The Multi-Ethnic New Zealand Study of Acute Coronary Syndromes (MENZACS): Design and Methodology
Abstract
1. Introduction
2. Methods
2.1. Study Design and Participants
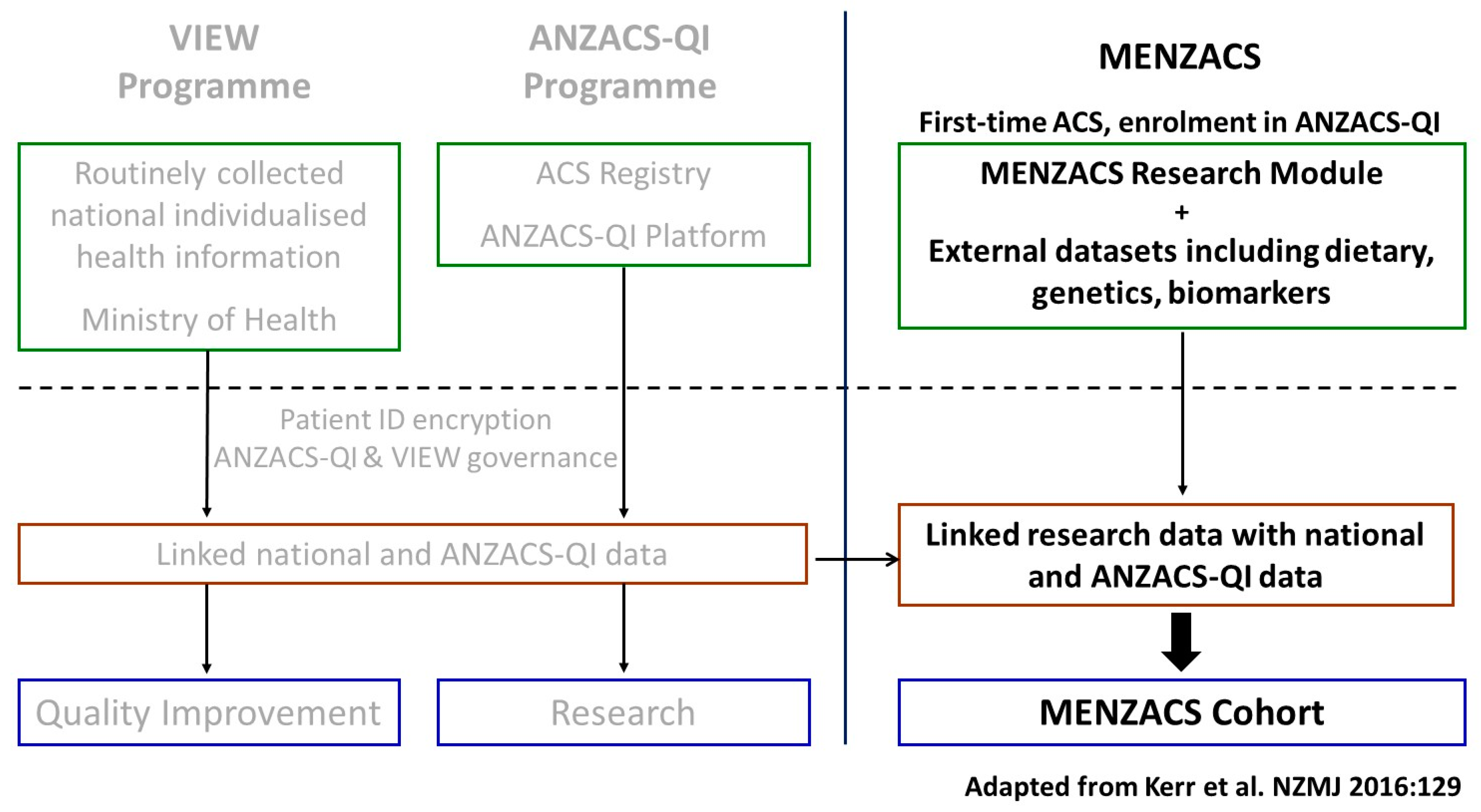
2.2. Study Infrastructure
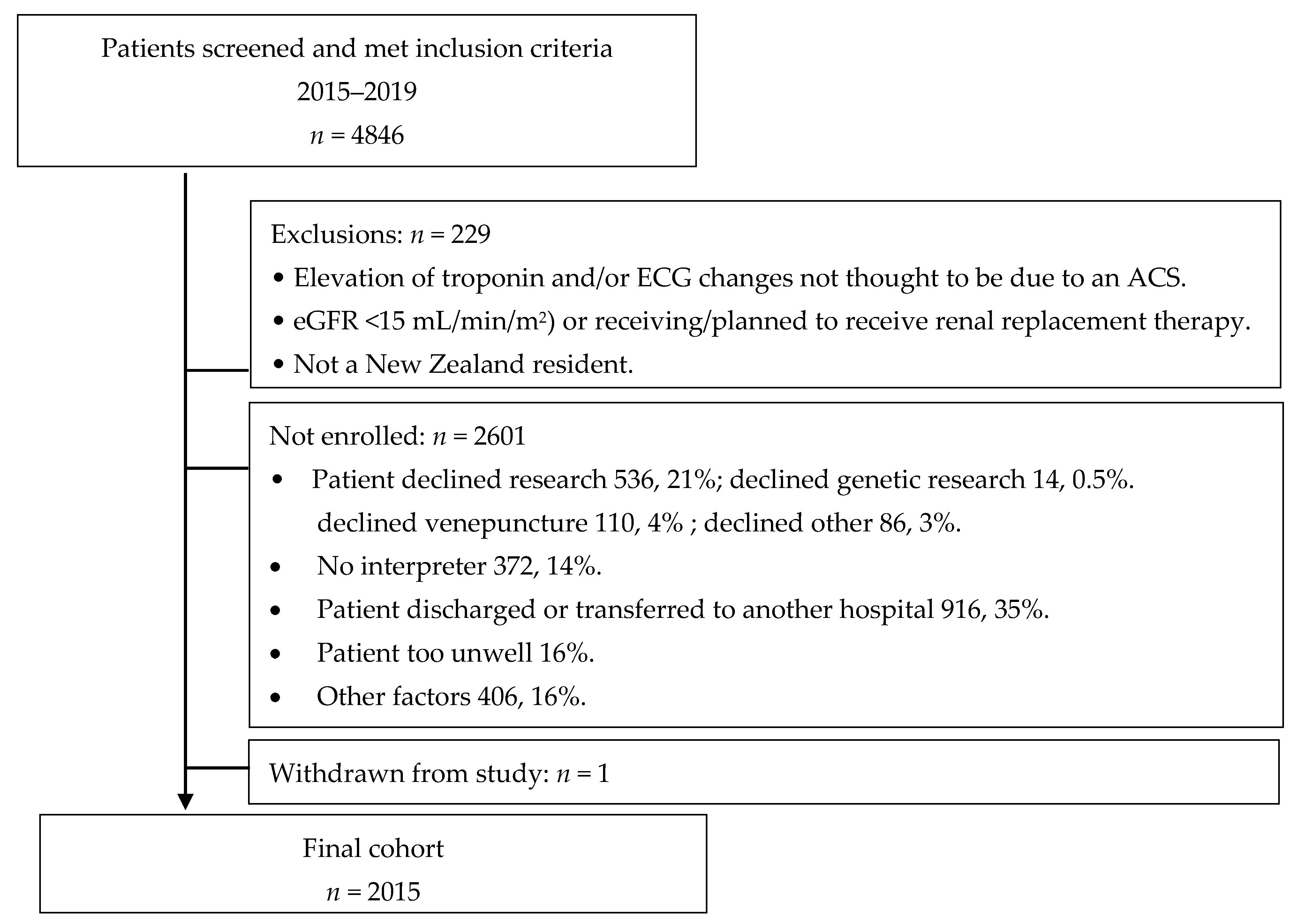
2.3. MENZACS Inclusion Criteria and Recruitment Pathway
2.4. MENZACS Research Data
2.5. Biological Samples
2.6. Biochemical and Genetic Analyses
2.7. Clinical Outcome Variables
2.8. Key Research Questions
- What dietary, lifestyle, and socio-economic factors are associated with first time ACS and subsequent outcomes;
- Can risk stratification be refined using clinical, biomarker, genetic, and epigenetic factors to build on existing secondary risk equations in New Zealand [27];
- Association studies of genetic variants with first time ACS and subsequent outcomes;
- The interaction of genomics, environmental factors, and biomarkers associated with certain phenotypes (e.g., diabetics, metabolic syndrome, hypertension, and obesity);
- Pharmacogenetic variability and the frequency of certain known variants across a diverse New Zealand population;
- Ethnic variation in genomic and genetic “signatures” related to cardiovascular risk factors.
2.9. Statistical Approach
3. Study Progress
3.1. Baseline Characteristics
3.2. Quality Assessment
3.2.1. Data Linkage
3.2.2. DNA Quality and Quantification
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ministry of Health. Mortality 2016 Data Tables. Annual Update of Key Results 2016. 2020. Available online: https://www.health.govt.nz/publication/mortality-2016-data-tables (accessed on 19 April 2020).
- Grey, C.; Jackson, R.; Schmidt, M.; Ezzati, M.; Asaria, P.; Exeter, D.J.; Kerr, A.J. One in four major ischaemic heart disease events are fatal and 60% are pre-hospital deaths: A national data-linkage study (ANZACS-QI 8). Eur. Heart J. 2017, 38, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Grey, C.; Jackson, R.; Wells, S.; Wu, B.; Poppe, K.; Harwood, M.; Sundborn, G.; Kerr, A.J. Trends in ischaemic heart disease: Patterns of hospitalisation and mortality rates differ by ethnicity (ANZACS-QI 21). N. Z. Med. J. 2018, 131, 21–31. [Google Scholar]
- Earle, N.; Poppe, K.; Doughty, R.; Rolleston, A.; Kerr, A.; Legget, M. Clinical Characteristics and Burden of Risk Factors Among Patients With Early Onset Acute Coronary Syndromes: The ANZACS-QI New Zealand National Cohort (ANZACS-QI 17). Heart Lung Circ. 2017, 27, 568–575. [Google Scholar] [CrossRef] [PubMed]
- Devlin, G. Mind the Gap: ANZACSQI and Inequality in New Zealand. Heart Lung Circ. 2016, 25, 768. [Google Scholar] [CrossRef] [PubMed]
- Grey, C.; Jackson, R.; Wells, S.; Marshall, R.; Mehta, S.; Kerr, A.J. Ethnic differences in case fatality following an acute ischaemic heart disease event in New Zealand: ANZACS-QI 13. Eur. J. Prev. Cardiol. 2016, 23, 1823–1830. [Google Scholar] [CrossRef] [PubMed]
- Grey, C.; Jackson, R.; Wells, S.; Randall, D.; Harwood, M.; Mehta, S.; Exeter, D.J.; Kerr, A.J. Ethnic Differences in Coronary Revascularisation following an Acute Coronary Syndrome in New Zealand: A National Data-linkage Study (ANZACS-QI 12). Heart Lung Circ. 2016, 25, 820–828. [Google Scholar] [CrossRef]
- Earle, N.J.; Kerr, A.J.; Legget, M.; Wu, B.P.; Doughty, R.N.; Poppe, K.K. Acute coronary syndrome registry enrolment status: Differences in patient characteristics and outcomes and implications for registry data use (ANZACS-QI 36). Eur. Heart J. Qual. Care Clin. Outcomes 2019, 2019, 6. [Google Scholar] [CrossRef]
- Pylypchuk, R.; Wells, S.; Kerr, A.; Poppe, K.; Riddell, T.; Harwood, M.; Exeter, D.; Mehta, S.; Grey, C.; Wu, B.P.; et al. Cardiovascular disease risk prediction equations in 400,000 primary care patients in New Zealand: A derivation and validation study. Lancet 2018, 391, 1897–1907. [Google Scholar] [CrossRef]
- Shah, S.H.; Newgard, C.B. Integrated metabolomics and genomics: Systems approaches to biomarkers and mechanisms of cardiovascular disease. Circ. Cardiovasc. Genet. 2015, 8, 410–419. [Google Scholar] [CrossRef]
- Martin, A.R.; Kanai, M.; Kamatani, Y.; Okada, Y.; Neale, B.M.; Daly, M.J. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 2019, 51, 584–591. [Google Scholar] [CrossRef]
- Rosenberg, N.A.; Huang, L.; Jewett, E.M.; Szpiech, Z.A.; Jankovic, I.; Boehnke, M. Genome-wide association studies in diverse populations. Nat. Rev. Genet. 2010, 11, 356–366. [Google Scholar] [CrossRef]
- Wu, Y.; Waite, L.L.; Jackson, A.U.; Sheu, W.H.-H.; Buyske, S.; Absher, D.; Arnett, D.K.; Boerwinkle, E.; Bonnycastle, L.L.; Carty, C.L.; et al. Trans-ethnic fine-mapping of lipid loci identifies population-specific signals and allelic heterogeneity that increases the trait variance explained. PLoS Genet. 2013, 9, e1003379. [Google Scholar] [CrossRef]
- Loos, R.J.F. CREBRF variant increases obesity risk and protects against diabetes in Samoans. Nat. Genet. 2016, 48, 976–978. [Google Scholar] [CrossRef] [PubMed]
- Minster, R.L.; Hawley, N.L.; Su, C.-T.; Sun, G.; Kershaw, E.; Cheng, H.; Buhule, O.D.; Lin, J.; Reupena, M.S.; Viali, S.; et al. A thrifty variant in CREBRF strongly influences body mass index in Samoans. Nat. Genet. 2016, 48, 1049–1054. [Google Scholar] [CrossRef] [PubMed]
- Gibson, C.M.; Yuet, W.C. Racial and Ethnic Differences in Response to Anticoagulation: A Review of the Literature. J. Pharm. Pract. 2019, 897190019894142. [Google Scholar] [CrossRef]
- Kerr, A.J.; Lee, M.; Jiang, Y.; Grey, C.; Wells, S.; Williams, M.; Jackson, R.; Poppe, K. High level of capture of coronary intervention and associated acute coronary syndromes in the all New Zealand acute coronary syndrome quality improvement cardiac registry and excellent agreement with national administrative datasets (ANZACS-QI 25). N. Z. Med. J. 2019, 132, 19–29. [Google Scholar]
- Kerr, A.; Williams, M.; Harding, S.; Doughty, R.; Nunn, C.; Devlin, G.; Grey, C.; Lee, M.; Flynn, C.; Rhodes, M.; et al. The All New Zealand Acute Coronary Syndrome Quality Improvement Programme: Implementation, Methodology and Cohorts (ANZACS-QI 9). N. Z. Med. J. 2016, 129, 23–36. [Google Scholar] [PubMed]
- Hudson, M.; Southey, K.; Uerata, L.; Beaton, A.; Milne, M.; Russell, K.; Smith, B.; Wilcoxet, P.; Toki, V.; Cheung, M.; et al. Key informant views on biobanking and genomic research with Maori. N. Z. Med. J. 2016, 129, 29–42. [Google Scholar]
- Thygesen, K.; Alpert, J.S.; Jaffe, A.S.; Chaitman, B.R.; Bax, J.J.; Morrow, D.A.; White, H.D.; The Executive Group on behalf of the Joint European Society of Cardiology (ESC)/American College of Cardiology (ACC)/American Heart Association (AHA)/World Heart Federation (WHF) Task Force for the Universal Definition of Myocardial Infarction Fourth Universal Definition of Myocardial Infarction. Fourth Universal Definition of Myocardial Infarction (2018). Circulation 2018, 138, e618–e651. [Google Scholar] [CrossRef]
- Anderson, J.L.; Adams, C.D.; Antman, E.M.; Bridges, C.R.; Califf, R.M.; Casey, D.E.; Chavey, W.E.; Fesmire, F.M.; Hochman, J.S.; Levin, T.N.; et al. ACC/AHA 2007 guidelines for the management of patients with unstable angina/non-ST-Elevation myocardial infarction: A report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines (Writing Committee to Revise the 2002 Guidelines for the Management of Patients with Unstable Angina/Non-ST-Elevation Myocardial Infarction) developed in collaboration with the American College of Emergency Physicians, the Society for Cardiovascular Angiography and Interventions, and the Society of Thoracic Surgeons endorsed by the American Association of Cardiovascular and Pulmonary Rehabilitation and the Society for Academic Emergency Medicine. J. Am. Coll. Cardiol. 2007, 50, e1–e157. [Google Scholar]
- Armstrong, T.; Bull, F. Development of the World Health Organization Global Physical Activity Questionnaire (GPAQ). J. Public Health 2006, 14, 66–70. [Google Scholar] [CrossRef]
- Stewart, R.A.H.; Wallentin, L.; Benatar, J.; Danchin, N.; Hagström, E.; Held, C.; Husted, S.; Lonn, E.; Stebbins, A.; Chiswell, K.; et al. Dietary patterns and the risk of major adverse cardiovascular events in a global study of high-risk patients with stable coronary heart disease. Eur. Heart J. 2016, 37, 1993–2001. [Google Scholar] [CrossRef] [PubMed]
- Mehta, S.; Jackson, R.; Exeter, D.; Wu, B.; Wells, S.; Kerr, A. Data Resource: Vascular Risk in Adult New Zealanders (VARIANZ) datasets. Int. J. Popul. Data Sci. 2019, 4, 1–23. [Google Scholar] [CrossRef]
- Danesh, J.; Saracci, R.; Berglund, G.; Feskens, E.; Overvad, K.; Panico, S.; Thompson, S.; Fournier, A.; Clavel-Chapelon, F.; Canonico, M.; et al. EPIC-Heart: The cardiovascular component of a prospective study of nutritional, lifestyle and biological factors in 520,000 middle-aged participants from 10 European countries. Eur. J. Epidemiol. 2007, 22, 129–141. [Google Scholar] [CrossRef]
- Sam, C.H.; Skeaff, S.; Skidmore, P.M. A comprehensive FFQ developed for use in New Zealand adults: Reliability and validity for nutrient intakes. Public Health Nutr. 2014, 17, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Poppe, K.K.; Doughty, R.N.; Wells, S.; Gentles, D.; Hemingway, H.; Jackson, R.; Kerr, A.J. Developing and validating a cardiovascular risk score for patients in the community with prior cardiovascular disease. Heart 2017, 103, 891–892. [Google Scholar] [CrossRef]
- Poppe, K.K.; Doughty, R.N.; Wells, S.; Wu, B.; Earle, N.J.; Richards, A.M.; Troughton, R.; Jackson, R.; Kerr, A.J. Development and validation of a cardiovascular risk score for patients in the community after acute coronary syndrome. Heart 2020, 106, 506–511. [Google Scholar] [CrossRef] [PubMed]
- Khera, A.V.; Kathiresan, S. Genetics of coronary artery disease: Discovery, biology and clinical translation. Nat. Rev. Genet. 2017, 18, 331–344. [Google Scholar] [CrossRef] [PubMed]
- Khera, A.V.; Chaffin, M.; Aragam, K.G.; Haas, M.E.; Roselli, C.; Choi, S.H.; Natarajan, P.; Lander, E.S.; Lubitz, S.A.; Ellinor, P.T.; et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 2018, 50, 1219–1224. [Google Scholar] [CrossRef] [PubMed]
- Mega, J.L.; Stitziel, N.O.; Smith, J.G.; Chasman, D.I.; Caulfield, M.J.; Devlin, J.J.; Nordio, F.; Hyde, C.L.; Cannon, C.P.; Sacks, F.M.; et al. Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: An analysis of primary and secondary prevention trials. Lancet 2015, 385, 2264–2271. [Google Scholar] [CrossRef]
- Vaara, S.; Tikkanen, E.; Parkkonen, O.; Lokki, M.-L.; Ripatti, S.; Perola, M.; Nieminen, M.S.; Sinisalo, J. Genetic Risk Scores Predict Recurrence of Acute Coronary Syndrome. Circ. Cardiovasc. Genet. 2016, 9, 172–178. [Google Scholar] [CrossRef]
- Aragam, K.G.; Natarajan, P. Polygenic Scores to Assess Atherosclerotic Cardiovascular Disease Risk: Clinical Perspectives and Basic Implications. Circ. Res. 2020, 126, 1159–1177. [Google Scholar] [CrossRef]
- Lin, A.; Devlin, G.; Lee, M.; Kerr, A.J. Performance of the GRACE scores in a New Zealand acute coronary syndrome cohort. Heart 2014, 100, 1960–1966. [Google Scholar] [CrossRef]
- Ellis, K.L.; Frampton, C.M.; Pilbrow, A.P.; Troughton, R.W.; Doughty, R.N.; Whalley, G.A.; Ellis, C.J.; Skelton, L.; Thomson, J.; Yandle, T.G.; et al. Genomic risk variants at 1p13.3, 1q41, and 3q22.3 are associated with subsequent cardiovascular outcomes in healthy controls and in established coronary artery disease. Circ. Cardiovasc. Genet. 2011, 4, 636–646. [Google Scholar] [CrossRef]
- Ellis, K.L.; Pilbrow, A.P.; Frampton, C.M.; Doughty, R.N.; Whalley, G.A.; Ellis, C.J.; Palmer, B.R.; Skelton, L.; Yandle, T.G.; Palmer, S.C.; et al. A Common Variant at Chromosome 9P21.3 Is Associated With Age of Onset of Coronary Disease but Not Subsequent Mortality. Circ. Cardiovasc. Genet. 2010, 3, 286–293. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.S.; Tragante, V.; Schmidt, A.F.; McCubrey, R.O.; Holmes, M.V.; Howe, L.J.; Direk, K.; Akerblom, A.; Leander, K.; Virani, S.S.; et al. Subsequent Event Risk in Individuals With Established Coronary Heart Disease. Circ. Genom. Precis. Med. 2019, 12, e002470. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.S.; Schmidt, A.F.; Tragante, V.; McCubrey, R.O.; Holmes, M.V.; Howe, L.J.; Direk, K.; Akerblom, A.; Leander, K.; Virani, S.S.; et al. Association of Chromosome 9p21 With Subsequent Coronary Heart Disease Events A GENIUS-CHD Study of Individual Participant Data. Circ. Genom. Precis. Med. 2019, 12, e002471. [Google Scholar] [PubMed]
- Chowdhury, R.; Cardiology Research Group; Alam, D.S.; Fakir, I.I.; Adnan, S.D.; Naheed, A.; Tasmin, I.; Monower, M.; Hossain, F.; Hossain, F.M.; et al. The Bangladesh Risk of Acute Vascular Events (BRAVE) Study: Objectives and design. Eur. J. Epidemiol. 2015, 30, 577–587. [Google Scholar] [CrossRef]
- Saleheen, D.; Zaidi, M.; Rasheed, A.; Ahmad, U.; Hakeem, A.; Murtaza, M.; Kayani, W.; Faruqui, A.; Kundi, A.; Zaman, K.S.; et al. The Pakistan Risk of Myocardial Infarction Study: A resource for the study of genetic, lifestyle and other determinants of myocardial infarction in South Asia. Eur. J. Epidemiol. 2009, 24, 329–338. [Google Scholar] [CrossRef]
- Dikilitas, O.; Schaid, D.J.; Kosel, M.L.; Carroll, R.J.; Chute, C.G.; Denny, J.A.; Fedotov, A.; Feng, Q.; Hakonarson, H.; Jarvik, G.P.; et al. Predictive Utility of Polygenic Risk Scores for Coronary Heart Disease in Three Major Racial and Ethnic Groups. Am. J. Hum. Genet. 2020, 106, 707–716. [Google Scholar] [CrossRef]
- Martin, A.R.; Gignoux, C.R.; Walters, R.K.; Wojcik, G.L.; Neale, B.M.; Gravel, S.; Daly, M.J.; Bustamante, C.D.; Kenny, E.E. Human Demographic History Impacts Genetic Risk Prediction across Diverse Populations. Am. J. Hum. Genet. 2017, 100, 635–649. [Google Scholar] [CrossRef]
- Merriman, T.R.; Wilcox, P.L. Cardio-metabolic disease genetic risk factors among Maori and Pacific Island people in Aotearoa New Zealand: Current state of knowledge and future directions. Ann. Hum. Biol. 2018, 45, 202–214. [Google Scholar] [CrossRef] [PubMed]
- Robertson, S.P.; Hindmarsh, J.H.; Berry, S.; Cameron, V.A.; Cox, M.P.; Dewes, O.; Doughty, R.N.; Gray, G.; Jacobsen, J.C.; Laurence, A.; et al. Genomic medicine must reduce, not compound, health inequities: The case for hauora-enhancing genomic resources for New Zealand. N. Z. Med. J. 2018, 131, 81–89. [Google Scholar] [PubMed]
- Henare, K.L.; Parker, K.E.; Wihongi, H.; Blenkiron, C.; Jansen, R.; Reid, P.; Findlay, M.P.; Lawrence, B.; Hudson, M.; Print, C.G. Mapping a route to Indigenous engagement in cancer genomic research. Lancet Oncol. 2019, 20, e327–e335. [Google Scholar] [CrossRef]
| n | Denominator # | |
|---|---|---|
| Age, years | 61 (53, 69) | 2015 |
| Male | 1589 (78.9) | 2015 |
| Ethnicity | 2015 | |
| Māori | 259 (12.8) | |
| Pacific | 104 (5.2) | |
| Indian | 94 (4.7) | |
| Chinese | 13 (0.6) | |
| European | 1499 (74.4) | |
| Other * | 46 (2.3) | |
| NZDep Index, quintile | 2015 | |
| 1 (least deprived) | 479 (23.8) | |
| 2 | 416 (20.6) | |
| 3 | 361 (17.9) | |
| 4 | 374 (18.6) | |
| 5 (most deprived) | 385 (19.1) | |
| Charlson comorbidity score | 2015 | |
| 0 (no comorbidity) | 1801 (89.4) | |
| 1–2 (moderate) | 189 (9.4) | |
| ≥3 (severe) | 25 (1.2) | |
| Diabetes on admission | 368 (18.5) | 1986 |
| COPD | 126 (6.4) | 1984 |
| Smoking status | 2014 | |
| Never smoked | 838 (41.6) | |
| Ex-smoker | 700 (34.8) | |
| Current smoker | 476 (23.6) | |
| BMI, kg/m2 | 29.5 ± 5.6 | 2015 |
| ≥30 kg/m2 | 788 (39.1) | |
| TC: HDL | 4.7 ± 1.7 | 1872 |
| ≥4 | 1209 (64.6) | |
| LDL cholesterol, mmol/L | 3.0 ± 1.4 | 1985 |
| eGFR, mL/min/1.73 m2 | 79 (68, 92) | 1967 |
| HbA1c, mmol/mol | 1610 | |
| Diabetes | 62 (50, 79) | 320 |
| No diabetes | 38 (35, 40) | 1290 |
| n (%) | Denominator | |
|---|---|---|
| Type of acute coronary syndrome | 2015 | |
| STEMI | 827 (41.0) | |
| Non-STEMI | 1083 (53.8) | |
| Unstable angina | 105 (5.2) | |
| Coronary angiography | 1968 (99.1) | 1985 |
| Ejection fraction assessed | 1824 (92.4) | 1974 |
| Ejection fraction <50% | 693 (35.1) | |
| GRACE in-hospital score | 1984 | |
| Low <1% | 580 (29.2) | |
| Intermediate 1–2% | 864 (43.6) | |
| High ≥3% | 540 (27.2) | |
| Management | 1971 | |
| PCI | 1408 (71.4) | |
| CABG | 371 (18.8) | |
| Medications on admission | 2015 | |
| Statin | 493 (24.5) | |
| Beta-blocker | 241 (12.0) | |
| ACEi/ARB | 566 (28.1) | |
| Aspirin | 324 (16.1) | |
| Medications on discharge | 1972 | |
| Statin | 1922 (97.5) | |
| Beta-blocker | 1652 (83.8) | |
| ACEi/ARB | 1550 (78.6) | |
| Aspirin | 1927 (97.7) | |
| Other antiplatelet agent | 1636 (83.0) | |
| Dual antiplatelet therapy | 1611 (81.7) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Legget, M.E.; Cameron, V.A.; Poppe, K.K.; Aish, S.; Earle, N.; Choi, Y.; Bradbury, K.E.; Wall, C.; Stewart, R.; Kerr, A.; et al. The Multi-Ethnic New Zealand Study of Acute Coronary Syndromes (MENZACS): Design and Methodology. Cardiogenetics 2021, 11, 84-97. https://doi.org/10.3390/cardiogenetics11020010
Legget ME, Cameron VA, Poppe KK, Aish S, Earle N, Choi Y, Bradbury KE, Wall C, Stewart R, Kerr A, et al. The Multi-Ethnic New Zealand Study of Acute Coronary Syndromes (MENZACS): Design and Methodology. Cardiogenetics. 2021; 11(2):84-97. https://doi.org/10.3390/cardiogenetics11020010
Chicago/Turabian StyleLegget, Malcolm. E., Vicky. A. Cameron, Katrina. K. Poppe, Sara Aish, Nikki Earle, Yeunhyang Choi, Kathryn. E. Bradbury, Clare Wall, Ralph Stewart, Andrew Kerr, and et al. 2021. "The Multi-Ethnic New Zealand Study of Acute Coronary Syndromes (MENZACS): Design and Methodology" Cardiogenetics 11, no. 2: 84-97. https://doi.org/10.3390/cardiogenetics11020010
APA StyleLegget, M. E., Cameron, V. A., Poppe, K. K., Aish, S., Earle, N., Choi, Y., Bradbury, K. E., Wall, C., Stewart, R., Kerr, A., Harrison, W., Devlin, G., Troughton, R., Richards, A. M., Porter, G., Gladding, P., Rolleston, A., & Doughty, R. N. (2021). The Multi-Ethnic New Zealand Study of Acute Coronary Syndromes (MENZACS): Design and Methodology. Cardiogenetics, 11(2), 84-97. https://doi.org/10.3390/cardiogenetics11020010






