLOS Comput. Biol. 2014, 10, e1003731. [Google Scholar] [CrossRef] [PubMed]
- Baindara, P.; Agrawal, S.; Mandal, S.M. Host-directed therapies: A potential solution to combat COVID-19. Expert Opin. Biol. Ther. 2020, 20, 1117–1120. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S.M.; Panda, S. Inhaler with electrostatic sterilizer and use of cationic amphiphilic peptides may accelerate recovery from COVID-19. Biotechniques 2020, 69, 206–210. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S. Peptide targets to SARS-CoV-2. J. Glob. Infect. Dis. 2020, 12, 234–235. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, T.; Baindara, P.; Mandal, S.M. LPD-12: A promising lipopeptide to control COVID-19. Int. J. Antimicrob. Agents 2020, 57, 106218. [Google Scholar] [CrossRef] [PubMed]




| AMPs/Factors | Distribution in Lung | References |
|---|---|---|
| Defensins (α and β defensins 1–4) | Lung epithelium, airways, neutrophils | [17,18,19] |
| Cathelicidins (LL37) | Lung epithelium, airways, respiratory sub-mucosal glands, neutrophils | [20,21] |
| Surfactant protein-A and surfactant protein-D (SP-A, SP-D) | Lung epithelium | [22,23] |
| Lysozyme | Tracheobronchial sub-mucosal glands, tracheal and bronchial surface epithelium, neutrophils, alveolar macrophages, monocytes | [24,25] |
| Secretory leukocyte proteinase inhibitor (SLPI) | Respiratory sub-mucosal glands, monocytes, alveolar macrophages, neutrophils | [26,27] |
| Lactoferrin | Tracheobronchial sub-mucosal glands, tracheal surface epithelium, neutrophils | [24] |
| Phospholipase A2 | Lung epithelium, neutrophils | [28,29] |
| Lactoperoxidase | Airways mucosal membrane | [30] |
| CCL20 | Airways epithelium | [31] |
| Gene | Protein | Molecular Function (UniProt) | Ensemble ID |
|---|---|---|---|
| SLPI | Antileukoproteinase | Antibiotic, Antimicrobial, Protease inhibitor, Serine protease inhibitor | ENSG00000124107 |
| CCL20 | C-C Motif Chemokine 20 | Antibiotic, Antimicrobial, Cytokine | ENSG00000115009 |
| STAT 1 | Signal transducer and activator of transcription 1-alpha/beta | Activator, DNA-binding | ENSG00000115415 |
| ZBP1 | Z-DNA-binding protein 1 | DNA-binding | ENSG00000124256 |
| TNFRSF1B | Tumor necrosis factor receptor superfamily member 1B | Receptor | ENSG00000028137 |
| TANK | TRAF family member-associated NF-kappa-β activation | Ubiquitin protein ligase binding, metal ion binding. | ENSG00000136560 |
| IL6 | Interleukin-6 | Cytokine, a Growth factor | ENSG00000136244 |
| CEBPA | CCAAT/enhancer-binding protein alpha | Activator, Developmental protein, DNA-binding | ENSG00000245848 |
| TLR5 | Toll-like receptor 5 | Receptor | ENSG00000187554 |
| LTF | Lactotransferrin | Antibiotic, Antimicrobial, DNA-binding, Heparin-binding, Hydrolase, Protease, Serine protease | ENSG00000012223 |
| LYZ | Lysozyme | Antimicrobial, Bacteriolytic enzyme, Glycosidase, Hydrolase, Milk protein | ENSG00000090382 |
| LPO | Lactoperoxidase | Antimicrobial, Oxidoreductase, Peroxidase | ENSG00000167419 |
| CAMP/LL37 | Cathelicidin antimicrobial peptide | Antibiotic, Antimicrobial | ENSG00000164047 |
| HBD-1 | Beta-defensin 1 | Antibiotic, Antimicrobial, Defensin | ENSG00000164825 |
| CC10/CCSP/SCGB1A1A | Uteroglobin | Phospholipase A2 inhibitor | ENSG00000149021 |
| STATH | Statgerin | Biomineralization | ENSG00000126549 |
| MSMB | Beta-microseminoprotein | Secretion of mucus, antimicrobial andanti-inflammatory peptides | ENSG00000263639 |
| BPIFA1 | BPI fold containing family A member 1 | Antibiotic, Antimicrobial | ENSG00000198183 |
| PI3 | Elafin | Kinase, Serine/threonine-protein kinase, Transferase | ENSG00000124102 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baindara, P.; Ganguli, S.; Chakraborty, R.; Mandal, S.M. Preventing Respiratory Viral Diseases with Antimicrobial Peptide Master Regulators in the Lung Airway Habitat. Clin. Pract. 2023, 13, 125-147. https://doi.org/10.3390/clinpract13010012
Baindara P, Ganguli S, Chakraborty R, Mandal SM. Preventing Respiratory Viral Diseases with Antimicrobial Peptide Master Regulators in the Lung Airway Habitat. Clinics and Practice. 2023; 13(1):125-147. https://doi.org/10.3390/clinpract13010012
Chicago/Turabian StyleBaindara, Piyush, Sriradha Ganguli, Ranadhir Chakraborty, and Santi M. Mandal. 2023. "Preventing Respiratory Viral Diseases with Antimicrobial Peptide Master Regulators in the Lung Airway Habitat" Clinics and Practice 13, no. 1: 125-147. https://doi.org/10.3390/clinpract13010012
APA StyleBaindara, P., Ganguli, S., Chakraborty, R., & Mandal, S. M. (2023). Preventing Respiratory Viral Diseases with Antimicrobial Peptide Master Regulators in the Lung Airway Habitat. Clinics and Practice, 13(1), 125-147. https://doi.org/10.3390/clinpract13010012







