Assessment of Novel Genetic Diversity Induced by Mutagenesis and Estimation of Genetic Parameters in Sesame M4 Mutant Lines
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Field Experiment
2.3. Data Collecting and Parametrs Studied
- Genotypic variance , where MSg is the genotype mean square, MSe is the mean square of experimental error, and r is the number of replicates.
- Phenotypic variance:
- Phenotypic coefficient of variation:
- Genotypic coefficient of variation:
- Broad sense heritability:
- Genetic advance:
- Genetic advance over mean: where is the grand mean and k = 2.06 is the intensity of selection at 5% [30].
2.4. Statistical Analyses
3. Results and Discussion
3.1. Effects of Genotype, Environment and Their Interaction
3.2. Mean Performances of Genotypes
3.2.1. Number of Days to Flowering
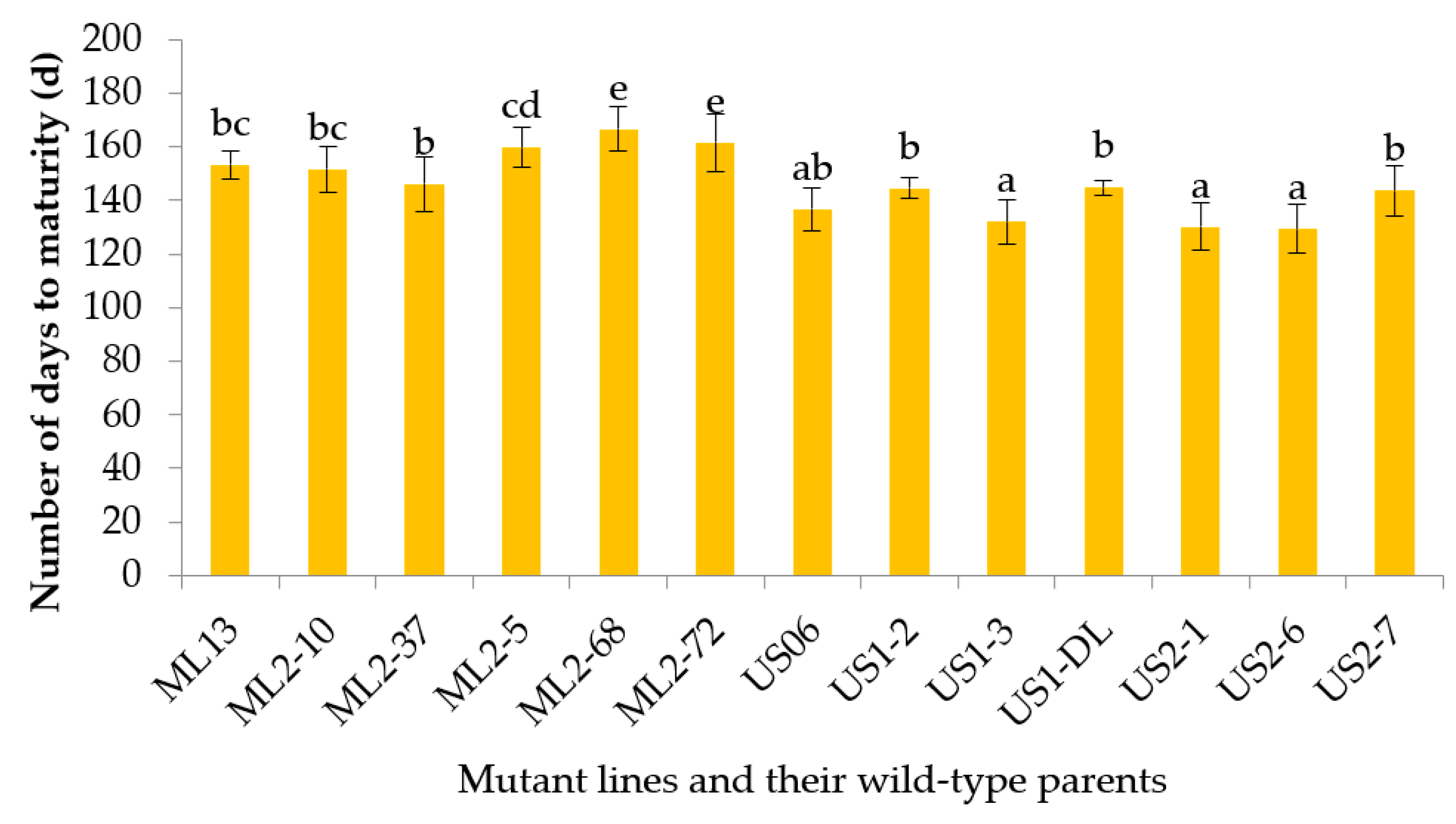
3.2.2. Number of Days to Maturity
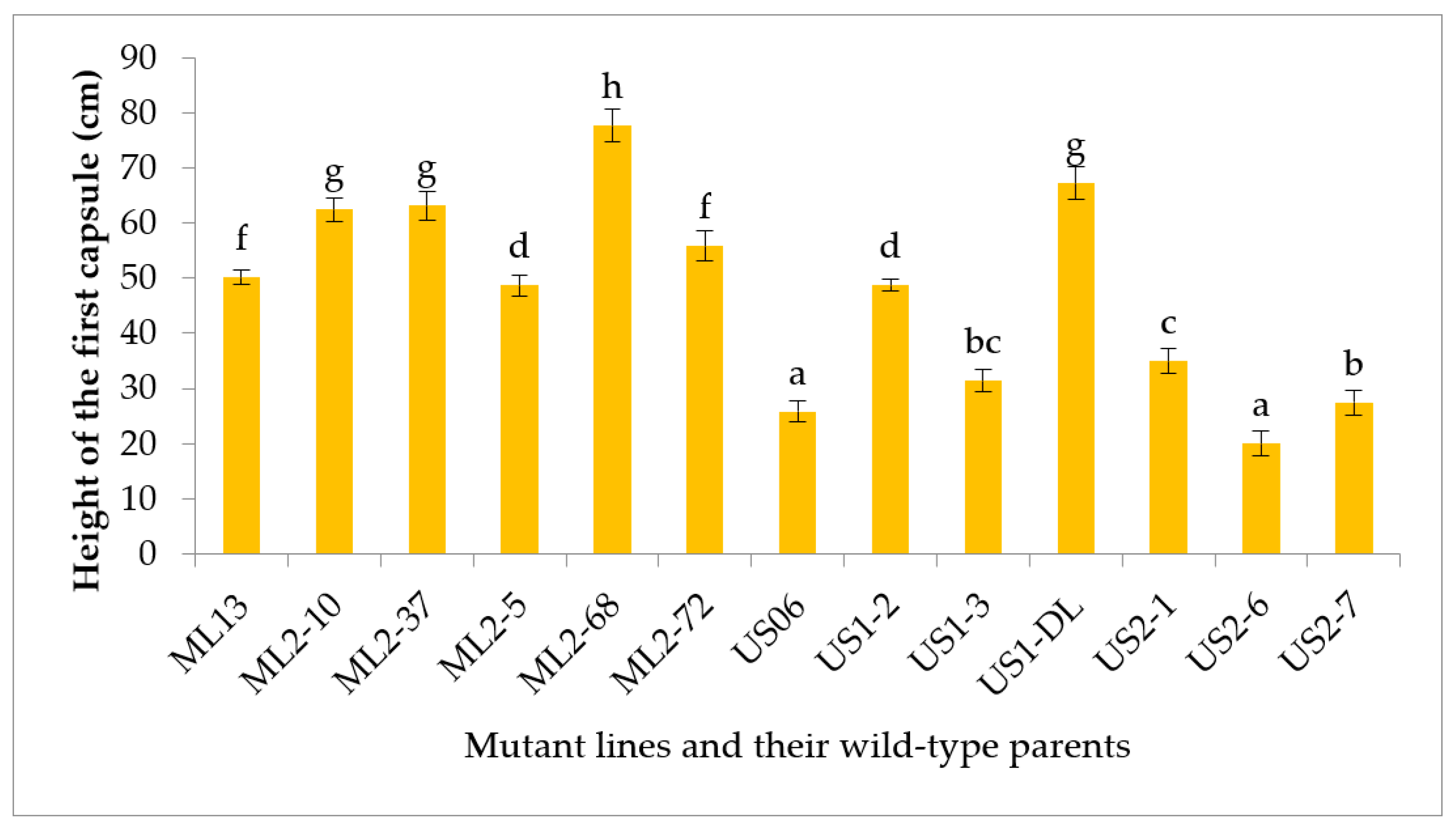
3.2.3. Height of the first Capsule
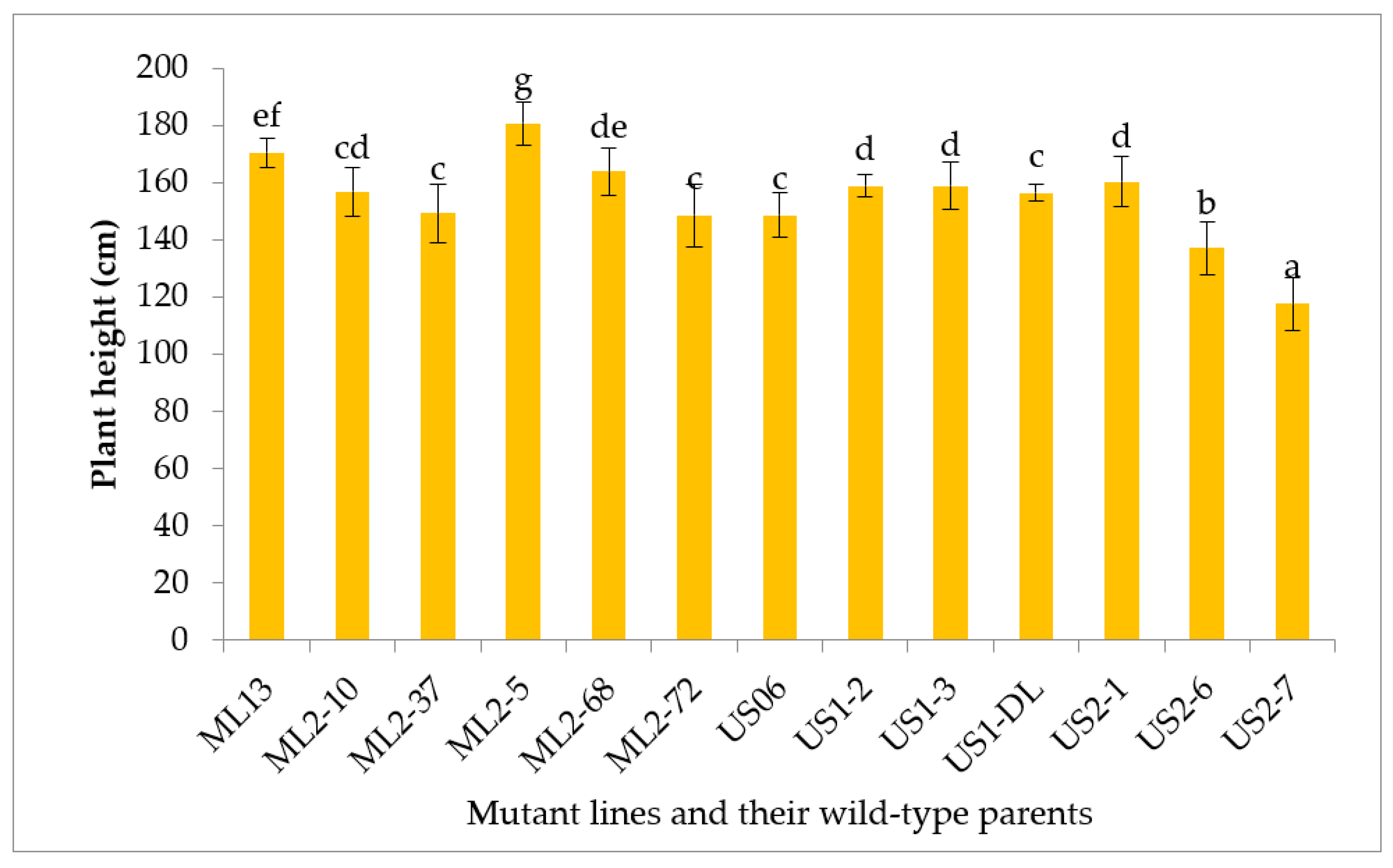
3.2.4. Plant Height
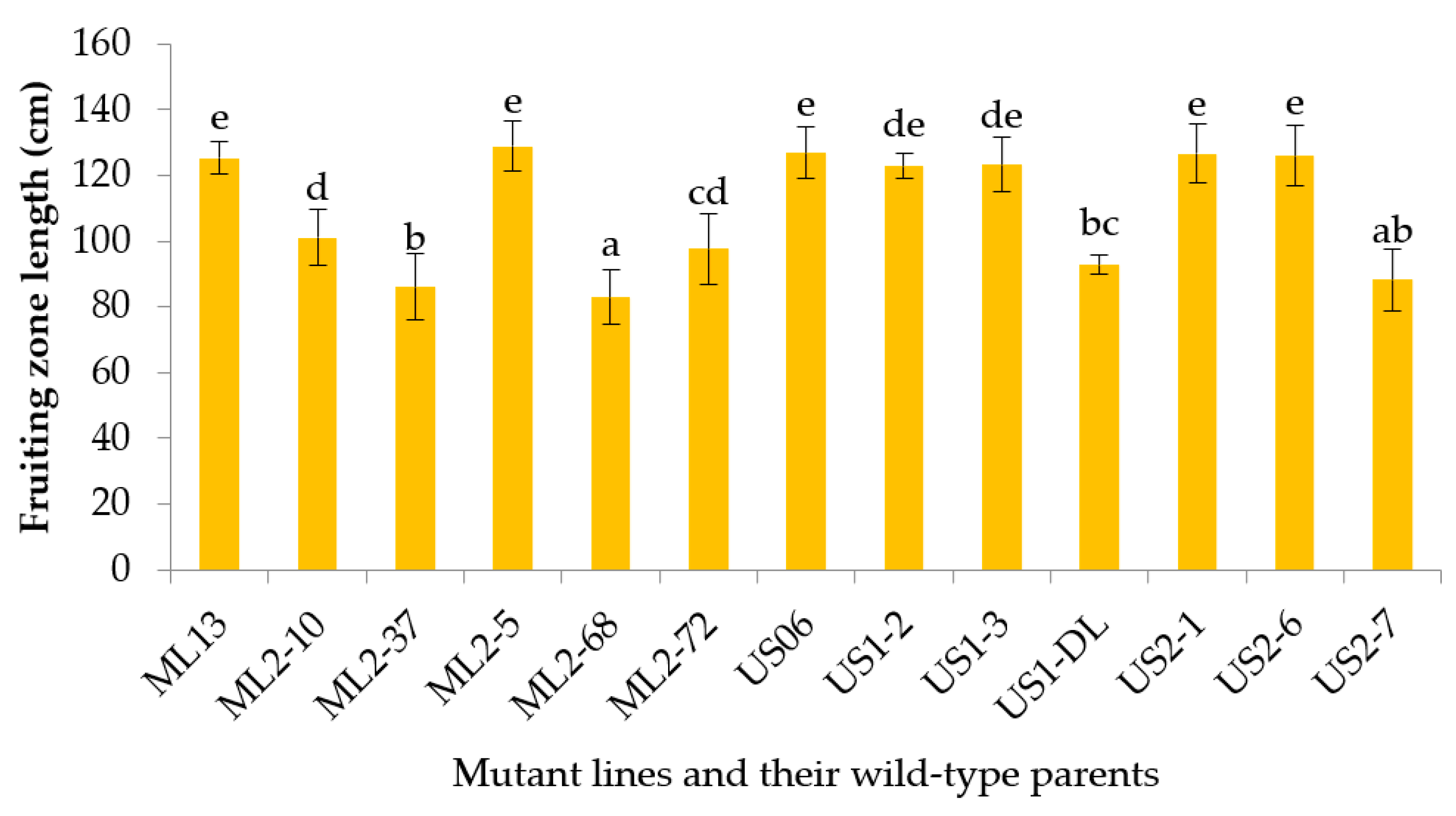
3.2.5. Fruiting Zone Length
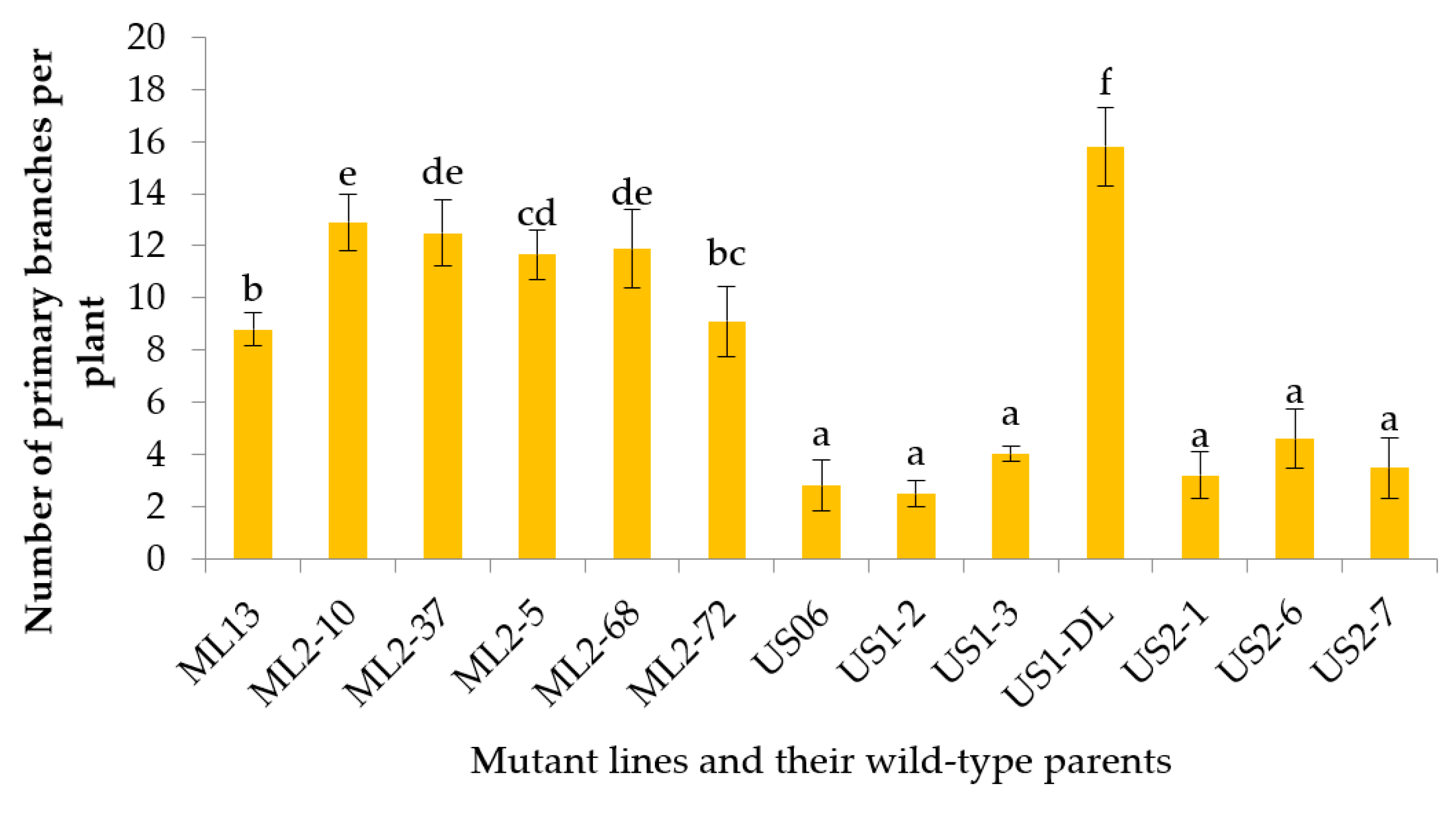
3.2.6. Number of Primary Branches per Plant
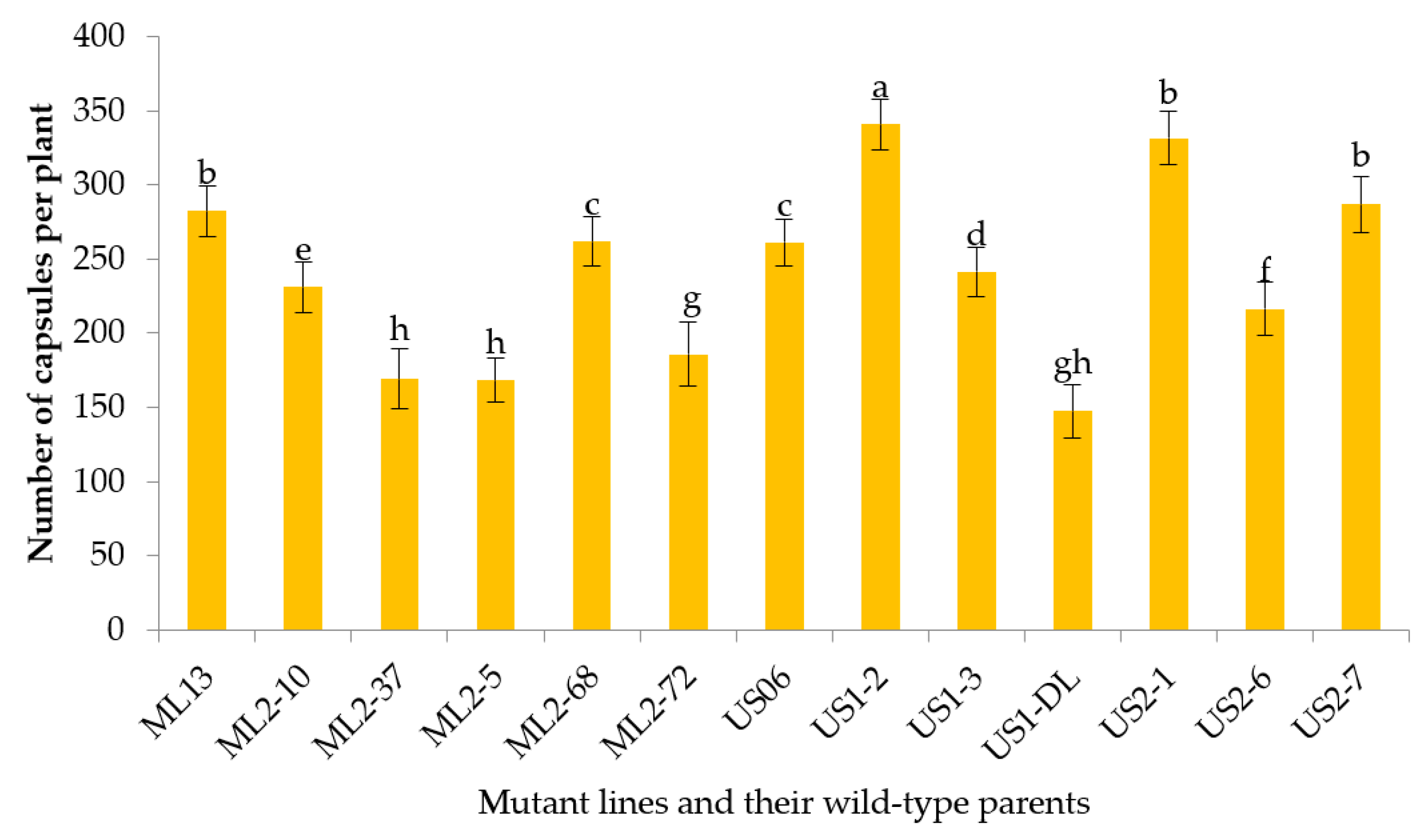
3.2.7. Number of Capsules per Plant
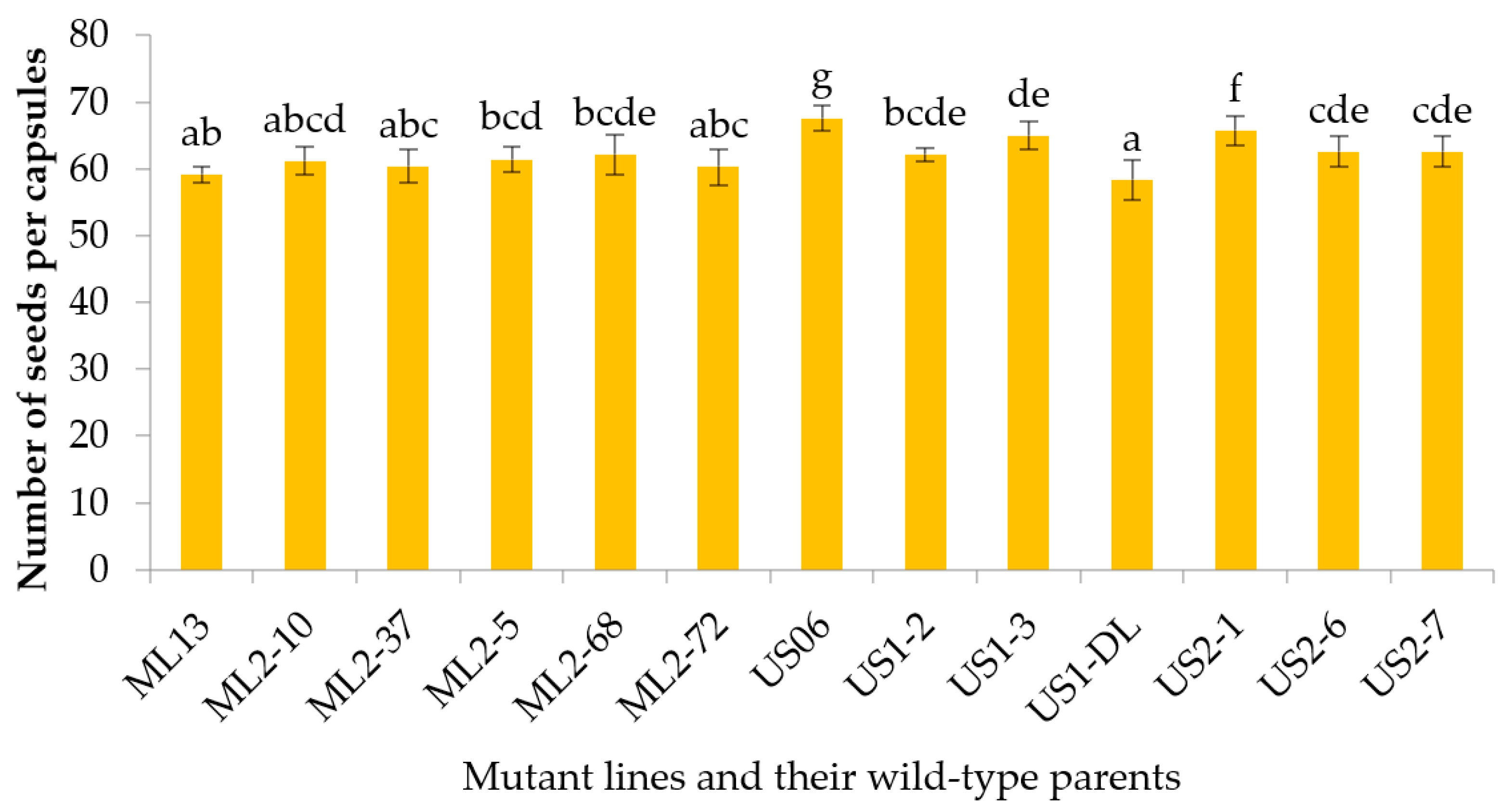
3.2.8. Number of Seeds per Capsule
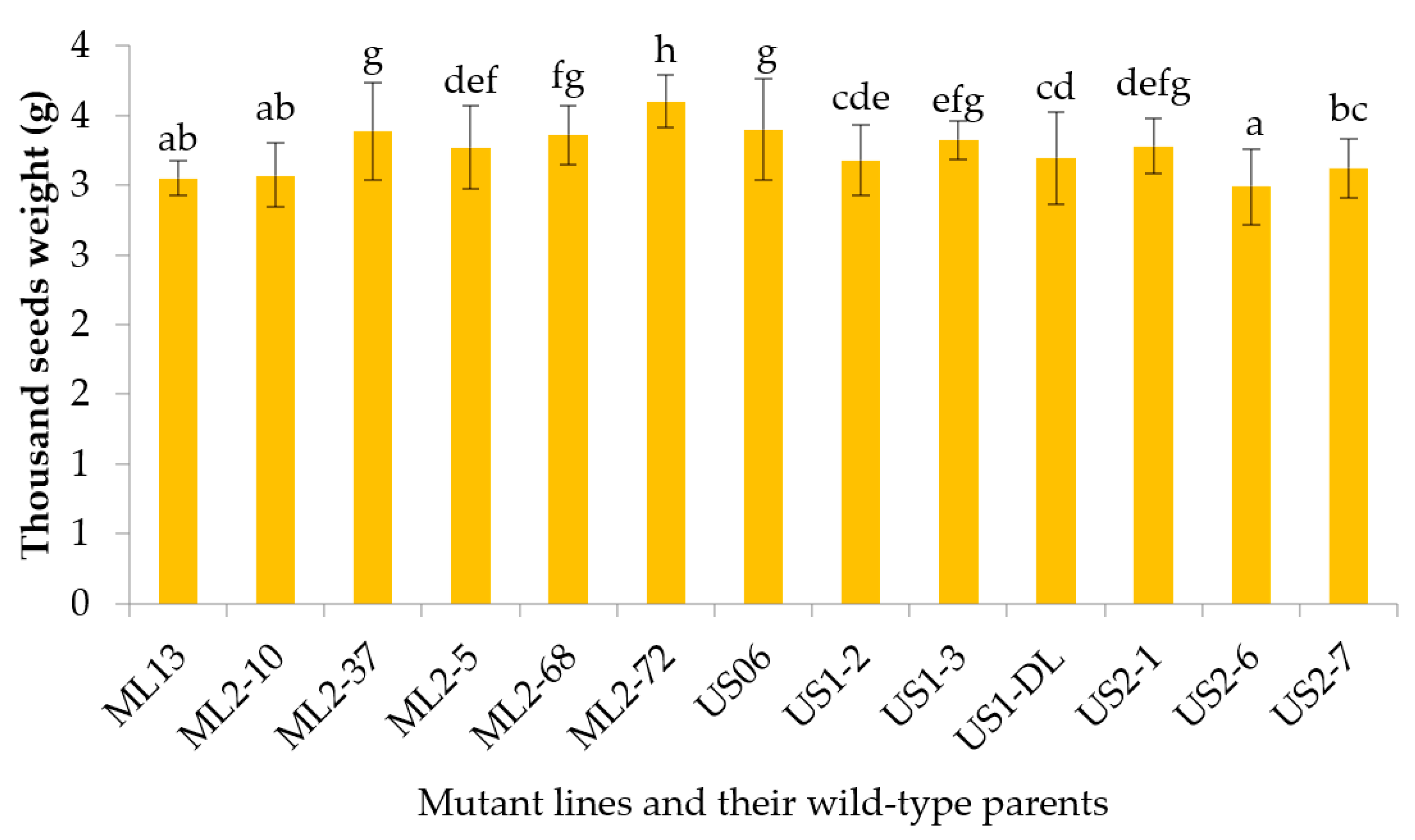
3.2.9. Thousand Seeds Weight
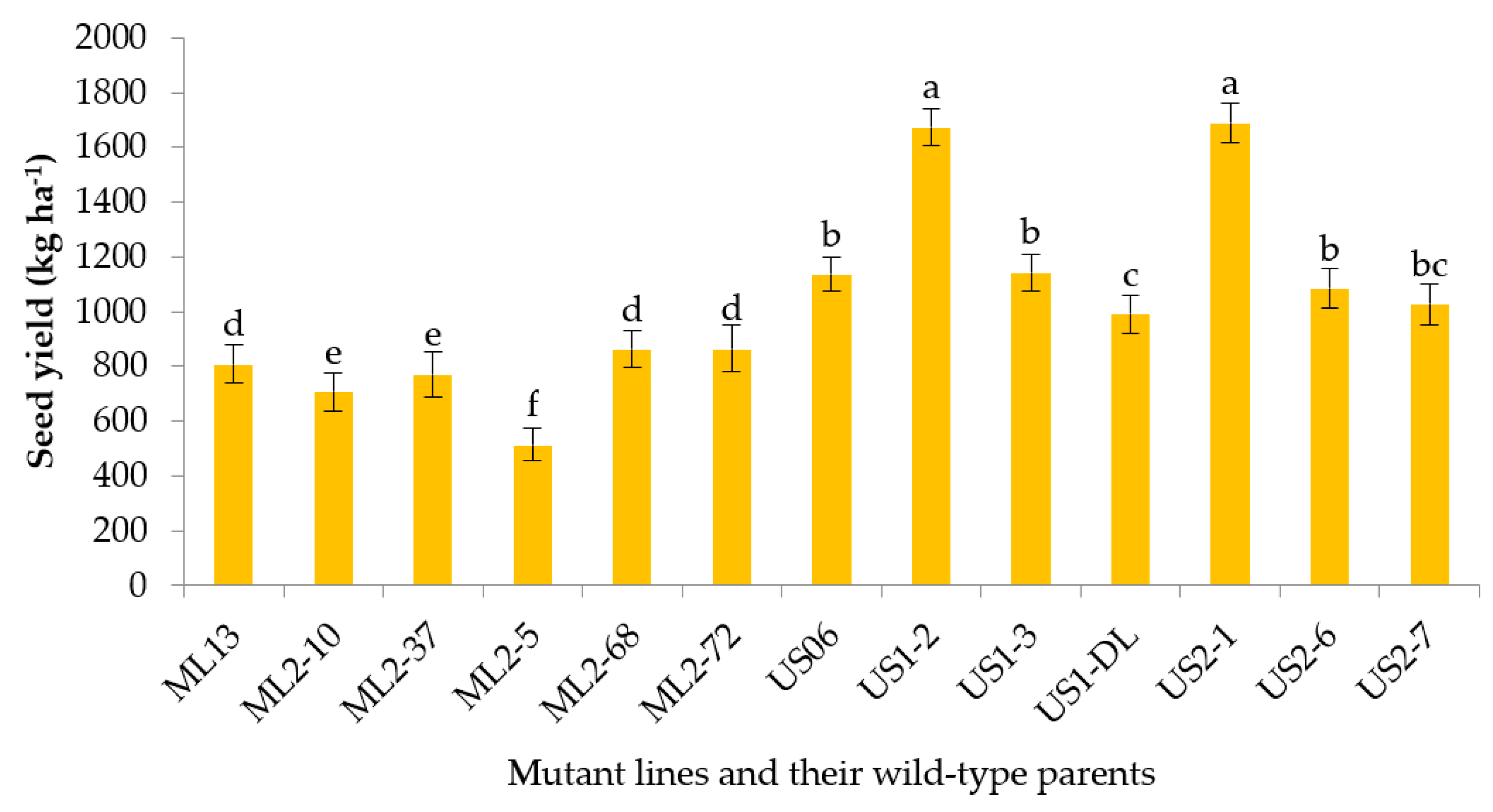
3.2.10. Seed Yield
3.3. Association among the Investigated Traits
3.4. Genetic Relatednes Based on Phenotypical Clustering
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dravie, E.E.; Kortei, N.K.; Essuman, E.K.; Tettey, C.O.; Boakye, A.A.; Hunkpe, G. Antioxidant, Phytochemical and Physicochemical Properties of Sesame Seed (Sesamum indicum L). Sci. Afr. 2020, 8, e00349. [Google Scholar] [CrossRef]
- Hama, J.R. Comparison of Fatty Acid Profile Changes between Unroasted and Roasted Brown Sesame (Sesamum indicum L.) Seeds Oil. Int. J. Food Prop. 2017, 20, 957–967. [Google Scholar] [CrossRef]
- Yaseen, G.; Ahmad, M.; Zafar, M.; Akram, A.; Sultana, S.; Ahmed, S.N.; Kilic, O. Sesame (Sesamum indicum L.). In Green Sustainable Process for Chemical and Environmental Engineering and Science; Elsevier: Amsterdam, The Netherlands, 2021; pp. 253–269. ISBN 978-0-12-821886-0. [Google Scholar]
- FAOSTAT. Available online: http://www.fao.org/faostat/fr/#data/QC (accessed on 10 May 2022).
- El Harfi, M.; Jbilou, M.; Hanine, H.; Rizki, H.; Fechtali, M.; Nabloussi, A. Genetic Diversity Assessment of Moroccan Sesame (Sesamum indicum L.) Populations Using Agro-Morphological Traits. J. Agric. Sci. Technol. A 2018, 8, 296–305. [Google Scholar]
- El Harfi, M.; Charafi, J.; Houmanat, K.; Hanine, H.; Nabloussi, A. Assessment of Genetic Diversity in Moroccan Sesame (Sesamum indicum) Using ISSR Molecular Markers. OCL 2021, 28, 3. [Google Scholar] [CrossRef]
- Kouighat, M.; Harfi, M.E.; Hanine, H.; Fechtali, M.E.; Nabloussi, A. Moroccan Sesame: Current Situation, Challenges, and Recommended Actions for Its Development. OCL 2022, 29, 27. [Google Scholar] [CrossRef]
- Venkataramana Bhat, K.; Babrekar, P.P.; Lakhanpaul, S. Study of Genetic Diversity in Indian and Exotic Sesame (Sesamum indicum L.) Germplasm Using Random Amplified Polymorphic DNA (RAPD) Markers. Euphytica 1999, 110, 21–34. [Google Scholar] [CrossRef]
- Dossa, K.; Wei, X.; Zhang, Y.; Fonceka, D.; Yang, W.; Diouf, D.; Liao, B.; Cissé, N.; Zhang, X. Analysis of Genetic Diversity and Population Structure of Sesame Accessions from Africa and Asia as Major Centers of Its Cultivation. Genes 2016, 7, 14. [Google Scholar] [CrossRef]
- Laurentin, H.E.; Karlovsky, P. Genetic Relationship and Diversity in a Sesame (Sesamum indicum L.) Germplasm Collection Using Amplified Fragment Length Polymorphism (AFLP). BMC Genet. 2006, 7, 10. [Google Scholar] [CrossRef]
- Tesfaye, T.; Tesfaye, K.; Keneni, G.; Ziyomo, C.; Alemu, T. Genetic Diversity of Sesame (Sesamum indicum L.) Using High Throughput Diversity Array Technology. J. Crop Sci. Biotechnol. 2022, 25, 1–13. [Google Scholar] [CrossRef]
- Roy, B.; Pal, A.K.; Basu, A.K. The Estimation of Genetic Variability and Genetic Divergence of Some Advance Lines of Sesame Based on Morphological Traits. Plant Sci. Today 2022, 9, 281–287. [Google Scholar] [CrossRef]
- Ashri, A. Increased Genetic Variability for Sesame Improvement by Hybridization and Induced Mutations; FAO Plant Production and Protection Papers (No. 29); Food and Agriculture Organization of the United Nations: Rome, Italy, 1981; pp. 141–145. [Google Scholar]
- Ashri, A. Modification and Adaptation of the Induced Determinate Sesame Mutant by Cross Breeding and Its Evaluation. In Mutation Breeding of Oil Seed Crops, Proceedings of a final Research Co-Ordination Meeting of an FAO/IAEA Co-Ordinated Research Programme Organized by the Joint FAO/IAEA Division of Nuclear Techniques in Food and Agriculture, Vienna, Austria, 11–15 January 1993; IAEA-TECDOC-781; International Atomic Energy Agency: Vienna, Austria, 1994; pp. 111–114. [Google Scholar]
- Begum, T.; Dasgupta, T. A Comparison of the Effects of Physical and Chemical Mutagens in Sesame (Sesamum indicum L.). Genet. Mol. Biol. 2010, 33, 761–766. [Google Scholar] [CrossRef]
- Boureima, S.; Eyletters, M.; Diouf, M.; Diop, T.A.; Van Damme, P. Sensitivity of Seed Germination and Seedling Radicle Growth to Drought Stress in Sesame Sesamum indicum L. Res. J. Environ. Sci. 2011, 5, 557. [Google Scholar] [CrossRef]
- Begum, T.; Dasgupta, T. Amelioration of Seed Yield, Oil Content and Oil Quality through Induced Mutagenesis in Sesame (Sesamum indicum L.). Bangladesh J. Bot. 2015, 44, 15–22. [Google Scholar] [CrossRef]
- Saha, A. EMS Induced Polygenic Mutation in Tilottama Cultivar of Sesamum indicum. Plant Arch. 2019, 19, 630–632. [Google Scholar]
- Gadri, Y.; Williams, L.E.; Peleg, Z. Tradeoffs between Yield Components Promote Crop Stability in Sesame. Plant Sci. 2020, 295, 110105. [Google Scholar] [CrossRef] [PubMed]
- Wacal, C.; Ogata, N.; Sasagawa, D.; Handa, T.; Basalirwa, D.; Acidri, R.; Ishigaki, T.; Yamamoto, S.; Nishihara, E. Seed Yield, Crude Protein and Mineral Nutrient Contents of Sesame during a Two-Year Continuous Cropping on Upland Field Converted from a Paddy. Field Crops Res. 2019, 240, 125–133. [Google Scholar] [CrossRef]
- Bedawy, I.M.; Mohamed, N.E. Phenotypic and Genotypic Variability in a Set of Sesame (Sesamum indicum L.) Genotypes. Egypt. J. Agron. 2018, 40, 193–205. [Google Scholar] [CrossRef]
- Patil, M.K.; Lokesha, R. Estimation of Genetic Variability, Heritability, Genetic Advance, Correlations and Path Analysis in Advanced Mutant Breeding Lines of Sesame (Sesamum indicum L.). J. Pharmacogn. Nat. Prod. 2018, 4, 151. [Google Scholar] [CrossRef]
- Begum, T.; Dasgupta, T. Induced Genetic Variability, Heritability and Genetic Advance in Sesame (Sesamum indicum L.). SABRAO J. Breed. Genet. 2014, 46, 21–33. [Google Scholar]
- Divya, K.; Rani, T.S.; Babu, T.K.; Padmaja, D. Assessment of Genetic Variability, Heritability and Genetic Gain in Advanced Mutant Breeding Lines of Sesame (Sesamum indicum L.). Int. J. Curr. Microbiol. App. Sci. 2018, 7, 1565–1574. [Google Scholar] [CrossRef]
- Laghari, J.; Mari, S.N.; Laghari, F. Screening of Sesame (Sesamum indicum L.) Heritability Estimates and Genetic Variability. EasyChair, 2022; preprint no. 7660. [Google Scholar]
- Kouighat, M.; Channaoui, S.; Labhilili, M.; El Fechtali, M.; Nabloussi, A. Novel Genetic Variability in Sesame Induced via Ethyl Methane Sulfonate. J. Crop Improv. 2021, 35, 654–665. [Google Scholar] [CrossRef]
- Langham, D.R. Growth and Development of Sesame; American Sesame Grower Association: San Antonio, TX, USA, 2008; p. 44. [Google Scholar]
- Singh, R.K.; Chaudhry, B.D. Biometrical Methods in Quantitative Genetic Analysis; Kalyani Publishers: New Dehli, India, 1985; pp. 38–54. [Google Scholar]
- Deshmukh, S.N.; Basu, M.S.; Reddy, P.S. Genetic Variability, Character Association and Path Coefficients of Quantitative Traits in Virginia Bunch Varieties of Groundnut. Indian J. Agric. Sci. 1986, 56, 816–821. [Google Scholar]
- Johnson, H.W.; Robinson, H.F.; Comstock, R.E. Estimates of Genetic and Environmental Variability in Soybeans 1. Agron. J. 1955, 47, 314–318. [Google Scholar] [CrossRef]
- Hoballah, A.A. Selection and Agronomic Evaluation of Induced Mutant Lines of Sesame. In Sesame Improvement by Induced Mutations; Technical Report IAEA-TECDOC-1195; IAEA: Vienna, Austria, 2001; pp. 71–84. [Google Scholar]
- Ali, N.; Javidfar, F.; Attary, A.A. Genetic Variability, Correlation and Path Analysis of Yield and Its Components in Winter Rapeseed (Brassica napus L.). Pak. J. Bot. 2002, 34, 145–150. [Google Scholar]
- El-Mohsen, A.; Dsoky, S.E.; Abdallah, A.M. Statistical Estimates of the Main Characteristics of Sesame Mutated Generations. J. Plant Prod. 2008, 33, 2449–2464. [Google Scholar] [CrossRef]
- Mahmud, F.; Rasul, M.G.; Rahim, M.A. Genetic Diversity Analysis in Some Advanced Lines of Brassica napus. Sci. Asia 2008, 34, 432–434. [Google Scholar] [CrossRef]
- Pavadai, P.; Girija, M.; Dhanavel, D. Effect of Gamma Rays on Some Yield Parameters and Protein Content of Soybean in M2, M3 and M4 Generation. J. Exp. Sci. 2010, 1, 8–11. [Google Scholar]
- Aliyu, R.E.; Aliyu, A.; Adamu, A.K. Inducing Phenotypic Variants in Sesame (Sesamun indicum L.) with Fast Neutron Irradiation. J. Res. For. Wildl. Environ. 2017, 9, 52–60. [Google Scholar]
- Rahman, A.; Das, M.L. Evolution of improved varieties of sesame through induced mutations. In Sesame Improvement by Induced Mutations; Technical Report IAEA-TECDOC-1195; IAEA: Vienna, Austria, 2001; pp. 157–170. [Google Scholar]
- Bhuiyan, M.S.H.; Malek, M.A.; Sarkar, M.M.A.; Islam, M.; Akram, M.W. Genetic Variance and Performance of Sesame Mutants for Yield Contributing Characters. Malays. J. Sustain. Agric. 2019, 3, 27–30. [Google Scholar] [CrossRef]
- Aristya, V.E.; Taryono, T.; Wulandari, R.A. Yield Components of Some Sesame Mutant Populations Induced by Gamma Irradiation. Bul. Tanam. Tembakau Serat Miny. Ind. 2019, 10, 64. [Google Scholar] [CrossRef]
- Zahran, H.A.; Abd-Elsaber, A.; Tawfeuk, H.Z. Genetic Diversity, Chemical Composition and Oil Characteristics of Six Sesame Genotypes. OCL 2020, 27, 39. [Google Scholar] [CrossRef]
- Akbar, F.; Rabbani, M.A.; Shinwari, Z.K.; Khan, S.J. Genetic Divergence in Sesame (Sesamum indicum L.) Landraces Based on Qualitative and Quantitative Traits. Pak. J. Bot. 2011, 43, 2737–2744. [Google Scholar]
- Van Zanten, L. Sesame Improvement by Induced Mutations: Results of the Co-ordinated Research Project and Recommendation for Future Studies. In Sesame Improvement by Induced Mutations; Technical Report IAEA-TECDOC-1195; Van Zanten, L., Ed.; IAEA: Vienna, Austria, 2001; pp. 1–12. [Google Scholar]
- Langham, D.R.; Wiemers, T. Progress in Mechanizing Sesame in the US through Breeding. In Trends in New Crops and New Uses, Proceedings of the Fifth National Symposium, Atlanta, GA, USA, 10–13 November 2001; ASHS Press: Alexandria, VA, USA, 2002; pp. 157–173. [Google Scholar]
- Imran, M.; Dash, M.; Das, T.R.; Kabi, M. Analysis of Induced Genetic Variability for Morphological and Floral Characters with Male Sterility in Sesame (Sesamum indicum L.). Electron. J. Plant Breed. 2018, 9, 801–807. [Google Scholar] [CrossRef]
- Zhang, H.; Miao, H.; Ju, M. Potential for Adaptation to Climate Change through Genomic Breeding in Sesame. In Genomic Designing of Climate-Smart Oilseed Crops; Springer: Cham, Switzerland, 2019; pp. 371–440. [Google Scholar]
- Ragab, A.I. Improving Some Economical Characters in Sesame (Sesamum indicum, L.) by Irradiation Induced Mutation and Hybridization. Ph.D. Thesis, Faculty of Agriculture, Cairo University, Giza, Egypt, 1982. [Google Scholar]
- Beech, D.F.; Imrie, B.C. Breeding for Mechanised Sesame Production in Australia. In Sesame Improvement by Induced Mutations; Technical Report IAEA-TECDOC-1195; Van Zanten, L., Ed.; IAEA: Vienna, Austria, 2001; pp. 63–70. [Google Scholar]
- Miao, H.; Zhang, H.; Kole, C. Compendium of Plant Genomes; Kole, C., Ed.; Springer International Publishing: Cham, Switzerland, 2021; ISBN 978-3-319-98097-3. [Google Scholar]
- Baydar, H. Breeding for the Improvement of the Ideal Plant Type of Sesame. Plant Breed. 2005, 124, 263–267. [Google Scholar] [CrossRef]
- Ahmad, M.; Khan, M.A.; Zafar, M.; Sultana, S. Environment-Friendly Renewable Energy from Sesame Biodiesel. Energy Sources Part Recovery Util. Environ. Eff. 2009, 32, 189–196. [Google Scholar] [CrossRef]


| Lines | Origin | Characteristics |
|---|---|---|
| ‘ML13′ | Local cultivar from Morocco | Parent (check cultivar), beige seeds |
| ‘ML2-10′ | Developed by EMS-mutagenesis from ML13 | Mutant, brown seeds, high branching, small capsules |
| ‘ML2-37′ | Mutant, brown seeds, high branching, low number of capsules per plant | |
| ‘ML2-5′ | Mutant, brown seeds, tall plant | |
| ‘ML2-68′ | Mutant, grey seeds, late maturity | |
| ‘ML2-72′ | Mutant, brown seeds, late flowering | |
| ‘US06′ | Accession from Mexico | Parent (check cultivar), white seeds |
| ‘US1-2′ | Developed by mutagenesis from US06 | Mutant, white seeds, high number of capsules per plant |
| ‘US1-3′ | Mutant, white seeds, early maturity | |
| ‘US1-DL’ | Mutant, white seeds, late flowering, high branching | |
| ‘US2-1′ | Mutant, white seeds, early maturity, high number of capsules per plant | |
| ‘US2-6′ | Mutant, black seeds, early flowering and maturity | |
| ‘US2-7′ | Mutant, white seeds, low branching |
| Afourare (E2) | Taoujdate (E1) | |||||||
|---|---|---|---|---|---|---|---|---|
| Temperature [°C] | Precipitation [mm] | Temperature [°C] | Precipitation [mm] | |||||
| Date | Min | Max | Mean | Sum | Min | Max | Mean | Sum |
| 2021-04 | 04.64 | 25.89 | 19.80 | 55.80 | 06.64 | 22.2 | 16.19 | 72.20 |
| 2021-05 | 03.65 | 35.64 | 28.75 | 36.80 | 05.93 | 23.6 | 20.32 | 23.60 |
| 2021-06 | 07.40 | 36.28 | 30.50 | 02.40 | 10.42 | 31.21 | 26.09 | 1 |
| 2021-07 | 11.23 | 45.44 | 35.73 | 0 | 13.37 | 42.97 | 33.27 | 0 |
| 2021-08 | 09.73 | 46.00 | 44.20 | 0 | 12.18 | 43.20 | 41.65 | 0.20 |
| 2021-09 | 11.04 | 42.41 | 41.36 | 09.60 | 13.91 | 31.6 | 29.74 | 01.60 |
| 2021-10 | 10.12 | 37.37 | 31.12 | 03.50 | 13.42 | 28.14 | 22.89 | 04.80 |
| Source of Variation | DF | Number of Days to Flowering (d) | Number of Days to Maturity (d) | Height of the First Capsule (cm) | Plant Height (cm) | Fruiting Zone Length (cm) | Number of Branches per Plant | Number of Capsules per Plant | Number of Seeds per Capsule | 1000-Seed Weight (g) | Seed Yield (kg·ha−1) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Afourare (E1) | |||||||||||
| Block | 2 | 30.11 * | 18.02 | 14.64 | 40.38 * | 20.84 | 0.10 | 69.46 * | 4.38 | 0.02 | 9.25 |
| Genotype | 12 | 2609.70 *** | 1854.62 *** | 2656.01 *** | 2282.14 *** | 2317.41 *** | 228.64 *** | 62,916.86 *** | 189.58 *** | 0.21 *** | 1998.83 *** |
| Error | 24 | 29.86 | 30.41 | 39.32 | 31.41 | 41.32 | 2.08 | 234.77 | 8.10 | 0.01 | 19.42 |
| Taoujdate (E2) | |||||||||||
| Block | 2 | 39.93 * | 14.18 * | 11.47 | 61.34 * | 23.27 | 1.45 | 67.41 * | 5.671 | 0.03 | 11.13 * |
| Genotype | 12 | 191.70 *** | 1162.15 *** | 2952.30 *** | 5318.92 *** | 7854.32 *** | 146.87 *** | 24,026.92 *** | 258.41 *** | 0.29 *** | 853.95 *** |
| Error | 24 | 23.56 | 100.63 | 47.09 | 18.28 | 77.53 | 5.40 | 349.30 | 8.42 | 0.09 | 33.62 |
| Combined | |||||||||||
| Environment (E) | 1 | 490.43 *** | 10,674.17 *** | 347.343 *** | 2829.14 *** | 20,129.17 ** | 74.818 *** | 49,025.88 *** | 64.77 *** | 0.09 | 20.12 *** |
| Block | 4 | 35.02 | 16.10 | 13.05 | 50.86 | 22.05 | 0.77 | 68.43 | 5.026 | 0.025 | 19.19 |
| Genotype (G) | 12 | 267.5 *** | 1463.3 *** | 6064.5 *** | 2312.8 *** | 3175.6 *** | 197.7 *** | 24,878.6 *** | 233.4 *** | 0.282 *** | 1330.6 *** |
| G × E | 12 | 48.3 * | 422.3 *** | 2063.60 *** | 2063.60 *** | 2043.10 *** | 30.90 *** | 17,198.10 *** | 11.90 | 0.062 *** | 18.8 *** |
| Error | 48 | 26.71 | 65.52 | 43.205 | 24.845 | 59.425 | 3.74 | 292.035 | 8.26 | 0.05 | 26.52 |
| Source of Variation | DF | Number of Days to Flowering (d) | Number of Days to Maturity (d) | Height of the First Capsule (cm) | Plant Height (cm) | Fruiting Zone Length (cm) | Number of Branches per Plant | Number of Capsules per Plant | Number of Seeds per Capsule | 1000-Seed Weight (g) | Seed Yield (kg·ha−1) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Mean square | |||||||||||
| Genotype | 12 | 2609.70 *** | 1854.62 *** | 2656.01 *** | 2282.14 *** | 2317.41 *** | 228.64 *** | 62,916.86 *** | 189.58 *** | 0.21 *** | 198.83 *** |
| Contrast value | |||||||||||
| ML13 parent vs. ML mutants | 1 | 0.90 | 10.30 * | 46.02 ** | 23.20 ** | 77.70 ** | 10.54 ** | 302.26 *** | 76.80 ** | 1.50 *** | 1.23 * |
| US06 parent vs. US mutants | 1 | 27.10 * | 34.40 * | 33.30 ** | 9.22 * | 27.70 * | 18.40 *** | 130.40 ** | 24.90 * | 0.95 *** | 2.40 ** |
| US genotypes vs. ML genotypes | 1 | 73.80 ** | 118.60 *** | 161.02 *** | 17.47 ** | 125.70 *** | 41.45 *** | 306.66 *** | 34.10 * | 0.59 *** | 3.43 *** |
| Component of Variance | Genetic Variability | Genetic Advance | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Characters | Range | Mean | Vg | Vp | Ve | PCV | GCV | H2 b.s% | GA | GAM% |
| NDF | 55.8–72.5 | 65.46 | 40.13 | 66.84 | 26.71 | 12.50 | 9.68 | 60.04 | 13.05 | 19.95 |
| NDM | 129.08–166.63 | 145.81 | 232.97 | 298.49 | 65.52 | 11.85 | 10.47 | 78.05 | 31.44 | 21.56 |
| HFC | 19.96–77.70 | 47.21 | 1003.56 | 1046.76 | 43.21 | 68.53 | 67.10 | 95.87 | 65.26 | 138.23 |
| PH | 117.6–180.67 | 154.43 | 381.34 | 406.18 | 24.85 | 13.05 | 12.65 | 93.88 | 40.23 | 26.05 |
| FZL | 82.92–128.81 | 110.05 | 519.37 | 578.80 | 59.43 | 21.70 | 20.55 | 89.73 | 46.95 | 42.34 |
| NBP | 2.5–15.8 | 7.94 | 32.34 | 36.08 | 3.74 | 74.62 | 70.65 | 89.63 | 11.72 | 145.53 |
| NCP | 169.10–341 | 248.08 | 4097.77 | 4389.80 | 292.04 | 27.20 | 26.28 | 93.35 | 131.87 | 54.13 |
| NSC | 58.30–67.60 | 62.96 | 37.53 | 45.79 | 8.26 | 10.72 | 9.70 | 81.96 | 12.62 | 19.99 |
| TSW | 2.99–3.60 | 3.25 | 0.04 | 0.09 | 0.05 | 9.19 | 6.07 | 43.61 | 0.41 | 12.50 |
| Yield | 516–1691 | 1052 | 217.35 | 243.87 | 26.52 | 196.19 | 185.21 | 89.13 | 30.37 | 381.54 |
| NDF | NDM | HFC | PH | FZL | NBP | NCP | NSC | TSW | Seed Yield | |
|---|---|---|---|---|---|---|---|---|---|---|
| NDF | 1 | |||||||||
| NDM | 0.786 *** | 1 | ||||||||
| HFC | 0.861 *** | 0.757 *** | 1 | |||||||
| PH | 0.328 | 0.380 | 0.451 * | 1 | ||||||
| FZL | −0.592 ** | −0.470 * | −0.624 *** | 0.371 | 1 | |||||
| NBP | 0.744 *** | 0.627 *** | 0.834 *** | 0.377 | −0.552 ** | 1 | ||||
| NCP | −0.342 | −0.282 | −0.346 | −0.084 | 0.333 | 0.728 *** | 1 | |||
| NSC | −0.614 *** | −0.568 ** | −0.656 *** | −0.165 | 0.459 * | −0.743*** | 0.495 ** | 1 | ||
| TSW | 0.348 | 0.304 | 0.243 | 0.115 | −0.238 | 0.068 | −0.239 | 0.195 | 1 | |
| Seed yield | −0.410 * | −0.628 *** | −0.410 * | 0.178 | 0.353 | −0.316 | 0.892 *** | 0.508 ** | 0.048 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kouighat, M.; Hanine, H.; Chetto, O.; Fakhour, S.; El Fechtali, M.; Nabloussi, A. Assessment of Novel Genetic Diversity Induced by Mutagenesis and Estimation of Genetic Parameters in Sesame M4 Mutant Lines. Int. J. Plant Biol. 2022, 13, 644-662. https://doi.org/10.3390/ijpb13040052
Kouighat M, Hanine H, Chetto O, Fakhour S, El Fechtali M, Nabloussi A. Assessment of Novel Genetic Diversity Induced by Mutagenesis and Estimation of Genetic Parameters in Sesame M4 Mutant Lines. International Journal of Plant Biology. 2022; 13(4):644-662. https://doi.org/10.3390/ijpb13040052
Chicago/Turabian StyleKouighat, Mohamed, Hafida Hanine, Oumaima Chetto, Samir Fakhour, Mohamed El Fechtali, and Abdelghani Nabloussi. 2022. "Assessment of Novel Genetic Diversity Induced by Mutagenesis and Estimation of Genetic Parameters in Sesame M4 Mutant Lines" International Journal of Plant Biology 13, no. 4: 644-662. https://doi.org/10.3390/ijpb13040052
APA StyleKouighat, M., Hanine, H., Chetto, O., Fakhour, S., El Fechtali, M., & Nabloussi, A. (2022). Assessment of Novel Genetic Diversity Induced by Mutagenesis and Estimation of Genetic Parameters in Sesame M4 Mutant Lines. International Journal of Plant Biology, 13(4), 644-662. https://doi.org/10.3390/ijpb13040052






