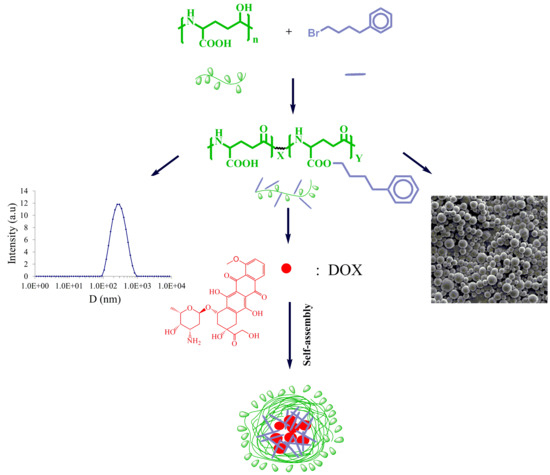
Hydrophobic Modification of Poly(γ-glutamic acid) by Grafting 4-Phenyl-butyl Side Groups for the Encapsulation and Release of Doxorubicin
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Characterization
2.3. Esterification of PGGA
2.4. Nanoparticle Preparation
2.5. Stability of Nanoparticles in Solution
2.6. Doxorubicin Loading and Releasing
3. Results and Discussion
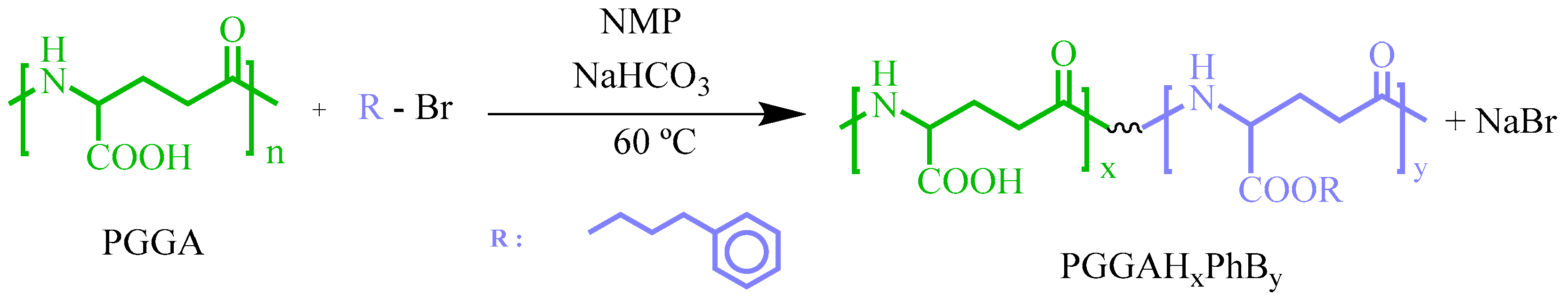
3.1. PGGA Esterification
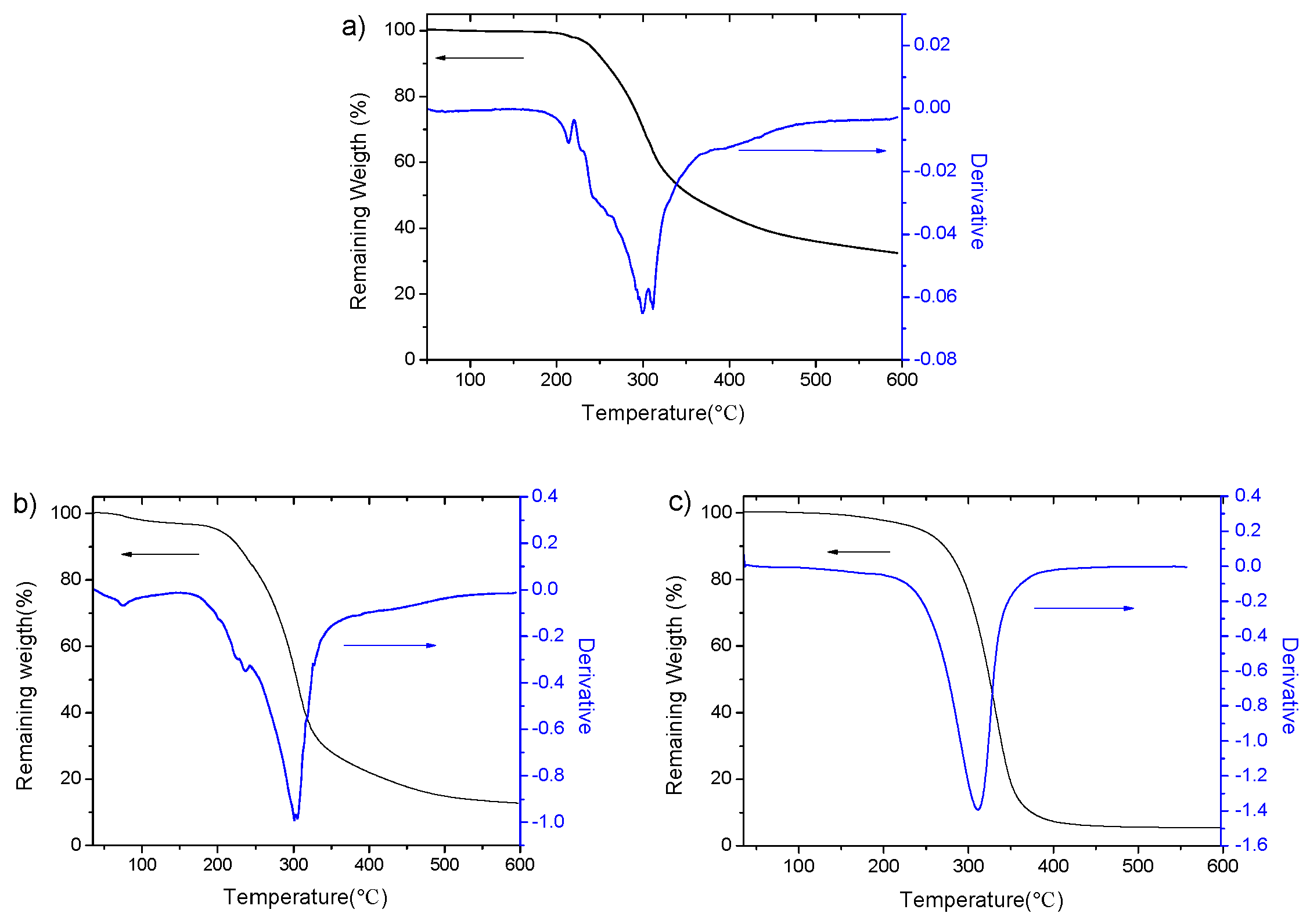
3.2. Characterization of Copolymers
3.3. Preparation, Characterization, Morphology, and Stability of PGGAHxPhBy Nanoparticles over Time
3.4. Doxorubicin Loading and Encapsulation Efficiency
3.5. In Vitro Drug Release Behavior of NPs
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chhikara, B.S.; Parang, K. Global Cancer Statistics 2022: The trends projection analysis. Chem. Biol. Lett. 2022, 10, 451. [Google Scholar]
- Watanabe, T.; Maeda, K.; Kondo, T.; Nakayama, H.; Horita, S.; Kusuhara, H.; Sugiyama, Y. Prediction of the Hepatic and Renal Clearance of Transporter Substrates in Rats Using in Vitro Uptake Experiments. Drug Metab. Dispos. 2009, 37, 1471–1479. [Google Scholar] [CrossRef]
- Basak, D.; Arrighi, S.; Darwiche, Y.; Deb, S. Comparison of Anticancer Drug Toxicities: Paradigm Shift in Adverse Effect Profile. Life 2022, 12, 48. [Google Scholar] [CrossRef]
- Goldstein, M.J.; Peters, M.; Weber, B.L.; Davis, C.B. Optimizing the Therapeutic Window of Targeted Drugs in Oncology: Potency-Guided First-in-Human Studies. Clin. Transl. Sci. 2021, 14, 536–543. [Google Scholar] [CrossRef]
- Hamidi, M.; Azadi, A.; Rafiei, P.; Ashrafi, H. A Pharmacokinetic Overview of Nanotechnology-Based Drug Delivery Systems: An ADME-Oriented Approach. Crit. Rev. Ther. Drug Carr. Syst. 2013, 30, 435–467. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.-S.; Chien, S. Chemotherapeutic engineering: Application and further development of chemical engineering principles for chemotherapy of cancer and other diseases. Chem. Eng. Sci. 2003, 58, 4087–4114. [Google Scholar] [CrossRef]
- Manocha, B.; Margaritis, A. Production and Characterization of γ-Polyglutamic Acid Nanoparticles for Controlled Anticancer Drug Release. Crit. Rev. Biotechnol. 2008, 28, 83–99. [Google Scholar] [CrossRef]
- Kuperkar, K.; Patel, D.; Atanase, L.I.; Bahadur, P. Amphiphilic Block Copolymers: Their Structures, and Self-Assembly to Polymeric Micelles and Polymersomes as Drug Delivery Vehicles. Polymers 2022, 14, 4702. [Google Scholar] [CrossRef] [PubMed]
- Serizawa, T.; Takehara, S.; Akashi, M. Transmission Electron Microscopic Study of Cross-Sectional Morphologies of Core−Corona Polymeric Nanospheres. Macromolecules 2000, 33, 1759–1764. [Google Scholar] [CrossRef]
- Ning, W.; Shang, P.; Wu, J.; Shi, X.; Liu, S. Novel Amphiphilic, Biodegradable, Biocompatible, Thermo-Responsive ABA Triblock Copolymers Based on PCL and PEG Analogues via a Combination of ROP and RAFT: Synthesis, Characterization, and Sustained Drug Release from Self-Assembled Micelles. Polymers 2018, 10, 214. [Google Scholar] [CrossRef]
- Lanz-Landázuri, A.; Martínez de Ilarduya, A.; García-Alvarez, M.; Muñoz-Guerra, S. Modification of microbial polymers by thiol-ene click reaction: Nanoparticle formation and drug encapsulation. React. Funct. Polym. 2016, 106, 143–152. [Google Scholar] [CrossRef]
- Akashi, M.; Kirikihira, I.; Miyauchi, N. Synthesis and polymerization of a styryl terminated oligovinylpyrrolidone macromonomer. Die Angew. Makromol. Chem. 1985, 132, 81–89. [Google Scholar] [CrossRef]
- Tuzar, Z.; Kratochvíl, P. Block and graft copolymer micelles in solution. Adv. Colloid Interface Sci. 1976, 6, 201–232. [Google Scholar] [CrossRef]
- Hiwatari, K.-i.; Serizawa, T.; Seto, F.; Kishida, A.; Muraoka, Y.; Akashi, M. Graft Copolymers Having Hydrophobic Backbone and Hydrophilic Branches XXXIV. Fabrication and Control of Honeycomb Structure Prepared from Amphiphilic Graft Copolymers. Polym. J. 2001, 33, 669–675. [Google Scholar] [CrossRef]
- Xiao, D.; Jia, H.-Z.; Zhang, J.; Liu, C.-W.; Zhuo, R.-X.; Zhang, X.-Z. A Dual-Responsive Mesoporous Silica Nanoparticle for Tumor-Triggered Targeting Drug Delivery. Small 2014, 10, 591–598. [Google Scholar] [CrossRef]
- Hans, M.L.; Lowman, A.M. Biodegradable nanoparticles for drug delivery and targeting. Curr. Opin. Solid State Mater. Sci. 2002, 6, 319–327. [Google Scholar] [CrossRef]
- Wu, J. The Enhanced Permeability and Retention (EPR) Effect: The Significance of the Concept and Methods to Enhance Its Application. J. Pers. Med. 2021, 11, 771. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; van der Meel, R.; Chen, X.; Lammers, T. The EPR effect and beyond: Strategies to improve tumor targeting and cancer nanomedicine treatment efficacy. Theranostics 2020, 10, 7921–7924. [Google Scholar] [CrossRef] [PubMed]
- Kataoka, K.; Harada, A.; Nagasaki, Y. Block copolymer micelles for drug delivery: Design, characterization and biological significance. Adv. Drug Deliv. Rev. 2001, 47, 113–131. [Google Scholar] [CrossRef]
- Davis, M.E.; Chen, Z.; Shin, D.M. Nanoparticle therapeutics: An emerging treatment modality for cancer. Nat. Rev. Drug Discov. 2008, 7, 771–782. [Google Scholar] [CrossRef]
- Lee, J.; Cho, E.C.; Cho, K. Incorporation and release behavior of hydrophobic drug in functionalized poly(D,L-lactide)-block–poly(ethylene oxide) micelles. J. Control. Release 2004, 94, 323–335. [Google Scholar] [CrossRef]
- Samadi, N.; van Steenbergen, M.J.; van den Dikkenberg, J.B.; Vermonden, T.; van Nostrum, C.F.; Amidi, M.; Hennink, W.E. Nanoparticles Based on a Hydrophilic Polyester with a Sheddable PEG Coating for Protein Delivery. Pharm. Res. 2014, 31, 2593–2604. [Google Scholar] [CrossRef]
- Gericke, M.; Schulze, P.; Heinze, T. Nanoparticles Based on Hydrophobic Polysaccharide Derivatives—Formation Principles, Characterization Techniques, and Biomedical Applications. Macromol. Biosci. 2020, 20, 1900415. [Google Scholar] [CrossRef]
- Deming, T.J. Synthesis of Side-Chain Modified Polypeptides. Chem. Rev. 2016, 116, 786–808. [Google Scholar] [CrossRef] [PubMed]
- Arkaban, H.; Barani, M.; Akbarizadeh, M.R.; Pal Singh Chauhan, N.; Jadoun, S.; Dehghani Soltani, M.; Zarrintaj, P. Polyacrylic Acid Nanoplatforms: Antimicrobial, Tissue Engineering, and Cancer Theranostic Applications. Polymers 2022, 14, 1259. [Google Scholar] [CrossRef]
- Hamaguchi, T.; Matsumura, Y.; Suzuki, M.; Shimizu, K.; Goda, R.; Nakamura, I.; Nakatomi, I.; Yokoyama, M.; Kataoka, K.; Kakizoe, T. NK105, a paclitaxel-incorporating micellar nanoparticle formulation, can extend in vivo antitumour activity and reduce the neurotoxicity of paclitaxel. Br. J. Cancer 2005, 92, 1240–1246. [Google Scholar] [CrossRef] [PubMed]
- Ogunleye, A.; Bhat, A.; Irorere, V.U.; Hill, D.; Williams, C.; Radecka, I. Poly-γ-glutamic acid: Production, properties and applications. Microbiology 2015, 161, 1–17. [Google Scholar] [CrossRef]
- Luo, Z.; Guo, Y.; Liu, J.; Qiu, H.; Zhao, M.; Zou, W.; Li, S. Microbial synthesis of poly-γ-glutamic acid: Current progress, challenges, and future perspectives. Biotechnol. Biofuels 2016, 9, 134. [Google Scholar] [CrossRef]
- Muñoz-Guerra, S.; García-Alvarez, M.; Portilla-Arias, J.A. Chemical Modification of Microbial Poly(gamma-glutamic acid). J. Renew. Mater. 2013, 1, 42–60. [Google Scholar] [CrossRef]
- Portilla-Arias, J.A.; Camargo, B.; Garcia-Alvarez, M.; Martinez de Ilarduya, A.; Munoz-Guerra, S. Nanoparticles Made of Microbial Poly(gamma-glutamate)s for Encapsulation and Delivery of Drugs and Proteins. J. Biomater. Sci.-Polym. Ed. 2009, 20, 1065–1079. [Google Scholar] [CrossRef]
- Cedrati, V.; Pacini, A.; Nitti, A.; Martínez de Ilarduya, A.; Muñoz-Guerra, S.; Sanyal, A.; Pasini, D. “Clickable” bacterial poly(γ-glutamic acid). Polym. Chem. 2020, 11, 5582–5589. [Google Scholar] [CrossRef]
- Khalil, I.R.; Burns, A.T.; Radecka, I.; Kowalczuk, M.; Khalaf, T.; Adamus, G.; Johnston, B.; Khechara, M.P. Bacterial-Derived Polymer Poly- γ -Glutamic Acid (γ -PGA)-Based Micro/Nanoparticles as a Delivery System for Antimicrobials and Other Biomedical Applications. Int. J. Mol. Sci. 2017, 18, 313. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Kubota, H.; Nambu, Y.; Endo, T. Convenient esterification of poly(γ-glutamic acid) produced by microorganism with alkyl halides and their thermal properties. J. Polym. Sci. Part A Polym. Chem. 1995, 33, 85–88. [Google Scholar] [CrossRef]
- del Rosario, L.S.; Demirdirek, B.; Harmon, A.; Orban, D.; Uhrich, K.E. Micellar Nanocarriers Assembled from Doxorubicin-Conjugated Amphiphilic Macromolecules (DOX–AM). Macromol. Biosci. 2010, 10, 415–423. [Google Scholar] [CrossRef]
- Price, D.J.; Khuphe, M.; Davies, R.P.W.; McLaughlan, J.R.; Ingram, N.; Thornton, P.D. Poly(amino acid)-polyester graft copolymer nanoparticles for the acid-mediated release of doxorubicin. Chem. Commun. 2017, 53, 8687–8690. [Google Scholar] [CrossRef]
- Rivankar, S. An overview of doxorubicin formulations in cancer therapy. J. Cancer Res. Ther. 2014, 10, 853–858. [Google Scholar] [CrossRef]
- Lu, B.; Lv, X.; Le, Y. Chitosan-Modified PLGA Nanoparticles for Control-Released Drug Delivery. Polymers 2019, 11, 304. [Google Scholar] [CrossRef]
- Li, M.; Song, W.; Tang, Z.; Lv, S.; Lin, L.; Sun, H.; Li, Q.; Yang, Y.; Hong, H.; Chen, X. Nanoscaled Poly(L-glutamic acid)/Doxorubicin-Amphiphile Complex as pH-responsive Drug Delivery System for Effective Treatment of Nonsmall Cell Lung Cancer. ACS Appl. Mater. Interfaces 2013, 5, 1781–1792. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Zhang, L.; Liu, M.; Le, Y.; Lv, S.; Wang, J.; Chen, J.-F. Dual-responsive star-shaped polypeptides for drug delivery. RSC Adv. 2016, 6, 6368–6377. [Google Scholar] [CrossRef]
- Du, Y.; Chen, W.; Zheng, M.; Meng, F.; Zhong, Z. pH-sensitive degradable chimaeric polymersomes for the intracellular release of doxorubicin hydrochloride. Biomaterials 2012, 33, 7291–7299. [Google Scholar] [CrossRef]
- Nukolova, N.V.; Oberoi, H.S.; Cohen, S.M.; Kabanov, A.V.; Bronich, T.K. Folate-decorated nanogels for targeted therapy of ovarian cancer. Biomaterials 2011, 32, 5417–5426. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Sun, L.; Liu, Y.; Yao, H.; Jiang, S.; Li, Y.; Zhang, Y. To reduce premature drug release while ensuring burst intracellular drug release of solid lipid nanoparticle-based drug delivery system with clathrin modification. Nanomedicine 2019, 15, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.-Y.; Hu, Z.-H.; Jin, T. Sustained-release microspheres of amifostine for improved radio-protection, patient compliance, and reduced side effects. Drug Deliv. 2016, 23, 3704–3711. [Google Scholar] [CrossRef]
- Kocak, G.; Tuncer, C.; Bütün, V. pH-Responsive polymers. Polym. Chem. 2017, 8, 144–176. [Google Scholar] [CrossRef]
- Jiang, T.-Y.; Wang, Z.-Y.; Tang, L.-X.; Mo, F.-K.; Chen, C. Polymer micellar aggregates of novel amphiphilic biodegradable graft copolymer composed of poly(aspartic acid) derivatives: Preparation, characterization, and effect of pH on aggregation. J. Appl. Polym. Sci. 2006, 99, 2702–2709. [Google Scholar] [CrossRef]
- Dong, S.; Tang, Y.; He, P.; Ma, S.; Song, W.; Deng, M.; Tang, Z. Hydrophobic modified poly(L-glutamic acid) graft copolymer micelles with ultrahigh drug loading capacity for anticancer drug delivery. Polym. Int. 2022, 71, 487–494. [Google Scholar] [CrossRef]
- Bruschi, M.L. 5—Mathematical models of drug release. In Strategies to Modify the Drug Release from Pharmaceutical Systems; Bruschi, M.L., Ed.; Woodhead Publishing: Sawston, UK, 2015; pp. 63–86. [Google Scholar]
- Yilmaz, E.; Yalinca, Z.; Yahya, K.; Sirotina, U. pH responsive graft copolymers of chitosan. Int. J. Biol. Macromol. 2016, 90, 68–74. [Google Scholar] [CrossRef]
- Yoo, H.S.; Park, T.G. Biodegradable polymeric micelles composed of doxorubicin conjugated PLGA-PEG block copolymer. J. Control. Release 2001, 70, 63–70. [Google Scholar] [CrossRef]
- Hovorka, O.; St’astný, M.; Etrych, T.; Subr, V.; Strohalm, J.; Ulbrich, K.; Ríhová, B. Differences in the intracellular fate of free and polymer-bound doxorubicin. J. Control. Release 2002, 80, 101–117. [Google Scholar] [CrossRef]
- Lee, Y.; Park, S.Y.; Mok, H.; Park, T.G. Synthesis, Characterization, Antitumor Activity of Pluronic Mimicking Copolymer Micelles Conjugated with Doxorubicin via Acid-Cleavable Linkage. Bioconj. Chem. 2008, 19, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Wen, X.; Liang, J.; Gu, Z.; Zhang, X.; Fan, Y. A pH-sensitive nano drug delivery system derived from pullulan/doxorubicin conjugate. J. Biomed. Mater. Res. Part B Appl. Biomater. 2009, 89, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Hrubý, M.; Konák, C.; Ulbrich, K. Polymeric micellar pH-sensitive drug delivery system for doxorubicin. J. Control. Release 2005, 103, 137–148. [Google Scholar] [CrossRef] [PubMed]





| Copolymer | Feed 1 | Esterification Degree 2 | Yield | Mw 3 | Ð |
|---|---|---|---|---|---|
| (mg:mL:mg) | (%) | (%) | (g·mol−1) | ||
| PGGAH95PhB5 | 500:0.029:340 | 4.6 | 92 | - | - |
| PGGAH89PhB11 | 500:0.135:340 | 11 | 55 | - | - |
| PGGAH70PhB30 | 500:0.270:340 | 29.9 | 75 | - | - |
| PGGAH54PhB46 | 500:0.810:340 | 46.2 | 77 | 23,550 | 1.9 |
| PGGAH37PhB63 | 500:0.675:639 | 63 | 90 | 22,900 | 1.9 |
| PGGAH25PhB75 | 500:1.080:1022 | 75 | 94 | 33,200 | 1.7 |
| PGGAH3PhB97 | 500:1.282:1210 | 97 | 97 | 34,550 | 1.9 |
| Copolymers | Td 1 (°C) | Td1/Td2/Td3 2 (°C) | RW 3 (%) |
|---|---|---|---|
| PGGA | 282 | 299/311 | 31.3 |
| PGGAH95PhB5 | 256 | 311/245/216 | 28.5 |
| PGGAH89PhB11 | 252 | 309/216 | 27.5 |
| PGGAH70PhB30 | 235 | 294/354/255 | 26.1 |
| PGGAH54PhB46 | 233 | 304/308/240 | 12.9 |
| PGGAH37PhB63 | 237 | 322/205 | 8.9 |
| PGGAH25PhB75 | 268 | 331 | 4.4 |
| PGGAH3PhB97 | 274 | 332 | 5.4 |
| Copolymers | Nanoprecipitation | Nanoemulsion | ||||
|---|---|---|---|---|---|---|
| D | Pd.I 1 | ζ-Pot | D | Pd.I 1 | ζ-Pot | |
| (nm) | (mV) | (nm) | (mV) | |||
| PGGAH95PhB5 | 374 ± 8.3 | 0.15 ± 0.03 | −48.4 ± 0.8 | - | - | - |
| PGGAH89PhB11 | 296 ± 9.8 | 0.17 ± 0.03 | −49.5 ± 1.1 | - | - | - |
| PGGAH70PhB30 | 180 ± 2.1 | 0.29 ± 0.02 | −31.7 ± 1.3 | - | - | - |
| PGGAH54PhB46 | 297 ± 1.5 | 0.22 ± 0.01 | −40.6 ± 1.0 | - | - | - |
| PGGAH37PhB63 | 184 ± 0.7 | 0.04 ± 0.01 | −35.6 ± 0.1 | 245 ± 4.2 | 0.24 ± 0.02 | −40.0 ± 1.4 |
| PGGAH25PhB75 | 89 ± 1.8 | 0.40 ± 0.01 | −35.5 ± 0.6 | 175 ± 1.6 | 0.16 ± 0.01 | −37.1 ± 1.6 |
| PGGAH3PhB97 | 146 ± 2.0 | 0.38 ± 0.01 | −30.1 ± 0.4 | 157 ± 4.6 | 0.10 ± 0.01 | −13.1 ± 1.2 |
| Copolymers | Week 1 | Week 2 | Week 3 | Week 4 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D (nm) | Pd.I | ζ-Pot (mV) | D (nm) | Pd.I | ζ-Pot (mV) | D (nm) | Pd.I | ζ-Pot (mV) | D (nm) | Pd.I | ζ-Pot (mV) | |
| PGGAH54PhB46 | 258 ± 1.3 | 0.16 ± 0.00 | −36.9 ± 2.4 | 261 ± 0.5 | 0.16 ± 0.00 | −35.9 ± 0.9 | 254 ± 3.6 | 0.13 ± 0.01 | −36.8 ± 0.9 | 258 ± 0.9 | 0.18 ± 0.00 | −35.0 ± 0.4 |
| PGGAH37PhB63 | 186 ± 0.6 | 0.04 ± 0.04 | −35.0 ± 1.5 | 184 ± 1.7 | 0.03 ± 0.01 | −36.9 ± 0.5 | 184 ± 1.7 | 0.08 ± 0.00 | −40.7 ± 1.0 | 183 ± 1.0 | 0.03 ± 0.02 | −41.6 ± 3.4 |
| PGGAH25PhB75 | 86 ± 0.4 | 0.28 ± 0.00 | −33.1 ± 9.1 | 82 ± 1.1 | 0.29 ± 0.00 | −30.4 ± 7.9 | 76 ± 1.7 | 0.27 ± 0.00 | −29.8 ± 0.7 | 76 ± 0.3 | 0.27 ± 0.00 | −20.5 ± 2.1 |
| Copolymers | Unloaded | Loaded Doxorubicin | ||||
|---|---|---|---|---|---|---|
| D (nm) | ζ-Pot (mV) | D (nm) | ζ-Pot (mV) | EE% 1 | DL% 2 | |
| PGGAH89PhB11 | 296 ± 9.8 | −49.5 ± 1.1 | 1594 ± 57.7 | −49.6 ± 12.0 | 27 | 6.0 |
| PGGAH70PhB30 | 180 ± 2.0 | −31.7 ± 1.3 | 216 ± 3.3 | −42.4 ± 2.9 | 36 | 7.1 |
| PGGAH54PhB46 | 163 ± 1.5 | −40.6 ± 1.0 | 170 ± 4.9 | −47.4 ± 1.7 | 46 | 10.1 |
| PGGAH3PhB97 | 146 ± 2.0 | −30.1 ± 0.4 | 102 ± 0.2 | −40.3 ± 2.0 | 40 | 8.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dorost, P.; García-Alvarez, M.; Martínez de Ilarduya, A. Hydrophobic Modification of Poly(γ-glutamic acid) by Grafting 4-Phenyl-butyl Side Groups for the Encapsulation and Release of Doxorubicin. Pharmaceutics 2023, 15, 1377. https://doi.org/10.3390/pharmaceutics15051377
Dorost P, García-Alvarez M, Martínez de Ilarduya A. Hydrophobic Modification of Poly(γ-glutamic acid) by Grafting 4-Phenyl-butyl Side Groups for the Encapsulation and Release of Doxorubicin. Pharmaceutics. 2023; 15(5):1377. https://doi.org/10.3390/pharmaceutics15051377
Chicago/Turabian StyleDorost, Porochista, Montserrat García-Alvarez, and Antxon Martínez de Ilarduya. 2023. "Hydrophobic Modification of Poly(γ-glutamic acid) by Grafting 4-Phenyl-butyl Side Groups for the Encapsulation and Release of Doxorubicin" Pharmaceutics 15, no. 5: 1377. https://doi.org/10.3390/pharmaceutics15051377
APA StyleDorost, P., García-Alvarez, M., & Martínez de Ilarduya, A. (2023). Hydrophobic Modification of Poly(γ-glutamic acid) by Grafting 4-Phenyl-butyl Side Groups for the Encapsulation and Release of Doxorubicin. Pharmaceutics, 15(5), 1377. https://doi.org/10.3390/pharmaceutics15051377